Abstract
The Yucatan Peninsula (YP) is a biotic province located in southeastern Mexico, delimited mainly by climatic variables. One endemic species of the YP is the Yucatan deer mouse Peromyscus yucatanicus. It is considered a member of the P. mexicanus species group, but some morphological characters and habitat preferences separate it from them. Herein, the DNA barcoding identification of P. yucatanicus, intraspecific relationships, and the level of genetic differentiation among its geographical distribution were examined. Sequences of the mitochondrial cytochrome c oxidase subunit 1 gene were used for the phylogenetic, demographic history, and genetic structure analysis. In addition, an ecological niche model was built and transferred to 6000 years ago in order to explore the current and past environmental suitability of the species. Results showed that P. yucatanicus was monophyletic and its phylogenetic relationships unresolved. Intraspecific analyses showed signatures of a scenario of demographic stability followed by population growth, and three genetic haplogroups were identified. Paleontological, paleoclimate, and the results presented here are useful to hypothesize that P. yucatanicus likely diverged in the Pleistocene and invaded the south of YP after the Last Glacial Period with the arrival of the current vegetation in the late Pleistocene—early-middle Holocene, and its demographic population was stable during the remaining Holocene epoch with slight growth at the late-middle Holocene resulting in the major precipitation changes that provided more plant coverage.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Current knowledge supports that the biological history of the Yucatan Peninsula (YP), located in southeastern Mexico, began 65 million of years ago (mya) after the collision of an asteroid at what is now the north of the YP, extinguishing all life for hundreds of kilometers (Vázquez-Domínguez and Arita 2010). After numerous geologic, climatic, and biotic processes, the recent subsequent Great American Biotic Interchange 2.8 mya marked the beginning of the modern flora and fauna of the region (MacFadden 2006).
The YP is a biotic province delimited mainly by climatic variables, flora and fauna distributions. Two geographical subprovinces are considered in the YP: the northern Yucatan Province and the southern Peten Province (Morrone 2005), but specific studies in plants and vertebrates expanded the number to 14 subunits (Duno-de Stefano et al. 2012). In this region, the climate is warm with a gradient of humidity from northwest to southeast (Folan et al. 1983). According to physiologic and phenological criteria, the principal types of vegetation are low, semi-low, and medium semi-deciduous forest; low, medium, and high semi-evergreen forest; savannas; palm groves; mangroves; coastal dunes; and reed beds (Islebe et al. 2015). On the other hand, the biodiversity in the YP is abundant. It is estimated to have 2300 species of vascular plants with almost 9% endemism, 5765 species of invertebrates, and 1551 vertebrates including 118 species of mammals (Durán-García et al. 2016). Only 15 species of small mammals inhabit the peninsula, including three species of cricetids: Oryzomys palustris, Otonyctomys hatti, and Peromyscus yucatanicus (Hernández-Betancourt et al. 2010; Zaragoza-Quintana et al. 2016).
The Yucatan deer mouse Peromyscus yucatanicus Allen and Chapman, 1897, is distributed throughout the semi-deciduous forests in the YP, but locally common in cornfields, banana plantations, secondary growth forests, and dense grass near towns (Young and Jones 1983; Zarza et al. 2003). It was considered a monotypic species (Lawlor 1965), but currently, two subspecies are recognized based only on the pelage coloration: P. y. yucatanicus Allen and Chapman, 1897, with ochraceous buff upperparts distributed in the northern part of the peninsula, and at south P. y. badius Osgood, 1904, with brownish buff upper parts (Young and Jones 1983; Zarza et al. 2003).
Peromyscus yucatanicus has been contemplated as a member of the P. mexicanus species group, but some morphological characters separate it from them (Huckaby 1980; Carleton 1989). Genetic information of allozymes identified similarity of P. yucatanicus with some species of the P. mexicanus species group (P. guatemalensis, P. gymnotis, P. mexicanus, and P. zarhynchus), and P. nudipes (Rogers and Ensgstrom 1992). However, because of its strict lowland habitat preferences, this species was not considered as a member of the P. mexicanus species group, which is of mountain habitat preferences (Ordóñez-Garza et al. 2010; Pérez-Consuegra and Vázquez-Domínguez 2015). Recently, phylogenetic analyses using the mitochondrial gene cytochrome b (Cytb) have confirmed the monophyly of the P. mexicanus species group, and the close phylogenetic relationships among P. tropicalis, P. nudipes, P. mexicanus, P. gymnotis, P. zarhynchus, P. gardneri, P. grandis, P. guatemalensis, P. salvadorensis, P. nicaraguae, and five lineages undescribed (Ordóñez-Garza et al. 2010; Bradley et al. 2016; Pérez-Consuegra and Vázquez-Domínguez 2016). Although these studies included other species, such as P. mayensis, P. megalops, P. stirtoni, P. melanophrys, P. melanocarpus, and P. perfulvus, confirming their close phylogenetic relationships (Bradley et al. 2007; Miller and Engstrom 2008; Castañeda-Rico et al. 2014; Platt et al. 2015; Sullivan et al. 2017), P. yucatanicus was not included and its phylogenetic relationships remains unclear.
The aim of this study is to explore the genetic affinity of P. yucatanicus and other closely related species. In addition, the intraspecific relationships within P. yucatanicus populations, demographic history, and the level of genetic differentiation were examined using the public DNA sequences of the mitochondrial cytochrome c oxidase subunit 1 gene (CO1) available in Bold Systems DataBase and GenBank.
Material and Methods
Character Sampling and Taxonomic Identity
Taxa belonging to the P. mexicanus species group and relatives were selected from the Bold Systems DataBase (Ratnasingham and Hebert 2007), which also are available in NCBI Genbank Database (https://www.ncbi.nlm.nih.gov/genbank/). A total of 810 sequences of the mitochondrial CO1 gene were downloaded with its specimen data, and initially included the following species and specimen number: P. grandis (67), P. guatemalensis (46), P. gymnotis (22), P. mayensis (14), P. megalops (3), P. melanocarpus (4), P. mexicanus (366), P. nudipes (34), P. stirtoni (22), P. zarhynchus (49), and P. yucatanicus (183). Two additional sequences were downloaded from GenBank of Neotomodon alstoni as outgroup. The specimen information and accession numbers from BOLD Systems and Genbank can be found in the Supporting Information Table S1.
Sequences were aligned using the MUSCLE algorithm implemented in the MSA v 1.16.0 package (Bodenhofer et al. 2015) in R software v 3.5.2 (R Foundation for Statistical Computing, Vienna, Austria). The nucleotide substitution saturation was analyzed by codon with the method of Xia et al. (2003), using 1000 replicates and considering the extreme 32 operative taxonomic units in DAMBE v 7.2.7 (Xia 2018). This method calculates an entropy-based index of substitution saturation (Iss) and its critical value (Issc). If this index of saturation is not smaller than the critical value, then sequences contain substantial saturation and poor phylogenetic signal (Xia et al. 2003). Finally, the best fitting partitioning scheme by codons, and nucleotide substitution model were selected using the branch lengths unlinked, greedy algorithm, and the Akaike Information Criterion (AIC) in PartitionFinder Version 1.1 (Lanfear et al. 2012).
Phylogenetic Inference
The phylogenetic inferences were carried out with Maximum Likelihood (ML) and Bayesian Inference (BI). The ML analysis was performed in RAXML v 8.2.8. (Stamatakis 2014) using the best model of nucleotide substitution selected, and 1000 rapid bootstraps. The BI was achieved in MRBAYES v 3.2.6 (Ronquist et al. 2012) using the best model of nucleotide substitution with two independent Metropolis Markov-chains Monte Carlo runs, 20 million generations, sampling every 10,000 trees, and a burn-in of 20%; a majority rule consensus tree with nodal support was constructed with the remaining trees. Stationarity, convergence, and visualization of the log-likelihood scores for the two runs were checked in RWTY v 1.0.2 package (Warren et al. 2017) in R software.
