Abstract
Heart development is a complex process regulated by multi-layered genetic as well epigenetic regulators many of which are still unknown. Besides their critical role during cardiac development, these molecular regulators emerge as key modulators of cardiovascular pathologies, where fetal cardiac genes’ re-expression is witnessed. MicroRNAs have recently emerged as a crucial part of signalling cascade in both development and diseases. We aimed to identify, validate, and perform functional annotation of putative novel miRNAs using chicken as a cardiac development model system. Novel miRNAs were obtained through deep sequencing of small RNAs extracted from chicken embryonic cardiac tissue of different developmental stages. After filtering out real pre-miRNAs, their expression analysis, potential target gene’s prediction and functional annotations were performed. Expression analysis revealed that miRNAs were differentially expressed during different developmental stages of chicken heart. The expression of selected putative novel miRNAs was further validated by real-time PCR. Our analysis indicated the presence of novel cardiac miRNAs that might be regulating critical cardiac development events such as cardiac cell growth, differentiation, cardiac action potential generation and signal transduction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cardiomyogenesis is the first sign of organogenesis during embryonic development. The process is strictly coordinated, timed, and calibrated through genetic and epigenetic regulation of the cardiac genome [1]. Most of the underlying pathways such as Wnt signalling, Insulin signaling and Hedgehog signaling are considered to be conserved from insects to vertebrates, making it convenient to extrapolate the observations made in any model organism to humans [2]. The fetal cardiac molecular events reactivate in case of pathological cardiomyocyte hypertrophy. These include hyperpolarization of cyclic nucleotide-gated calcium channel, elevated expression of β-myosin heavy chain and the suppression of α-myosin heavy chain [3, 4]. Therefore, identification and characterization of underlying molecular mechanisms regulating cardiogenesis will provide better understanding of fetal cardiac gene programming as well as adaptive responses during heart failure.
Although a high number of genes expressed during cardiac development have been identified, their regulation through different genetic as well as epigenetic factors is still not clear. MicroRNAs (miRNAs) are one such epigenetic entity that regulates the gene expression at post-transcriptional level via seed pairing to the 3’UTR of mRNA leading to its degradation or translational repression [5]. Diverse families of miRNAs, such as miR-1, miR-133 and miR-199 targeting multiple genes, were found to regulate cardiac development and cardiomyocyte proliferation in mice, zebrafish and rats [6]. Knockout mice lacking DGCR8/Dicer (miRNA processing enzyme) were found to develop defects in cardiac development and premature death with signs of cardiac arrest [7]. Our previous study also identified and validated the differential expression of highly conserved known miRNAs during different developmental stages of cardiac development [8]. However, the total count of miRNAs identified from developing heart is far less than the total genes present in each organism, indicating that a myriad of miRNAs and their implications on cardiac development are yet to be discovered.
The conservation of vertebrate heart development between chicken, mice, zebrafish and human is well known. It begins with transformation of mesoderm into myocardium, tract formation, looping, trabeculation, septation, valvulogenesis and growth, leading to the formation of four-chambered heart [9]. They share a large number of common miRNAs such as let-7, miR-30, miR-22 and miR-140 as well as transcription factors such Nkx2.5, GATA and tbx5 [10]. However, several transcriptional and post-transcriptional regulatory factors involved during cardiac morphogenesis are still unclear. In the present study, we performed a comprehensive identification and characterization of novel miRNAs using chicken model for their role in cardiac development. Chicken model for developmental studies provides additional advantage as its heart has the closest anatomy to human heart during fetal development and post-fetal life in non-mammals [11]. From the myriad of putative novel miRNAs, we identified real novel miRNAs, validated them using real time PCR, predicted their potential target genes and developed a network of miRNA-targeting genes’ interactions.
Materials and methods
Putative novel miRNA sequences obtained from chicken cardiac miRNA datasets
In total, six small RNA libraries were prepared and sequenced from different stages (4th, 6th, 8th, 10th, 12th and 14th days) of developing embryonic chicken heart tissue using Next generation sequencing (NCBI, GEO accession: GSE69663) [8]. In brief, the small RNA libraries were searched on miRBase (v.19.0) for identification of miRNAs. Precursor miRNA sequences were predicted using Gallus gallus genome (v.2.1). The total counts of each miRNA were used as an input in DeSeq module to identify the differential expressions of miRNAs across the input samples.
