Abstract
Mammary epithelial cells (MEC) secrete fat in the form of milk fat globules (MFG) which are found in milk in diverse sizes. MFG originate from intracellular lipid droplets, and the mechanism underlying their size regulation is still elusive. Two main mechanisms have been suggested to control lipid droplet size. The first is a well-documented pathway, which involves regulation of cellular triglyceride content. The second is the fusion pathway, which is less-documented, especially in mammalian cells, and its importance in the regulation of droplet size is still unclear. Using biochemical and molecular inhibitors, we provide evidence that in MEC, lipid droplet size is determined by fusion, independent of cellular triglyceride content. The extent of fusion is determined by the cell membrane’s phospholipid composition. In particular, increasing phosphatidylethanolamine (PE) content enhances fusion between lipid droplets and hence increases lipid droplet size. We further identified the underlying biochemical mechanism that controls this content as the mitochondrial enzyme phosphatidylserine decarboxylase; siRNA knockdown of this enzyme reduced the number of large lipid droplets threefold. Further, inhibition of phosphatidylserine transfer to the mitochondria, where its conversion to PE occurs, diminished the large lipid droplet phenotype in these cells. These results reveal, for the first time to our knowledge in mammalian cells and specifically in mammary epithelium, the missing biochemical link between the metabolism of cellular complex lipids and lipid-droplet fusion, which ultimately defines lipid droplet size.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lipid droplets are intracellular neutral lipid depots that play a pivotal role in energy storage and lipid trafficking, and as a source for membrane polar lipids [1]. Essentially all cell types can store excess fatty acids as triglycerides in lipid droplets. However, specific cells, such as enterocytes, hepatocytes and mammary epithelial cells (MEC), can also secrete lipid droplets as lipid–protein assemblies (e.g., very low density lipoproteins (VLDL) or milk fat globules (MFG)). In general, the size of the secreted or intracellular lipid–protein assemblies vary within a narrow size range, as was demonstrated in VLDL [2], human neuroblastoma and medulloblastoma cells, as well as in human hepatocyte cell line and Drosophila S2 cells [3,4,5]. Deviation from the normal size range is usually associated with tissue dysfunction, as has been demonstrated in muscle cell lipid storage myopathy [6], hepatocytes in fatty liver [7, 8], or human monocytes induced to secrete proinflammatory agents [9]. The mammary gland is an exception, as healthy mammals secrete lipid–protein assemblies, also known as MFG, in a size range that spans over three orders of magnitude [10]. A tight association between MFG size and its proteome and lipidome has been demonstrated [10,11,12]. Therefore, revealing the underlying regulatory mechanisms of MFG size has important implications for maternal and infant health in the context of human lactation, as well as nutritional, health and organoleptic properties of milk and dairy products [13,14,15,16]. MFG size is determined prior to or during its secretion by the MEC [17] and therefore, mechanisms contributing to the regulation of cytoplasmic lipid droplets are relevant to the MFG’s size properties.
Two major pathways have been suggested to regulate intracellular lipid droplet size. The first has been well-documented, suggesting that lipid droplet size is a function of the triglyceride amount stored in the cell. Metabolic signals control triglyceride storage by autophagy [18] or by regulating the localization of members of PAT (perilipin, adipophilin, and tail-interacting protein of 47 kDa, a.k.a. perilipin 3(protein family on the lipid droplet surface. Consequently, the lipolytic activity on the surface of the droplet modulates cellular triglyceride content [19]. Accordingly, in MEC, deficiency in one of the PAT proteins, adipophilin (a.k.a. perilipin 2(, resulted in failure to produce large lipid droplets [20]. Taken together, it is reasonable to assume that metabolic conditions that change the cell’s energy requirements will affect lipid droplet size. Nevertheless, this relationship does not always hold. For example, in yeast, the presence of supersized lipid droplets was not necessarily correlated with greater triglyceride content [21]. In addition, in Drosophila, knockdown of Cct1, a rate-limiting enzyme in the phosphatidylcholine (PC)-synthesis pathway, resulted in the formation of very large lipid droplets, but did not affect triglyceride content [22]. Similarly, in milk and MEC, differences in lipid droplet size are not necessarily associated with altered triglyceride synthesis or yield [14, 23,24,25,26]. Taken together, the regulation of lipid droplet size cannot be explained merely by the cellular triglyceride content, suggesting that an additional mechanism contributes to the size regulation.
A second possible mechanism involved in regulation of lipid droplet size is fusion of intracellular lipid droplets. Fusion was demonstrated in various systems, like liposomes and artificial lipid droplets [5, 27,28,29], and in organisms like yeast [21], nematodes [30, 31] and Drosophila [22]. Fusion was even demonstrated in vivo in murine mammary gland [32], however, the regulatory mechanisms involved in this process were not assessed. Fusion may be regulated by specific proteins on the surface of intracellular lipid droplets as was shown for murine MEC in vivo [33] and for NIH-3T3 cells [34], or by membrane phospholipid composition which was studied essentially in liposomes. In liposomes and synthetic systems, a pivotal role was demonstrated for PC and phosphatidylethanolamine (PE) and their mass ratio in determining membrane stability and the extent of fusion [5, 27,28,29]. In yeast, phosphatidic acid was also found to play a role in fusion [21], however it is only a minor component in mammalian cell membranes, therefore its role in fusion in mammalian cells is less established. In living cells and organisms, a direct proof of the involvement of phospholipid composition in lipid droplet fusion and size has been documented in nematodes [30, 31], Drosophila [22] and yeast [21], and even in mouse macrophages [5]. However, those studies were performed on mutant strains of yeast, RNAi functional genomic screening in Drosophila, and recombinant mouse strains, all characterized by the formation of extremely large lipid droplets that are not necessarily present under physiological conditions. In mammalian cells, the potential role of the cellular content of PC and PE in regulating lipid droplet size was poorly studied, although the synthesis and degradation pathways of PC and PE have been well documented [35, 36]. Moreover, the contribution of fusion to regulation of lipid droplet size, especially in mammalian cells, remains to be elucidated.