Demographic History and Genetic Differentiation of P. yucatanicus
A total of 183 sequences available of P. yucatanicus from 17 localities were used for the posterior analyses. The number of haplotypes, haplotype diversity, segregating sites, and nucleotide diversity were estimated for all specimens in the PEGAS v 0.11 package (Paradis 2010) in R software. Estimation of the number of total haplotypes was performed with the “chaoHaplo” function in SPIDER v 1.5.0 package (Brown et al. 2012) after removing the sites with gaps in the matrix. To assess the historical demographic change, such as expansion population, the Tajima’s D (Tajima 1989) and R2 (Ramos-Onsins and Rozas 2002) were calculated using 10,000 coalescent simulations. Lastly, the mismatch distributions were computed using the function “MMD” in PEGAS package v 0.11 in R software, and in order to infer demographic population fluctuations over time, the Bayesian Skyline Plot was constructed in BEAST v 2.5.2 (Bouckaert et al. 2019) using the Coalescent Bayesian Skyline prior that does not depend on a pre-specified parametric model of demographic history (Drummond et al. 2005), the substitution rate was estimated by BEAST, the GTR + I + G model of nucleotide substitution was used, with a relaxed lognormal molecular clock, 20 million of generations sampling each 1000 iterations, and a final burn-in of 20%; the demographic plot was performed in TRACER v 1.7.1 (Rambaut et al. 2018).
The haplotype network was constructed using the Minimum Spanning Network algorithm implemented in POPART v 1.7 (Leigh and Bryant 2015). The genetic differentiation among the haplogroups resulting from the haplotype network was used as a-priori groups and the genetic variation among them was examined with a Discriminant Analyses of Principal Components (DAPC; Jombart et al. 2010) conducted in the ADEGENET v 2.1.1 package (Jombart 2008) in R software. To retain the optimal number of principal components (PC) and avoid over-fitting in the discriminant functions, the “optim.a.score” function was used with 1000 simulations. This score assesses the trade-off between the power of discrimination and overfitting by using too many PC in the analyses (Jombart 2008). In order to evaluate the distribution of the genetic variation among the haplogroups detected from the haplotype network, an analysis of molecular variance (AMOVA) was conducted using the pairwise distances corrected with the Kimura 2-parameter (K2p), using 10,000 permutations in PEGAS v 0.11 package in R software. Mean genetic distances among the resulting genetic groups were estimated using the K2p model of nucleotide substitution in MEGA X (Kumar et al. 2018). Finally, isolation by distance was evaluated; a Mantel Test was performed using all specimens with the Euclidian geographic distance matrix pairwise and the K2p genetic distance matrix using a Pearson correlation and 10,000 permutations in the ADE4 v 1.7–13 package (Dray and Dufour 2007) in R software.
Ecological Niche Modeling and Spatial Connectivity
In order to assess the current and past environmental suitability of the P. yucatanicus, an ecological niche modeling (ENM) analysis was performed. The occurrences of P. yucatanicus were downloaded from the Global Biodiversity Information Facility (GBIF; http://www.gbif.org/), filtered by only specimens deposited in any biological collection in Mexico, the United States, and Canada. Specimens without geographic coordinates and insufficient locality information were discarded. To reduce sampling biases, the occurrences were spatially thinned at 10 km in the SPTHIN v 0.1.0.1 package (Aiello-Lammens et al. 2015) in R software. Then, the training dataset was generated with this thinned data, and the remaining occurrences were used to test the niche models (Supporting Information Table S2).
As potential predictors of niche models, 19 bioclimatic layers for current conditions were downloaded from WorldClim v 1.4 (www.worldclim.org; Hijmans et al. 2005) at a resolution of 30 s. The ENM analysis was delimited to the YP polygon downloaded from the United States Department of Agriculture (https://www.fs.fed.us/rm/ecoregions/products/map-ecoregions-north-america/#) and hypothesized as the accessible area to P. yucatanicus because of geographic restriction of this species to this area. To minimize redundancy in the bioclimatic layers, values of the 19 layers were extracted and two sets of layers were implemented according to the Pearson’s correlation threshold of 0.75 and 0.85 in the NTBOX v 0.1.4.5 package (Osorio-Olvera et al. 2016) in R software.
Niche models were built under the maximum entropy algorithm (MAXENT v 3.4.1; Phillips et al. 2017) using the KUENM v 1.1.1 package (Cobos et al. 2019) in R software. For each environmental dataset, 10,000 background points were used, and some level of niche model complexity was estimated by varying the regularization multiplier values from 0.5 to 5, every 0.5 and feature classes: linear (L), quadratic (Q), product (P), threshold (T), and hinge (H) in five combinations: L, LQ, LQP, LQPT, and LQPTH. A total of 100 candidate niche model performances was evaluated based on statistical significance of the partial Receiver Operating Characteristic (pROC; Peterson et al. 2008) with 20% of random point and 500 iterations, 5% omission rates (OR; Anderson et al. 2003) as predictive power, and the Akaike information criterion corrected for small sample sizes (AICc; Warren et al. 2010) for the model complexity. The best model was selected considering the statistically significant models, and the lowest values of AICc and OR. Finally, the niche model selected with the best parameters was built and projected in the YP using ten replicates of bootstrap and the logistic output format. Finally, for the paleoclimate reconstruction, the environmental layers from the Holocene 6000 years ago (ya; Beijing Climate Center Climate System Model version 1.1; BCC-CSM1.1) were downloaded from WorldClim v 1.4. The best model calibrated was transferred to the past Holocene climate conditions using three methods of extrapolation: free extrapolation, extrapolation and clamping, and no extrapolation.
In order to assess spatial connectivity among the localities of the genetic samples of P. yucatanicus, the electronic circuit theory algorithm was used in CIRCUITSPACE v 4.0.5 (McRae 2006). This algorithm quantifies movement across multiple possible paths in a landscape, not just a single least-cost path or corridor to predict patterns of movement, gene flow, and genetic differentiation among populations in heterogeneous landscapes (Dickson et al. 2018). The best ecological niche model of the present and the three models projected to the past were used as habitat map, wherein each cell value was assigned as conductance. The higher values of ecological suitability correspond to greater ease of movement and apply a connection scheme that allow gene flow among the four nearest cells (Ruiz-Sanchez and Specht 2014). Lastly, in order to determinate isolation by resistance in present and past (early-middle Holocene), a Mantel Test was performed with the resistance pairwise distances matrix output of Circuitscape and the K2p genetic distance matrix using a Pearson correlation and 10,000 permutations in the ADE4 v 1.7–13 package in R software.
Data Availability
All data analyzed during this study are included in this published article, and its supplementary information files, and are available from the corresponding author on reasonable request. All nucleotide sequences are available in Bold Systems DataBase (http://www.boldsystems.org/) and NCBI GenBank repository (https://www.ncbi.nlm.nih.gov/genbank) under the accession numbers included in the Supporting Information Table S1. The geographic occurrences are available in Global Biodiversity Information Facility (GBIF; http://www.gbif.org/) and the ID of each one included in the Supporting Information Table S2. The subspecies and haplogroup affinity of each genetic sample are available in the Supporting Information Table S3.
Results
The final matrix consisted of 657 nucleotide characters and 812 terminals. The nucleotide substitution saturation test revealed that the Iss value was significantly lower than the Issc in all cases (Table 1), excepting the non-significant third codon assuming an extreme asymmetrical tree, which is not the further resulting tree. Therefore, little substitution saturation is assumed in sequences and useful for phylogenetic analyses. The best scheme of partition was using the first and second codon, and other partition with the third codon (−lnL = 3694.85). The nucleotide substitution models were: TrN + I for the first and second codon, and GTR + G for the third codon.
