Abstract
Glutathione-S-transferase enzymes (GSTs) are essential components of the phase II detoxification system and protect organisms from oxidative stress induced by xenobiotics and harmful toxins such as 1-chloro-2,4-dinitrobenzene (CDNB). In Tetrahymena thermophila, the TtGSTm34 gene was previously reported to be one of the most responsive GST genes to CDNB treatment (LD50 = 0.079 mM). This study aimed to determine the kinetic features of recombinantly expressed and purified TtGSTm34 with CDNB and glutathione (GSH). TtGSTm34-8xHis was recombinantly produced in T. thermophila as a 25-kDa protein after the cloning of the 660-bp full-length ORF of TtGSTm34 into the pIGF-1 vector. A three-dimensional model of the TtGSTm34 protein constructed by the AlphaFold and PyMOL programs confirmed that it has structurally conserved and folded GST domains. The recombinant production of TtGSTm34-8xHis was confirmed by SDS‒PAGE and Western blot analysis. A dual-affinity chromatography strategy helped to purify TtGSTm34-8xHis approximately 3166-fold. The purified recombinant TtGSTm34-8xHis exhibited significantly high enzyme activity with CDNB (190 µmol/min/mg) as substrate. Enzyme kinetic analysis revealed Km values of 0.68 mM with GSH and 0.40 mM with CDNB as substrates, confirming its expected high affinity for CDNB. The optimum pH and temperature were determined to be 7.0 and 25 °C, respectively. Ethacrynic acid inhibited fully TtGSTm34-8xHis enzyme activity. These results imply that TtGSTm34 of T. thermophila plays a major role in the detoxification of xenobiotics, such as CDNB, as a first line of defense in aquatic protists against oxidative damage.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Many xenobiotic substances, which are released into freshwater bodies, activate detoxifying enzyme systems in aquatic organisms. Hydrophobic-electrophilic xenobiotics easily pass through the cell membranes of aquatic organisms and bind to hydrophobic sites on macromolecules such as DNA, RNA, and proteins to cause irreversible damage [1]. Detoxifying enzymes, which are capable of transforming hydrophobic-electrophilic regions of xenobiotics into hydrophilic regions that are less harmful, are widely recognized for their broad specificity among similar substrate structures [2]. Detoxification of xenobiotics proceeds in three phases: modification, conjugation, and excretion [3]. In conjugation (phase II), GSTs catalyze the conjugation of one or more polar tripeptides (Glu-Cys-Gly), known as glutathione (GSH), to the electrophilic sulfhydryl group in the hydrophobic sites of xenobiotics to transform them into nontoxic hydrophilic molecules.
GST isoenzymes are widely distributed in organisms as diverse as bacteria [4], insects [5], ciliated protists [6], plants [7], fish [8], birds [9], and mammals [10]. Organisms have different compositions of GST subfamilies with similar conserved functions and enzyme structures. GSTs have two substrate-binding sites, the G site (which binds only to GSH) and the H site (which binds to groups of xenobiotics with similar structures). Cytosolic, microsomal, and mitochondrial GSTs are the three primary categories of GSTs, distinguished by their cellular location. Based on their varied tertiary structures, protein sequences, and active site configurations, cytosolic GSTs are further classified into beta, alpha, pi, mu, tau, sigma, delta, zeta, phi, epsilon, theta, and omega subfamilies [11].
Recently, there are numerous studies aimed at elucidating the roles of GST enzymes in detoxification of various agents. For example FcδGST, a novel delta class GST gene, was cloned from the marine invertebrate Fenneropenaeus chinensis (the chinese white shrimp), and was the significantly up-regulated in hemocytes and hepatopancreas upon stimulation with Vibrio anguillarum or white spot syndrome virus (WSSV). This report indicated that FcδGST plays a critical role in the innate immune responses to F. chinensis [12]. In another study, the regulation of CpGST-Mu, a mu class GST from Cristaria plicata (the freshwater cockscomb pearl mussel), was studied to reveal its expression and resistance to microcystins (cyanotoxins) generated hydrogen peroxide, which emphasizes the relationship of CpGST-Mu with the Nrf2/Keap1 signaling pathway [13]. In Panonychus citri, the omega-family gene PcGSTO1 has been cloned and characterized for its significant upregulation in the cyetpyrafen-resistant Artropoda strains, in which cyetpyrafen works an insecticide and acaricide [14]. However, in ciliated Protista, the role of GSTs in xenobiotic-induced oxidative stress is largely unknown despite their ecological importance in the food chain [15]. First cytosolic TtGST enzyme was purified and characterized from the ciliated protist T. thermophila as a functional monomeric GST without its gene sequence [16]. However, TtGSTzeta of T. thermophila has been succesfully expressed in E. coli as a recombinant TtGSTz-6xHis protein [17]. Recently, the 70 members of T. thermophila GST enzyme family were divided into four cytoplasmic subfamilies: 49 in mu (TtGSTm), seven in omega (TtGSTo), five in theta (TtGSTt), two in zeta (TtGSTz), in addition to four in MAPEG membrane-associated proteins (TtMAPEG) and three in eukaryotic protein elongation factor (TtEF1G) subfamily [6].
In our previous study, the most responsive GST genes were identified as TtGSTm19 and TtGSTm34 in CDNB-exposed T. thermophila using a combination of glutathione (GSH) affinity purification, 2D-SDS‒PAGE, and MALDI-TOF MS analysis. These genes exhibited significant transcriptional and posttranslational upregulation in response to nontoxic doses of CDNB (LD50 = 0.079 mM) [18]. The approach could reveal a GSTm enzyme, which may have better enzyme activity towards CDNB and affinity features to GSH compared to commercial enzymes. Hypothetically, the ability of T. thermophila GST enzymes to react quickly to xenobiotics in the environment may enhance organismal survival. However, the enzymatic properties of the TtGSTm34 enzyme with the CDNB as substrate have not been studied. Therefore, in this study, TtGSTm34 was recombinantly expressed in T. thermophila, and the purified recombinant TtGSTm34 was subjected to enzymatic characterization.
2 Materials and Methods
2.1 Cloning of the TtGSTm34 Gene into the pIGF-1 Vector
The TtGSTm34 (Glutathione-S-transferase mu 34, GST34; GenBank Accession number: XP_001020129.1; Tetrahymena Genome Database gene prediction number: TTHERM_00661650) coding sequence has two ATG codons for translation initiation. However, the conserved “AAAATGG” Kozak sequence [19] of TtGSTm34 is positioned only in the second methionine. Therefore, the second methionine was chosen as the start codon, and the 660 bp cDNA coding sequence was cloned and inserted into the pIGF-1 vector (Fig. 1B) without codon adaptation.
