Abstract
Biocompatibility, low toxicity and high selectivity towards bacterial cells has been the hallmark of peptide-based antibiotics. The innate immune system has been employing such molecular systems against invading pathogens as a successful defense strategy. In this study, we attempt to develop topologically constrained antimicrobial peptides with syndiotactic stereochemical arrangement, by incorporating L and D amino acids successively in its amino acid sequence. Acetylated versions of the designed peptides were also examined for its influence on bactericidal potency, against Gram-positive and Gram-negative bacteria. Syndiotactic stereochemical arrangement of the polypeptide main chain mimics stereochemistry of Gramicidin, a naturally occurring antimicrobial peptides. Gramicidin is a class of penta-deca-peptides isolated from soil bacteria Bacillus brevis, but their utility as antibiotic was limited to topical use due to high levels of hemotoxicity. Activity profiles of the four de novo designed peptide variants show higher specificity towards Gram-positive bacteria than Gram-negative variants, matching earlier reports on the therapeutic potential of gramicidin as a broad spectrum antibiotic. Significantly, our hemolytic assay confirms very low (<1%) levels of toxicity for the designed peptides unlike gramicidin. Earlier reports confirm that incorporation of D amino acids effectively negates the possibility of proteolytic degradation, thus pointing to the potential utility of de novo designed peptides with diversified stereochemistry as a promising new approach in the generation of novel antibiotic peptides.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Higher organisms use peptides as a defense mechanism against invading pathogens (Dosler and Karaaslan 2014; Ravichandran et al. 2017; Czihal and Hoffmann 2009; Saikia et al. 2017), Defensins (Lehrer 2004), cecropins (Moore et al. 1996), magainins (Berkowitz et al. 1990) and cathelicidins (Zanetti 2005) constitute major classes of “host defense peptide”. There is a remarkable increase in peptide based antibiotics, with nearly two dozen molecules are either completed or under clinical trials as antimicrobial or immunomodulatory agents (Felicio et al. 2017). However, cost, uncertain toxicology profiles and lability to proteolytic susceptibility hampers the otherwise promising approach, especially in the light of the phenomenal increase in antibiotic resistance against conventional therapeutics (Li et al. 2015; Koh et al. 2015). Decreasing the size of peptides and machine learning approach have more or less addressed the first two issues, while the problem of proteolytic degradation may be sorted out by the use of D-stereoisomers and unnatural amino acids in the sequence (Hamamoto et al. 2002). Designed antimicrobial peptides, in general, are cationic amphipathic peptide sequences with distinct membrane permeabilizing ability (Hazam et al. 2017; Wimley 2010). Though various mechanisms have been proposed like carpet (Tene et al. 2016), detergent (Bechinger and Lohner 2006), barrel stave (Chang et al. 2008), toroidal pore (Brogden 2005) etc., a consensus picture is yet to emerge.
We designed two amphipathic peptide sequences and their N-terminal acetylated variants mimicking the stereochemical sequence of Gramicidin, a penta-deca-peptide isolated from soil bacterial species Bacillus brevis (Urry 1971). Antibiotic profile of Gramicidin reports that they are particularly effective against Gram-positive bacteria. However, their utility is rather limited to topical use as a lotion or ointment in the treatment of infected surface wounds and throat infections (Wang and Ishii 2012; Wishart et al. 2008). Reported mechanism of action for Gramicidin suggests that it inserts itself into bacterial membranes resulting in membrane disruption and permeabilization, leading to a sequence of events such as the loss of intracellular solutes, dissipation of trans-membrane potential, inhibition of respiration and reduction in ATP pools, culminating in cell death (Burkhart et al. 1998). Naturally-occurring Gramicidin or Gramicidin D exists in isoforms with a variation of valine or isoleucine at first position. Gramicidin molecule is a mixture of Gramicidin A, B and C with a variation at the 11th position containing tryptophan, phenylalanine and tyrosine respectively (Burkhart et al. 1999). In this paper we present a design philosophy to synthesize cationic, amphipathic peptide molecules incorporating D-amino acids in the design alphabet. To promote the membrane permeabilizing ability, we design our sequence around Gramicidin with gramicidin helix (6.3 helix) as the target structure. Gramicidins form cation selective trans-membrane ion channel in typical model membranes (Hladky and Haydon 1972). Even though, Gramicidin acquires various folding states, two of the major conformations have been predominant in different environments; first is the channel forming single stranded helical dimer and the second type is a non-channel forming intertwined helix (Urry 1971; LoGrasso et al. 1988). Even though conformations of gramicidin are solvent dependent, single stranded β (6.3) dimer is the thermodynamically most stable structure (LoGrasso et al. 1988; Killian et al. 1988; Kelkar and Chattopadhyay 2007).