In this study, PE and PC metabolism was modulated by metabolic and molecular means to study the role of polar lipid composition in lipid droplet size and fusion in bovine MEC. The results provide first evidence of extensive fusion events in MEC which control lipid droplet size, regardless of the total cellular fat content.
Results
Lipid Droplet Size is Associated with Phospholipid Composition
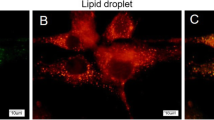
In the current study, we found that treating cells with free oleic acid (C18:1) increased the number of large lipid droplets sixfold compared to free palmitic acid (C16:0) treatment (Fig. 1a, b). The distribution of the cells among the different phenotypic size groups differed significantly between treatments (Fig. 1c, P ≤ 0.0001). The prevalence of cells with large lipid droplets (≥ 2.5 µm) was 4% for the palmitic acid and 22% for the oleic acid treatment. Triglyceride content in cells and medium was 24% and 100% higher in the oleic acid vs. palmitic acid treatment, respectively (Fig. 1d, P = 0.0021; Fig. 1e, P = 0.016). Concomitant with the change in lipid droplet size, phospholipid composition differed between the treatments (Fig. 1f). Specifically, compared with palmitic acid, treatment with oleic acid increased the content of PE by 63% (P < 0.0001), while those of phosphatidylserine (PS) and PC decreased by 10% (P = 0.02) and 14% (P < 0.0001), respectively.
Free fatty acids alter lipid droplet size concomitant with phospholipid composition. MEC were treated with 100 µM free fatty acids for 24 h. a Morphology of lipid droplets in representative cells. Cells treated with oleic acid showed larger lipid droplets relative to palmitic acid treatment. Neutral lipids were stained with Nile red (red) and nuclei were stained with DAPI (blue). Scale bars, 20 μm. b Number of large lipid droplets (> 2 µm) per image. Oleic acid-treated cells demonstrated approximately sixfold increase in the number of large lipid droplets relative to palmitic acid treatment. Droplets were counted in 45–56 cells in each treatment. All lipid droplets visualized within the cell were measured. c Greater percentage of cells with large droplet phenotype in MEC treated with oleic compared to palmitic acid, as determined by Chi square test (large droplets = larger than 2.5 µm, P < 0.05). 80–100 cells were analyzes for each replicate (240–310 cells/treatment). d Cellular triglyceride amount differs slightly between palmitic and oleic acid treatments. e Triglyceride concentration in the medium was twofold higher in oleic acid vs. palmitic acid treatment. f Phospholipid composition (weight %) differs between palmitic and oleic acid treatments. MEC treated with oleic acid had greater PE and lower PC content compared to palmitic acid treatment. Data are presented as mean ± SD. *P < 0.05
Oleic Acid Treatment Increases Fusion Rates
To determine whether the increased size of the lipid droplets under oleic acid treatment is a consequence of fusion, MEC stained with Nile red were followed by time-lapse confocal microscopy. In general, analysis of movies (2 h) revealed lipid-droplet fusion events; each fusion of two lipid droplets, visualized as a flattened z-stack movie, was recorded and verified in a 3D-reconstructed movie (Fig. 2a and Online Resource 1). In the oleic acid treatment, the number of fused droplets was 7.25-fold higher than with the palmitic acid treatment (Fig. 2b, P = 0.0002). In addition, lipid droplet number was 1.74-fold higher in the oleic acid treatment (Fig. 2c, P = 0.06). Thus, to normalize the extent of fusion between treatments, the percentage of lipid droplets engaged in fusion out of the total number of lipid droplets was calculated (Fig. 2d). Approximately 20% of all droplets were involved in the fusion process in the oleic acid treatment, compared to less than 5% in the palmitic acid treatment (Fig. 2d, P = 0.006). As oleic acid was shown to increase PE content, these results imply that PE content has an important role in enhancing lipid droplet fusion rate and consequently increase lipid droplet size.
Fusion of lipid droplets is enhanced by oleic acid treatment relative to palmitic acid treatment. a Representative fusion event of two lipid droplets (indicated by arrows), observed in MEC treated with 100 µM free palmitic acid + 10 µM DZA and stained for lipid droplets with Nile red. MEC imaged by time-lapse system. Scale bar, 5 µm. b Number of droplets engaged in fusion process was 7.25-fold higher in oleic acid relative to palmitic acid treatment. MEC treated with 100 µM palmitic acid or 360 µM oleic acid and stained with Nile red. Images were taken for 2 h, starting 1.5 h after the beginning of treatment. c Total lipid droplet number was higher in oleic acid relative to palmitic acid treatment. d Proportion of droplets that were involved in fusion out of total droplets (%) was fourfold greater in oleic acid relative to palmitic acid treatment. Data are presented as mean ± SD. *P < 0.05. n = 3 movies for each treatment, 100–150 lipid droplets in each movie. See also Online Resource 1
Inhibition of PE Degradation Through the Phosphatidylethanolamine N-methyltransferase (PEMT) Pathway Increases the Number of Large Lipid Droplets
To elucidate PE’s role in the regulation of lipid droplet size, we intervened with its degradation pathway via PEMT. Inhibition of PE degradation is expected to increase PE content and destabilize the membrane, therefore increasing lipid droplet fusion and size. This altered membrane properties will be best visualized under treatment that induces the formation of small droplets, a.k.a cellular exposure to palmitic acid. Cells were exposed to palmitic acid in the presence or absence of 3-deazaadenosine (DZA), a PEMT inhibitor, for 24 h (Fig. 3a). To reduce PC synthesis even further, we used a choline-deficient medium (in both control and DZA treatments) [37, 38], which reduces the ability to synthesize PC de novo. Exposing the cells to palmitic acid + DZA resulted in a higher number of large lipid droplets compared to treatment with palmitic acid alone (Fig. 3b, c). No toxic effects were recorded (Online Resource 2). Treatment of palmitic acid + DZA did not change the amount of triglycerides in the medium (Fig. 3d, P = 0.8), whereas it elevated the amount of triglycerides in the cells 3.5-fold compared to palmitic acid alone (Fig. 3e, P ≤ 0.0001). Determination of phospholipid composition revealed higher amounts of PC, PE and PS (P = 0.04, P = 0.08, P = 0.08, respectively) in the palmitic acid + DZA treatment compared with palmitic acid alone. The greatest change between treatments was found in PE level, which was 29% higher in the palmitic acid + DZA treatment (Fig. 3f).