Structural elements of the pIGF-1-TtGSTm34 extrachromosomal protein expression vector of T. thermophila. A: Protein and nucleotide sequences of TtGSTm34, the Kozak sequence (bold and underlined), B: pIGF-1-TtGSTm34 vector. The construct was formed by replacing the GFP coding sequence in the pIGF-I-sfGFP vector with the TtGSTm34-8xHis coding sequence [20]. The promotor MTT1 is induced by cadmium. A terminating sequence for rpL29 ends transcription. The rDNA gene locus with 3′ NTS and 5′ rDNA flanking sequences, including 5′ CBS1 and CBS2, is necessary for rDNA origin function and rDNA minichromosome construction during conjugative transformation. The 17S RNA gene sequence carries a mutation to confer puromycin resistance
Total RNA was isolated from T. thermophila SB210 (Tetrahymena Stock Center, Cornell University, USA) by using an RNeasy Mini Kit (Qiagen Cat No: 74104) and treated with DNase (Promega Cat No: M6101). cDNA synthesis kit was used to synthesize cDNAs from ~ 2.5 µg of total RNA(Fermentas Cat. No: K1621). Subsequently, the TtGSTm34 protein-coding cDNA sequence region was first amplified with the FGSTmu34pV2 (5′ CGCGTTTAAACCTCGAGATGACAACTCTTGGTTACTGGGGCATAG 3′) and R1GSTmu34pV2 (3′ CCTCAAAGAGCTACCTGGTCTGGTCCTTAAGACGACGACAAA 5′) primer sets. The amplified cDNA template was separated on an agarose gel, and the resulting DNA band was excised. The first PCR product from the gel was purified with a GeneJet Gel Extraction Kit (Thermo Cat No: K0691) and used as the template DNA sequence in the second PCR, in which the FGSTmu34pV2 and R2GSTmu34pV2 (3′GGTCCT TAAGACGACGACGACAAACACCACCACCACCACCACCACTGATGGCCC5′) primers were used to add the 8xHis affinity tag. The amplified cDNA fragments were rerun on an agarose gel. The 725 bp DNA fragment was cut, purified and digested with the restriction enzymes PmeI (NEB, Cat No: R0560S) and ApaI (NEB, Cat No: R0114S) to obtain sticky ends. In parallel, the pIGF-1-sfGFP vector was also cut with PmeI and ApaI to obtain sticky ends by removing the sfGFP coding sequence from the cadmium-inducible MTT1-sfGFP-rpl29 gene cassette [20]. The digested and purified TtGSTm34 gene fragment and pIGF-1 vector were subjected to ligation with T4 DNA ligase (NEB, Cat No: M0202S) and transformed into E. coli XL1-Blue competent cells (Stratagene Cat. No: 200249) using the standard heat shock protocol. Positive colonies were selected from the LB agar-Amp plates and scanned for insert sizes by using the colony PCR method with the FGSTmu34pV2 and R1GSTmu34pV2 primer sets. Using PmeI and ApaI restriction enzyme digestion, the recovered plasmids were found to be present in the positive colonies. The constructed vector carrying the cadmium-inducible MTT1-TtGSTm34-8xHis-rpl29 gene cassette was named pIGF-1-GSTm34 (16,369 bp).
2.2 Transformation of the pIGF-1-GSTm34 Vector into Conjugative Tetrahymena Cells by Electroporation and Antibiotic Selection of Positive Transformants
Using the Bio-Rad Gene Pulser device, the pIGF-1-GSTm34 vector, which has a high copy number (~ 9000), was transformed by biolistic gun transformation into conjugating T. thermophila B2086 and CU428 cells, which were freshly removed from liquid nitrogen, based on the methods of Gaertig and Gorovsky with minor modifications [21]. After transformation, the cells were transferred to SPP medium containing 5 ml of 250 μg/ml penicillin‒streptomycin (pen-strep). After 12–24 h of incubation, the transformant cells were exposed to 60 μg/ml paromomycin on the first day, 100 μg/ml on the third day, and 120 μg/ml on the fifth day of conjugation. Positive transformant cells were selected and frozen in a liquid nitrogen tank.
2.3 Induction of Transformation and Total Protein Isolation
Transformant cells were induced with CdCl2 (Sigma, 439800) to a final concentration of 5 µg/ml for 36 h. The samples were collected at different time intervals, and total protein isolation was performed with modified T100B lysis buffer (25 mM Tris–HCl, 1% Triton X-100, 300 mM NaCl, pH 7.5, 2 mM PMSF, and protease inhibitor cocktail (ROCHE Cat no: 04693159001)) [22, 23]. Protein isolation was performed by adding approximately 3 ml of T100B lysis buffer to approximately 25 million cells.
2.4 SDS‒PAGE and Western Blot Analyses of Glutathione Affinity-Purified Proteins
The TtGSTm34-8xHis protein was purified based on the intrinsic glutathione affinity of TtGSTm34 using GSH-Sepharose 4B beads (GE Healthcare Cat No: 52–2303-00 AK). The Bradford method was utilized to determine the protein concentrations (Pierce™ Coomassie Plus Bradford Assay Reagent, Cat no: 23238) using a standard curve constructed with bovine serum albumin. Unbound proteins, washed-off proteins, and the final eluate (40 µg of each) were separated via 12.5% SDS‒PAGE and transferred to nitrocellulose membranes (Millipore, Billerica, MA) for Western blot analysis. After blocking with 5% nonfat milk dissolved in PBS (pH 7.4) plus 0.1% Tween 20, the membranes were incubated with a monoclonal primary mouse anti-6xHis antibody (Rockland Cat No: 200–301-382S) at 4 °C for overnight. The membranes were washed and incubated with the polyclonal secondary goat anti-mouse IgG antibody HRP (GenScript Cat No: A00160) (Supplementary File 1). TMB colorimetric solution was used to visualize the protein bands (Sigma‒Aldrich T0565).
2.5 Dual-Affinity Purification of TtGSTm34-8xHis with Ni–NTA and GSH Beads
TtGSTm34-8xHis was first purified using Ni–NTA agarose beads to collect the recombinant proteins according to the manufacturer’s protocol (Protino Cat No: 745400.25). The beads were washed three times with 20 mM imidazole and eluted three times with 250 mM imidazole. The stepwise collected samples were subjected to SDS‒PAGE for confirmation. The presence of contaminating proteins other than recombinant TtGSTm34-8xHis proteins in the eluted samples necessitated a second purification with GSH beads (Fig. 2). The buffer of the pooled elutions (total of 60 ml) was replaced with PBS (pH 7.4) using ultrafiltration to remove the imidazole present in the elution buffer (Amicon® Ultra15 10 K, Millipore Cat No: UFC901008). Since the ultrafiltration tubes had a 15 ml capacity, the centrifugation process was repeated four times. Thus, the inhibitory effect of imidazole on the binding of the recombinant protein to the GSH beads was eliminated for the next purification step.