Like all natural proteins, most of the therapeutic peptides are sequences of asymmetric amino acids having L-chiral stereo-chemical structure. In other words, they are stereo-regular “isotactic” hetero-polymers. The isotactic stereochemistry of the backbone, coupled with planarity of peptide bond and steric effects limits the conformational space of a polypeptide. Even then, the predictive ability of a given amino acid sequence to fix the structure, folding intermediates and pathways are still weak, despite making enormous advancement in the basic understanding of this unsolved problem (Ramakrishnan et al. 2012; Khoury et al. 2014). This limits the success of a protein de novo design experiment in realizing its translational objectives.
Polymer sequences with alternating l and d-chiral monomers (amino acids) are referred to as syndiotactic polymers and with random distribution of l and d are heterotactic or atactic polymer (Billmeyer 1957; Durani 2008). Gramicidin is a 15 amino acid long trans membrane helical molecule with alternating l and d-chiral amino acids in its sequence and can hence be termed as a syndiotactic peptide (Nanda et al. 2007). In this study we tried to develop topologically constrained antimicrobial peptide molecules with syndiotactic stereo-chemical arrangements after making an objective assessment about their relative stability by molecular dynamics simulations. Acetylated variants of both the designed peptides, were also examined to test their possible differential efficacy, against Gram-positive and Gram-negative bacterial species. The activity profiles were considerably similar, indicating limited significance of acetylation in syndiotactic sequences. The bactericidal potency of the designed peptides was higher against Gram-positive than in Gram-negative bacteria, though the design approach may be further modified in future designs in the generation of broad spectrum antibiotics.
Materials and Methods
Design and Simulation
The peptides molecules used in this study were designed by using an in-house software PDB make and a modified version of Ribosome Software from G.D. Rose laboratory (Srinivasan 2013). Molecular dynamics (MD) simulations were performed using GROMACS (version: 4.6.5) (Pronk et al. 2013) with GROMOS 96 43a1 force field (Scott et al. 1999). The peptide structures were energy minimized in vacuum over 2000 steps using the steepest descent algorithm in a cubic box, with each face separated at 1.5 nm from the protein molecule. Subsequently, water molecules (SPC) were added to the simulation box and the system was further energy minimized. A 10 ns production run under NVT conditions was performed with an integration step of 2 fs. Initial velocities were taken from Maxwell distribution at 300 K, using Berendsen thermostat for temperature coupling with coupling relaxation step setting at 0.1 ps. The cut-off limit for non-bonded interactions was set at 0.8 nm. All through the simulation run, bond length was constrained with geometric accuracy of 10−4.
Electrostatic Profiling
Electrostatic potential for the designed peptides was calculated by solving Finite Difference Poisson–Boltzmann equation using Delphi software (Li et al. 2012). These electrostatic potential maps of candidate structures were further compared to identify distinct electrostatic potential signatures for each structure. The electrostatic potentials were represented in a 3D graph using MATLAB.
Reagents
Rink Amide resin, Fmoc amino acids, N, N, N, N-tetramethyl-O-(1H-benzotriazol-1-yl) uronium hexafluorophosphate (HBTU), N, N-dimethylformamide (DMF), m-Cresol, Glutaraldehyde, Dimethylsulphoxide (DMSO), and Ethanol were purchased from Merck. 1-Hydroxybenzotriazole (HOBt), diethyl ether, 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), sodium phosphate monobasic anhydrous, acetonitrile (ACN), ethylene diamine tetra acetic acid (EDTA) and Piperidine were obtained from SRL laboratories. N, N-diisopropylethylamine (DIPEA), thioanisole, 1, 2 ethane dithiol (EDT), Trifluoroactic acid (TFA), 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT), was purchased from Sigma-Aldrich. Nutrient Broth and Agar was purchased from Himedia laboratories. All other reagents used were of the highest purity (≥99%).
