Abstract
This study was aimed to construct classification and regression tree (CART) model of glycosaminoglycans (GAGs) for the differential diagnosis of Mucopolysaccharidoses (MPS). Two-dimensional electrophoresis and liquid chromatography–tandem mass spectrometry (LC–MS/MS) were used for the qualitative and quantitative analysis of GAGs. Specific enzyme assays and targeted gene sequencing were performed to confirm the diagnosis. Machine learning tools were used to develop CART model based on GAG profile. Qualitative and quantitative CART models showed 96.3% and 98.3% accuracy, respectively, in the differential diagnosis of MPS. The thresholds of different GAGs diagnostic of specific MPS types were established. In 60 MPS positive cases, 46 different mutations were identified in six specific genes. Among 31 different mutations identified in IDUA, nine were nonsense mutations and two were gross deletions while the remaining were missense mutations. In IDS gene, four missense, two frameshift, and one deletion were identified. In NAGLU gene, c.1693C > T and c.1914_1914insT were the most common mutations. Two ARSB, one case each of SGSH and GALNS mutations were observed. LC–MS/MS-based GAG pattern showed higher accuracy in the differential diagnosis of MPS. The mutation spectrum of MPS, specifically in IDUA and IDS genes, is highly heterogeneous among the cases studied.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Mucopolysaccharidoses (MPS) are a group of inherited lysosomal storage disorders (LSDs) caused by the deficiency of specific enzymes that catalyze the stepwise degradation of glycosaminoglycans (GAGs) leading to accumulation of fragments of GAGs in the body and subsequent widespread tissue damage [1]. The clinical manifestations in these patients include coarse facial features, spinal deformities, skeletal deformities (dysostosis multiplex), joint contractures, short stature, corneal clouding, inguinal/umbilical hernias, hepatosplenomegaly, recurrent respiratory infections and psychomotor retardation [2,3,4]. Due to overlapping clinical presentation, it is difficult to differentiate within the MPS subtypes as well as from other LSDs and rheumatologic disorders [5]. Establishment of diagnosis purely on clinical observations is rarely conclusive; biochemical diagnosis or genetic confirmatory testing is always required. Enzyme assays are considered the most definitive tests, though mutation analysis is often equally effective [6].
The differential diagnosis of MPS remains a clinical challenge as analyzing all the screen-positive cases for 11 different enzymes is cumbersome and unaffordable; hence, several researchers focused on biochemical characterization of GAGs for the differential diagnosis of MPS. One-dimensional electrophoresis showed a high degree of false negative and false positive results limiting their clinical utility. We have recently proposed the utility of two-dimensional electrophoresis (2-DE) in the differential diagnosis of MPS [7]. However, the spot distribution limits its utility for densitometric measurement and the information is more or less qualitative in nature.
With the advent of mass spectrometric tools, specific quantification of GAGs is possible [8]. However, there is a need to integrate this information using machine learning tools in order to improve its clinical utility in the differential diagnosis. Specific enzyme assays and molecular analysis are well-established modes of confirmatory diagnosis of MPS.
There have been recent advances in the treatment of certain subtypes of MPS with bone marrow or hematopoietic stem cell transplantation (HSCT) and enzyme replacement therapy (ERT) [9,10,11,12,13,14,15]. The prognosis is better if the treatment is started early. It is, therefore, crucial to diagnose as early as possible in order to provide targeted treatment options for MPS patients.
In the current study, an attempt was made to capture liquid chromatography–tandem mass spectrometry (LC–MS/MS) and 2-DE GAG profiles and translate them into classification and regression trees (CART) using machine learning tools. This enables quick decision making in referring specific enzyme assays and targeted mutation analysis.