SDS‒PAGE analysis of TtGSTm34-8xHis production and Ni–NTA purification. The transformant cells were induced with CdCl2 for 30 h. TP: total protein; FT: unbound proteins; W1 and W3: proteins that were released after the wash steps; E1-E3: first, second and third elutions that contained purified TtGSTm34-8xHis; NegE: Ni–NTA elution of uninduced transformant cells harboring pIGF-TtGSTm34; Control: Commercial GST enzyme as a positive control (Cayman Glutathione S-Transferase Assay Kit, Item No: 703302)
Second, GSH affinity purification was carried out as described previously with purified concentrated TtGSTm34-8xHis after the ultrafiltration step. The elution step was performed twice. The dual-affinity recombinant TtGSTm34-8xHis protein was purified with Ni–NTA and GSH beads and analyzed via 12.5% SDS‒PAGE. The recombinant protein was stored in 50% glycerol at − 80 °C. Every stage of purification was carried out at room temperature.
2.6 Enzyme Kinetics of the Purified Recombinant TtGSTm34-8xHis Protein
The activity of dual-affinity purified TtGSTm34-8xHis was determined spectrophotometrically according to published methods [24], with minor modifications, using 1-chloro-2,4-dinitrobenzene (CDNB) as a substrate. The reaction was set up by mixing 20 µl of the purified enzyme with 180 μl of assay cocktail (10 mM reduced glutathione, 10 mM CDNB in PBS, pH 6.5). The enzyme activities were measured with a BioTek Synergy ™ HT ELISA device using Gen-5 software. The enzyme activities were calculated from OD values taken at 2-min intervals for 30 min at 340 nm using the kinetics program of the BioTek The Synergy™ HT ELISA Reader. The activity unit was defined as the amount of enzyme that catalyzed 1 μmol of substrate at 25 °C (CDNB extinction coefficient340 nm = 9.6 mM−1 cm−1). The specific activity was expressed as micromoles of product per min per milligram of protein. The enzyme activity assays were repeated three times, and the mean values are shown in the graphs.
2.7 Determining Ionic Strength, Optimum pH and Temperature Of Enzyme Activity
To determine the optimum pH for the recombinant TtGSTm34-8xHis enzyme, the enzyme activity was measured with 0.1 M potassium phosphate buffer (pH 5.0–7.5) and 0.1 M Tris–HCl buffer (pH 8.0–10) at 25 °C. The final concentrations of 1 mM CDNB and 1 mM GSH were kept constant for the activity measurements.
To determine the ionic strength, at which the recombinant TtGSTm34-8xHis enzyme exhibited optimum activity, enzyme activity was measured with potassium phosphate buffer at pH 7.0 and with different ionic strengths (0.00625 M, 0.0125 M, 0.025 M, 0.05 M, 0.1 M, 0.15 M, 0.2 M, and 0.3 M) at 25 °C. The final concentrations of 1 mM CDNB and 1 mM GSH were kept constant for the activity measurements.
The optimum temperature of the recombinant TtGSTm34-8xHis enzyme was determined at the optimum ionic strength and pH, which were 0.2 M potassium phosphate and pH 7.0. For measuring the enzyme activity, the reaction mixture was incubated for five min in a well that helped to maintain the enzymes at the desired temperature (5 °C, 15 °C, 20 °C, 25 °C, 30 °C, 35 °C, 40 °C, 50 °C, and 60 °C). A final concentration of 1 mM CDNB and 1 mM GSH was used for the activity measurements.
2.8 k cat , V max , k cat, and Specificity Constant (k cat /K m ) of GSH and CDNB
Kinetic parameters were obtained from the Lineweaver‒Burk plot and then recalculated by nonlinear regression using Excel Solver software [25, 26]. The initial velocity parameters (Km and Vmax) of the recombinant TtGSTm34-8xHis enzyme were analyzed by varying the concentrations of GSH (0.1125 mM, 0.225 mM, 0.45 mM, 0.9 mM, 1.8 mM, 2.7 mM and 3.6 mM) or those of CDNB (0.1125 mM, 0.225 mM, 0.45 mM, 0.9 mM, 1.8 mM, 2.7 mM and 3.6 mM), whereas the concentrations of the other components of the reaction mixture were held constant, such as CDNB at 0.9 mM and GSH at 0.9 mM, respectively. Activity measurements were performed in 0.2 M potassium phosphate buffer (pH 7.0) at 25 °C.
The kcat value, which reflects the enzyme turnover number, was calculated using the kcat = Vmax/Et formula after the total enzyme amount (Et) was determined by a quantitative Bradford protein assay. One way of comparing the catalytic effects of enzymes encoded by paralogous or orthologous genes from different species or by the conversion of different substrates to a product with the same enzyme is to determine the specificity constant (V0), which was calculated by the V0 = kcat/Km formula.
2.9 Effects of Inhibitors
In vitro inhibition experiments of the TtGSTm34-8xHis enzyme were carried out by making minor changes to the method used by Trute et al. (2007) [27]. First, the enzyme (20 μl) was incubated with different concentrations of the inhibitor in 0.2 M potassium phosphate buffer (pH 7.0) for 15 min at 25 °C in 96-well plates. After the incubation, the reactions were initiated by adding 80 µl of 0.2 M potassium phosphate buffer (pH 7.0) containing 1 mM CDNB and 1 mM GSH. The initial rates of GST-CDNB activity were measured as described above.
2.10 Enzyme Stability Test
Changes in the activity of the TtGSTm34-8xHis enzyme stored at different temperatures (25 °C, 4 °C, and –20 °C in a deep freezer) for different durations (2 days, 10 days, 6 months, and 12 months) were determined under optimum enzyme reaction conditions (25 °C, 0.2 M potassium phosphate buffer, pH 7.0) with 1 mM CDNB and 1 mM GSH.
2.11 3D Modeling of the TtGSTm34 Protein
Molecular structural modeling was performed with the automated SWISS-MODEL homology modeling program [28]. The peptide sequence of TtGSTm34 was submitted to the SWISS-MODEL server to identify structures with sufficient similarity for homology modeling. Mus musculus GSTmu7 (Protein Data Bank Code: 2dc5), which has significant homology to TtGSTm34 (36.4% sequence identity), was used to construct the protein model. The model was drawn using BIOVIA’s Discovery Studio 2021 software. We also performed protein structure prediction via the AlphaFold machine learning program [29]. A ribbon diagram was generated using PyMol molecular graphics software.
3 Results
Previously, TtGSTm19 and TtGSTm34 were reported to be highly responsive genes after sublethal CDNB dose treatment of T. thermophila based on the results of the MTT test, GST activity, and real-time PCR analyses [18]. These enzymes seem to be major players in T. thermophila cell survival because they detoxify sublethal CDNB doses. Therefore, the enzymatic kinetics of TtGSTmu34 were studied with the CDNB substrate to determine how TtGSTmu34 could contribute to T. thermophila survival.