Peptide Synthesis and Characterization
The peptides were synthesized employing standard Solid phase peptide synthesis protocol (Fmoc chemistry). Post synthesis, the peptides were cleaved from the resin using a cleavage cocktail (m-Cresol: Thioanisole: EDT: TFA :: 2:2:1:20) for 12 h in dark. The peptides were purified by means of semi-preparative reversed-phase liquid chromatography. A chromatographic run of 10% ACN to 100% ACN with 0.1% TFA, was used for the gradient elution of peptides at a flow rate of 1 mL/min. The eluents were monitored at 210 nm and verified by electrospray ionization mass spectrometry (Wang et al. 2012; Falcao et al. 2016; Chaudhary and Nagaraj 2011).
Antibacterial Assay
Staphylococcus aureus (NCTC 8530) and Pseudomonas aeruginosa (NCTC6750) bacterial strains were used for the assay. Mid logarithmic phase bacterial cells were washed and resuspended in sodium phosphate buffer (10 mM, pH 7.4). The resulting suspension was diluted to a net absorbance value of 0.2 at 600 nm and 50 μL of the inoculum was treated with required concentrations of peptide (Net incubation broth is 100 µL), incubated for 2 h. Post incubation, 100 µL of MTT solution (0.5 mg/mL) was added to the broth and incubated for 4 h at 37 °C. Subsequently, the reaction broth was centrifuged and the bacterial pellets were mixed with DMSO. The difference in the absorbance at 570 and 660 nm was calculated and percent bacterial cell lysis was reported, relative to untreated bacterial cells (Rufian-Henares and Morales 2011; Piaru et al. 2012; Eloff 1998). A relative inhibition of 80% growth with reference to growth control was considered to be as Minimal Inhibitory Concentration, as per relative reported standard procedure (Wu et al. 2014).
Field Emission-Scanning Electron Microscopy (FE-SEM)
Post incubation, glutaraldehyde (4%) was added to peptide-bacteria mixture, followed by an incubation of 30 min. The entire broth was centrifuged (1500×g, 5 min) and the bacterial pellet was placed over a glass slide. The dried samples were washed (gradient, 30–100% ethanol), coated with gold and images were recorded using scanning electron microscopy (Abd-El-Aziz et al. 2015).
Hemolytic Assay
Fresh human blood was collected and mixed with 1.5 mg EDTA per milliliter and mixed thoroughly. These red blood cells were re-suspended and repeatedly washed with buffer (5 mM HEPES buffer saline, pH 7.4) by centrifugation at 800×g for 5 min, until the supernatant was colorless. A 10% cell suspension in buffer was prepared and 50 µL of the suspension was mixed with peptide solution, incubated for 2 h at 37 ± 2 °C. After incubation, the suspension was centrifuged at 800×g for 5 min and the absorbance of supernatant was recorded at 540 nm (Balaji and Trivedi 2012; Lee et al. 2014). Percent cell lysis was calculated in comparison to the extent of hemolysis recorded with 100% lysis of blood cells in water.
Results and Discussion
Design Philosophy
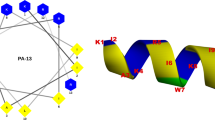
We designed two pairs of syndiotactic sequences mimicking Gramicidin 6.3 helix as model systems, with a starting (φ, ψ) dihedral angle pair shown in Table 1. Gramicidin has a right handed helical structure with (i to i + 7) and (i to i − 5) (Ramakrishnan et al. 2005) hydrogen bonding pattern, and a pitch 4.7 Å (Katsaras et al. 1992), with 6.3 residues per turn (Urry 1971). Theoretically such helical structures can assume two structural variants depending on their ‘handedness’; either right or left handed (Fig. 1). Therefore, we have designed two variants right handed (RH) and left handed (LH) for each sequence resulting in eight possible variants, though only four variants are synthetically possible.