Materials and methods
Study population
Control urine samples (n = 153) from healthy individuals were collected to establish GAG reference values with age range of 0.1–41 (6.0 ± 6.8) years and male-to-female ratio of 99: 54. Eighty-nine patients (62 males and 27 females) were diagnosed with different types of MPS type I (n = 28), II (n = 17), III (n = 23), IV (n = 12), VI (n = 6), and VII (n = 3) with an age range: 0.1–25.4 (5.6 ± 4.4) years formed an experimental group. The non-specific quantitative urinary total GAG, qualitative 2-DE, and the three urinary GAGs levels (DS, HS, and KS) on LC–MS/MS were evaluated. The diagnosis of specific MPS type was confirmed by specific enzyme assays in leukocytes and by identification of a pathogenic mutation. None of the patients had received ERT or HSCT before entering this study.
Urinary GAG quantitative and qualitative estimation
Urine samples were used for estimation of urinary GAG quantification and 2-DE. Urinary GAG quantification was carried out by the standard dimethylmethylene blue (DMB) dye method, and 2-DE was performed with Alcian blue reagent method [16]. GAG/creatinine ratio (milligrams of GAG per millimole of creatinine) was used as a measure of the urinary excretion of GAG. Creatinine estimation was carried out by the standard method [17].
LC–MS/MS analysis
The LC–MS/MS method for GAG disaccharide analysis of chondroitin sulfate (CS), heparan sulfate (HS), dermatan sulfate (DS), and keratan sulfate (KS) was done by the method reported by Auray-Blais et al., Chuang et al. [18, 19]. GAGs were first precipitated from urine using the Alcian Blue (AB) reagent containing sodium acetate. Then, sodium chloride and methanol were used to dissolve MPS-AB complex. Sequentially, sodium carbonate was added to dissociate the MPS complex and AB. Finally, ethanol was used to re-precipitate MPS. After evaporating to dryness, the precipitate was dissolved in the water based on DMB values in mg/l, followed by methanolysis. In 1.5 ml microcentrifuge tubes 2 µl of precipitated GAG samples were taken and 200 µl of 3N Methanolic HCl and 10 µl of 2, 2-dimethoxy propane were added. Samples were dried under N2 stream at 65 °C for 75 min. To this, 100 µl of acetonitrile was added and further dried under N2 stream. Samples were reconstituted with 100 µl of NH4OAC (10 mM). Internal standards, [2H6] DS, [2H6] CS, and [2H6] HS were prepared in-house by deuteriomethanolysis of CS, DS, and HS [20]. Disaccharides were separated on an Atlantis T3 C18 column (3 µm, 2.1 × 50 mm, Waters Corporation) over 6.5 min at a flow rate of 450 µl/min. Before injection, all of the clinical samples were passed through a disposable 0.2 µm filter to minimize the ion suppression due to endogenous contaminants. Data acquisition was performed by selected reaction monitoring (SRM) using the protonated molecular ion transition mass-to-charge ratio (m/z) 384.2 > 161.9 for HS derived disaccharides and m/z 426.1 > 236.2 for DS and CS-derived disaccharides. The m/z values of [2H6] HS-derived disaccharides were 390.2 > 168, and m/z values for [2H6] CS and [2H6] DS disaccharides were 432 > 239. Quantifications were achieved using peak areas that were processed using Analyst 1.5.2™ software (AB Sciex).
Enzyme analysis of MPS types
Four milliliters of peripheral blood was collected in a sodium heparin vacutainer. Plasma was separated, and leukocytes were isolated from sodium heparin whole blood. Leukocytes were stored at − 80 °C till process. All MPS disorders were confirmed by respective enzyme analysis assays. These assays were carried out by using artificial fluorigenic (4-methylumbelliferyl) substrates, except arylsulfatase B which was assayed by photometric method using para-nitrocatechol sulfate K2 as substrate [21,22,23,24,25,26,27,28,29,30]. The fluorescence of the 4-methylumbelliferyl was measured on multimode reader at λEx of 362 nm and λEm of 448 nm. The absorbance of para-nitrocatechol was measured at 515 nm.
Mutation analysis
Genomic DNA was prepared from peripheral blood leukocytes by high salt extraction. PCR amplification of genomic DNA in MPS patients was carried out using oligonucleotide primers (Table 1). PCR products were purified and sequenced using a DNA sequencer. All amplified fragments flanking the exons were analyzed to identify variations. The resultant sequences were imported into Codon-code aligner software for alignment, editing, and mutation analysis.