3.1 Recombinant Protein Expression, Affinity Purification, and Activity Analysis
The TtGSTm34 gene has a 660-bp long open reading frame, starting from ATG having Kozak sequence (Fig. 1), and encoding a polypeptide of 219 amino acids with a calculated molecular weight of 25 kDa and a theoretical pI of 7.7. The presence of rare codons in the open reading frame of the gene was examined according to the methods of Salim et al., (2008) [30], and no rare codons were found. Therefore, the sequence was not codon optimized for recombinant protein expression.
The constructed pIGF-1-TtGSTm34-8xHis vector was transformed into conjugative Tetrahymena cells by electroporation. Isolated positive transformants under antibiotic selection were induced with CdCl2 for 36 h, and total protein was isolated from the cells collected at different time intervals. The protein isolates were loaded into an SDS‒PAGE gel (Supplementary File 1). The amount of the expected 25 kDa protein band gradually increased from 6 to 36 h.
The soluble TtGSTm34-8xHis enzyme was first purified by using glutathione (GSH) affinity chromatography. This approach could help to purify recombinant TtGSTm34-8xHis along with all native GST proteins from transformant cells, but an anti-His antibody would detect only the 8xHis-tagged recombinant TtGSTm34. The transformant cells produced the 25 kDa recombinant TtGSTm34-8xHis protein, which was successfully purified by glutathione (GSH) affinity chromatography, according to Western blot analysis performed with anti-His antibodies (Supplementary File 2).
Alternatively, the soluble TtGSTm34-8xHis enzyme was purified from 40 ml of total protein lysate using 10 ml of Ni–NTA resin. SDS‒PAGE analyses also showed that Ni‒NTA affinity chromatography successfully purified the TtGSTm34-8xHis protein (Fig. 2, lanes 5, 6, and 7). Most of the TtGSTm34-8xHis protein was bound to the resins according to the flow-through data, whereas during the first wash step, some proteins were removed due to weak binding (Fig. 2, lanes 3 and 4). TtGSTm34-8xHis (25 kDa) was purified after elution in three steps with 20 ml of elution buffer containing 250 mM imidazole (Fig. 2; lanes 5, 6, and 7). As expected, Ni–NTA elution from uninduced T. thermophila transformant cells harboring the pIGF-TtGSTm34 vector did not produce 25 kDa protein bands, revealing no leaky recombinant protein production in the negative control (Fig. 2, lane 8).
The results suggest that the TtGSTm34-8xHis protein can be purified using either Ni–NTA or GSH affinity chromatography. However, TtGSTm34-8xHis was also dual-affinity purified with much higher purity (up to 95%) by sequential Ni–NTA and GSH bead purification. In this approach, it is important to note that the 250 mM imidazole in the Ni–NTA elution buffer could prevent the binding of TtGSTm34-8xHis to GSH resins, which might lead a decrease not only in the purified enzyme amount but also in its activity [31]. Therefore, it is crucial to remove the imidazole from the elution buffer in the end of Ni–NTA purification. The dual-affinity purification started with purified total soluble protein from transformant cells and continued with Ni–NTA bead purification to exclude native GST enzymes. The Ni–NTA elution buffer was replaced with a GSH binding buffer during the concentration of the eluted proteins by ultrafiltration. In the second affinity purification step, TtGSTm34-8xHis eluent in the GSH binding buffer was further purified with GSH beads. At the end of this sequential Ni–NTA and GSH (dual) affinity purification strategy, the purified proteins from each step were analyzed with SDS-PAGE (Fig. 3). TtGSTm34-8xHis elution protein was purified from Ni–NTA beads (Fig. 3, lane 2), while the flowthrough of ultrafiltration contained a very low amount of the nonspecific ~ 60 kDa protein band (Fig. 3, lanes 3 and 4). The soluble part of the TtGSTm34-8xHis elute was retained, and some precipitated proteins in the ultrafiltration membrane were redissolved in GSH binding buffer. However, the concentrated TtGSTm34-8xHis protein eluate still contained a low amount of contaminating background proteins (Fig. 3, lane 5). The flowthrough from the GSH affinity column showed that there were some unbound TtGSTm34-8xHis proteins plus 30 kDa and 70 kDa nonspecific contaminating proteins (Fig. 3, lane 6). The GSH affinity column showed a major band corresponding to the 25 kDa TtGSTm34-8xHis protein with approximately 95% purity and very low contamination (Fig. 3, lanes 7 and 8). Table 1 lists the results of each step of the dual-affinity purification of recombinant TtGSTm34. The enzyme activity of TtGSTm34 was determined under nonoptimized reaction conditions, namely, PBS buffer (pH 7.3), room temperature, 1 mM GSH, and 1 mM CDNB. The yield, purification fold change, and specific activity of the dual-affinity-purified TtGSTm34-8xHis enzyme from 40 ml of total protein lysate were determined to be 24.78%, 3166-fold, and 190 U/mg, respectively.
Coomassie-stained SDS‒PAGE gel of purified TtGSTMu34-8xHis protein after dual-affinity purification with ultrafiltration. pIGF-TtGSTm34-expressing transformant cells were induced with CdCl2. 1: Marker (Bio-Rad Cat No: 1610363); 2: first elution after purification with Ni–NTA beads; 3 and 4: soluble proteins that passed through the filter after the first and second centrifugation during ultrafiltration; 5: proteins that were concentrated by the ultrafiltration filter, 6: proteins that were not bound to GSH affinity beads (flow-through); 7 and 8: first and second elutions of proteins that were purified using GSH affinity beads
3.2 Enzyme Kinetics Assays
Using CDNB and GSH substrates, the enzymatic characteristics of the dual-affinity-purified TtGSTm34-8xHis were determined. In the studied pH range of 5.0 to 10.0, the optimum pH was found to be 7.0 (Fig. 4A). Following the determination of the optimum pH, the optimum ionic strength of the enzyme was determined by analyzing its activity at different phosphate buffer concentrations ranging from 6.25 mM to 300 mM at pH 7. The specific activity results indicated that the enzyme exhibited the highest activity at an ionic strength of 200 mM potassium phosphate buffer (Fig. 4B). On the other hand, the optimum temperature for the enzyme was found to be 25 °C when 5-min enzyme assays ranging from 5 °C to 60 °C were used (Fig. 4C). Incubating the enzyme at 5 °C and 60 °C resulted in a loss of activity of approximately 66% and 94%, respectively.
The Km values of TtGSTm34-8xHis for CDNB and GSH (0.47 and 0.54 mM, respectively) were first obtained by utilizing the Michaelis–Menten model, and these values were latter recalculated using the Lineweaver–Burk approach (Fig. 5) via the Solver tool of Microsoft Excel software. These recalculated values for GSH were Km = 0.68 mM and Vmax = 493 U/mg. For CDNB, the recalculated values were as Km = 0.40 mM and Vmax = 443 U/mg. Lineweaver–Burk plots for CDNB and GSH are shown in Fig. 5, and the calculated finalized values of Km, Vmax, kcat (catalytic constant), and kcat/Km (catalytic efficiency) for the CDNB and GSH substrates are given in Table 2.