Backbone structures of the designed hetero-chiral peptides having syndiotactic stereochemistry. All the peptides can either be in left or right handed orientation, resulting in a maximum of eight theoretical backbone combinations possible for four peptides. a (01LH) and b (01RH) are the left handed and right handed back bone of sequence 01, whereas c (01Ac-LH) and d (01Ac-RH) are the left and right handed backbone of 01Ac. Likewise, e (02LH) and f (02RH) are left handed and right handed back bone of sequence 02 along with g (02Ac-LH) and h (02Ac-RH) are left and right handed version of 02Ac
The designed molecules are both cationic and amphipathic, which may play an important role in membrane lysis mediated by electrostatic interactions. Electrostatic potential maps of the designed peptides were generated by solving Finite Difference Poisson Boltzmann equation using Delphi software. The topology resulting from the side-chain orientation and the electrostatic potential environment it creates, were visibly distinct in all four models (Fig. 2). Precisely, the designed molecules display distinct topological and electrostatic fingerprint, which may get translated to their antibacterial potency.
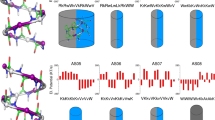
Electrostatic potential map of the designed peptides, using Delphi Software solving Finite Difference Poisson Boltzmann equation. a, b are the potential maps of left and right handed conformations of the peptide 01. Similarly, c, d are the potential maps of the left handed and right handed peptide 02 respectively. Though two conformations are theoretically possible, only one sequence can be experimentally verified. Differential color coding of amino acid residues, points to their electrostatic potential with reference to the color bar. The color map shows the potential distribution in KT/e units. (Color figure online)
The stereochemistry and sequences of 01 and 01-Ac pair and 02 and 02-Ac pair are the same, except that 01-Ac and 02-Ac being N-terminal acetylated (Table 1). To verify the stability of the designed peptides in a 6.3 helical conformation, similar to gramicidin helix, a 10 ns MD run at 298 K under NVT condition using GROMOS 96 force field, was performed for each structural variant. The cluster analysis of the theoretical models suggest that the structures are quite stable with the percent sampling of most populous structure being more than 95% in all eight models (Supplementary Fig. 1; Supplementary Table 1).
Further, the structural integrity of the molecules was verified by measuring Root Mean Square Deviation (RMSD) from the starting structure (Fig. 3). Radius of Gyration has been measured to verify whether the designed structures remain folded throughout the simulation (Fig. 3). Detailed examination of average structure forming largest cluster, however shows that hydrogen bonding pattern has been changed during the course of MD simulations (Supplementary Figs. 2 and 3; Supplementary Table 2). In our earlier studies, we have shown that gramicidin 6.3 helical structures with alternate l and d stereo-chemical sequence have their short-range and long range backbone electrostatic interactions complementing each other, while in poly L sequences they are in conflict (Ramakrishnan et al. 2006; Kumar et al. 2009; Ranbhor et al. 2006).
The resultant modification of energy landscape leading to extra stability due to complimentary local electrostatics is the key for stereo-chemical engineering of peptide sequences. We have further shown that countervailing short range–long range interaction holds the key for side-chain sequences dictating the overall conformation of a given polypeptide sequence. Since local electrostatics of gramicidin helical backbone is complimenting and stable, the fold is much more adaptable to varied sequence solutions (Ramakrishnan et al. 2005). Our MD results on all eight variants support earlier observations and this design strategy. Since amphipathicity and cationicity has been the hallmark of antibacterial activity of natural and designed peptides, we have designed all peptides retaining amphipathicity, at the same time presenting differential electrostatic potential arising from cationic side-chains.
Antibacterial Assay and Toxicity Studies
The in vitro antimicrobial activity of the synthesised peptides was tested against Gram-positive (S. aureus) and Gram-negative (P. aeruginosa) bacteria. All the peptides were showing better activity against Gram-positive bacteria, in comparison with Gram-negative bacteria (Fig. 4; Table 1).