Analysis of novel miRNAs characteristics and expression pattern using in silico analysis
Common novel miRNA candidates with cut-off at ten counts were selected from each stage of cardiac development. To authenticate each putative novel sequence, following criteria were set: (a) the precursor miRNA should form hairpin loop structure with mature miRNA present in one arm without large loops, (b) the minimum free energy (MFE) of binding should be < − 20 kcal/mol, (c) the hairpin should be derived from intragenic or intronic regions, and (d) the precursor miRNA should not align with any known miRNAs or any other non-coding RNAs. For validating these criteria, we used four different miRNA tool/databases: (a) MirEval 2.0 [12], (b) MiPred [13], RNA fold [14] and (c) miRBase version 22 [15]. The putative novel sequences fulfilling all the set criteria under different database were filtered and selected for functional characterization. Expression profiling of filtered novel miRNAs’ throughout embryonic heart development was carried out, and heat map was generated using log2 RPM (Reads per million) by Cimminer (https://discover.nci.nih.gov/cimminer/oneMatrix.do).
Potential target gene prediction and functional annotation
Potential target genes of all the validated novel miRNAs were predicted under custom prediction of miRDB database with prediction score of > 80 (most likely to be real) [16]. To predict the role of target genes, gene ontology was performed using PANTHER 14.1, a web-based tool for connecting genes to their cellular, molecular and biological processes [17]. Similarly, target genes were subjected to pathway enrichment analysis by DAVID 6.8 [18]. Interaction network between novel miRNAs, their targets genes and pathways was constructed using Cytoscape [19].
Conservation analysis
Seed-paralogue and homology were searched using BLASTN under mirBase database with default E-value cut-off. Hairpin or precursor sequence was used for multiple sequence alignment on UCSC genome browser (ICGSC Gallus-gallus 4.0/galGal4). In addition, BLAT (UCSC) was used to find perfect sequence matches of mature miRNAs in Gallus-gallus and to reconfirm the alignment of intronic as well as intergenic sequences.
miRNA isolation, cDNA preparation and expression analyses by RT-PCR
Small RNAs were isolated from cardiac tissues of different developmental stages as described previously [8]. The purities and concentrations of RNAs were confirmed using NanoDrop ND–1000 spectrophotometer (Nanodrop Technologies, USA). MicroRNA stem-loop primers were designed for five novel miRNAs as described previously [20, 21] (Table 1). The stem-loop reverse transcription was carried out using the RevertAid First Strand cDNA Synthesis Kit (Thermo Scientific, USA) as per the manufacturer’s instructions. Real-time PCR was performed using SYBR green fluorescence quantitative PCR reagent kit (Thermofischer Scientific, USA) and Piko Real-Time 96 (Thermofischer Scientific, USA). The 10 μL reaction mixture constituted 5 μL SYBR green real-time PCR Master Mix, 0.5 μL forward and reverse primers each (1 μM), 1 μL stem-loop cDNA product, and 3 μL nuclease-free water. The PCR reaction conditions were as follows: initial denaturation at 95 °C for 30 s, followed by 40 cycles of 95 °C for 15 s and 55 °C for 30 s. All the reactions were performed in triplicates. miRNAs crossing the threshold in < 35 cycles were considered to be expressed [22]. The relative expression of miRNA was measured using 2−ΔΔCq after normalization to U6 snRNA used as internal control. To verify the specificity of amplification, the PCR products were visualized on 4% agarose gel.
Statistical analysis
Significant p-values were obtained by Fisher’s exact test from DAVID database to determine the probability that each biological, cellular and molecular function assigned to the dataset is due to chance alone. Enriched targets and their pathways (including KEGG pathways) with p-values < 0.05, and enrichment score of > 1.5 were considered. Each experiment was conducted thrice, and data were expressed as mean ± SEM. The relative differential expression of miRNAs between two cardiac development stages were compared using Student t test. p values of < 0.05 were considered to be statistically significant.