Lineweaver‒Burk double-reciprocal plots of substrate concentration versus GST activity. A: Effect of different concentrations of CDNB (0.1125 mM–3.6 mM) as a substrate in the presence of 0.9 mM GSH in the final volume of 200 mM potassium phosphate. B: Effect of different concentrations of GSH (0.1125 mM–3.6 mM) in the presence of 0.9 CDNB as a substrate in the final volume of 200 mM potassium phosphate
kcat values for GSH and CDNB, known as the turnover number/rate and defined as the amount of substrate converted into a product by a single active site of an enzyme per unit time, were determined to be 186.9 and 195.4, respectively (Table 2). The kcat/Km ratios for TtGSTm34-8xHis were found to be 274, 8 and 488.5 for GSH and CDNB, respectively.
Different concentrations of CuSO4, CdCl2, and ethacrynic acid (ECA) were investigated to determine whether they had effects on the ability of the TtGSTm34-8xHis enzyme to conjugate GSH to CDNB, and the results are shown in Fig. 6. Even at the maximum dose of 15 µM, CdCl2 caused an activity loss of approximately 23% compared to the control group. However, CuSO4 exhibited a greater percentage of dose-dependent inhibition; the three highest concentrations of CuSO4 inhibited TtGSTm34-8xHis activity, lowering the initial activity to 62%. The highest used, 1 mM, concentration of ECA completely inhibited the enzyme activity.
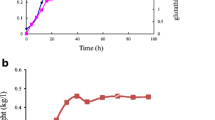
The shelf life or stability of the TtGSTm34-8xHis enzyme was also tested under different storage/usage conditions at the time points defined within the scope of the experiments (Table 3). When the enzyme was stored in glycerol and phosphate buffer at -20 °C, the lost enzyme activity was approximately 23% after six months and 39% after twelve months. Furthermore, the enzyme retained 67% of its activity even after being kept in a + 4 °C freezer for 10 days. After incubating for 2 days at room temperature, the TtGSTm34-8xHis enzyme was able to retain approximately 82% of its activity.
3.3 Homology-Based 3D Modeling of the TtGSTm34 Protein
Homology modeling by SWISS-MODEL based upon the crystal structure of Mus musculus GSTMu7 (PDB ID: 2dc5) was used to predict the protein structure of TtGSTm34. The homology model of TtGSTm34 includes a thioredoxin fold β1-α1-β2-α2-β3-β4-α3 in the N-terminal domain and a six-alpha helix fold in the C-terminal domain, similar to the predicted 3D structures of other Tetrahymena GSTs (Fig. 7A) [6]. The alignment of TtGSTm34 and Mus musculus GSTMu7 amino acid sequences revealed 36.4% identity and 49% similarity (Fig. 7B). The introduced gaps were in the loop regions of GSTMu7, so they are unlikely to affect the secondary structure formation. In TtGSTm34, the * marked amino acids, which are locating in the active sites and GSH substrate binding sites, are also well conserved. The beta sheets and alpha helixes were also positioned correctly in the GSH binding domain in the N-terminal. However, the identity (red boxes) and similarity (yellow boxes) were decreasing in the substrate binding domain of the C-terminal because of the expected differentiation on the substrate selectivity (amino acids marked with + and #) (Fig. 7B). We also constructed the three-dimensional structure of TtGSTm34 using AlphaFold, which has remarkable similarities to the homology model made by SWISS-MODEL (Fig. 7C), indicating that the overall protein’s tertiary structure was consistent and reliably predicted.
Structural modeling of TtGSTm34. A: Ribbon representation of structural 3D model of TtGSTm34 was constructed based on the Mus musculus GSTmu7 (pdb ID: 2dc5) protein using SWISS-MODEL protein modeling program. In the TtGSTm34 protein, the N-terminal domain is the glutathione binding site, while the C-terminal domain has xenobiotics binding. Proteins function as homodimeric structures, but only monomeric structures are given in the figure. B: The amino acid sequences of the M. musculus GSTmu7 and TtGSTm34 were aligned using ClustalW, outcome of which was evaluated with ESPript 3 program using the GSTm7 3D structure information. Alpha helices and beta strands are represented as helices and arrows, respectively, and strict beta turns are marked with TT. The catalytic Tyr7 (Y) residue in the active site of Mu class GSTs was also conserved as Tyr6 in TtGSTm34, marked with a red asterisk. Periods indicate the introduced gaps to optimize the alignment. Identities (in red boxes) and similarities (in yellow boxes) between the TtGSTm34 and GSTm7 sequences are shown by colored shadowing. The GSH-binding amino acid residues (G-site) in the N-terminal domain marked with “*”. The substrate binding amino acid residues (H-site) in the C-terminal domain were shown with “#”. The interacting interface residues of the N-terminal domain with the C-terminal domain are marked with “ + ”. C: Ribbon 3D structural models of TtGSTm34 and Mus musculus GSTmu7 were constructed based on the Alphafold program. The beta-sheets are shown in green, and the alpha-helices are shown in red or pink
4 Discussion
In this study, the TtGSTm34 gene was cloned and expressed in T. thermophila, and the protein was affinity purified. The dual affinity method was chosen for the purification of recombinant enzyme with the aim of eliminating endogenous GST enzymes. In this method, initially, all proteins bound to the column via the histidine tag were purified using Ni–NTA affinity. While non-specifically bound proteins to nickel were also collected in the elution, in the second glutathione affinity purification step, only the recombinant enzyme remained bound to the column. In our previous study, gel images, which obtained after the purification of endogenous GST enzymes from Tetrahymena thermophila, showed approximately 23 kDa protein bands [6]. The length and size of TtGSTm34 were similar to those of other GST genes, such as those of Hyalomma rufipes (233 a.a., 25.62 kDa) [32], Hemaphysalis longicornis (223 a.a., 25.7 kDa) [33] and Litopenaeus vannamei (215 a.a., 25.18 kDa) [34]. In this study, the expected size of TtGSTm34 is 23 kDa but reaching 25.5 kDa with the approximately 2.5 kDa 8 × histidine tag attachment in the recombinant TtGSTm34-8xHis protein, as observed after the column purification (Fig. 3). The enzymatic properties of recombinant TtGSTm34 were subsequently characterized with CDNB and GSH, which are universal GST substrates.
The hypothetical origin of the GST superfamily was ancient, and the superfamily evolved from a thioredoxin-like ancestor, which is also supported by the predicted TtGSTm34 structure [35]. In GST enzymes, the C-terminal region participates in substrate recognition, the N-terminal domain has the GSH binding activity [36]. In TtGSTm34, the GSTm subfamily specific motif 2, which has FPNLPY(L/I)I(D/H)GD consensus sequence, was also present as FPNIPYLIDGD in the β3 sheet (Fig. 7B) [37]. In higher eukaryotes, the mu loop between β2 strand and α2 helix with GDAPDYDRSQ sequence increases substrate affinity and localization [38]. However, the absence of mu loop in the TtGSTm34 caused a prediction about the lower enzyme’s affinity against CDNB or other xenobiotic substrates but the results in this study showed that it is otherwise.