This observation is consistent with the earlier reports on gramicidin. Gramicidin has a reported MIC value of 2.2 µM (Xiong et al. 2005) against S. aureus and is inactive against Gram-negative bacteria. Our peptides show similar trend, though the sequence similarity between gramicidin and the designed peptides are minimal. Our designed peptides primarily rely on cationicity to induce membranolytic activity. Acetylation of synthesized peptides is aimed at lowering the net positive charge by blocking the free N-terminus. We, however also could not find any notable difference in antimicrobial potency between acetylated and non-acetylated peptide variants.
Hemolytic Activity
Hemolytic assay against mammalian RBC was performed to estimate the extent of toxicity. All four peptides exhibited negligible cytotoxicity against mammalian cells at 100 µM concentration following standard protocols. Gramicidin has a reported toxicity estimate of 50% hemolysis at 5 µM concentration (Wang et al. 2012). Unlike gramicidin, toxicity levels of all four reported peptides are well within the accepted safe threshold of 2% lysis at a very high concentration of 100 µM; and three out of the four peptide variants are showing a value less than 1% (Table 2).
Microscopic Analysis
A qualitative evaluation of membrane rupturing potential of designed antimicrobial peptide against bacteria was performed using FE-SEM analysis. The cationic peptides caused significant membrane damage which was inferred after a comparative analysis between peptide treated (ruptured) and untreated (intact) bacterial cells. The bacterial cells were treated with 100 µM peptide resulting in deformed bacterial membrane architecture. (Fig. 5).
FE-SEM images of Gram-positive and Gram-negative bacteria, treated with individual peptides after 2 h of incubation. Comparison of untreated S. aureus (Gram-positive) and P. aeruginosa (Gram-negative) cells against ruptured bacterial membrane treated with 01, 01 Ac, 02 and 02 Ac at 100 µM concentration. The scale bars correspond to 200 nm
Qualitative pictures indicate that the membrane rupturing ability of the amphipathic cationic peptides are responsible for the desired activity profile of the designed peptides, though detailed mechanism investigation is beyond the scope of this paper.
Conclusions
Almost all natural proteins are isotactic with chain stereochemistry (poly L). An isotactic peptide molecule roughly acquires 21% allowed conformational space in Ramachandran plot. If the stereochemical sequence is a variable, it increases the designable space manifold, resulting in an informed walk across Ramachandran φ, ψ space. We employed a peptido-mimetic approach, modeling around Gramicidin adopting the simplest stereo-chemical sequence with syndiotactic stereochemistry. Gramicidin is a natural antibiotic peptide with Syndiotactic backbone (alternating L and D) having good bactericidal activity against Gram-positive bacteria, but could only be used topically because of very high levels of toxicity. We attempt to re-design the amino acid sequence (side-chain sequence) space retaining its stereo-chemical (main-chain) sequence, with an objective to minimize its toxicity, while retaining antimicrobial activity. The two designed cationic amphipathic peptides with alternating L and D chiral amino acid sequences, and their acetylated variants exhibit negligible levels of toxicity, well within the acceptable limits. Our best peptide sequence is showing an MIC value of 12.5 µM against a reported value of 2.2 µM (Xiong et al. 2005) for gramicidin D. Importantly, this design strategy offers an effective means to develop possible sequence variation in future designs. Incorporation of stereochemistry as an additional design variable significantly expands the design space of a peptide sequence. In a previous study (Hazam et al. 2017), we have systematically varied stereo-chemistry of the designed peptides resulting in different structure and electrostatic interaction profiles while they approach a target. However, our efforts to directly relate the structure and electrostatic profiles in defining activity and toxicity of a given peptide, is not yet conclusive.
Role of acetylation in naturally occurring proteins has not been precisely deduced but studies indicate that it improves the potency of peptides, as it mimics the native protein itself (Drazic et al. 2016). As indicated in literature, it enhances the overall potency, metabolic stability and resistance against enzymatic degradation (Di 2015; Strömstedt et al. 2009). In our study, we could not find any significant difference in activity or toxicity profiles of our designed peptides, which was also an objective of this investigation.