Development of classification and regression models
For the development of CART model, pattern of dermatan sulfate, heparan sulfate, keratan sulfate was used as input variable to derive the type of MPS. From the given set of input variables, the most significant classifier formed the apex of the tree. It bifurcates into two classes based on a threshold value. Further branching continues until clear discrimination between classes is established based on ‘if’ and ‘then’ rules. At each node, there will be one input variable branching into two based the threshold value. Finally, a decision is made whose performance can be verified by computing a different combination of input variables to derive the output. In order to implement such a model in a diagnostic set-up, the performance characteristics, namely overall accuracy, sensitivity, specificity, positive predictive value, and negative predictive values, were assessed.
In silico analysis
In order to cross-verify identified mutations and their impact on the protein, ww.mutalyzer.nl was used. Further, the sequence change was computed in mutation taster to retrieve the functional impact of the mutation and also to check the mutation frequency in 1000 genome and ExAc databases.
Statistical analysis
Fisher exact test was used to calculate the performance characteristics of the model. In parallel, receiver operating characteristic (ROC) curves were also plotted to assess the overall diagnostic utility of the models.
Results
Urinary quantitative GAG analysis
The reference ranges for GAG by DMB method were determined from 153 healthy control persons (males 99 and females 54) with different age groups (i.e., 0–6 months, 6–12 months, 1–2 years, 2–4 years, 4–6 years, 6–8 years, 8–10 years, and > 10 years). High DMB ratio is observed in neonatal and infantile age group, i.e., 26.3 ± 14.6 mg/mM creatinine and very less DMB ratio in adult age group, i.e., 3.7 ± 1.3 mg/mM creatinine. The average concentration of urinary CS found in normal healthy controls is 35.5 ± 27.9 µg/ml with very low concentrations of KS (4.6 ± 3.0 µg/ml) HS (0.3 ± 0.2 µg/ml) and DS (0.2 ± 0.1 µg/ml) (Table 2).
CART model of 2-DE-GAG
The 2-DE of GAG identifies DS, HS, and KS as bands. The presence or absence of the band was computed to construct classification and regression tree (CART) using purity criteria. The branching of tree stops when it can clearly classify them into certain groups based on ‘if’ and ‘then’ rules.
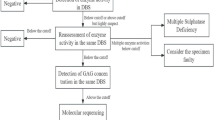
Positivity for DS and HS was characteristics of MPS I, MPS II and MPS VII. DS positivity and HS negativity were observed in MPS VI. DS negative MPS were segregated based on the presence or absence of KS band. KS positivity is characteristic of MPS IVA and MPS IVB. HS band alone was observed in MPSIIIA, MPS IIIB and MPS IIIC. Based on these criteria, the overall accuracy of this model in differential diagnosis was 96.3% (Fig. 1a).
CART model of 2-DE-GAG and LC–MS/MS-GAG. a The two-dimensional electrophoresis (2-DE) of glycosaminoglycans (GAG) identifies dermatan sulfate (DS), heparan sulfate (HS), and keratan sulfate (KS) as bands. The presence or absence of the band was computed to construct classification and regression tree (CART) using purity criteria. The branching of tree stops when it can clearly classify them into certain groups based on ‘if’ and ‘then’ rules. b Tandem mass spectrometric (LC–MS/MS) quantification of glycosaminoglycans (GAG) was used to establish thresholds of dermatan sulfate (DS), heparan sulfate (HS), and keratan sulfate (KS) that can classify the groups based on ‘if’ and ‘then’ rules based on purity criterion. The classification and regression tree (CART) segregated the classes in tree-like fashion starting from apex to branches
CART model of LC–MS/MS-GAG
Construction of CART model established thresholds of HS, DS, and CS for the differential diagnosis of MPS. HS is the key predictor in differential diagnosis with a threshold value of 4.93 µg/ml. HS > 4.93 µg/ml and DS > 10.22 µg/ml were characteristics of MPS I, MPS II, and MPS VII. HS > 4.93 µg/ml and DS < 10.22 µg/ml was observed in MPSIIIA, MPS IIIB, and MPS IIIC. HS < 4.93 µg/ml and DS < 0.47 µg/ml were ruling out the possibility of MPS. HS < 4.93 µg/ml, DS > 0.47 µg/ml, and KS > 10 µg/ml are characteristic of MPS IVA and MPS IVB. HS < 4.93 µg/ml, DS > 0.47 µg/ml, and KS < 10 µg/ml are characteristic of MPS VI (Fig. 1b). The overall accuracy of this model in the differential diagnosis of MPS was 98.3%. As shown in Fig. 2, HS levels are significantly higher in MPS IIIA, MPS IIIB and MPS IIIC. DS levels were higher in MPS I, MPS II, MPS VI, and MPS VII. KS levels were elevated in MPS IVA and IVB.