The Ni–NTA purification step of the double-affinity purification protocol was highly successful, with 38-fold greater purification of the TtGSTm34 protein than 4.8-fold greater purification of the recombinant Thais clavigera GSTmu gene from E. coli [39]. TtGSTm34 also has much greater specific activity than GSTs from the other species listed in Table 4. However, the high amount of recombinant TtGSTm34 protein was lost due to precipitation under a low volume of water during the ultrafiltration step between the Ni–NTA and glutathione affinity purification. Therefore, this step needs to be replaced with dialysis to increase the yield. These findings suggest that in aquatic environments, TtGSTm34 may be crucial in protecting Tetrahymena cells from oxidative stress caused by xenobiotic substances like CDNB [18].
In enzyme kinetics studies, the Km values provide basic information about the affinity of an enzyme for a substrate. For example, the Km of the recombinant GSTpi enzyme with CDNB in the blue mussel Mytilus edulis has been reported to be 0.68 mM, with a Vmax of 103 μmol min−1 mg protein−1 [44]. The Km values of recombinant Ruditapes philippinarum GSTμ with glutathione and CDNB were determined to be 1.03 mM and 0.56 mM, respectively [45]. For GSH and CDNB, the kcat/Km ratios for TtGSTm34-8xHis were found to be 345.3 and 407.7, respectively. The lower Km values (0.68 mM and 0.40 mM) of the recombinant TtGSTm34 enzyme signify high substrate affinity with GSH and CDNB substrates compared to other GST enzymes in aquatic organisms (Table 4).
The optimum temperature, pH, and ionic strength of the TtGSTm34 protein were determined to be approximately 25 °C, pH 7.0, and 0.15–0.2 M potassium phosphate buffer, respectively. The optimum pH values of the GST enzymes obtained from Schistosoma japonicum, Plasmodium vivax, and Sus scrofa were 7.0, 7.0–7.5, and 8.0, respectively [46,47,48]. These temperature and pH values revealed that the stability and capability of GSTs evolved to function more closely to the extracellular and intracellular conditions of the studied organisms.
Metal cations are generally not necessary for the catalytic activity of GSTs. On the other hand, it has been demonstrated that Cu2+ and Cd2+ can reduce the catalytic activity of certain cytosolic GSTs, such as human GSTP [49]. An inhibition assay of TtGSTm34 activity by CuSO4 and CdCl2 showed that recombinant TtGSTm34 activity is significantly inhibited by divalent metal cations, such as Cu2+. This inhibition could occur through sulfhydryl oxidation, which is catalyzed by Cu2+. In the present study, 12.5 µM, 25 µM, and 50 µM CuSO4 decreased the TtGSTm34 enzyme activity by approximately 32%, 36%, and 38%, respectively. Similarly, the enzyme activity of the Locusta migratoria-African locust GST was highly sensitive and inhibited by 50 µM CuSO4 by approximately 80% [50]. Copper also significantly reduced the affinity of GST for GSH (Vmax ↓, Km ↑), while the substrate affinity for xenobiotics remained unchanged [51]. Thus, the decrease in the Vmax and catalytic efficiency of TtGSTm34 following the addition of CuSO4 could be explained by the decrease in the affinity of TtGSTm34 for GSH. Moreover, the highest dose of CdCl2 (15 µM) inhibited TtGSTm34 about 27%, whereas the much higher dose of 200 µM CdCl2 inhibited GST activity by approximately 37% in the rat liver [52]. It was already reported that cadmium depletes GSH levels through the formation of the Cd–glutathione complex and/or oxidation of GSH to its oxidized form [53]. These observed and reported differences in inhibition could be explained by the experimental conditions (substrate concentrations, source, concentration of GSTs, and GST isoforms), species-specific GST enzyme differentiation, and the GSTs usage from different GST subfamilies.
Recombinant TtGSTm34 was found to be very sensitive to ECA, which is a common GST inhibitor. A statistically significant inhibitory effect was observed at 0.01 mM, 0.1 mM, and 1 mM ECA, which resulted in 76%, 92%, and 98% inhibition, respectively. The potency of ECA as a GST inhibitor on CDNB has been observed in rat and human GSTs [54]. However, the inhibitory effect of ECA on the activity of TtGSTm34 is probably attributed to its direct binding to the SH group of two cysteines on the enzyme. Additionally, ECA has a ketone moiety that can also bind to glutathione [55]. Thermodynamically, the conjugation of ECA to GSH is highly preferred over the enzymatic conjugation of GSH to CDNB. Therefore, ECA likely acts not only as a depleting agent for GSH but also as a direct inhibitor of TtGSTm34, resulting in total inhibition of TtGSTm34 enzyme activity.
In our previous study, TtGSTm34 was the gene with the highest responsiveness at the transcriptional and posttranscriptional levels when T. thermophila cells were exposed to CDNB [18]. The low Km value and high turnover rate of the TtGSTm34-8xHis enzyme also supported our previous published results. The defined enzymatic characteristics of recombinant TtGSTm34 provide evidence that this enzyme might indeed contribute to the cell’s first defense mechanism against oxidative stress driven by xenobiotics. Moreover, the high affinity of TtGSTm34 for GSH revealed that this protein could be used as a native GST tag in biotechnological studies of recombinant protein production to increase purification efficiency. Additionally, the high GST activity obtained from the TtGSTm34 protein has revealed its potential use in commercial GST Enzyme Assay kit when the activities of commercially available enzymes was compared with TtGSTm34-8xHis (Table 5). We concluded that the TtGSTm34-8xHis enzyme could have a commercial value with its high enzyme activity and affinitic feature.
Data Availability
No datasets were generated or analysed during the current study.