Apart from increasing the overall design space considerably, incorporation of D amino acids effectively negates the possibility of proteolytic degradation. This point to the potential utility of de novo designed peptides with diversified stereochemistry as a promising new approach in the generation of antibiotic peptides. Such attempts are important to effectively manage the broad antibiotic spectrum and pharmacological profile with increased instances of antibiotic resistance reported in recent times.
Abbreviations
- MD:
-
Molecular dynamics
- RMSD:
-
Root mean square deviation
- Rg:
-
Radius of gyration
- FE-SEM:
-
Field emission scanning electron microscopy
- MS:
-
Mass spectrometry
- CFU:
-
Colony forming unit
References
Abd-El-Aziz AS, Agatemor C, Etkin N, Overy DP, Lanteigne M, McQuillan K (2015) Antimicrobial organometallic dendrimers with tunable activity against multidrug-resistant bacteria. Biomacromolecules 16(11):3694–3703
Balaji S, Trivedi V (2012) Extracellular Methemoglobin mediated early ROS spike triggers osmotic fragility and RBC destruction: an Insight into the enhanced hemolysis during malaria. Indian J Clin Biochem 27(2):178–185
Bechinger B, Lohner K (2006) Detergent-like actions of linear amphipathic cationic antimicrobial peptides. Biochim Biophys Acta 1758(9):1529–1539
Berkowitz BA, Bevins CL, Zasloff MA (1990) Magainins: a new family of membrane-active host defense peptides. Biochem Pharmacol 39(4):625–629
Billmeyer FW (1957) Textbook of polymer science, 3rd edn. Inter-science Publishers, New york
Brogden KA (2005) Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3(3):238–250
Burkhart BM, Li N, Langs DA, Pangborn WA, Duax WL (1998) The conducting form of gramicidin A is a right-handed double-stranded double helix. Proc Natl Acad Sci U S A 95(22):12950–12955
Burkhart BM, Gassman RM, Langs DA, Pangborn WA, Duax WL, Pletnev V (1999) Gramicidin D conformation, dynamics and membrane ion transport. Biopolymers 51(2):129–144
Chang WK, Wimley WC, Searson PC, Hristova K, Merzlyakov M (2008) Characterization of antimicrobial peptide activity by electrochemical impedance spectroscopy. Biochim Biophys Acta 1778(10):2430–2436
Chaudhary N, Nagaraj R (2011) Impact on the replacement of Phe by Trp in a short fragment of Abeta amyloid peptide on the formation of fibrils. J Pept Sci 17(2):115–123
Czihal P, Hoffmann R (2009) Mapping of apidaecin regions relevant for antimicrobial activity and bacterial internalization. Int J Pept Res Ther 15:157–164
Di L (2015) Strategic approaches to optimizing peptide ADME properties. AAPS J 17(1):134–143
Dosler S, Karaaslan E (2014) Inhibition and destruction of Pseudomonas aeruginosa biofilms by antibiotics and antimicrobial peptides. Peptides 62:32–37
Drazic A, Myklebust LM, Ree R, Arnesen T (2016) The world of protein acetylation. Biochim Biophys Acta 1864(10):1372–1401
Durani S (2008) Protein design with L- and D-alpha-amino acid structures as the alphabet. Acc Chem Res 41(10):1301–1308
Eloff J (1998) A sensitive and quick microplate method to determine the minimal inhibitory concentration of plant extracts for bacteria. Planta Med 64(08):711–713
Falcao LL, Silva-Werneck JO, Ramos AdR, Martins NF, Bresso E, Rodrigues MA (2016) Antimicrobial properties of two novel peptides derived from Theobroma cacao osmotin. Peptides 79:75–82
Felicio MR, Silva ON, Gonçalves S, Santos NC, Franco OL (2017) Peptides with dual antimicrobial and anticancer activities. Front Chem 5(5):1–9