Distribution of heparan sulfate, dermatan sulfate, and keratan sulfate in different MPS types. As illustrated, HS levels are higher in MPS IIIA, MPS IIIB and MPS IIIC. DS levels were higher in MPS I, MPS II, MPS VI, and MPS VII. KS levels were higher in MPS IVA and MPS IVB. Error bars represent Mean ± SD. The statistical significance (p value) between different groups being compared were labeled as ****(p < 0.0001) and ***(p < 0.001)
Table 3 compared the advantage of LC–MS/MS over the 2-DE in GAG characterization and differential diagnosis of MPS. Total GAG was inversely associated with the age of onset [r = − 0.76, p < 0.0001]. All GAG types (CS, DS, HS, KS) were inversely associated with age of onset in MPS II [DS: r = − 0.64, p < 0.005; HS: r = − 0.59, p < 0.01; KS: r = − 0.57, p < 0.01]. Figure 3 showed the representative 2-DE data and LC–MS/MS spectra used to generate the machine learning tools.
Representative two-dimensional electrophoresis and LC–MS/MS mass spectra used to generate machine learning tools. a Two-dimensional electrophoresis of GAGs. b LC–MS/MS mass spectra of GAGs a, b mass spectrum from methanolysate of CS, DS and deuteriomethanolysis product of CS, DS; c, d mass spectrum from methanolysate of HS and deuteriomethanolysis product of HS; e, f mass spectrum from methanolysate of KS and deuteriomethanolysis product of KS
Specific enzyme assays
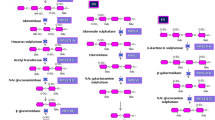
Ten specific enzyme assays were performed to confirm the diagnosis of MPS type. All the assays were fluorometric except for MPS VI. Total GAG excretion was highest in MPS I, MPS II, and MPS VII. Using the cases doubly confirmed through biochemical and molecular approaches, thresholds of specific enzyme activity diagnostic for each type of MPS were also established. β-glucuronidase and N-Acetyl glucosamine 6-sulfate sulfatase activities were < 1.0% of mean normal; β-galactosidase and glucosamine-N-acetyl transferase activities were < 2.0% of mean normal; α-N-acetylglucosaminidase and galactose 6-sulfate sulfatase activities were < 3.0% of mean normal and remaining MPS types showed activities < 5.0% of mean normal in homozygous mutants (Table 4).
Mutation spectrum
Among IDUA mutations, c.1469T > C was the most common followed by c.784delC, c.532G > A, c.908T > C, c.1759C > T. Out of the 31 different mutations identified in IDUA, 9 were non-sense mutation resulting in premature termination, which contribute to 32.3% cases, i.e., c.436A.T (p.K146*), c.606C > G (p.Y202*), c.895 G > T (p.E299*), c.1029C > A (p.Y343*), c.1750C > T (p.Q584*), c.1759 C > T(p.Q587*), c.1855C > T (p.R619*), c.1861C > T (p.R621*), c. 1882 C > T (p.R628*). Two gross deletions were also reported, i.e., ex8_ex14del and ex9_ex14del. IDS mutation spectrum was highly heterogeneous involving 7 different mutations in 8 patients. There were 4 missense mutations, i.e.,c.253G > A (p.A85T), c.263G > A (p.R88H), c.329G > A (p.R110K), c.1402C > T (p.R468W); two frameshift mutations, i.e., c.1467_1468insG (p.Y490Vfs*9) and c.474_474delT (p.H159Ifs*54); one large deletion, i.e., c.982_996del (p. I328_T332-DEl) in IDS gene. In MPS IIIB, NAGLU c.1693C > T (p.R565W) and c.1914_1915insT (p.E639*) are the most common accounting for 6 cases out of 9. The other mutations observed were c.1694G > T (p.R565L) and c.2209C > G (p.R737G). SGSH c.1129C > T (p.R377C) was observed in only one case diagnosed to be MPS IIIA. In MPS VI cases (n = 2), two different ARSB mutations were identified, i.e., c.479G > A (p.R160Q) and c.1208_1208delC (p.S403Y). Only one case of MPS IVA had GALNS c.647T > C (p.F216S) mutation (Fig. 4).