Results
Identification of novel miRNA from chicken fetal heart
We extracted putative novel miRNA sequences from NGS datasets of chicken fetal heart and detected 55 novel miRNAs with abundance > 10 between 4th, 6th, 8th, 10th, 12th, and 14th day of embryonic development. All the novel miRNAs were validated using MirEval, MiPred, RNA fold and MiRBase databases. Out of 55 putative novel sequences, sixteen novel miRNAs fulfilled all the criteria set for validation and were further processed. The precursor sequence of novel miRNAs possesses MFE < − 20 kcal/mol. Their chromosomal distribution was non-clustered, while their p value of randomization was < 0.05 (Table 1). All the sequences formed miRNA like hairpin structures with mature sequence lying on one of the arms (Fig. 1a). None of the sequences were found to be nearby any of the known miRNA or other non-coding RNAs. Thus, provisional names were assigned using gga for gallus gallus, followed by additional number as per the miRBase nomenclature [23]. Functional annotations of all the 16 novel miRNA target genes were carried out; however, novel miRNAs (namely: gga-mir-N2, gga-mir-N4, gga-mir-N10, gga-mir-N12 and gga-mir-N16) targeting > 30 genes involved in cardiovascular development and diseases were considered for further evaluation.
Precursor and mature nucleotide sequence of novel miRNA. a In silico structure prediction through MirEval and RNA fold depicts hairpin loop structure with mature miRNA sequence lying in one of the arms (red rectangular box). b Multiple sequence alignment representing the nearest known miRNA sequence from miRBase. (Color figure online)
Novel chicken miRNAs were found to be species specific
To learn whether the novel miRNAs share sequence similarities among other vertebrates, multiple sequence alignments of precursor sequence were carried out using UCSC genome browser. The sequences of all the five miRNAs showed 100% alignment with chicken as well as turkey genome. However, we observed no significant similarities between novel miRNAs and any other vertebrates. Sequence search to predict potential miRNA families revealed that sixteen base pairs of gga-mir-N12 (mature sequence) were aligned with another chicken miRNA gga-miR-6560-3p. Importantly, the seed sequences (positions 2–7 inclusive of those from the 5′ end) which determines the commonality among the targeted mRNAs were found to be similar between the two miRNAs. The sequence of gga-mir-N4 was found to be closest to nve-miR-9434 (Nematostella vectensis), while gga-mir-N10 was found to be closest to ath-miR858a (Arabidopsis thaliana) with fifteen nucleotides aligning with each other, respectively. Likewise, closest miRNA for gga-mir-N2 and gga-mir-N16 were found to be hsa-miR-346 and mmu-miR-3073a-5p, respectively; however, the two did not share seed sequence (Fig. 1b). Lastly, we analysed the nucleotide bias of miRNAs by calculating the percentage of four nucleotides appearing at different position and frequency of a particular nucleotide at a particular position. We detected that U was the most-biased nucleotide at the start and the end positions of miRNAs (75%), while U and A dominated the initial and the end positions of miRNAs. However, high percentage of G and C (58%) was present in the mid-region of miRNAs.
Novel miRNAs depicted differential expression during different developmental stages
Since, the stem-loop RT-PCR is considered as one of the most sensitive and specific method to detect miRNA, selected novel miRNAs obtained after in silico analysis were validated using stem-loop RT-PCR. Three novel miRNAs (gga-mir-N2 and gga-mir-N12, and gga-mir-N16) out of five were found to depict amplification suggesting that they were stably expressed in the fetal heart. PCR product of ~ 65 bp size was verified by gel electrophoresis (Fig. 2a). After normalization the expression of miRNAs with U6 snRNA, the relative expression of gga-mir-N12 was found to be significantly higher on 6th and 10th day of development as compared to 4th day (p-value < 0.05). However, after the 10th day, the expression of gga-mir-N12 reduced to its initial level. The expression of gga-mir-N2 showed similar pattern as gga-novel-mir-12; however, the elevation in expression was not statistically significant (p-value > 0.05). Contrasting expression patterns were observed for gga-novel-mir-16. Its expression got significantly elevated during later stages (14th day) of development compared to initial stages (the 4th and 6th days) (Fig. 2b). Remaining two miRNAs (gga-novel-mir-4 and gga-novel-mir-10) were not detected in any of the samples up to 35 cycles of amplification or on agarose gel. Expression analysis through RT-PCR validated the differential expression of miRNAs obtained by cluster analysis of NGS data (Fig. 3).