Phylogenetic Inference
Phylogenetic ML and IB trees were similar (Fig. 1). Both analyses recovered 16 well-supported monophyletic lineages: P. mayensis, P. stirtoni, P. yucatanicus, P. melanocarpus, P. megalops, P. gymnotis, P. mexicanus, P. nudipes, P. zarhynchus, P. guatemalensis, P. grandis, and another four lineages. These four lineages were specimens previously identified as P. mexicanus, but in this study were labeled according to the findings of Pérez-Consuegra and Vázquez-Domínguez (2016; Fig. 2) as P. tropicalis, lineage “L,” P. salvadorensis, P. nicaraguae, and lineage “O.” Additionally, five specimens identified initially as P. nudipes from Nicaragua and four as P. gymnotis from Costa Rica were closely related to the P. nicaraguae. All specimen data with the previous taxonomic assignment and the posterior to phylogenetic analyses are available as Supporting Information Table S1.
Phylogenetic relationships of Peromyscus yucatanicus based on the mitochondrial cytochrome c oxidase subunit 1 gene. At the left-side Maximum likelihood (ML) topology, and right-side Bayesian Inference (BI) topology. Numbers above the branches indicate bootstrap nodal support for ML, and the posterior probability nodal support for BI, asterisks represent values of 100% and 1 respectively. Only some few terminals are presented for better visualization, the final number of sequences analyzed by species are in parenthesis and its information are available in the Supporting Information Table S1. Lineages "O" and "L" were labeled according to the findings of Pérez-Consuegra and Vázquez-Domínguez (2016)
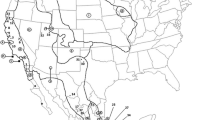
Geographic distribution in Mexico and Central America of the mitochondrial sequences downloaded from Bold Systems DataBase and used for the phylogenetic analyses. Information on localities and accession numbers are presented as Supporting Information Table S1
The monophyly among the P. mexicanus species group was supported by 87% bootstrap value and 0.93 posterior probability. The remaining species with low nodal support, such as P. megalops, P. yucatanicus, P. melanocarpus, and P. stirtoni. The clade of P. mexicanus and P. gymnotis, and the clade of P. guatemalensis, lineage “L,” and P. salvadoensis were well supported. Peromyscus mayensis was the sister group of all species in both ML and BI trees with high nodal support. Lastly, all specimens of P. yucatanicus were monophyletic (Fig. 1).
Demographic History and Genetic Differentiation of P. yucatanicus
The 183 sequences analyzed showed a haplotypic diversity of 0.9619, 108 segregate sites, and 0.0105 nucleotide diversity. The expected number of haplotypes by the chao estimator was 82. Neutrality and/or expansion test was −0.2594 (P < 0.0239) for Tajima’s D, and 0.0268 (P < 0.0186) for R2. The mismatch distribution analysis produced a slightly unimodal distribution (Fig. 3), the Bayesian Skyline Plot showed a constant equilibrium of effective population size with an increase around 5000–1000 ya, and a recent decreasing towards the present (Fig. 3).
Demographic history analyses for specimens of Peromyscus yucatanicus. Mismatch distribution (above) with the expected pattern showed by the dashed line, and the observed pattern by the continuous line. The Bayesian Skyline Plot (below) with the number of effective size (Ne) mean estimate in dashed line and 95% highest posterior density limits, over the time in millions of years are shown
The haplotype network showed 56 haplotypes organized in three main haplogroups: the first haplogroup distributed at center-south of the YP (haplogroup I), the second haplogroup at the north and center of the peninsula (haplogroup II), and the third haplogroup at the south-west of the peninsula (haplogroup III; Fig. 4). Although the three haplogroups are visually identifiable in the haplotype network, some haplotypes from the three haplogroups are shared in localities at the center of the peninsula, without geographic structure (Fig. 4). According to the geographic distribution of the subspecies of P. yucatanicus stated by Hall (1981), all 84 specimens of the haplogroup I belonged to P. y. badius. For the 59 samples of the haplogroup II were 17 of P. y. badius and 42 of P. y. yucatanicus. The 40 samples of the haplogroup III were P. y. badius (Supporting Information Table S3). The three haplogroups were supported by the DAPC analysis, the first two principal components contained 57.8% of the conserved variance, and two linear discriminant functions were used for the visualization of the samples in the multivariate space. Based on the discriminant functions, the genetic variation among the three haplogroups was clear (Fig. 5). The AMOVA for the three haplogroups revealed significance (P < 0.001) at all levels, the variation among the haplogroups was 76.28%, while within localities was 41.35% and a Fct = 0.7628 index fixation (Table 2). The mean K2p genetic distances were: 1.16% between haplogroup I and II; 1.54% between haplogroup I and III; and 1.72% between haplogoup II and III. Finally, according to the Mantel test, the correlation between the genetic and geographic distances was positive and significant (r = 0.375, P < 0.001).
Geographic distribution of Peromyscus yucatanicus samples in the Yucatan Peninsula, Mexico. The haplotype network of cytochrome oxidase subunit 1 and geographic distribution of samples of the three haplogroups are shown. Haplogroup I by diamonds, haplogroup II by circles, and haplogroup III by squares. Polygons show the geographic distribution of the subspecies recognized by Hall (1981), P. y. yucatanicus above and P. y. badius below
Discriminant Analyses of Principal Components scatter plot based on the mitochondrial cytochrome c oxidase subunit 1 gene. The first and second discriminant functions axes are shown, polygons enclose the dispersion of individual scores for each haplogroup, and the dashed line denotes the distance of each individual to the centroid. Haplogroup I by diamonds, haplogroup II by circles, and haplogroup III by triangles
Ecological Niche Modeling and Spatial Conectivity
A total of 1005 specimen geographical records were registered in biological collections with 123 different localities, and after spatial filtering, 79 occurrences were used to build the niche model, and the remaining 44 occurrences for testing the models (Supporting Information Table S2). The two environmental datasets evaluated were: 1) bio1 (annual mean temperature), bio2 (mean diurnal range), bio3 (isothermality), bio4 (temperature seasonality), bio9 (mean temperature of driest quarter), bio12 (annual precipitation), bio14 (precipitation of driest month), and bio18 (precipitation of warmest quarter), according to the 0.75 Pearson’s correlation threshold; and dataset 2) bio1, bio2, bio3, bio4, bio5 (max temperature of warmest month), bio6 (min temperature of coldest month), bio8 (mean temperature of wettest quarter), bio9, bio12, bio14, bio15 (precipitation seasonality), bio16 (precipitation of wettest quarter), bio18, and bio19 (precipitation of coldest quarter), following the 0.85 Pearson’s correlation threshold.
According to the evaluation of the 100 candidate models, three niche models with the set two of environmental layers were the most appropriate as stated in the criteria selected (Table 3). The best model with the lowest OR and AICc was selected to project it. The ecological niche of P. yucatanicus was mainly influenced by the following variable layers and percentage of contribution: bio12 (62.5%), bio 19 (16.6%), and bio 3 (6.4%). This niche model at the present showed that the highest environmental suitability was heterogeneous throughout the YP, higher suitability values mainly at northern and coastal areas (Fig. 6). On the other hand, the past projections (6000 ya) were similar without strong differences; the no extrapolation model showed cero values of suitability and was not shown, whereas the free extrapolation (Fig. 6a) and extrapolation and clamping (Fig. 6b) models showed low values of suitability.
Geographic projection of the best ecological niche model, the current niche suitability is shown by scale colors. The crosses indicate the training occurrences, and the genetic groups: Haplogroup I by diamonds, haplogroup II by circles, and haplogroup III by squares. In the right-side, the ecological niche model transferred to the early-middle Holocene (6000 years ago) using free extrapolation (a), and extrapolation and clamping (b)
The climatic spatial resistance surfaces of the present model showed high spatial connectivity among near localities, mainly at the center and northeast of the YP, while localities without near neighbors were isolated. On the other hand, during the early-middle Holocene the free extrapolation, and extrapolation and clamping models showed a lack of spatial connectivity, except for localities immediately adjacent (Fig. 7). The isolation by resistance test showed no significant correlation between genetic distances and resistance distances in the present (r = 0.126, P = 0.169) and the past using the raster input of free extrapolation (r = −0.145, P = 0.881), and extrapolation and clamping (r = −0.136, P = 0.186).