Identified mutations in IDUA and IDS genes. a In the current study, except for exon 1, 2 and 12, all other exons had IDUA mutations. Out of the identified mutations, nine were nonsense mutations. b In the current study, four missense, two frame shift, and one large deletion (exon 7) were identified in IDS gene
Discussion
The utility of machine learning tools in diagnosing Gaucher disease type I based on trabecular bone microarchitecture has been demonstrated in a recent study [31]. However, till date no such tools were used for the differential diagnosis of MPS. To the best of our knowledge, this is the first study to demonstrate the application of machine learning tools for differential diagnosis of MPS based on GAG pattern. We compared 2-DE and LC–MS/MS-based methodologies to achieve this objective. Both were informative, the first being qualitative in nature and the second being quantitative. The performance of LC–MS/MS-based GAG pattern was better than the 2-DE pattern. This application will have direct utility for the physician in deciding which specific enzyme assays to be performed to arrive at a conclusive diagnosis. Further, this study indicates the percentage of specific enzyme activity that can be considered diagnostic of specific MPS.
Although there are several studies that have utilized LC–MS/MS-based GAG characterization, there were no efforts to translate this data into the specific type of MPS. Auray-Blais et al. [6] applied LC–MS/MS data to investigate changes in GAG during treatment, mainly in MPS I, II, and VI. Our data are consistent with Li et al. in classifying MPS into MPS I, II, III, IV, and VI based on GAG pattern on LC–MS/MS. However, minor cutoff differences were observed between their study and ours suggesting that there could be population-level differences in GAG that needs to be considered before applying the proposed CART model. Mashima et al. [32] have demonstrated that GAG profile can also be used to differentiate subtypes of MPS, e.g., attenuated versus severe forms of MPS II. Our study is consistent with their observation and indicated an inverse association of HS and DS concentrations with age of onset in MPS II. In other terms, GAG pattern also serves as severity index. However, no such association is observed in MPS I. Although the accuracy of the machine learning tools appears to be 96.3–98.3%, there is still occasional misdiagnosis. The machine learning tools are based on classification and regression models, and hence, thresholds of GAGs were used as purity criteria for branching and splitting of the tree. In view of this, it is likely that few cases with borderline values of GAGs might be misdiagnosed. Since this model is dynamic in nature, we can further improvise with larger data sets.
Among the studies from India, our results are consistent with Utterilli et al. [33] in demonstrating a high frequency of c. 1469T > C, c.784delC mutations. However, we observed a total of 31 different mutations of IDUA in 40 patients, indicating a high degree of allelic heterogeneity in MPS I. Similar to Utterilli et al., we observed c.253G > A, c. 263G > A, and c.[1402C > T] mutations in IDS gene. In addition, we observed c.474_delT, c.982_996del, c.1467_1464insG, and c.263G > A mutations in the IDS, which were not reported in their study. The current study is the first reporting the NAGLU mutations from India. We observed three missense mutations and one insertion mutation in NAGLU gene. Out of the two ARSB mutations observed in the current study, c.479G > A was reported by Mathew et al. [34] while c.1208_1208delC was reported by Utterilli et al. 2016 [33]. The first SGSH mutation identified in Indian subject is c.613G > C mutation, while we report another mutation, i.e., c.1129C > T in one of our patients. We have only one case of GALNS c.647T > C mutation, which was already reported earlier in the Asian Indians [35].