Real-time PCR analysis. a miRNAs were found expressed differentially between different days of development (*p value < 0.05). b miRNAs PCR products of expected size were visualized on 4% agarose gel. The difference in band intensities verified the differential expressions of miRNAs. (L: 50 bp ladder; 4, 6, 8, 10, 12 and 14 represent days; NC: Non-template control; U6: U6 snRNA)
Critical biological, cellular and molecular processes regulated by miRNAs
To recognize the biological importance of novel miRNAs, their potential target genes were predicted using miRDB (prediction score > 80) and Target Scan (score ≤ − 0.45). Common potential target genes obtained from both the databases were selected. Gene ontology (GO) analysis classified the conserved target genes according to their biological process (BP), molecular function (MF), cellular component (CC), and protein class (PC). The overrepresented biological processes were associated with clusters related to cell proliferation, cell migration, metabolic and developmental processes. The molecular functions were associated with binding, catalytic activity and transcription regulator activity. The cellular components were related to cell junction, cell membrane, and extracellular matrix. Lastly, the overrepresented protein class were calcium binding proteins, signalling molecules, extracellular matrix proteins, and enzyme modulators (Fig. 4a).
Gene ontology and KEGG pathway analysis of potential genes targeted by miRNAs enriched at different cardiac developmental stages. a GO classification representing percentage of target genes (Y axis) involved in diverse biological role on the basis of biological process, cellular component and molecular function (X axis). b Interactions’ network depicting significantly enriched GO and KEGG pathways (orange colour box), target genes (green colour box) and novel miRNA (Blue colour box). (Color figure online)
To predict the involvement of these miRNAs in cardiac development, we carried out KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway analysis. We observed that target genes of gga-mir-N12 were significantly enriched in insulin signalling pathway (p-value: 0.028, Fold Enrichment: 5.8). In addition, target genes of gga-mir-N2 were significantly associated with cardiac muscle cell action potential (p-value: 0.013, Fold Enrichment: 156.3). Lastly, potential target genes of gga-mir-N16 were found to be significantly involved in Wnt signalling pathway (p-value: 0.009, Fold Enrichment: 8.7). Interaction network was plotted using intersected target genes between novel miRNAs, GO terms and enriched KEGG pathways (Fig. 4b).
Apart from gene ontology and function annotation, DAVID also provides disease association for a group of genes when analysed for humans. Potential target genes of all the novel miRNAs were searched for their association with human diseases. Target genes of gga-mir-N2 were found to be significantly associated with cardiovascular diseases (p-value: 0.014, Fold Enrichment: 1.6). Similarly, target genes of gga-mir-N2 miRNA were found to be significantly associated with developmental disorders (p-value: 0.014, Fold Enrichment: 2).
Discussion
Cardiac development is one of the most complex and critical phenomena, as it is the first organ to be developed during embryogenesis. Complete development comprises of several stages such as specification, proliferation and migration of precardiac cells (cardiac progenitors), cardiac remodelling, tract formation, chamber distribution and generation of cardiac action potential. In the chicken embryo, these observations were classified into several stages by Hamburger and Hamilton, also known as HH-stage series. According to HH series, atrial septation, valve development, cardiac looping, and cardiac conduction system begin from HH 21–23 (3.5–4 days) and get completed well before hatching (14th day) [24]. The present study was designed to characterize putative novel miRNAs sequences obtained from chicken fetal cardiac tissue at 4th, 6th, 8th, 10th, 12th and 14th days of development so as to obtain maximum miRNA population present during entire cardiogenesis process. We observed miRNA gga-mir-N2, gga-mir-N12 and gga-mir-N16 to be the real novel miRNAs depicting differential expression during cardiac development. Previously, several miRNAs such as miR-30, miR-338, miR-126-3p, miR-126-5p and miR-140 were reported to express differentially between chicken embryo and adult as well as among different tissue types [25]. These observations suggest that similar to gene expression, their regulator (miRNAs) also depicts differential expressions during development and adult stages. Our findings further extrapolate these observations by establishing that miRNAs not only depict differential expression between embryonic and post embryonic life, but they also express differentially within different stages of embryonic development.