Geographic projection of the current climatic spatial resistance surfaces, blue to red colors indicates low to high density current surface values, respectively. The genetic groups are depicted: haplogroup I by diamonds, haplogroup II by circles, and haplogroup by squares. In the right-side, the climatic spatial resistance surfaces of the early-middle Holocene (6000 years ago) using as input the ecological niche models of free extrapolation (a), and extrapolation and clamping (b)
Discussion
DNA barcoding in mammals has been useful for species identification, detection of new lineages, and taxonomy, contributing new information on biology, distribution, and conservation (Galimberti et al. 2015). The use of CO1 has been able to discriminate cryptic species in rodents (Álvarez-Castañeda et al. 2012; Galan et al. 2012; Müller et al. 2013; Li et al. 2015; Pinto et al. 2018; Da Cruz et al. 2019; Ramatla et al. 2019). In the present study, the CO1 marker was useful to recover monophyletic species and lineages, but poor phylogenetic signal to infer well supported relationships. Therefore, the present genetic study with P. yuctanicus has important findings; hypotheses and their limitations will be discussed.
Phylogenetic Inference
The CO1 marker was useful for detecting monophyletic species and lineages of P. mexicanus group, which were concordant with the lineages using the mitochondrial gene Cytb (Ordóñez-Garza et al. 2010; Bradley et al. 2016; Pérez-Consuegra and Vázquez-Domínguez 2016), but only some phylogenetic relationships were well supported, for example, the clade of P. mexicanus and P. gymnotis, and the clade of P. guatemalensis, lineage “L,” and P. salvadoensis (Fig. 1). In general, CO1 was useful for detecting several misidentified specimens mainly determined previously as P. mexicanus, which belonged to others recently detected lineages (Pérez-Consuegra and Vázquez-Domínguez 2016), but they must be confirmed by morphological revisions. The phylogenetic relationships of P. yucatanicus cannot be solved using only CO1; therefore, more molecular characters must be included, such as cytochrome b and other nuclear markers in order to increase the phylogenetic signal, and make it comparable to other studies. However, at species level the CO1 marker shows sufficient genetic variation for genetic identification of the monophyletic P. yucatanicus, which is strictly from lowland habitats (Young and Jones 1983). Other lowland species, such as P. stirtoni, were included in this study, but it is necessary to add more closely related lowland species, such as P. melanophrys, P. mekisturus, and P. perfulvus, known as the P. melanophrys species group, which were monophyletic with the Cytb mitochondrial marker (Bradley et al. 2007; Castañeda-Rico et al. 2014). These species are closely related to P. stirtoni, P. mayensis, P. megalops, P. melanocarpus, and P. mexicanus species group (Bradley et al. 2007). However, CO1 sequences were not available for including in this exploratory phylogenetic analysis; therefore, the inclusion of these species is mandatory in posterior analyses in order to clarify the phylogenetic relationships with P. yucatanicus, and for testing the species group boundary among P. mexicanus, P. megalops, and P. melanophrys groups. For example, in Ordóñez-Garza et al. (2010) and Bradley et al. (2016), P. melanophrys and P. perfulvus were included using the mitochondrial gene Cytb, but not samples of P. yucatanicus.
Demographic History, Genetic Differentiation, Ecological Niche, and Spatial Connectivity of P. yucatanicus
All demographic analyses were performed using all samples, rather than the genetic groups found, in order to avoid the violation of panmixia assumption (Grant 2015). The high haplotypic and nucleotide diversity suggested demographic equilibrium, but significant negative Tajima’s D value and significant low R2 value demonstrated a tendency to demographic growth. On the other hand, these results combined with the slightly unimodal performance of the mismatch distribution analysis, and the relatively stable curve of the Skyline analysis, suggested signatures of a likely scenario of demographic stability followed by slight population growth and decreasing towards the present. This scenario may be supported with the paleontological information, which suggested that the general climate did not changed drastically after the late Pleistocene to the present (Arroyo-Cabrales and Alvarez 2003). Nevertheless, recent paleoclimatic information showed that during the Holocene, the YP presented oscillations in precipitation, increasing gradually from the early-middle Holocene (subdivisions according to Walker et al. 2012) with a remarkable drop in the late-middle Holocene around 4200 ya, and an increase of precipitation around 2500 ya, which was higher than the present. These changes of precipitation modified the forest distribution, suggesting closely covered vegetation during the highest precipitation and posteriorly a dramatic reduction during the Late Preclassic Period (Hengstum et al. 2010; Carrillo-Bastos et al. 2012; Roy et al. 2017; Vela-Pelaez et al. 2018).
These hypotheses of major covered vegetation and high precipitation 2500 ya matched with the time estimates of slightly demographic growth of the skyline plot and the slight decrease in the actuality (Fig. 3). The paleoclimate model presented in this study in the early-middle Holocene (6000 ya) provided evidence to support this hypothesis of climate changes in the past for P. yucatanicus. Some studies with extreme climate conditions as the Last Glacial Period (LGP; 22,000 ya) showed that to compare the different sets of environmental layers available and methods of extrapolations is a good practice in order to avoid dramatic differences in the paleodistributions and avoid biologically unrealistic model estimations (Guevara et al. 2018; Guevara and León-Paniagua 2019). The paleoclimate models constructed for P. yucatanicus were similar without strong differences, and may be useful for inferences. These paleoclimate models suggested that the niche ecological suitability had lower suitability values than the present, suggesting that P. yucatanicus perhaps invaded the YP more recently with the major precipitation and covered vegetation. These models support the hypothesis of major environmental suitability after the early-middle Holocene. The climatic spatial resistance surfaces for the present showed spatial connectivity among nearest localities, and for the past (6000 ya) showed no connectivity among localities; all isolation by resistance tests were not significant indicating that there is no a clear association between genetic distances and resistance distances. However, it is likely that this result was related with the limited sample design, and it is necessary to include specimens of the southern and western portion of the YP where no localities were tested for spatial connectivity, and probably high spatial connectivity is present in that areas.
Studies suggested that deeper climate change oscillations of the LGP in the YP gradually increased the temperature, resulting in high rates of ecological change during the late Pleistocene to Holocene (Leyden et al. 1993; Correa-Metrio et al. 2012; Hodell et al. 2012). The present vegetation in the YP was absent in the Pleistocene, but it was during the Pleistocene—early-middle Holocene transition when the current flora invaded the south of YP in response to warmer and wetter conditions (Brenner et al. 2002; Cruz et al. 2016). This scenario is concordant with the hypothesis of divergence of P. yucatanicus during the Pleistocene (Lawlor 1965). This idea is supported by a phylogeographic study of the lowland rodent Ototylomys phyllotis by Gutierrez-García and Vázquez-Domínguez (2012), which stated that this rodent colonized the YP 800,000 ya with only individuals at southern Belize, and likely the faunas of northern YP originated by dispersion from the south. A recent phylogeographic study with the cycad Zamia prasina in the YP (Montalvo-Fernández et al. 2019) hypothesized that this species could colonize the YP from the southeast or from Campeche during the late Pleistocene, and the populations expanded during the warmest periods of the middle Holocene. This hypothesis is concordant with the evolutionary history of P. yucatanicus inferred herein.
Although scarce small mammal fossil records are found in the YP, P. yucatanicus fossils have been found during the Pleistocene—early-middle Holocene, but absent in the Pleistocene substrates at the northern YP (Arroyo-Cabrales and Alvarez 2003). More fossil evidence is necessary to confirm the evolutionary history of P. yucatanicus, but information presented herein is useful to hypothesize that P. yucatanicus likely diverged in the Pleistocene and invaded the south of YP after the Last Glacial Period with the arrival of the current vegetation in the late Pleistocene—early-middle Holocene, and its demographic population was stable during the remaining Holocene epoch with slight growth at the late-middle Holocene resulted in the major precipitation changes that provided more plat coverage.