The major strengths of our study are: (i) Three-tier evaluation of MPS cases in terms of GAG characterization, specific enzyme assay and targeted mutation analysis; (ii) construction and application of machine learning based CART models for early diagnosis while referring specific enzyme assays; (iii) establishing the diagnostic threshold of specific enzyme assays to differentiate affected vs. unaffected; and (iv) reporting of Indian specific mutation spectrum, mainly for MPS I and MPS II.
Conclusion
To conclude GAG characterization when coupled with machine learning tools facilitate quick decision making in referring specific enzyme assays to differentially diagnose the type of MPS, with improved accuracy. Hence, the proposed CART models can be adapted as first tier testing in GAG positive-MPS types. Compared to 2-DE, LC–MS/MS-based GAG analysis suggested to have a higher clinical utility in the differential diagnosis of MPS. In specific enzyme assays, residual enzyme activity < 5% is found to be diagnostic of a particular type of MPS. In view of highly heterogeneous mutation spectrum of MPS-related genes such as IDUA and IDS among the South Indians, the CART models of GAG play a significant role in the differential diagnosis of MPS as demonstrated in the current study.
References
Byers S, Rozaklis T, Brumfield LK, Ranieri E, Hopwood JJ (1998) Glyosaminoglycan accumulation and excretion in the Mucopolysaccharoses: characterization and basis of a diagnostic test for MPS. Mol Genet Meta 65:282–290
Wraith JEd (2006) Mucopolysaccharidoses and oligosaccharidoses. In: Fernandes J, Saudubray JM, van den Berghe G, Walter JH (eds) Inborn metabolic diseases. Springer, Heidelberg, pp 495–507
Coutinho MF, Lacerda L, Alves S (2012) Glycosaminoglycan storage disorders: a review. Biochem Res Int 16
Neufeld EF, Muenzer J (2001) The mucopolysaccharidoses, Scriver CR, Beaudet AL, Sly WS, Valle D (eds), The metabolic and molecular bases of inherited diseases. 8th edn, vol 3. McGraw-Hill, New York
Lehman TJ, Miller N, Norquist B, Underhil L, Keutzer J (2011) Diagnosis of the mucopolysaccharidoses. Rheumatology 50:41–48
Auray-Blais C, Bherer P, Gagnon R, Young SP, Zhang HH, An Y, Charke JT, Millington DS (2011) Efficient analysis of urinary glycosaminoglycans by LC-MS/MS in mucopolysaccharidoses type I, II and VI. Mol Genet Metab 102:49–56
Kadali S, Patlotla RD, Kolusu A, Undamatla KT, Gummadi MR, Undamatla J (2016) The utility of two dimensional electrophoresis in diagnosis of mucopolysaccharidosis disorders. Clin Chim Acta 457:36–40
Kubaski F, Osago H, Manson RW, Yamaguchi S, Kobayashi H, Tsuchiya M, Orii T, Tomatsu S (2017) Glycosaminoglycans detection methods: applications of mass spectrometry. Mol Genet Metab 120:67–77
Plastores GM (2008) Laronidase (Aldurazyme): enzyme replacement therapy for Mucopolysaccharidosis type I. Expert Opin Biol Ther 8:1003–1009
Jameson E, Jones S, Remmington T (2016) Enzyme replacement therapy with laronidase (Aldurazyme) for treating Mucopolysaccharidosis type I. Cochrane Database Syst Rev
De Silva EM, Strufaldi MW, Andriolo RB, Silva LA (2011) Enzyme replacement therapy with idursulfase for Mucopolysaccharidosis type II (Hunter syndrome). Cochrane Database Syst Rev
Kubaski F, Yabe H, Suzuki Y, Seto T, Hamazaki T, Mason RW, Xie L, Onsten TGH, Leistner-segal S, Giugliani R, Dung VC, Nqo CBT, Yamaguchi S, Montano AM, Orii KE, Fukao T, Shintaku H, Orii T, Tomatsu S (2017) Hemotopoietic stem cell transplantation for patients with Mucopolysaccharidosis II. Biol Blood Marrow Transplant 23:1795–1803