Hamamoto K, Kida Y, Zhang Y, Shimizu T, Kuwano K (2002) Antimicrobial activity and stability to proteolysis of small linear cationic peptides with D-amino acid substitutions. Microbiol Immunol 46(11):741–749
Hazam PK, Jerath G, Kumar A, Chaudhary N, Ramakrishnan V (2017) Effect of tacticity-derived topological constraints in bactericidal peptides. Biochim Biophys Acta 1859(2017):1388–1395
Hladky SB, Haydon DA (1972) Ion transfer across lipid membranes in the presence of gramicidin A. I. Studies of the unit conductance channel. Biochim Biophys Acta 274(2):294–312
Katsaras J, Prosser RS, Stinson RH, Davis JH (1992) Constant helical pitch of the gramicidin channel in phospholipid bilayers. Biophys J 61(3):827–830
Kelkar DA, Chattopadhyay A (2007) The gramicidin ion channel: a model membrane protein. Biochim Biophys Acta 1768(9):2011–2025
Khoury GA, Smadbeck J, Kieslich CA, Floudas CA (2014) Protein folding and de novo protein design for biotechnological applications. Trends Biotechnol 32(2):99–109
Killian JA, Prasad KU, Hains D, Urry DW (1988) The membrane as an environment of minimal interconversion. A circular dichroism study on the solvent dependence of the conformational behavior of gramicidin in diacylphosphatidylcholine model membranes. BioChemistry 27(13):4848–4855
Koh J-J, Lin S, Aung TT, Lim F, Zou H, Bai Y (2015) Amino acid modified xanthone derivatives: novel, highly promising membrane-active antimicrobials for multidrug-resistant gram-positive bacterial infections. J Med Chem 58(2):739–752
Kumar A, Ramakrishnan V, Ranbhor R, Patel K, Durani S (2009) Homochiral stereochemistry: the missing link of structure to energetics in protein folding. J Phys Chem B 113(51):16435–16442
Lee M-R, Raman N, Gellman SH, Lynn DM, Palecek SP (2014) Hydrophobicity and helicity regulate the antifungal activity of 14-helical β-peptides. ACS Chem Biol 9(7):1613–1621
Lehrer RI (2004) Primate defensins. Nat Rev Microbiol 2(9):727–738
Li L, Li C, Sarkar S, Zhang J, Witham S, Zhang Z (2012) DelPhi: a comprehensive suite for DelPhi software and associated resources. BMC Biophys 5:9
Li J, Liu S, Koh J-J, Zou H, Lakshminarayanan R, Bai Y (2015) A novel fragment based strategy for membrane active antimicrobials against MRSA. Biochim Biophys Acta 1848(4):1023–1031
LoGrasso PV, Moll F, Cross TA (1988) Solvent history dependence of gramicidin A conformations in hydrated lipid bilayers. Biophys J 54(2):259–267
MATLAB version 7.10.0. (2010) Natick Massachusetts: The MathWorks Inc
Moore AJ, Beazley WD, Bibby MC, Devine DA (1996) Antimicrobial activity of cecropins. J Antimicrob Chemother 37:1077–1089
Nanda V, Andrianarijaona A, Narayanan C (2007) The role of protein homochirality in shaping the energy landscape of folding. Protein Sci 16(8):1667–1675
Piaru SP, Mahmud R, Perumal S (2012) Determination of antibacterial activity of essential oil of Myristica fragrans Houtt. Using tetrazolium microplate assay and its cytotoxic activity against vero cell line. Int J Pharmacol 8(6):572–576
Pronk S, Pall S, Schulz R, Larsson P, Bjelkmar P, Apostolov R (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29(7):845–854
Ramakrishnan V, Ranbhor R, Durani S (2005) Simulated folding in polypeptides of diversified molecular tacticity: implications for protein folding and de novo design. Biopolymers 78(2):96–105
Ramakrishnan V, Ranbhor R, Kumar A, Durani S (2006) The link between sequence and conformation in protein structures appears to be stereochemically established. J Phys Chem B 110(18):9314–9323
Ramakrishnan V, Srinivasan SP, Salem SM, Matthews SJ, Colón W, Zaki M (2012) GeoFold: topology-based protein unfolding pathways capture the effects of engineered disulfides on kinetic stability. Proteins 80(3):920–934