The haplotype network, DAPC, and AMOVA suggested that three genetic haplogroups can be identified in P. yucatanicus, but there is no clear geographic pattern among these genetic groups at the center of the YP. Most of the haplotypes of each genetic group are not shared, mainly these of greatest geographic distance, which are concordant with the significant isolation by distance results. These three genetic groups are not concordant with the geographic distribution of the two subspecies recognized by Hall (1981; Fig. 4). Only all samples of the haplogroups I and III were P. y. badius, but samples of haplogroup II were of P. y. yucatanicus and P. y. badius at the center of the YP. These two subspecies are recognized mainly by pelage coloration, the “pale” northern P. y. yucatanicus and the “dark” southern P. y. badius (Hall 1981; Young and Jones 1983). However, some studies in mammals suggested that pelage coloration can vary in response to different causes, for instance, specific environmental conditions (Stoner et al. 2003; Hoekstra et al. 2005; Sandoval-Salinas et al. 2017). This environmental condition likely can be related to the precipitation gradient in the YP, being more humid in the south and dryer in the north, determining differences in type of vegetation (Carnevali et al. 2010). On the other hand, Lawlor (1965) did not recognize subspecies, mainly because: the area of intergradation would be larger than the area occupied by any one subspecies; there is clinal discordance of size and color; there is local reversal of clinal variation; and no other significant variation is apparent. In addition, the genetic distances among the haplogroups were low (< 1.8%), which is less than at the level of species based on the genetic species concept (Baker and Bradley 2006). In general, the genetic data suggest that there is not phylogeographic structure and no clear separation of subspecies, therefore supporting the idea of Lawlor (1965) who did not recognize subspecies based only on pelage coloration. However, more samples must be included, and this hypothesis tested with morphologic analyses, and genetic variation with other molecular markers.
There is not a clear relationship among each haplogroup with any kind of vegetation, microclimate, geological, or environmental features in the YP, and consequently their existence can be related to other biotic characteristics. For instance, the genetic structure found in Z. prasina was influenced by precipitation gradient in the YP (Montalvo-Fernández et al. 2019); therefore, analyzing these differences in precipitation, specific vegetation type, and microenvironmental conditions could offer new insights about spatial connectivity and specific requirements in different populations within the YP. The use of microsatellites would be appropriate to confirm more accurately the recent landscape features that may cause the genetic structure observed with sequences of CO1. Although few ecological studies have been performed for P. yucatanicus, a population ecology study performed at the north of the YP showed small and stable populations with a high rate of recaptures and low rate of young immigrants (MacSwiney et al. 2012). If this kind of behavior is maintained throughout the YP, then the haplogroups detected in this study could be explained by the low immigration to other populations, which is supported by the significant isolation by distance model, in which the localities more distanced not shared haplotypes, and the localities at the YP center shared haplotypes present from closer localities of each haplogroup (Fig. 4), wherein there are more spatial connection (Fig. 7).
The geographic distribution of the three genetic groups in P. yucatanicus is similar to the biogeographical patterns of faunistic groups that recognized three major areas in the continental YP: north, center, and south, but with different patterns for each biological group (Cortés-Ramírez et al. 2012). For example, in other rodent Oryzomys couesi were found genetic structuration in the Quintana Roo populations from the Yucatán and Chiapas populations (Vázquez-Domínguez et al. 2009). There is strong evidence of genetic differentiation in other biological groups in the YP, such as: the fish Mayaheros urophthalmus with genetic structure between the north and south of the YP (Barrientos-Villalobos and Schmitter-Soto 2019), four genetic groups of Triatoma dimidiata found at southern of YP (Monteiro et al. 2013), different haplotypic composition at center-north of the YP in Artibeus jamaicensis (Ruiz et al. 2013), the north and south distinct haplogroups of Sceloporus serrifer serrifer (Martínez-Méndez et al. 2015), low genetic differentiation, but distinctive haplogroups of the three subspecies of Colinus nigrogularis at north, center, and south of the YP (Williford et al. 2015), genetic differentiation, and isolation by distance in close populations of Ruellia nudiflora at northern of the YP (Vargas-Mendoza et al. 2015), and the two genetic groups of Hydropuntia cornea/Hydropuntia usneoides complex in the north and south coastal areas of the YP (Núñez-Resendiz et al. 2016).
Another kind of evidence with ecological niche modeling has demonstrated that there are similar heterogeneous patterns of suitability as in P. yucatanicus in species of birds in the YP, such as Campylorhynchus yucatanicus, Pyrilia haematotis, Pionus senilis, Amazona farinosa, Amazona autumnalis, and Amazona xantholora (Plasencia-Vázquez and Escalona-Segura 2014; Plasencia-Vázquez et al. 2014; Serrano-Rodríguez et al. 2017). These studies, as that presented herein, manifest that the evolutionary history of several groups of the YP is complex and closely related to the effects of the climatic changes and oscillations during the Pleistocene and Holocene, but mainly highlights the need to study the evolutionary history of several groups distributed in the YP in order to discover common patterns that help to improve our understanding about this region.
References
Aiello-Lammens ME, Boria RA, Radosavljevic A, Vilela B, Anderson RP (2015) spThin: an R package for spatial thinning of species occurrence records for use in ecological niche models. Ecography 38:541–545. https://doi.org/10.1111/ecog.01132
Álvarez-Castañeda ST, Lorenzo C, Ríos E, Cortés-Calva P, Elías M, Ortega J, Cervantes FA (2012) DNA barcoding of mammals in Mexico: implications for biodiversity. Open Zool J 5:18–26. https://doi.org/10.2174/1874336601205010018
Anderson RP, Lew D, Peterson AT (2003) Evaluating predictive models of species’ distributions: criteria for selecting optimal models. Ecol Model 162:211–232. https://doi.org/10.1016/S0304-3800(02)00349-6
Arroyo-Cabrales J, Alvarez T (2003) A preliminary report of the late Quaternary mammal fauna from Loltún cave, Yucatán, México. In: Shubert BW, Mead JI, Graham RWm (eds) Ice Age Cave Faunas of North America. Indiana University Press and Denver Museum of Nature and Science, Bloomington and Indianapolis
Baker RJ, Bradley RD (2006) Speciation in mammals and the genetic species concept. J Mammal 87(4):643–662. https://doi.org/10.1644/06-MAMM-F-038R2.1
Barrientos-Villalobos J, Schmitter-Soto JJ (2019) Phylogeography of the Mayan cichlid Mayaheros urophthalmus (Teleostei: Cichlidae) in the Yucatan peninsula based on mitochondrial markers CYTB and COI. Environ Biol Fish 102(12):1461–1472. https://doi.org/10.1007/s10641-019-00920-4