Increased size of lipid droplets in MEC treated with palmitic acid and methylation inhibitor (DZA). a PE-to-PC conversion was inhibited by administration of DZA, a PEMT inhibitor, to cells treated with palmitic acid and incubated in choline-deficient medium. b MEC shows larger lipid droplets in palmitic acid + DZA treatment relative to palmitic acid alone. Nile red staining for lipid droplets (red) and DAPI staining for nuclei (blue) of MEC treated with 10 µM DZA or control, in the presence of 100 µM palmitic acid for 24 h. Scale bars, 20 μm. c MEC treated with palmitic acid + DZA demonstrated approximately threefold increase in the number of large lipid droplets relative to palmitic acid alone. Lipid droplets were measured in 45–69 cells in each treatment. All lipid droplets visualized within the cell were measured. d Triglyceride concentration in the medium was similar for palmitic and palmitic acid + DZA treatments. e Cellular triglyceride amount was greatly increased in the palmitic acid + DZA treatment. f Cellular phospholipid amount analyzed by HPLC–ELSD shows that palmitic acid + DZA treatment increased the amount of PE, PS and PC. The greatest change between treatments was found in the amount of PE, with 29% higher content in the palmitic acid + DZA treatment. Data are presented as mean ± SD. *P < 0.05. See also Online Resource 2
Disruption of Phosphatidylserine Translocation to the Mitochondria Inhibited the Formation of Large Lipid Droplets
To reduce PE synthesis through the phosphatidylserine decarboxylase (PSD) pathway, we used sodium azide and sodium fluoride (NaN3 + NaF; Fig. 4a), as previously demonstrated in baby hamster kidney (BHK)-21 cells [39]. Treating the cells for 24 h with oleic acid + 5 mM NaN3 + 20 mM NaF diminished the large lipid droplet phenotype; 100% of the cells under this treatment presented only small lipid droplets (Online Resource 3a). In addition, the phospholipid composition was changed, characterized by lower PE and higher sphingomyelin contents in cells cultured with oleic acid + NaN3 + NaF relative to those cultured with oleic acid alone (Online Resource 3b). Given that pronounced cell death (approximately 50%) was recorded in the oleic acid + NaN3 + NaF-treated group (Online Resource 3c), another set of experiments was performed with a shorter incubation time and lower concentrations of NaN3 + NaF. Incubating cells with oleic acid + 2.5 mM NaN3 + 10 mM NaF for 2 h significantly reduced lipid droplet size (Fig. 4b,c). In particular, an eightfold decrease in the number of large droplets was observed after 2 h of treatment with oleic acid + NaN3 + NaF compared to oleic acid alone (P = 0.02). No effect was observed on cell viability (Online Resource 3d).
Inhibition of PSD pathway by NaN 3 + NaF reduces the number of large lipid droplets. a PE synthesis was inhibited by addition of NaN3 + NaF to oleic acid-treated cells. b MEC treated with 360 µM oleic acid + 2.5 mM NaN3 + 10 mM NaF or with oleic acid alone, show smaller lipid droplets in oleic acid + NaN3 + NaF after 2 h of treatment. Neutral lipids were stained with Nile red (red) and nuclei were stained with DAPI (blue). Scale bars, 20 μm. c Number of large lipid droplets (> 2 µm) per image. Cells treated with oleic acid + NaN3 + NaF exhibited a sharp, eightfold decrease in the number of large lipid droplets relative to oleic acid alone (*P ≤ 0.05). Droplets were counted in 54–72 cells in each treatment. All lipid droplets visualized within the cell were measured. d Treatment duration affected cellular triglyceride amount in MEC with no effect of treatment in itself. Graph indicates the triglyceride concentration in MEC treated with oleic acid + NaN3 + NaF or with oleic acid alone, for 2 and 4 h (*P ≤ 0.05 between oleic acid and oleic acid + NaN3 + NaF treatments, #P ≤ 0.05 between 2 and 4 h of treatment (n = 4 for each replicate, 2 replicates). e Cellular phospholipid amount analyzed by HPLC–ELSD showed that treatment with oleic acid + NaN3 + NaF decreases the amount of PE and PC compared to oleic acid alone (*P ≤ 0.05 between oleic acid and oleic acid + NaN3 + NaF treatments, #P ≤ 0.05 between 2 and 4 h of treatment). f Triglyceride concentration in the medium was reduced twofold when MEC were treated with 360 µM oleic acid + 2.5 mM NaN3 + 10 mM NaF relative to oleic acid alone, after 2 h of treatment. Data are presented as mean ± SD. See also Online Resource 3 (Figure) and Online Resources 4 and 5 (Movies)
To determine whether the decrease in lipid droplet size was due to reduction in triglyceride amounts, we examined its cellular content. Interestingly, although lipid droplet size was reduced by oleic acid + NaN3 + NaF, no effect on cellular triglyceride content was found after 2 h of treatment (Fig. 4d, P = 0.8). Nevertheless, after 4 h, a slight increase in triglyceride concentration was observed in this group (Fig. 4d, P = 0.0017), which continued to present the small droplet phenotype, compared to cells treated with oleic acid alone (Online Resource 3e). These findings support our assumption that the reduced lipid droplet size observed in cells treated with oleic acid + NaN3 + NaF cannot be attributed to changes in cellular triglyceride concentration.