References
Lundgren B, DePierre JW (1990) The metabolism of xenobiotics and its relationship to toxicity/genotoxicity: Studies with human lymphocytes. In: Acta Physiol Scand Suppl 592:49–59
Rostami-Hodjegan A, Tucker GT (2007) Simulation and prediction of in vivo drug metabolism in human populations from in vitro data. Nat Rev Drug Discov 6:140–148. https://doi.org/10.1038/nrd2173
Xu C, Li CY-T, Kong A-NT (2005) Induction of phase I, II and III drug metabolism/transport by xenobiotics. Arch Pharm Res 28:249–268. https://doi.org/10.1007/BF02977789
Allocati N, Federici L, Masulli M, Di Ilio C (2009) Glutathione transferases in bacteria. FEBS J 276:58–75. https://doi.org/10.1111/j.1742-4658.2008.06743.x
Enayati AA, Ranson H, Hemingway J (2005) Insect glutathione transferases and insecticide resistance. Insect Mol Biol 14:3–8. https://doi.org/10.1111/j.1365-2583.2004.00529.x
Üstüntanır Dede AF, Arslanyolu M (2019) Genome-wide analysis of the Tetrahymena thermophila glutathione S-transferase gene superfamily. Genomics 111:534–548. https://doi.org/10.1016/j.ygeno.2018.11.034
Gullner G, Komives T, Király L, Schröder P (2018) Glutathione S-transferase enzymes in plant-pathogen interactions. Front Plant Sci 871:1–19. https://doi.org/10.3389/fpls.2018.01836
Ogunmoyole T, Adewale IO, Fodeke AA, Afolayan A (2020) Catalytic studies of glutathione transferase from Clarias gariepinus (Burchell) in dilute and crowded solutions. Comp Biochem Physiol - C: Toxicol Pharmacol 228:108648. https://doi.org/10.1016/j.cbpc.2019.108648
Dasari S, Ganjayi MS, Meriga B (2017) Bird glutathione S-transferases: Endogenous and exogenous toxic insults. Adv Anim Vet Sci 5:388–394. https://doi.org/10.17582/journal.aavs/2017/5.9.388.394
Landi S (2000) Mammalian class theta GST and differential susceptibility to carcinogens: A review. Mutat Res Rev Mutat Res 463:247–283. https://doi.org/10.1016/S1383-5742(00)00050-8
Frova C (2006) Glutathione transferases in the genomics era: New insights and perspectives. Biomol Eng 23:149–169. https://doi.org/10.1016/j.bioeng.2006.05.020
Li J, Wang Y, Hu J et al (2023) Molecular identification and biochemical characteristics of a delta class glutathione S-transferase gene (FcδGST) from Chinese shrimp Fenneropenaeus chinensis. J Oceanol Limnol 41:1940–1953. https://doi.org/10.1007/s00343-022-2271-2
Feng M, Hu Y, Yang L et al (2023) GST-Mu of Cristaria plicata is regulated by Nrf2/Keap1 pathway in detoxification microcystin and has antioxidant function. Aquat Toxicol Oct 263:106708. https://doi.org/10.1016/j.aquatox.2023.106708
Ding LL, Yu SJ, Lei S et al (2024) Identification and functional characterization of an omega-class glutathione s-transferase gene PcGSTO1 associated with cyetpyrafen resistance in panonychus citri (McGregor). J Agric Food Chem. https://doi.org/10.1021/acs.jafc.4c00732
Weisse T, Sonntag B (2016) Ciliates in Planktonic Food Webs: Communication and Adaptive Response. In: Witzany G, Nowacki M (eds) Biocommunication of Ciliates. Springer, Cham. https://doi.org/10.1007/978-3-319-32211-7_19
Overbaugh JM, Lau EP, Marino VA, Fall R (1988) Purification and preliminary characterization of a monomeric glutathione S-transferase from Tetrahymena thermophila. Arch Biochem Biophys 261:227–234. https://doi.org/10.1016/0003-9861(88)90336-0
Öziç C, Arslanyolu M (2012) Characterization of affinity tag features of recombinant Tetrahymena thermophila glutathione-S-transferase zeta for Tetrahymena protein expression vectors. Turk J Biol 36:513–526. https://doi.org/10.3906/biy-1110-2
Kapkaç HA, Arslanyolu M (2021) Identification of glutathione-S-transferase m19 and m34 among responsive GST genes against 1-chloro-2,4-dinitrobenzene treatment of Tetrahymena thermophila. Eur J Protistol 81:125838. https://doi.org/10.1016/J.EJOP.2021.125838
Brunk CF, Sadler LA (1990) Characterizaton of the promoter region of Tetrahymena genes. Nucleic Acids Res 18:323–329. https://doi.org/10.1093/nar/18.2.323
Yilmaz G, Arslanyolu M (2015) Efficient expression of codon-adapted affinity tagged super folder green fluorescent protein for synchronous protein localization and affinity purification studies in Tetrahymena thermophila. BMC Biotechnol 15:1–9. https://doi.org/10.1186/s12896-015-0137-9
Gaertig J, Gorovsky MA (1992) Efficient mass transformation of Tetrahymena thermophila by electroporation of conjugants. Proc Natl Acad Sci U S A 89:9196–9200. https://doi.org/10.1073/pnas.89.19.9196
Aslan E, Arslanyolu M (2015) Identification of neutral and acidic deoxyribonuclease activities in Tetrahymena thermophila life stages. Eur J Protistol 51:173–185. https://doi.org/10.1016/j.ejop.2015.02.004
Zhang X, Thompson GA (1997) An apparent association between glycosylphosphatidylinositol-anchored proteins and a sphingolipid in Tetrahymena mimbres. Biochem J 323:197–206. https://doi.org/10.1042/bj3230197
Habig WH, Pabst MJ, Jakoby WB (1974) Glutathione S transferases. The first enzymatic step in mercapturic acid formation. J Biol Chem. https://doi.org/10.14026/j.cnki.0253-9705.2010.23.013
Marasović M, Marasović T, Miloš M (2017) Robust nonlinear regression in enzyme kinetic parameters estimation. Hindawi J Chem 2017:1–12. https://doi.org/10.1155/2017/6560983
Brown AM (2001) A step-by-step guide to non-linear regression analysis of experimental data using a Microsoft Excel spreadsheet. Comput Methods Programs Biomed 65:191–200. https://doi.org/10.1016/S0169-2607(00)00124-3
Trute M, Gallis B, Doneanu C et al (2007) Characterization of hepatic glutathione S-transferases in coho salmon (Oncorhynchus kisutch). Aquat Toxicol 81:126–136. https://doi.org/10.1016/j.aquatox.2006.11.009
Guex N, Peitsch MC, Schwede T (2009) Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: A historical perspective. Electrophoresis 30:162–173. https://doi.org/10.1002/elps.200900140
Jumper J, Evans R, Pritzel A et al (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589. https://doi.org/10.1038/s41586-021-03819-2
Salim HMW, Ring KL, Cavalcanti ARO (2008) Patterns of codon usage in two ciliates that reassign the genetic code: Tetrahymena thermophila and Paramecium tetraurelia. Protist 159:283–298. https://doi.org/10.1016/j.protis.2007.11.003
Caramia S, Gatius AGM, dal Piaz F et al (2017) Dual role of imidazole as activator/inhibitor of sweet almond (Prunus dulcis) β-glucosidase. Biochem Biophys Rep 10:137–144. https://doi.org/10.1016/J.BBREP.2017.03.007