Although, chicken shares most of its miRNAs with other vertebrates, several miRNAs such as gga-miR-757 and gga-miR-1609 are found to be chicken specific [25]. Previous studies have reported that species-specific or non-conserved miRNAs depict low levels of expression. In addition, a few of them may only get detected through deep sequencing technology [26]. Interestingly, all the three novel miRNAs validated in our study were found to be species specific. Further observations derived from nucleotide bias analysis were in agreement with the characteristic of a typical miRNA, where the nucleotide at 5’end is usually U [27]. High percentage of U and A at the ends of miRNA establishes the fact that AU base pairing positively effects mRNA target identification [28].
Generation of cardiac action potential and its propagation are highly essential for the initiation of coordinated contraction of the atrial and ventricular chambers during cardiac development. The cardiac pace making and conduction system which develops throughout all developmental stages is still not fully understood [29]. Few miRNAs such as miR-1 and miR-223-3p have been reported to regulate cardiac action potentials by targeting Kcnd2 transcript [30, 31]. We observed that target genes of gga-mir-N2 such as CACNB2 and CACNA1D were significantly associated with generation of cardiac action potential. Our data also suggest strong association between gga-mir-N12 and insulin signalling via PPP1CC, CALM1 and EIF4E genes. Several studies have reported the regulation of insulin signalling pathway by miR-15b, miR-128a, miR-222, and miR-424-5p [32,33,34,35]. It is one of the most conserved pathways that coordinately regulate cardiac growth (growth, mitogenesis and differentiation) via phosphatidylinositol 3-kinase (PI3 K) and MAPK pathways [36]. In adults, under prolonged excessive glucose (type II diabetes mellitus), the insulin signalling pathway becomes maladaptive leading to cardiac cell enlargement (cardiac hypertrophy) [37]. All the three target genes including PPP1CC, CALM1 and EIF4E are known as critical regulators of insulin pathway; however, none is reported as a target gene of any miRNA obtained from cardiac tissue [38,39,40]. Lastly, we observed a significant association between gga-mir-N16 (the most overexpressed miRNA at the end stage of development) and one of the complex and conserved signalling pathways known as Wnt pathway. Wnt signalling regulates multiple aspects of cardiogenesis, including cardiac specification, cardiomyocyte proliferation and valve formation [41]. Similar observations were reported for overexpressed miR-19b for regulating cardiac development in zebrafish embryo and mice model by suppressing CTNNB1 and GSK3β of Wnt signalling pathway, respectively [42, 43].
As previously discussed, fetal cardiac miRNAs and their target genes get redeployed during cardiac hypertrophy. Switch between MHC gene (Myosin heavy chain) isoforms depicts a classical example of fetal gene re-expression. In embryo, β-MHC along with BRG1 and PARP genes expresses predominantly, while in adult heart, their expression turns off. Later, in hypertrophic heart re-expression of these genes occurs by epigenetic mechanisms to compensate hypertrophic cardiomyopathy [44]. mir-1, mir-208 and mir-133 are believed to be regulating these genes, and they follow similar expression patterns in both embryonic and failing heart [45]. Interestingly, several of the potential targets such as CACNB2, ADAMTS17, PPP1CC and IGFBP2 obtained from our data have been very well reported during cardiovascular diseases. CACNB2 which encodes a subunit of cardiac voltage gated calcium channel is implicated in atrial fibrillation [46]. ADAMTS, a family of protein involved in remodelling of extracellular matrix, has been reported to play an important role in atherosclerosis [47]. Similarly, overexpressions of PPP1CC (a serine/threonine phosphatase that affects cardiac contractile function) and IGFBP2 have been reported to reduce cardiac function and increase the risk of cardiovascular diseases respectively [48, 49].
In conclusion, we have identified and validated differential expression of three novel miRNAs from chicken fetal heart during different development stages. We postulate that these miRNAs might be regulating critical mRNAs expressed during fetal cardiac development. Our study provides new information for the future research of small RNAs for their role in regulating fetal cardiac gene programme and as a new emerging epigenetic agent for plausible future cardiovascular therapeutics. Further validation of mRNA targets will shed more light on the fine-tuning mechanism that underpins cardiac development and fetal gene reprogramming during diseases conditions in vertebrates.