The effect of treatment on phospholipid composition was also investigated. In general, the amount of all phospholipids was reduced by either time or treatment, with no significant effect for the time-by-treatment interaction (Fig. 4e, P > 0.15). PE was the only phospholipid whose concentration was affected by treatment alone (P = 0.01), and its content was reduced by 19% in the oleic acid + 2.5 mM NaN3 + 10 mM NaF treatment compared to that with oleic acid alone. PC concentration was affected by both treatment (P = 0.0003) and time (P = 0.03) and was reduced by 30% in the oleic acid + NaN3 + NaF vs. oleic acid treatment. PS, phosphatidylinositol (PI) and sphingomyelin concentrations were not affected by the treatment (P = 0.17, 0.17 and 0.09, respectively).
Given that metabolic inhibitors such as NaN3 + NaF may reduce the cell’s secretion capacity, triglyceride concentration was determined in the culture medium. Our findings revealed 50% lower triglyceride concentration in the medium when the cells were treated with oleic acid + NaN3 + NaF compared to oleic acid alone (Fig. 4f). This effect was reflected by the slower movement (0.005 ± 0.003 µm/min) of lipid droplets in this treatment (Online Resource 4) compared to that with oleic acid alone (0.020 ± 0.023 µm/min, P = 0.01, Online Resource 5). It should be noted that lipid droplet size was smaller, although triglyceride amount was greater, in the presence of NaN3 + NaF, demonstrating the absence of a positive correlation between lipid droplet size and triglyceride content in the cells.
Taken together, the results imply a central role for PE in regulating lipid droplet size. Nevertheless, NaN3 + NaF might also affect droplet fusion directly; droplet movement on the cytoskeleton, which induces fusion, has been shown to be ATP-dependent [34]. NaN3 + NaF might inhibit ATP production and therefore affect droplet fusion directly, rather than through its inhibitory effect on PE synthesis.
PSD Knockdown Inhibits Biosynthesis of Large Lipid Droplets
Knockdown by siRNA designed to reduce PSD gene expression was employed as an additional strategy to reduce the conversion of PS to PE. Specifically, cells were incubated with oleic acid to induce large lipid droplet formation, and transfected with siRNA designed to knock down PSD expression or with negative control siRNA (Fig. 5a). PSD expression was reduced by 25% (Fig. 5b, P = 0.005), resulting in a threefold lower number of large lipid droplets (Fig. 5c, d, P = 0.056). In addition, a 35% decrease in cells presenting large lipid droplets was found when PSD was knocked down compared to controls (Fig. 5e, P < 0.0001). In terms of phospholipid composition (Fig. 5f), treatment increased PS content by 20% compared to controls (P = 0.08). None of the other phospholipid concentrations were affected by the treatment. The reduced lipid droplet size in the PSD siRNA treatment was not associated with changes in triglyceride content in the cells (Fig. 5g, P = 0.06).
PSD knockdown inhibits large lipid droplet biosynthesis in MEC. a PSD siRNA transfection of MEC treated with oleic acid was used to inhibit PS-to-PE conversion by PSD. b PSD gene expression decreased by 25% following transfection of PSD siRNA compared to control siRNA. c Reduced lipid droplet size in MEC transfected with PSD siRNA compared to negative control siRNA. Neutral lipids were stained with Nile red (red) and nuclei were stained with DAPI (blue). Scale bars, 20 μm. d Number of large lipid droplets (> 2 µm) per image. Cells exhibited a threefold decrease in the number of large lipid droplets following PSD knockdown compared to control siRNA. 70–160 cells were counted for each replicate in each treatment, total of 3 replicates. e Distribution of MEC phenotype was determined by Chi square test (P ≤ 0.05). MEC presenting large lipid droplets (> 2 µm) decreased by 35% in PSD siRNA treatment relative to control. f Phospholipid composition (weight %) analyzed by HPLC–ELSD showed that PSD knockdown increases PS content relative to control. g Cellular triglyceride amount analyzed by HPLC–ELSD was not altered due to PSD knockdown. Data are presented as mean ± SD. *P ≤ 0.05. See also Online Resource 8
Discussion
Biosynthesis of large lipid droplets can be attributed to greater accumulation of triglycerides in the cell [20, 31, 40], or to greater susceptibility to fusion [21, 22, 34, 41, 42]. The physiological importance of fusion in regulating lipid droplet size is unknown, especially in MEC. In the present study, we provide first evidence for the role of fusion in regulating lipid droplet size in bovine MEC, while distinguishing between this effect and that of cellular triglyceride content.
Treating MEC with free palmitic or oleic acid resulted in substantial differences in lipid droplet size and phospholipid composition (Fig. 1), while the differences in intracellular triglyceride content between treatments was only marginal, albeit statistically significant, (Fig. 1).This result suggests that elevated triglyceride accumulation in the cells cannot in and of itself explain the altered lipid droplet size. Alternatively, the altered PC and PE contents were expected to alter the lipid droplet’s susceptibility to fusion [27], as has been demonstrated in liposomes [28, 29].
The hypothesis that phospholipid composition is engaged in regulating lipid droplet size through fusion was supported by our finding of a much greater degree of fusion between lipid droplets in the oleic acid treatment (Fig. 2). Demonstrating lipid droplet fusion is a challenge, both technically and from a biological standpoint. Hence, only a few studies have demonstrated this process in living cells. While some of these showed fusion in wild-type 3T3 cells [34, 41], others used fusogenic reagents to induce fusion [42], or mutant yeast strains that produce supersized lipid droplets, without concomitant demonstration in the wild type [21]. This study is the first to show lipid droplet fusion in bovine MEC in culture. This joins the very recent demonstration of this process in murine mammary gland in vivo [32]. In this regard, it should be noted that capturing fusion in primary culture of MEC differs from terminally differentiated cells in vivo since MEC in culture have reduced ability to assemble and secrete milk components in optimal quantities relative to copious milk secretion in vivo [43]. In support of this, lipid droplet migration rate demonstrated herein was slower compared to that described for in vivo lipid droplet fusion in MEC [32]. The slower movement may be attributed to random movements in non-polarized cells in culture compared with directed movement towards the apical pole of the cell in the in vivo model.