Bodenhofer U, Bonatesta E, Horejs-Kainrath C, Hochreiter S (2015) msa: an R package for multiple sequence alignment. Bioinformatics 31:3997–3999. https://doi.org/10.1093/bioinformatics/btv494
Bouckaert R, Vaughan TG, Barido-Sottani J, Duchêne S, Fourment M, Gavryushkina A, Heled J, Jones G, Kühnert D, Maio ND, Matschiner M, Mendes FK, Müller NF, Ogilvie HA, Plessis L, Popinga A, Rambaut A, Rasmussen D, Siveroni I, Suchard MA, Wu CH, Xie D, Zhang C, Stadler T, Drummond AJ (2019) BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLoS Comput Biol 4(15):e1006650. https://doi.org/10.1371/journal.pcbi.1006650
Bradley RD, Durish ND, Rogers DS, Miller JR, Engstrom MD, Kilpatrick CW (2007) Toward a molecular phylogeny for Peromyscus: evidence from mitochondrial cytochrome b sequences. J Mammal 88:1146–1159. https://doi.org/10.1644/06-MAMM-A-342R.1
Bradley RD, Nuñez-Tabares M, Soniat TJ, Kerr S, Raymond RW, Ordóñez-Garza N (2016) Molecular systematics and phylogeography of Peromyscus nudipes (Cricetidae: Neotominae). In: Manning RW, Goetze JR, Yancey II FD (eds) Contributions in Natural History: A Memorial Volume in Honor of Clyde Jones. Spec Publ Mus Tex Tech 65:201–213
Brenner M, Rosenmeier M, Hodell D, Curtis J (2002) Paleoclima de la región maya: síntesis del conocimiento basado en registros paleolimnológicos. In: Islebe G (ed) Los Investigadores de la Cultura Maya, Tomo 1. Universidad Autónoma de Campeche, Campeche
Brown SDJ, Collins RA, Boyer S, Lefort MC, Malumbres-Olarte J, Vink CJ, Cruickshank RH (2012) SPIDER: an R package for the analysis of species identity and evolution, with particular reference to DNA barcoding. Mol Ecol Resour 12:562–565. https://doi.org/10.1111/j.1755-0998.2011.03108.x
Carleton MD (1989) Systematics and evolution. In: Kirkland GL, Layne JN (eds) Advances in the Study of Peromyscus (Rodentia). Texas Tech University Press, Lubbock
Carnevali FCG, Tapia-Muñoz JL, Duno de Stefano R, Ramírez-Morillo I (2010) Flora ilustrada de la Península de Yucatán: listado florístico. Centro de Investigación Científica de Yucatán, Yucatán
Carrillo-Bastos A, Islebe GA, Torrescano-Valle N (2012) Geospatial analysis of pollen records from the Yucatán peninsula, Mexico. Veg Hist Archaeobot 21:429–437. https://doi.org/10.1007/s00334-012-0355-1
Castañeda-Rico S, León-Paniagua L, Vázquez-Domínguez E, Navarro-Sigüenza AG (2014) Evolutionary diversification and speciation in rodents of the Mexican lowlands: the Peromyscus melanophrys species group. Mol Phylogenet Evol 70:454–463. https://doi.org/10.1016/j.ympev.2013.10.004
Cobos ME, Peterson AT, Barve N, Osorio-Olvera L (2019) kuenm: an R package for detailed development of ecological niche models using Maxent. PeerJ 7:e6281. https://doi.org/10.7717/peerj.6281
Correa-Metrio A, Bush MB, Cabrera KR, Sully S, Brenner M, Hodell DA, Ecobar J, Guilderson T (2012) Rapid climate change and no-analog vegetation in lowland Central America during the last 86,000 years. Quat Sci Rev 38:63–75. https://doi.org/10.1016/j.quascirev.2012.01.025
Cortés-Ramírez G, Gordillo-Martínez A, Navarro-Sigüenza AG (2012) Patrones biogeográficos de las aves de la Península de Yucatán. Rev Mex Biodivers 83:530–542.
Cruz JA, Arroyo-Cabrales JA, Reynoso VH (2016) Reconstructing the paleoenvironment of Loltún Cave, Yucatán, Mexico, with Pleistocene amphibians and reptiles and their paleobiogeographic implications. Rev Mex Cienc Geol 33:342–354.
Da Cruz MOR, Weksler M, Bonvicino CR, Bezerra AMR, Prosdocimi F, Furtado C, Geise L, Catzeflis F, Thoisy B, de Oliveira LFB, Silva C, de Oliveira JA (2019) DNA barcoding of the rodent genus Oligoryzomys (Cricetidae: Sigmodontinae): mitogenomic-anchored database and identification of nuclear mitochondrial translocations (Numts). Mitochondr DNA Part A 30(5):702–712. https://doi.org/10.1080/24701394.2019.1622692
Dickson BG, Albano MC, Anantharaman R, Beier P, Fargione J, Graves TA, Gray ME, Hall KR, Lawler JJ, Leonard PB, Littlefield CE, McClure ML, Novembre J, Schloss CA, Schumaker NH, Shah VB, Theobald DM (2018) Circuit-theory applications to connectivity science and conservation. Conserv Biol 0:1–11. https://doi.org/10.1111/cobi.13230
Dray S, Dufour AB (2007) The ade4 Package: implementing the duality diagram for ecologists. J Stat Softw 22:1–20. https://doi.org/10.18637/jss.v022.i04
Drummond AJ, Rambaut A, Shapiro B, Pybus OG (2005) Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol 22:1185–1192. https://doi.org/10.1093/molbev/msi103
Duno-de Stefano R, Can-Itza LL, Rivera-Ruiz A, Calvo-Irabién LM (2012) Regionalización y relaciones biogeográficas de la Península de Yucatán con base en los patrones de distribución de la familia Leguminosae. Rev Mex Biodivers 83:1053–1072. https://doi.org/10.7550/rmb.27824
Durán-García R, Méndez-González M, Larqué-Saavedra A (2016) The biodiversity of the Yucatan Peninsula: a natural laboratory. In: Cánovas F, Lüttge U, Matyssek R (eds) Progress in Botany. Springer, Cham. https://doi.org/10.1007/124_2016_8
Folan WJ, Gunn-Eaton J, Patch YR (1983) Paleoclimatological patterning in southern Mesoamerica. J Field Archaeol 10:453–468. https://doi.org/10.1179/009346983791504219
Galan M, Pagès M, Cosson JF (2012) Next-generation sequencing for rodent barcoding: species identification from fresh, degraded and environmental samples. PLoS ONE 7(11):e48374. https://doi.org/10.1371/journal.pone.0048374
Galimberti A, Sandionigi A, Bruno A, Bellati A, Casiraghi M (2015) DNA barcoding in mammals: what’s new and where next? Hystrix 26(1):13–24. https://doi.org/10.4404/hystrix-26.1-11347
Grant WS (2015) Problems and cautions with sequence Mismatch Analysis and Bayesian Skyline Plots to infer historical demography. Heredity 106:333–346. https://doi.org/10.1093/jhered/esv020
Guevara L, León-Paniagua L (2019) How to survive a glaciation: the challenge of estimating biologically realistic potential distributions under freezing conditions. Ecography 42:1–9. https://doi.org/10.1111/ecog.04202
Guevara L, León-Paniagua L, Rios J, Anderson RP (2018) Variation among Global Circulation Models for reconstructions of geographic distributions at the Last Glacial Maximum: relevance for phylogeography. Ecosistemas 27:62–76. https://doi.org/10.7818/ECOS.1443
Gutierrez-García TA, Vázquez-Domínguez E (2012) Biogeographically dynamic genetic structure bridging two continents in the monotypic Central American rodent Ototylomys phyllotis. Biol J Linnean Soc 107(3):593–610. https://doi.org/10.1111/j.1095-8312.2012.01966.x
Hall ER (1981) The North American Mammals, second edition. John Wiley and Sons, New York
Hengstum PJ, Reinhardt EG, Beddows PA, Gabriel JJ (2010) Linkages between Holocene paleoclimate and paleohydrogeology preserved in a Yucatan underwater cave. Quat Sci Rev 29:19–20. https://doi.org/10.1016/j.quascirev.2010.06.034
Hernández-Betancourt S, Sélem-Salas C, Cimé-Pool A, Chablé-Santos J (2010) Diversidad de pequeños roedores de la Península de Yucatán. Bioagrociencias 8:28–31
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25(15):1965–1978. https://doi.org/10.1002/joc.1276
Hodell DA, Turchyn AV, Wiseman CJ, Escobar J, Curtis JH, Brenner M, Gilli A, Mueller AD, Anselmetti F, Ariztegui D, Brown ET (2012) Late glacial temperature and precipitation changes in the lowland Neotropics by tandem measurements of δ18O in biogenic carbonate and gypsum hydration water. Geochim Cosmochim Acta 77:352–368. https://doi.org/10.1016/j.gca.2011.11.026