Ranbhor R, Ramakrishnan V, Kumar A, Durani S (2006) The interplay of sequence and stereochemistry in defining conformation in proteins and polypeptides. Biopolymers 83(5):537–545
Ravichandran G, Kumaresan V, Bhatt P, Arasu MV, Al-Dhabi NA, Arockiaraj J (2017) A cumulative strategy to predict and characterize antimicrobial peptides (AMPs) from protein database. Int J Pept Res Ther 23:281–290
Rufian-Henares JA, Morales FJ (2011) Microtiter plate-based assay for screening antimicrobial activity of melanoidins against E. coli and S. aureus. Food Chem 111(4):1069–1074
Saikia K, Sravani YD, Ramakrishnan V, Chaudhary N (2017) Highly potent antimicrobial peptides from N-terminal membrane-binding region of E. coli MreB. Sci Rep 7:42994
Scott WRP, Hünenberger PH, Tironi IG, Mark AE, Billeter SR, Fennen J (1999) The GROMOS biomolecular simulation program package. J Phys Chem A 103(19):3596–3607
Srinivasan R (2013) Ribosome–program to build coordinates for peptides from sequence. http://folding.chemistry.msstate.edu/~raj/Manuals/ribosome.html
Strömstedt AA, Pasupuleti M, Schmidtchen A, Malmsten M (2009) Evaluation of Strategies for improving proteolytic resistance of antimicrobial peptides by using variants of EFK17, an internal segment of LL-37. Antimicrob Agents Chemother 53(2):593–602
Tene N, Bonnafe E, Berger F, Rifflet A, Guilhaudis L, Ségalas-Milazzo I (2016) Biochemical and biophysical combined study of bicarinalin, an ant venom antimicrobial peptide. Peptides 79(:):103–113
Urry DW (1971) The gramicidin A transmembrane channel: a proposed pi(L, D) helix. Proc Natl Acad Sci U S A 68(3):672–676
Wang S, Ishii Y (2012) Revealing protein structures in solid-phase peptide synthesis by 13 C solid-state NMR: evidence of excessive misfolding for alzheimer’s β. J Am Chem Soc 134(6):2848–2851
Wang F, Qin L, Pace CJ, Wong P, Malonis R, Gao J (2012) Solubilized gramicidin A as potential systemic antibiotics. Chembiochem 13(1):51–55
Wimley WC (2010) Describing the mechanism of antimicrobial peptide action with the interfacial activity model. ACS Chem Biol 5(10):905–917
Wishart DS, Knox C, Guo AC, Cheng D, Shrivastava S, Tzur D (2008) Drug Bank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 36(Database issue):D901–D906
Wu X, Wang Z, Li X, Fan Y, He G, Wan Y (2014) In vitro and in vivo activities of antimicrobial peptides developed using an amino acid-based activity prediction method. Antimicrob Agents Chemother 58(9):5342–5349
Xiong YQ, Mukhopadhyay K, Yeaman MR, Adler-Moore J, Bayer AS (2005) Functional interrelationships between cell membrane and cell wall in antimicrobial peptide-mediated killing of Staphylococcus aureus. Antimicrob Agents Chemother 49(8):3114–3121
Zanetti M (2005) The role of cathelicidins in the innate host defenses of mammals. Curr Issue Mol Biol 7(2):179–196
Acknowledgements
Authors acknowledge Central Instrument Facility IIT Guwahati, Ms. Sajitha Sashidharan, Ms. Ruchika Goyal for their contributions in this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding Sources
This study was jointly funded by Department of Biotechnology, Govt. of India (Grant No. BT/350/NE/TBP/2012) and Indian Institute of Technology Guwahati, India.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hazam, P.K., Jerath, G., Chaudhary, N. et al. Peptido-mimetic Approach in the Design of Syndiotactic Antimicrobial Peptides. Int J Pept Res Ther 24, 299–307 (2018). https://doi.org/10.1007/s10989-017-9615-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-017-9615-3