The hypothesis that regulation of phospholipid composition is sufficient to induce phenotypic changes in lipid droplet size was further tested by disrupting PE- and PC-synthesis and degradation pathways. To reverse the phenotype of small lipid droplets under the palmitic acid treatment, which was characterized by high PC content, we used the PEMT inhibitor, DZA, which inhibits cellular methylation activity [44] (Fig. 3). Although the PEMT pathway has been shown to account for only 30% of PC synthesis in hepatocytes [45], and even less in other cell types [46], it may be significant in regulating the content of its substrate, PE. Moreover, in cell culture, PE- to- PC conversion may play a significant role in determining PE content since ethanolamine—which is required for de novo synthesis of PE—is absent from commercial culture media. In this experiment we also used choline-deficient medium, to prevent de novo synthesis of PC [37, 38] and further increase the effect of the PEMT inhibition on PC content.
The larger lipid droplets in the DZA treatment (Fig. 3b,c) is in agreement with the formation of supersized lipid droplets in a yeast mutant deleted for the cho2 gene (functional homologue with mammalian PEMT) [21] and with previous studies showing biosynthesis of large lipid droplets associated with lower cellular PC content [5, 22, 31].
Nonetheless, the use of DZA as a methylation inhibitor had some limitations. For example, the increased PE content was accompanied by elevated concentrations of PS and PC, as precursors for the PE-synthesis cycle (Fig. 3f). These results are in agreement with a previous study showing coordinated regulation of PE and PS content in adipose tissue of PEMT-knockout mice [47]. In addition, the effect of DZA was not specific, as adding it to the culture media elevated triglyceride content in the cells (Fig. 3e), which probably contributed to the formation of larger droplets.
Therefore, we used another approach to modify PE biosynthesis, this time on the background of oleic acid treatment, aiming to increase membrane stability and reduce the extent of the large lipid droplet phenotype in MEC. PE synthesis by the PSD pathway is inhibited by NaN3 + NaF, which inhibits the transport of PSD substrate, PS, into the mitochondria [39]. Indeed, treatment with NaN3 + NaF reduced PE content, which eliminated the large lipid droplet phenotype from MEC culture (Fig. 4b,c,e and Online Resource 3a). That treatment also decreased PC content, in agreement with a previous study that showed a direct effect of NaN3 + NaF on choline metabolism in PC12 (neuron-like) cells [48]. However, while the results of the present study show that reduced PC content was associated with decreased lipid droplet size, various studies demonstrated the opposite association [5, 21, 22, 31]. This inconsistency can be attributed to either inhibited fusion attributed to the reduction in PE content under the NaN3 + NaF treatment (Fig. 4e) or reduced ATP synthesis capacity by NaN3 + NaF as was shown in BHK-21 cells [39]. ATP shortage may reduce binding of lipid droplets to SNAP receptor (SNARE) proteins which are required for lipid droplet fusion [34]. This possible inhibition is supported by our findings of slower movement of the lipid droplets of the oleic acid + NaN3 + NaF treatment compared with oleic acid alone, as visualized in the time-lapse microscopy (Online Resources 4 and 5, respectively). Reduced ATP synthesis capacity does not necessarily lead to the formation of small droplets, as was demonstrated in liver cells [49]. Under starvation and sub optimal ATP synthesis two distinct mechanisms can change lipid droplet size - lipolysis and autophagy. While lipolysis is known to reduce lipid droplet size, it was shown in starved mouse embryonic fibroblasts that autophagy increases lipid droplet size [18]. Taken together, these data suggest that lipid droplet size is determined by a complex regulatory mechanisms including, but not exclusive to, cellular energy requirements [18], expression of proteins on the surface of lipid droplets as PAT and CIDEA proteins [19, 50], phospholipid membrane composition and the interplay between them.
To affect PE synthesis more specifically, we used PSD knockdown which resulted in the biosynthesis of smaller droplets (Fig. 5) and was not associated with change in triglyceride content in the cell (Fig. 5g). Therefore, this methodology enabled us to distinguish between the metabolic pathways responsible for phospholipid composition in the cell and the cellular triglyceride content. In addition, these results provide evidence that triglyceride content in the cell is only a part of the regulatory mechanism underlying the regulation of lipid droplet size. Other crucial regulators of lipid droplet size are proteins located on the envelope of the lipid droplet or recruited from the cytoplasm [33]. For example, CIDE proteins regulate lipid droplet fusion and growth in various tissues including adipose tissue, liver, skin and mammary glands (reviewed in [50]). Specifically, in murine mammary glands of CideA deficient mice, smaller cytosolic lipid droplets and decreased milk lipid secretion was found [51]. Additional example for the role of proteins in regulation of size is xanthine oxidoreductase which has a major role in regulation of lipid droplet and MFG size as was shown in mammary-specific xanthine oxidoreductase knockout mice [33]. Though the role of these proteins in regulating fusion is established, the role of membrane lipidome, as additional level of regulation in these study models, was not addressed.
Regulation of lipid droplet size in MEC has important implications for maternal and infant health in the context of human lactation. Lipid droplet size is also important for the quality of milk and dairy products, which are major sources of lipids in the western diet. The size of the intracellular lipid droplet formed by MEC prior to its secretion into the milk-collecting ducts will determine the size of the secreted MFG. The size of the MFG is tightly linked to its lipid, fatty acid [11] and protein [12] contents, and therefore determines the composition of the milk. In particular, the content of phospholipids in milk is determined by MFG size, with greater content in milk enriched with small globules [11]. Higher phospholipid content in the general population’s diet is desirable due to its health effects. In young animals, the polar lipid envelope of the MFG has been found to affect neurodevelopment [52]. It has also been found to provide health and cognitive benefits at 12 months of age in human infants [53, 54]. With respect to maternal health, mastitis has been found to be associated with MFG size and therefore, the ability to control MFG size may represent an important therapeutic target to ameliorate the prevalence of mastitis [13].