Hoekstra HE, Krenz JG, Nachman MW (2005) Local adaptation in the rock pocket mouse (Chaetodipus intermedius): natural selection and phylogenetic history of populations. Heredity 94:217–228. https://doi.org/10.1038/sj.hdy.6800600
Huckaby DG (1980) Species limits in the Peromyscus mexicanus group (Mammalia: Rodentia: Muroidea). Contrib Sci Nat Hist Mus Los Angeles County 326:1–24
Islebe GA, Calmé S, León-Cortés JL, Schmook B (2015) Biodiversity and Conservation of the Yucatán Peninsula. Springer, Cham
Jombart T (2008) adegenet: an R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics 11:94. https://doi.org/10.1186/1471-2156-11-94
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lanfear R, Calcott B, Ho SYW, Guindon S (2012). PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701. https://doi.org/10.1093/molbev/mss020
Lawlor TE (1965) The deer mouse, Peromyscus yucatanicus. Univ Kans Publ Mus Nat Hist 16:421–438
Leigh JW, Bryant D (2015) PopART: full-feature software for haplotype network construction. Methods Ecol Evol 6(9):1110–1116. https://doi.org/10.1111/2041-210X.12410
Leyden BW, Brenner M, Hodell DA, Curtis JH (1993) Late Pleistocene climate in the Central American lowlands. Am Geophys Union Geophys Monogr 78:165–178. https://doi.org/10.1029/GM078p0165
Li J, Zheng X, Cai Y, Zhang X, Yang M, Yue B, Li J (2015) DNA barcoding of Murinae (Rodentia: Muridae) and Arvicolinae (Rodentia: Cricetidae) distributed in China. Mol Ecol Res 15(1):153–167. https://doi.org/10.1111/1755-0998.12279
MacFadden BJ (2006) Extinct mammalian biodiversity of the ancient New World tropics. Trends Ecol Evol 21:157–165. https://doi.org/10.1016/j.tree.2005.12.003
MacSwiney MC, Hernández-Betancourt SF, Panti-May JA, Pech-Canché JM (2012) Ecología poblacional del ratón yucateco Peromyscus yucatanicus (Rodentia: Cricetidae) en las selvas de Quintana Roo, México. In: Cervantes FA, Ballesteros-Barrera C (eds) Estudios sobre la biología de roedores silvestres mexicanos. Universidad Nacional Autónoma de Mexico and Universidad Autónoma Metropolitana, Mexico City
Martínez-Méndez N, Mejía O, Méndez de la Cruz FR (2015) The past, present and future of a lizard: the phylogeography and extinction risk of Sceloporus serrifer (Squamata: Phrynosomatidae) under a global warming scenario. Zool Anz 254:86–98. https://doi.org/10.1016/j.jcz.2014.12.004
McRae BH (2006) Isolation by resistance. Evolution 60:1551–1561. https://doi.org/10.1111/j.0014-3820.2006.tb00500.x
Miller JR, Engstrom MD (2008) The relationships of major lineages within peromyscine rodents: a molecular phylogenetic hypothesis and systematic reappraisal. J Mammal 89:1279–1295. https://doi.org/10.1644/07-MAMM-A-195.1
Montalvo-Fernández G, Sánchez-Teyer LF, Carnevali G, Vovides AP, Gaytán-Legaria R, Ortiz-García MM, Muñoz-López JA, Martínez-Castillo J (2019) Impact of late Pleistocene-Holocene climatic fluctuations on the phylogeographic structure and historical demography of Zamia prasina (Cycadales: Zamiaceae). Bot Sci 97(4):588–608. https://doi.org/10.17129/botsci.2195
Monteiro FA, Peretolchina T, Lazoski C, Harris K, Dotson EM, Abad-Franch F, Tamayo E, Pennington PM, Monroy C, Condon-Rosales C, Salazar-Schettino PM, Gómez-Palacio A, Grijalva MJ, Beard CB, Marcet PL (2013) Phylogeographic pattern and extensive mitochondrial DNA divergence disclose a species complex within the Chagas disease vector Triatoma dimidiata. PLoS ONE 8(8):e70974. https://doi.org/10.1371/journal.pone.0070974
Morrone JJ (2005) Hacia una síntesis biogeográfica de México. Rev Mex Biodivers 76:207–252
Müller L, Gonçalves G L, Cordeiro-Estrela P, Marinho JR, Althoff SL, Testoni AF, González EM, Freitas TRO (2013) DNA barcoding of sigmodontine rodents: identifying wildlife reservoirs of zoonoses. PLoS ONE 8(11):e80282. https://doi.org/10.1371/journal.pone.0080282
Núñez-Resendiz ML, Zuccarello GC, Dreckman KM, Sentíes A (2016) Phylogeography of Hydropuntia cornea/Hydropuntia usneoides complex (Gracilariales, Rhodophyta) in the Yucatan Peninsula. Phycologia 56:14–20. https://doi.org/10.2216/16-46.1
Ordóñez-Garza N, Matson JO, Strauss RE, Bradley RD, Salazar-Bravo J (2010) Patterns of phenotypic and genetic variation in three species of endemic Mesoamerican Peromyscus (Rodentia: Cricetidae). J Mammal 91:848–859. https://doi.org/10.1644/09-MAMM-A-167.1
Osorio-Olvera L, Barve V, Barve N, Soberón J (2016) Nichetoolbox: from getting biodiversity data to evaluating species distribution models in a friendly GUI environment. R package version 0.2.0.0. https://github.com/luismurao/nichetoolbox/
Paradis E (2010) pegas: an R package for population genetics with an integrated-modular approach. Bioinformatics 26:419–420. https://doi.org/10.1093/bioinformatics/btp696
Pérez-Consuegra SG, Vázquez-Domínguez E (2015) Mitochondrial diversification of the Peromyscus mexicanus group in Mesoamerica: taxonomical and biogeographical implications. J Zool Syst Evol Res 53:300–311. https://doi.org/10.1111/jzs.12155
Pérez-Consuegra SG, Vázquez-Domínguez E (2016) Intricate evolutionary histories in montane species: a phylogenetic window into craniodental discrimination in the Peromyscus mexicanus species group (Mammalia: Rodentia: Cricetidae). J Zool Syst Evol Res 55:57–72. https://doi.org/10.1111/jzs.12155
Peterson AT, Papeş M, Soberón J (2008) Rethinking receiver operating characteristic analysis applications in ecological niche modeling. Ecol Model 213:63–72. https://doi.org/10.1016/j.ecolmodel.2007.11.008
Phillips SJ, Anderson RP, Dudík M, Schapire RE, Blair ME (2017) Opening the black box: an open-source release of Maxent. Ecography 40:887–893. https://doi.org/10.1111/ecog.03049
Pinto CM, Ojala-Barbour R, Brito J, Menchaca A, Carvhalo ARG, Weksler M, Amato G, Lee TE Jr (2018) Rodents of the eastern and western slopes of the Tropical Andes: phylogenetic and taxonomic insights using DNA barcodes. Therya 9(1):15–27. https://doi.org/10.12933/therya-18-430
Plasencia-Vázquez AH, Escalona-Segura G (2014) Caracterización del área de distribución geográfica potencial de las especies de aves psitácidas de la Península de Yucatán, México. Rev Biol Trop 62:1509–1522
Plasencia-Vázquez AH, Escalona-Segura G, Esparza-Olguín LG (2014) Modelación de la distribución geográfica potencial de dos especies de psitácidos neotropicales utilizando variables climáticas y topográficas. Acta Zool Mex 30:471–490
Platt RN II, Amman BR, Keith MS, Thompson CW, Bradley RD (2015) What is Peromyscus? evidence from nuclear and mitochondrial DNA sequences suggests the need for a new classification. J Mammal 96:708–719. https://doi.org/10.1093/jmammal/gyv067
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarisation in Bayesian phylogenetics using Tracer 1.7. Syst Biol syy032. https://doi.org/10.1093/sysbio/syy032
Ramatla T, Mphuthi N, Gofaone K, Taioe MO, Thekisoe OMM, Syakalima M (2019) Identification of rodent species that infest poultry houses in mafikeng, north west province, South Africa. Int J Zool 1:1–8. https://doi.org/10.1155/2019/1280578
Ramos-Onsins SE, Rozas J (2002) Statistical properties of new neutrality tests against population growth. Mol Biol Evol 19:2092–2100 https://doi.org/10.1093/oxfordjournals.molbev.a004034.