Hendriksz CJ, Burton B, Fleming TR, Harmatz P, Hughes D, Jones SA, Lin SP, Mengel E, Scarpa M, Valayannopoulos V, Giugliani R, Investigatotors STRIVE, Slasor P, Lounsbury D, Dummer W (2014) Efficacy and safety of enzyme replacement therapy with BMN 110 (elosulfase alfa) for Morquio A syndrome (Mucopolysaccharidosis IVA): a phase 3 randomised placebo-controlled study. J Inherit Metab Dis 37:979–990
Harmatz P, Whitley CB, Waber L, Pais R, Steiner R, Plecko B, Kaplan P, Simon J, Butensky E, Hopwood JJ (2004) Enzyme replacement therapy in Mucopolysaccharidosis VI (Maroteaux-Lamy syndrome). J Pediatr 144:575–580
Herskhovitz E, Young E, Rainer J, Hall CM, Lidchi V, Chong K, Vellodi A (1999) Bone marrow transplantation for Maroteaux-lamy syndrome (MPS VI): long-term follow-up. J Inherit Metab Dis 22:50–62
Chuang CK, Lin SP, Chung SF (2001) Diagnostic screening for Mucopolysaccharidoses by the Dimethylmethylene blue method and two dimensional electrophoresis. Zhonghua Yi Xue Za Zhi (Taipei) 64:15–22
Jaffe M (1886) About the precipitation caused by pikrinic acid in normal urine and about a new reaction of creatinine Physiol Chem 10:391
Auray-Blais C, Lavoie P, Tomatsu S, Valayannopoulos V, Mitchell JJ, Raiman J, Beaudoin M, Maranda B, Charkem JT (2016) UPLC-MS/MS detection of disaccharides derived from glycosaminoglycans as biomarkers of muopolysaccharidoses. Anal Chim Acta 939:139–148
Chuang CK, Lin HY, Wang TJ, Tsai CC, Liu HL, Lin SP (2014) A modified liquid chromatography/tandem mass spectrophotometry method for predominant disaccharide units of urinary glycosaminoglycans in patients with mucopolysaccharidoses. Orphanet J Rare Dis 936:139–148
Zhang H, Young SP, Auray-Blais C, Orchard PJ, Tolar J, Millington DS (2011) Analysis of glycosaminoglycans in cerebrospinal fluid from patients with mucopolysaccharidoses by isotope-dilution ultra-performance liquid chromatography–tandem mass spectrometry. Clin Chem 57:1005–1012
Hopwood JJ, Muller V, Smithson A, Baggett N (1979) A fluorometric assay using 4-methylumbelliferyl α-l-iduronide for the estimation of α-l-iduronidase activity and the detection of Hurler and Scheie syndrome. Clin Chim Acta 92:257–265
Voznyi YV, Keulemans JL, Van Diggelen OP (2001) A fluorimetric enzyme assay for the diagnosis of MPS II (Hunter disease). J Inherit Metab Dis 24:675–680
Karpova EA, Voznyi YaV, Keulemns JL, Hoogeveen AT, Winchester B, Tsyetkova IV, van Diggelen OP (1996) A fluorimetric enzyme assay for the diagnosis of Sanfilippo disease type A (MPS IIIA). J Inherit Metab Dis 19:278–285
Mauri V, Lotfi P, Segatori L, Sardiello M (2013) A rapid and sensitive method for measuring N-Acetylglucosaminidase activity in cultured cells. PLoS ONE 8(6), e68060
Voznyi YV, Karpova EA, Dudukina TV, Tsyetkova IV, Boer AM, Janse HC, van Diggelen OP (1993) A fluorimetric enzyme assay for the diagnosis of Sanfilippo disease type C (MPS IIIC). J Inherit Metab Dis 16:465–472
He W, Voznyi YaV, Boer AM, Kleijer WJ, van Diggelen OP (1993) A fluorimetric enzyme assay for the diagnosis of Sanfilippo disease type D (MPS IIID). J Inherit Metab Dis 16:935–941
van Diggelen OP, Zhai H, Kleijer WJ, Janse HC, Poorthuis BJ, van Pelt J, Kamerling JP, Galiaard H (1990) A fluorimetric enzyme assay for the diagnosis of Morquio disease type A (MPS IV A). Clin Chim Acta 187:131–139