In summary, our findings reveal a defining role for lipid droplet fusion in determining lipid droplet size in MEC. We identified a regulatory mechanism of lipid droplet size that is independent of cell triglyceride content. This mechanism involves the metabolic pathways and intracellular compartments responsible for controlling specific phospholipid content in the cellular membranes (Fig. 6). These pathways can now be targeted by a variety of molecular and metabolic means. The fact that changes in lipid droplet size can be achieved independently of cellular triglyceride content implies that lipid droplet size can be manipulated without reducing milk fat yield. This mechanism thus points directly to new approaches in which milk composition and quality are controlled without compromising its overall value.
Simplified model of coupling of phospholipid metabolic pathways in mammary cells and regulation of lipid droplet size. Phospholipid biosynthetic pathways in mammalian cells. The three major phospholipids found in mammalian membranes, PE, PS and PC, are linked through an enzymatic sequence. We incubated MEC in a choline- and ethanolamine-deficient medium to prevent de novo synthesis of PC and PE. Therefore, we isolated the activity of PEMT and PSD as determinants of PC and PE cellular content. Further, we modulated MEC phospholipid composition by metabolic and molecular means to elucidate the phospholipid contribution to the fusion process, and hence to lipid droplet size. Lipid droplet (LD) size was found to be regulated by fusion, through phospholipid envelope composition, regardless of triglyceride concentration in the cell. The final outcome of this fusion process is the cellular lipid droplet size phenotype
Materials and Methods
Materials
The basic culture medium used for the experiments was DMEM/F12 (Biological Industries, Beit Haemek, Israel), which is ethanolamine deficient. DMEM/F12 without choline, fetal bovine serum (FBS), penicillin, streptomycin, amphotericin B, L-glutamine solution and trypsin–EDTA solution C were purchased from Biological Industries (Beit Haemek, Israel). Bovine insulin, hydrocortisone, ovine prolactin, bovine serum albumin (BSA) solution, hyaluronidase, DNase I, heparin, sodium azide, sodium fluoride and DZA were purchased from Sigma Aldrich Israel Ltd. (Rehovot, Israel). Collagenase type II was purchased from Worthington Biochemical Corporation (Lakewood, NJ).
Primary Culture
Primary culture of MEC was isolated from mammary biopsies according to a protocol established in our laboratory [23] with minor modifications. Briefly, udder tissue was collected from lactating cows in a commercial slaughterhouse and immediately submerged in ice-cold growth medium with 1000 U/ml penicillin, 1 mg/ml streptomycin, 2.5 μg/ml amphotericin B mixture, and 0.02 mg/ml heparin supplementation to prevent cell clotting after digestion. Study protocols were in compliance with the regulations of the Israeli Ministry of Health.
After transfer to the laboratory, tissue was minced and digested by shaking in growth medium supplemented with collagenase (1 mg/ml), hyaluronidase (1 mg/ml) and 0.02 mg/ml heparin, at 100 rpm for 3 h at 37 °C. After incubation, the suspension was filtered through a metal mesh (250 μm) and the filtrate was centrifuged at 350 g for 5 min. The sediment was treated with trypsin–EDTA and 0.04% (w/v) DNase. The cells were then washed with growth medium supplemented with heparin and treated with DNase alone, filtered through a 100-μm cell strainer (BD Falcon, Bedford, MA) and washed with the growth medium.
Cells were grown in plastic culture dishes with DMEM/F12 supplemented with 10% (w/v) FBS, 100 U/ml penicillin, 100 μg/ml streptomycin, 0.25 μg/ml amphotericin B, 1 μg/ml insulin and 0.5 μg/ml hydrocortisone.
Experimental Design
Primary MEC were plated at 150,000 cells per 60-mm plastic dish for cellular lipid extraction and RNA extraction, at 50,000 cells per well in 6-well plates on glass cover slips for Nile red staining, or at 4,250 cells per plate in an 8-well chambered coverglass system (Nunc, Roskilde, Denmark) for live cell imaging. After overnight incubation, the medium was replaced with DMEM/F12 without serum, containing 0.15% (w/v) free fatty acids–free BSA and insulin (1 μg/ml), hydrocortisone (0.5 μg/ml) and prolactin (1 μg/ml) for 48 h to induce milk lipid and protein synthesis. Treatment medium included free palmitic acid (C16:0) or free oleic acid (C18:1), palmitic acid + DZA, oleic acid + NaN3 + NaF, or oleic acid + siRNA, all dissolved in DMEM/F12 supplemented with 0.5% (w/v) free fatty acids–free BSA, insulin (1 μg/ml), hydrocortisone (0.5 μg/ml) and prolactin (1 μg/ml).
The experiments described herein required multiple preparations of MEC primary culture. Due to individual variations between the animal origins of the MEC, the culture might differ in responsiveness to free fatty acid concentrations in relation to lipid droplet size (see Online Resource 6). For standardization of primary culture responsiveness to the basic treatments, dose-dependent studies with 100 to 1000 µM of either palmitic or oleic acid were carried out, followed by Nile red staining and visualization under a fluorescence microscope. Concentration of free fatty acids in the treatment medium was set according to the lipid droplet size phenotype induced upon treatment with free palmitic or oleic acid, and is indicated for each experiment.
Phospholipid-synthesis Inhibition
To inhibit PS to PE conversion, through the mitochondria PSD pathway, MEC were incubated with 2.5 mM NaN3 + 10 mM NaF for 2 or 4 h (modified from [39]) in the presence of 360 µM oleic acid. To reduce PC synthesis, we used the PEMT inhibitor DZA and choline-free medium. MEC were incubated for 24 h with 10 µM DZA (modified from [44]) in the presence of 100 µM palmitic acid in medium without choline.