References
Srivastava D, Olson EN (2000) A genetic blueprint for cardiac development. Nature 407:221–226
Abu-Issa R, Kirby ML (2007) Heart field: from mesoderm to heart tube. Annu Rev Cell Dev Biol 23:45–68
Nandi SS, Mishra PK (2015) Harnessing fetal and adult genetic reprograming for therapy of heart disease. J Nat Sci 1:4–6
Barry SP, Davidson SM, Townsend PA (2008) Molecular regulation of cardiac hypertrophy. Int J Biochem Cell Biol 40:2023–2039
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Fuller A, Qian L (2014) MiRiad roles for MicroRNAs in cardiac development and regeneration. Cells 3:724–750
Rao PK, Toyama Y, Chiang HR et al (2009) Loss of cardiac microRNA-mediated regulation leads to dilated cardiomyopathy and heart failure. Circ Res 105:585–594
Rustagi Y, Jaiswal HK, Rawal K et al (2015) Comparative characterization of cardiac development specific microRNAs: fetal regulators for future. PLoS ONE 10:e0139359
Epstein JA (2010) Cardiac development and implications for heart disease. N Engl J Med 363:1638–1647
Katz MG, Fargnoli AS, Kendle AP et al (2016) The role of microRNAs in cardiac development and regenerative capacity. Am J Physiol 310:H528–H541
Wittig J, Münsterberg A (2016) The early stages of heart development: insights from chicken embryos. J Cardiovasc Dev Dis 3:12
Gao D, Middleton R, Rasko JEJ et al (2013) MiREval 2.0: a web tool for simple microRNA prediction in genome sequences. Bioinformatics 29:3225–3226
Jiang P, Wu H, Wang W et al (2007) MiPred: classification of real and pseudo microRNA precursors using random forest prediction model with combined features. Nucleic Acids Res 35:W339–W344
Gruber AR, Lorenz R, Bernhart SH et al (2008) The Vienna RNA websuite. Nucleic Acids Res 36:W70–W74
Kozomara A, Griffiths-Jones S (2014) MiRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 42:D68–D73
Wong N, Wang X (2015) miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Res 43:D146–D152
Mi H, Muruganujan A, Thomas PD (2013) PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res 41:D377–D386
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Varkonyi-Gasic E, Wu R, Wood M et al (2007) Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods 3:12
Yang LH, Wang SL, Tang LL et al (2014) Universal stem-loop primer method for screening and quantification of microRNA. PLoS ONE 9:e115293
Laganà A, Veneziano D, Spata T et al (2015) Identification of general and heart-specific miRNAs in sheep (Ovis aries). PLoS ONE 10:e0143313
Ambros V, Bartel B, Bartel DP et al (2003) A uniform system for microRNA annotation. RNA 9:277–279
Martinsen BJ (2005) Reference guide to the stages of chick heart embryology. Dev Dyn 233:1217–1237
Xu H, Wang X, Du Z, Li N (2006) Identification of microRNAs from different tissues of chicken embryo and adult chicken. FEBS Lett 580:3610–3616
Ji Z, Wang G, Xie Z et al (2012) Identification and characterization of microRNA in the dairy goat (Capra hircus) mammary gland by Solexa deep-sequencing technology. Mol Biol Rep 39:9361–9371
Ge X, Zhang Y, Jiang J et al (2012) Identification of microRNAs in Helicoverpa armigera and Spodoptera litura based on deep sequencing and homology analysis. Int J Biol Sci 9:1–15
Islam MT, Ferdous AS, Najnin RA et al (2015) High-throughput sequencing reveals diverse sets of conserved, nonconserved, and species-specific miRNAs in jute. Int J Genom 2015:125048
Kim GH (2013) MicroRNA regulation of cardiac conduction and arrhythmias. Transl Res 161:381–392
Yang B, Lin H, Xiao J et al (2007) The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2. Nat Med 13:486–491
Liu X, Zhang Y, Du W et al (2016) MiR-223-3p as a novel MicroRNA regulator of expression of voltage-gated K + channel Kv4.2 in acute myocardial infarction. Cell Physiol Biochem 39:102–114