Nowakowski RW, Thompson JN, Baker HJ (1988) Diagnosis of feline GM1 gangliosidosis by enzyme assay of cultured conjunctival cells. Invest Ophthalmol Vis Sci 29:487–490
Christomanou H, Sandhoff K (1977) A sensitive fluorescence assay for the simultaneous and separate determination of arylsulphatases A and B. Clin Chim Acta 79:527–531
Natowicz MR, Isman F, Prence EM, Cedrone P, Allen JJ (2003) Rapid prenatal testing for human beta-glucuronidase deficiency (MPS VII). Genet Test 7:241–243
Sharma GB, Robertson DD, Laney DA, Gambello MJ, Terk M (2016) Machine learning based analytics of micro-MRI trabecular bone microarchitecture and texture in type I Gaucher disease. J Biomech 49:1961–1968
Mashima R, Sakai E, Tanaka M, Kosagu M, Okuyama T (2016) The levels of urinary glycosaminoglycans of patients with attenuated and severe type of mucopolysaccharidosis II determined by liquid chromatography-tandem mass spectrometry. Mol Genet Metab Rep 7:87–91
Uttarilli A, Ranganath P, Matta D, Md Nurul Jain J, Prasad K, Babu AS, Girish KM, Verma IC, Phadke SR, Mandal K, Puri RD, Aggarwal S, Danda S, Sankar VH, Kappor S, Bhat M, Gowrishankar K, Hasan AQ, Nair M, Nampoothiri S, Dalal A (2016) Identification and characterization of 20 novel pathogenic variants in 60 unrelated Indian patients with Mucopolysachharidoses type I and type II. Clin Genet 90(6):496–508
Mathew J, Jagadeesh SM, Bhat M, Udhaya Kumar S, Thiyagarajan S, Srinivasa S (2015) Mutations in ARSB in MPS VI patients in India. Mol Genet Metab rep 4:53–61
Bidchol AM, Dalal A, Shah H, Nampoothiri SS, Kabra S, Gupta M, Danda N, Gowrishankar S, Phadke K, Kapoor SR, Kamate S, Verma M, Puri IC, Sankar RD, Devi VH, Patil AR, Ranganath SJ, Jain P, Agarwal SJ, Singh M, Mishra A, Tamhankar P, Gopinath PM, Nagarajam PM, Satyamoorthy HA, Girish K KM (2014) GALNS mutations in Indian patients with mucopolysaccharidosis IVA. Am J Med Genet A 164A:2793–2801
Acknowledgements
We thank Mr. Rajeev Sindhi, MD, Sandor Lifesciences Pvt. Ltd. for providing necessary infrastructure for the study. We thank all the families who had participated in the study.
Author information
Authors and Affiliations
Contributions
SK participated in the study design, analysis, interpretation of data, and drafting the manuscript. SMN performed statistical analysis, interpretation of the data, and drafting of the manuscript. ARRD participated in recruitment of the patients, interpretation of clinical reports, and counseled the affected families. VLB carried out the conception and design of the study, interpretation of data, and final approval of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests (financial or non-financial) in the present study.
Ethical approval
The study was approved by the Institutional ethical committee for biomedical research, Institute of Genetics and Faculty of Science, Osmania University. This study complied with the ethical principles outlined in the Declaration of Helsinki.
Informed consent
Informed written consents were obtained from patients/guardians along with detailed clinical history during their enrollment for the study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kadali, S., Naushad, S.M., Radha Rama Devi, A. et al. Biochemical, machine learning and molecular approaches for the differential diagnosis of Mucopolysaccharidoses. Mol Cell Biochem 458, 27–37 (2019). https://doi.org/10.1007/s11010-019-03527-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-019-03527-6