Lipid Extraction and Analysis
After treatment, cells were harvested with trypsin (0.05%), washed with 0.9% (w/v) NaCl and stored at -20 °C until lipid extraction. Total lipids were extracted from the intact cells by cold-extraction procedure developed by Folch et al. [55] with few adaptations [23]. Briefly, a 5-ml aliquot of methanol:chloroform solution (2:1, v/v) was added to each sample. After incubation at room temperature, 1 ml of double-distilled water was added. After overnight incubation at 4 °C, the upper phase was discarded and the lower phase was filtered through a Pasteur pipette with glass wool. Samples were then dried under a nitrogen stream at 65 °C, diluted in 100 μl of chloroform:methanol (97:3, v/v) and stored at -20 °C until injection for HPLC. Separation of polar and neutral lipids was performed on a silica column (Zorbax RX-SIL, 4.6 × 250 mm, Agilent Technologies) by HPLC (HP 1200, Agilent Technologies, Santa Clara, CA) with an evaporative light-scattering detector (1200 series ELSD, Agilent Technologies). The separation protocol consisted of a gradient of dichloromethane, methanol:ammonium mix (99:1, v/v), and double-distilled water (see Online Resource 7). The separation process was managed by ChemStation software (Agilent Technologies) for the acquisition of data from the ELSD detector. The separated lipids were identified using external standards (Sigma Aldrich). Quantification was performed against external standard curves and expressed as µg/106 live cells or as weight % out of total phospholipids (µg) in the sample. Live cell number was determined with a hemocytometer after Trypan blue staining.
Triglyceride Quantification in the Culture Medium
After treatment, medium was collected and centrifuged at 500 g for 10 min. The supernatant was transferred to a fresh tube and kept at 4 °C until analysis. Medium triglyceride content was quantified with a Triglyceride Quantification Kit (Abcam, Cambridge, UK). The concentration of triglycerides in the medium was calculated per 106 cells and presented as fold change between treatments.
Lipid Droplet Staining
Cells grown on glass cover slips were rinsed three times with PBS and fixed with 4% paraformaldehyde in PBS for 20 min at room temperature. Then the cover slips were rinsed four times with PBS and stained with Nile red (200 nM, Sigma, St. Louis, MO) for 15 min. Cover slips were then rinsed three times with PBS and stained with DAPI (Sigma) for 5 min. Finally, cover slips were rinsed four times with PBS and mounted with fluorescent mounting medium (Dako, North America Inc., Carpinteria, CA).
Fluorescence Microscopy and Lipid Droplet Size Measurements
Slides were visualized with an Olympus BX40 fluorescence microscope equipped with an Olympus DP73 digital camera using CellSens Entry software (version 1.7, Olympus). Lipid droplet diameter was measured using ImageJ software (version 1.48, NIH, Bethesda, MD). Cells with at least one lipid droplet larger than 2.5 μm (or 2 μm in Fig. 5) were designated “large lipid droplets”. Cells with droplets at maximum diameter of 2.5 μm were designated “small lipid droplets”. Cells with no visualized lipid droplets were designated “no lipid droplets”. Values represent the mean size of all droplets in the cells.
Live Cell Imaging and Analysis of Droplet Fusion
Nile red was added directly to the treatment medium at a concentration of 64 ng/ml. Then cells were placed in an environmental chamber set at 37 °C and supplied with CO2 on a Leica TCS SP8 confocal microscope. The microscope was equipped with a hybrid detector and resonant scanner. Cells with several clear droplets were targeted. Pictures of 14–18 confocal planes through the cell (z stack of 6–7.5 µm) with a step size of 0.42 µm were taken with a 63X oil-immersion objective lens every 5 min for 2–3 h, using Leica LAS AF software. The laser wavelength was set to 488 nm and emission to 600 (585–625) nm. Flattened z-stack movies of MEC stained with Nile red were viewed by two independent examiners and lipid droplets that were suspected of fusion were marked. Then a 3D reconstruction of the relevant frames was conducted and visualized to validate fusion.
PSD Knockdown
Cells were transfected with PSD siRNA (sense: 5′-GCCUGUACAUCUGGACUUU[dT][dT]-3′) or negative control siRNA (SIC001, Sigma) at 30 nM concentration with the N-TER™ Nanoparticle siRNA Transfection System (Sigma) in the presence of 150 µM oleic acid for 24 h; siRNA sequences were designed by Sigma–Aldrich experts and the oligos were purchased from Sigma as well.
Gene-expression Analysis
To estimate PSD knockdown, real-time PCR analysis was performed on RNA isolated from primary mammary cells by GenElute™ Mammalian Total RNA Miniprep Kit (Sigma). Total RNA (1 μg) was reverse-transcribed using the qScript cDNA synthesis kit (Quanta Biosciences). Primers used for real-time PCR analysis were synthesized by Sigma (Rehovot, Israel), with the following sequences: PSD gene (NM_001024475.1) F – ACATCTGGACTTTCGGGGTG, R – GAGGGGCTGATCACGCTG (giving a 142-bp PCR product) (designed by Primer-BLAST software, National Center of Biotechnology Information [NCBI], http://www.ncbi.nlm.nih.gov/tools/primer-blast/index, based on cDNA sequences published by the NCBI database and validated by PCR product sequencing, Online Resource 8); UXT (normalizing) gene (BQ676558) F – TGTGGCCCTTGGATATGGTT, R – GGTTGTCGCTGAGCTCTGTG (101-bp product) [56]; β2-microglobulin (normalizing) gene (NM_173893) F – CATCCAGCGTCCTCCAAAGAT, R – CCCCATTCTTCAGCAAATCG (131-bp product) [57]. Real-time PCR was performed with platinum SYBR Green qPCR supermix–UDG without ROX (Invitrogen Corporation, Carlsbad, CA) in a LightCycler96 Real-Time PCR System (Roche, Basel, Switzerland). Analysis was performed by LightCycler96 software version 1.1.0.1320 (Roche). The efficiency of the reaction and the initial mRNA quantity in the sample were determined using DART-PCR software version 1.0. Expression data were normalized by geometrical means of the two housekeeping genes UXT and β2-microglobulin, and presented as fold change relative to control.