Yang WM, Jeong HJ, Park SW, Lee W (2015) Obesity-induced miR-15b is linked causally to the development of insulin resistance through the repression of the insulin receptor in hepatocytes. Mol Nutr Food Res 59:2303–2314
Motohashi N, Alexander MS, Shimizu-Motohashi Y et al (2013) Regulation of IRS1/Akt insulin signaling by microRNA-128a during myogenesis. J Cell Sci 126:2678–2691
Min KH, Yang WM, Lee W (2018) Saturated fatty acids-induced miR-424–5p aggravates insulin resistance via targeting insulin receptor in hepatocytes. Biochem Biophys Res Commun 503:1587–1593
Ono K, Igata M, Kondo T et al (2018) Identification of microRNA that represses IRS-1 expression in liver. PLoS ONE 13:e0191553
DeBosch BJ, Muslin AJ (2008) Insulin signaling pathways and cardiac growth. J Mol Cell Cardiol 44:855–864
Jia G, Whaley-Connell A, Sowers JR (2018) Diabetic cardiomyopathy: a hyperglycaemia- and insulin-resistance-induced heart disease. Diabetologia 61:21–28
Boucher J, Kleinridders A, Ronald Kahn C (2014) Insulin receptor signaling in normal and insulin-resistant states. Cold Spring Harb Perspect Biol 6:a009191
Chen Z, Ding L, Yang W et al (2017) Hepatic activation of the FAM3C-HSF1-CaM pathway attenuates hyperglycemia of obese diabetic mice. Diabetes 66:1185–1197
Park J, Ahn S, Jayabalan AK et al (2016) Insulin signaling augments eIF4E-dependent nonsense-mediated mRNA decay in mammalian cells. Biochim Biophys Acta 1859:896–905
Tian Y, Cohen ED, Morrisey EE (2010) The importance of Wnt signaling in cardiovascular development. Pediatr Cardiol 31:342–348
Li M, Hu X, Zhu J et al (2014) Overexpression of miR-19b impairs cardiac development in zebrafish by targeting ctnnb1. Cell Physiol Biochem 33:1988–2002
Ye X, Hemida MG, Qiu Y et al (2013) MiR-126 promotes coxsackievirus replication by mediating cross-talk of ERK1/2 and Wnt/β-catenin signal pathways. Cell Mol Life Sci 70:4631–4644
Han P, Li W, Lin CH et al (2014) A long noncoding RNA protects the heart from pathological hypertrophy. Nature 514:102–106
Tian J, An X, Niu L (2017) Role of microRNAs in cardiac development and disease. Exp Ther Med 13:3–8
Ling TY, Wang XL, Chai Q et al (2017) Regulation of cardiac CACNB2 by microRNA-499: potential role in atrial fibrillation. BBA Clin 7:78–84
Rienks M, Barallobre-Barreiro J, Mayr M (2018) The emerging role of the ADAMTS family in vascular diseases. Circ Res 123:1279–1281
Liu R, Correll RN, Davis J et al (2015) Cardiac-specific deletion of protein phosphatase 1β promotes increased myofilament protein phosphorylation and contractile alterations. J Mol Cell Cardiol 87:204–213
Hoeflich A, David R, Hjortebjerg R (2018) Current IGFBP-related biomarker research in cardiovascular disease: we need more structural and functional information in clinical studies. Front Endocrinol 9:388
Acknowledgements
This study was supported by DST-SERB, Government of India (File No: EMR/2016/005914); and CSIR, Government of India (File No: 09/1132 (0004)/18-EMR-I).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11010_2019_3618_MOESM1_ESM.tif
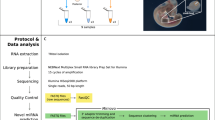
Supplementary material 1—Flowchart depicting the steps taken for identification and validation of novel miRNAs and generation of interaction network between enriched GO terms and KEEG pathways. (TIFF 148 kb)
Rights and permissions
About this article
Cite this article
Saxena, S., Mathur, P., Shukla, V. et al. Differential expression of novel MicroRNAs from developing fetal heart of Gallus gallus domesticus implies a role in cardiac development. Mol Cell Biochem 462, 157–165 (2019). https://doi.org/10.1007/s11010-019-03618-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-019-03618-4