Zhao M, Gao Z, Ji X et al (2024) The diverse functions of Mu-class Glutathione S-transferase HrGSTm1 during the development of Hyalomma rufipes with a focus on the detoxification metabolism of cyhalothrin. Parasit Vectors 17(1):1–12. https://doi.org/10.1186/s13071-023-06084-6
Hernandez EP, Kusakisako K, Talactac MR et al (2018) Characterization and expression analysis of a newly identified glutathione S-transferase of the hard tick Haemaphysalis longicornis during blood-feeding. Parasit Vectors 11:1–16. https://doi.org/10.1186/s13071-018-2667-1
Contreras-Vergara CA, Valenzuela-Soto E, García-Orozco KD et al (2007) A mu-class glutathione S-transferase from gills of the marine shrimp Litopenaeus vannamei: Purification and characterization. J Biochem Mol Toxicol 21:62–67. https://doi.org/10.1002/jbt.20162
Sheehan D, Meade G, Foley VM, Dowd CA (2001) Structure, function and evolution of glutathione transferases : implications for classification of non-mammalian members of an ancient enzyme superfamily. Biochem J 360:1–16. https://doi.org/10.1042/0264-6021:3600001
Marrs KA (1996) The functions and regulation of glutathione S-transferases in plants. Annu Rev Plant Physiol Plant Mol Biol 47:127–158. https://doi.org/10.1146/ANNUREV.ARPLANT.47.1.127
Rosa De Lima MF, Sanchez Ferreira CA, Joaquim De Freitas DR et al (2002) Cloning and partial characterization of a Boophilus microplus (Acari: Ixodidae) glutathione S-transferase. Insect Biochem Mol Biol 32:747–754. https://doi.org/10.1016/S0965-1748(01)00157-6
Hearne JL, Colman RF (2006) Contribution of the mu loop to the structure and function of rat glutathione transferase M1–1. Protein Sci 15:1277–1289. https://doi.org/10.1110/ps.062129506
Rhee JS, Raisuddin S, Hwang DS et al (2008) A Mu-class glutathione S-transferase (GSTM) from the rock shell Thais clavigera. Comp Biochem Physiol - C Toxicol Pharmacol 148:195–203. https://doi.org/10.1016/j.cbpc.2008.05.011
Adewale IO, Afolayan A (2005) Purification and catalytic properties of glutathione transferase from the hepatopancreas of crayfish Macrobrachium vollenhovenii (Herklots). J Biochem Mol Toxicol 18:332–344. https://doi.org/10.1002/jbt.20044
Hubatsch I, Ridderström M, Mannervik B (1998) Human glutathione transferase A4–4: An Alpha class enzyme with high catalytic efficiency in the conjugation of 4-hydroxynonenal and other genotoxic products of lipid peroxidation. Biochem J 330:175–179. https://doi.org/10.1042/bj3300175
Hoarau P, Garello G, Gnassia-Barelli M et al (2002) Purification and partial characterization of seven glutathione S-transferase isoforms from the clam Ruditapes decussatus. Eur J Biochem 269:4359–4366. https://doi.org/10.1046/j.1432-1033.2002.03141.x
Yamamoto K, Usuda K, Kakuta Y et al (2012) Structural basis for catalytic activity of a silkworm Delta-class glutathione transferase. Biochim Biophys Acta Gen Subj 1820:1469–1474. https://doi.org/10.1016/j.bbagen.2012.04.022
Yang HL, Zeng QY, Li EQ et al (2004) Molecular cloning, expression and characterization of glutathione S-transferase from Mytilus edulis. Comp Biochem Physiol - B Biochem Mol Biol 139:175–182. https://doi.org/10.1016/j.cbpc.2004.06.019
Bathige SDNK, Umasuthan N, SaranyaRevathy K et al (2014) A mu class glutathione S-transferase from Manila clam Ruditapes philippinarum (RpGSTμ): Cloning, mRNA expression, and conjugation assays. Comp Biochem Physiol - C: Toxicol Pharmacol 162:85–95. https://doi.org/10.1016/j.cbpc.2014.03.007
Na BK, Kang JM, Kim TS, Sohn WM (2007) Plasmodium vivax: Molecular cloning, expression and characterization of glutathione S-transferase. Exp Parasitol 116:414–418. https://doi.org/10.1016/j.exppara.2007.02.005
Yang XQ, Zhang YL (2015) Characterization of glutathione S-transferases from Sus scrofa, Cydia pomonella and Triticum aestivum: Their responses to cantharidin. Enzyme Microb Technol 69:1–9. https://doi.org/10.1016/j.enzmictec.2014.11.003
Lo WJ, Chiou YC, Hsu YT et al (2007) Enzymatic and nonenzymatic synthesis of glutathione conjugates: Application to the understanding of a parasite’s defense system and alternative to the discovery of potent glutathione S-transferase inhibitors. Bioconjug Chem 18:109–120. https://doi.org/10.1021/bc0601727
Mm A-A, Pj D (1990) In vitro interaction of mercury, copper (II) and cadmium with human glutathione transferase π. Res Commun Chem Pathol Pharmacol 69:99–102
Qin G, Jia M, Liu T et al (2013) Characterization and Functional Analysis of Four Glutathione S-Transferases from the Migratory Locust, Locusta migratoria. PLoS One 8:1–11. https://doi.org/10.1371/journal.pone.0058410
Dobritzsch D, Grancharov K, Hermsen C et al (2020) Inhibitory effect of metals on animal and plant glutathione transferases. J Trace Elem Med Biol 57:48–56. https://doi.org/10.1016/J.JTEMB.2019.09.007
Dierickx PJ (1982) In vitro inhibition of the soluble glutathione S-transferases from rat liver by heavy metals. Enzyme 27:25–32. https://doi.org/10.1159/000459018
Ullah H, Khan MF, Jan SU, Hashmat F (2015) Cadmium-glutathione complex formation in human t-cell and b-cell lymphocytes after their incubation with organo-cadmium diacetate. Pak J Pharm Sci 28:2075–2081
Ploemen JHTM, van Ommen B, de Haan A et al (1993) In vitro and in vivo reversible and irreversible inhibition of rat glutathione S-transferase isoenzymes by caffeic acid and its 2-S-glutathionyl conjugate. Food Chem Toxicol 31:475–482. https://doi.org/10.1016/0278-6915(93)90106-9
Wongtrakul J, Sramala I, Prapanthadara LA, Ketterman AJ (2005) Intra-subunit residue interactions from the protein surface to the active site of glutathione S-transferase AdGSTD3-3 impact on structure and enzyme properties. Insect Biochem Mol Biol 35:197–205. https://doi.org/10.1016/j.ibmb.2004.11.003
Acknowledgements
This work was funded by the Anadolu University Scientific Research Projects Commission (AUBAP) with grant number 1001F45, which was given to Muhittin Arslanyolu. This project was awarded a European Cooperation in Science and Technology (COST) Grant under the Action BM1102 by the Scientific and Technological Research Council of Türkiye (TÜBİTAK) and AUBAP.
Author information
Authors and Affiliations
Contributions
HAK carried out each experiment, examined the results, and prepared the manuscript. MA conceptualized and supervised the project and revised the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kapkaç, H.A., Arslanyolu, M. Molecular Cloning, Expression and Enzymatic Characterization of Tetrahymena thermophila Glutathione-S-Transferase Mu 34. Protein J 43, 613–626 (2024). https://doi.org/10.1007/s10930-024-10204-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-024-10204-1