Ratnasingham S, Hebert PDN (2007) BOLD: the barcode of life data system (www.barcodinglife.org). Mol Ecol Notes 7:355–364. https://doi.org/10.1111/j.1471-8286.2006.01678.x
Rogers DS, Ensgstrom MD (1992) Evolutionary implications of allozymic variation in tropical Peromyscus of the mexicanus species group. J Mammal 73:55–69. https://doi.org/10.2307/1381866
Ronquist F, Teslenko M, Van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012). MRBAYES 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Roy PD, Torrescano-Valle N, Islebe GA, Gutiérrez-Ayala LV (2017) Late Holocene hydroclimate of the western Yucatan Peninsula (Mexico). J Quat Sci 32:1112–1120. https://doi.org/10.1002/jqs.2988
Ruiz EA, Vargas-Miranda B, Zúñiga G (2013) Late-Pleistocene phylogeography and demographic history of two evolutionary lineages of Artibeus jamaicensis (Chiroptera: Phyllostomidae) in Mexico. Acta Chiropterol 15(1):19–33. https://doi.org/10.3161/150811013X667830
Ruiz-Sanchez E, Specht CD (2014) Ecological speciation in Nolina parviflora (Asparagaceae): lacking spatial connectivity along of the trans-Mexican volcanic belt. PLoS ONE 9(6):e98754. https://doi.org/10.1371/journal.pone.0098754
Sandoval-Salinas ML, Barquez RM, Colombo EM, Sandoval JD (2017) Intra-specific pelage color variation in a South American small rodent species. Braz J Biol 77:1–11. https://doi.org/10.1590/1519-6984.00615
Serrano-Rodríguez A, Escalona-Segura G, Plasencia Vázquez AH, Iñigo-Elias EE, Ruiz-Montoya L (2017) Distribución potencial y conectividad del paisaje: criterios para reevaluar el grado de amenaza de Campylorhynchus yucatanicus (Aves: Troglodytidae). Rev Biol Trop 65:1554–1568. https://doi.org/10.15517/rbt.v65i4.26599
Stamatakis A (2014) Raxml version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stoner CJ, Bininda-Emonds ORP, Caro T (2003) The adaptive significance of coloration in lagomorphs. Biol J Linnean Soc 79:309–328. https://doi.org/10.1046/j.1095-8312.2003.00190.x.
Sullivan KAM, Platt RN II, Bradley RD, Ray DA (2017) Whole mitochondrial genomes provide increased resolution and indicate paraphyly in deer mice. BMC Zool 2:11. https://doi.org/10.1186/s40850-017-0020-3
Tajima F (1989) Statistical-method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Vargas-Mendoza CF, Ortegón-Campos I, Marrufo-Zapata D, Herrera CM, Parra-Tabla V (2015) Genetic diversity, outcrossing rate, and demographic history along a climatic gradient in the ruderal plant Ruellia nudiflora (Acanthaceae). Rev Mex Biodivers 86:508–520. https://doi.org/10.1016/j.rmb.2015.04.034
Vázquez-Domínguez E, Arita HT (2010) The Yucatan Peninsula: biogeographical history 65 million years in the making. Ecography 33:212–219. https://doi.org/10.1111/j.1600-0587.2009.06293.x
Vázquez-Domínguez E, Mejía-Puente A, Vega R (2009). Oryzomys couesi en el sureste de México: estimaciones genéticas y filogeográficas. In: Cervantes FA, Hortelano-Moncada Y, Vargas-Cuenca J (eds) 60 años de la colección nacional de mamíferos del Instituto de Biología, UNAM: aportaciones al conocimiento y conservación de los mamíferos mexicanos. Universidad Nacional Autónoma de México, Instituto de Biología, Mexico, pp 197–211
Vela-Pelaez AA, Torrescano-Valle N, Islebe GA, Mas JF, Weissenberger H (2018) Holocene precipitation changes in the Maya forest, Yucatán Peninsula, Mexico. Palaeogeogr Palaeoclimatol Palaeoecol 505:42–52. https://doi.org/10.1016/j.palaeo.2018.05.024
Walker MJC, Berkelhammer M, Björck S, Cwynar LC, Fisher DA, Long AJ, Lowe JJ, Newnham RM, Rasmussen SO, Weiss H (2012) Formal subdivision of the Holocene Series/Epoch: a discussion paper by a working group of INTIMATE (Integration of ice-core, marine and terrestrial records) and the subcommission on Quaternary Stratigraphy (International Commission on Stratigraphy). J Quat Sci 27:649–659. https://doi.org/10.1002/jqs.2565
Warren DL, Geneva AJ, Lanfear R (2017) RWTY (R We There Yet): an R package for examining convergence of Bayesian phylogenetic analyses. Mol Biol Evol 34:1016–1020. https://doi.org/10.1093/molbev/msw279
Warren DL, Glor RE, Turelli M (2010) ENMTools: a toolbox for comparative studies of environmental niche models. Ecography 33:607–611. https://doi.org/10.1111/j.1600-0587.2009.06142.x
Williford D, Deyoung RW, Honeycutt RL, Brenan LA, Hernández F (2015) Phylogeography of the bobwhite (Colinus) quails. Wildlife Monogr 193:1–49. https://doi.org/10.1002/wmon.1017
Xia X (2018) DAMBE7: new and improved tools for data analysis in molecular biology and evolution. Mol Biol Evol 35:1550–1552. https://doi.org/10.1093/molbev/msy073
Xia XH, Xie ZH, Salemi M, Chen L, Wang Y (2003) An index of substitution saturation and its application. Mol Phylogenet Evol 26:1–7. https://doi.org/10.1016/S1055-7903(02)00326-3
Young CJ, Jones JK (1983) Peromyscus yucatanicus. Mammal Species 196:1–3. https://doi.org/10.2307/3503978
Zaragoza-Quintana EP, Pech-Canché JM, Sosa-Escalante JE, Hernández-Betancourt SF, León-Paniagua LS, MacSwiney MCG (2016) Small rodents in the Yucatan Peninsula: knowledge and perspectives in 114 years of research. Therya 7:299–314. https://doi.org/10.12933/therya-16-367
Zarza H, Medellín RA, Pérez S (2003) First record of the Yucatan deer mouse, Peromyscus yucatanicus (Rodentia: Muridae) from Guatemala. Southwest Nat 48:310–312. https://doi.org/10.1894/0038-4909(2003)048<0310:FROTYD>2.0.CO;2
Acknowledgments
I thank Flor del Carmen Rodríguez-Gómez, Tania Anaid Gutiérrez-García, Eduardo Ruiz-Sánchez, and Antonio González-Rodríguez for their teaching, conceptualization, and analytic framework in the Phylogeography course in the Universidad Autónoma de Guadalajara, Mexico. Flor del Carmen Rodríguez-Gómez, Mario A. Sandoval-Molina, two anonymous reviewers, and the editor-in-chief John R. Wible provided comments and suggestions that improved this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing Interests
The author declare that there are not any competing interests.
Rights and permissions
About this article
Cite this article
León-Tapia, M.Á. DNA Barcoding and Demographic History of Peromyscus yucatanicus (Rodentia: Cricetidae) Endemic to the Yucatan Peninsula, Mexico. J Mammal Evol 28, 481–495 (2021). https://doi.org/10.1007/s10914-020-09510-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10914-020-09510-z