Statistical Analysis
All statistical procedures were performed using JMP software version 12.0.1 (SAS Institute, Cary, NC). All reported data are means ± SD. All dependent variables were checked for homogeneous variance by unequal variances in JMP software and if the variance was not homogeneous, a Welch–ANOVA test was performed. Comparisons were made by Student t test. The distribution of cell phenotypes based on lipid droplet size categories was compared by Chi square test. In the NaN3 + NaF experiment, the effects of time, treatment and their interaction were determined using the mixed ANOVA model of JMP. Treatment was analyzed as a fixed effect and time as a continuous effect. Time-by-treatment interaction was tested and found not significant, and therefore excluded from the model. Significance probe was set to 0.05.
References
Walther TC, Farese RV Jr. The life of lipid droplets. Biochim Biophys Acta. 2009;1791(6):459 – 66.
Wojczynski MK, Glasser SP, Oberman A, Kabagambe EK, Hopkins PN, Tsai MY, et al. High-fat meal effect on LDL, HDL, and VLDL particle size and number in the Genetics of Lipid-Lowering Drugs and Diet Network (GOLDN): an interventional study. Lipids Health Dis. 2011;10:181.
Pan X, Wilson M, McConville C, Arvanitis TN, Kauppinen RA, Peet AC. The size of cytoplasmic lipid droplets varies between tumour cell lines of the nervous system: a 1H NMR spectroscopy study. MAGMA. 2012;25(6):479 – 85.
Nunn AD, Scopigno T, Pediconi N, Levrero M, Hagman H, Kiskis J, et al. The histone deacetylase inhibiting drug Entinostat induces lipid accumulation in differentiated HepaRG cells. Sci Rep. 2016;6:28025.
Krahmer N, Guo Y, Wilfling F, Hilger M, Lingrell S, Heger K, et al. Phosphatidylcholine synthesis for lipid droplet expansion is mediated by localized activation of CTP:phosphocholine cytidylyltransferase. Cell Metab. 2011;14(4):504 – 15.
Liang WC, Nishino I. State of the art in muscle lipid diseases. Acta Myol. 2010;29(2):351–6.
Cohen JC, Horton JD, Hobbs HH. Human fatty liver disease: old questions and new insights. Science. 2011;332(6037):1519–23.
Reddy JK, Rao MS. Lipid metabolism and liver inflammation. II. Fatty liver disease and fatty acid oxidation. Am J Physiol Gastrointest Liver Physiol. 2006;290(5):G852–G8.
den Hartigh LJ, Connolly-Rohrbach JE, Fore S, Huser TR, Rutledge JC. Fatty acids from very low-density lipoprotein lipolysis products induce lipid droplet accumulation in human monocytes. J Immunol. 2010;184(7):3927–36.
Argov N, Lemay DG, German JB. Milk fat globule structure & function; nanoscience comes to milk production. Trends Food Sci Technol. 2008;19(12).
Mesilati-Stahy R, Mida K, Argov-Argaman N. Size-dependent lipid content of bovine milk fat globule and membrane phospholipids. J Agric Food Chem. 2011;59(13):7427–35.
Lu J, Argov-Argaman N, Anggrek J, Boeren S, van Hooijdonk T, Vervoort J, et al. The protein and lipid composition of the membrane of milk fat globules depends on their size. J Dairy Sci. 2016;99(6):4726–38.
Mizuno K, Hatsuno M, Aikawa K, Takeichi H, Himi T, Kaneko A, et al. Mastitis is associated with IL-6 levels and milk fat globule size in breast milk. J Hum Lact. 2012;28(4):529 – 34.
Logan A, Auldist M, Greenwood J, Day L. Natural variation of bovine milk fat globule size within a herd. J Dairy Sci. 2014;97(7):4072–82.
O’Mahony JA, Auty MA, McSweeney PL. The manufacture of miniature Cheddar-type cheeses from milks with different fat globule size distributions. J Dairy Res. 2005;72(3):338 – 48.
Michalski MC, Gassi JY, Famelart MH, Leconte N, Camier B, Michel F, et al. The size of native milk fat globules affects physico-chemical and sensory properties of Camembert cheese. Lait. 2003;83:131–43.
Smoczyński M. Role of Phospholipid Flux during Milk Secretion in the Mammary Gland. J Mammary Gland Biol Neoplasia. 2017;22(2):117–29.
Rambold AS, Cohen S, Lippincott-Schwartz J. Fatty acid trafficking in starved cells: regulation by lipid droplet lipolysis, autophagy, and mitochondrial fusion dynamics. Dev Cell. 2015;32(6):678 – 92.
Bickel PE, Tansey JT, Welte MA. PAT proteins, an ancient family of lipid droplet proteins that regulate cellular lipid stores. Biochim Biophys Acta. 2009;1791(6): 419 – 40.
Russell TD, Schaack J, Orlicky DJ, Palmer C, Chang BH, Chan L, et al. Adipophilin regulates maturation of cytoplasmic lipid droplets and alveolae in differentiating mammary glands. J Cell Sci. 2011;124(Pt 19):3247–53.
Fei W, Shui G, Zhang Y, Krahmer N, Ferguson C, Kapterian TS, et al. A role for phosphatidic acid in the formation of “supersized” lipid droplets. PLoS Genet. 2011;7(7):e1002201.
Guo Y, Walther TC, Rao M, Stuurman N, Goshima G, Terayama K, et al. Functional genomic screen reveals genes involved in lipid-droplet formation and utilization. Nature. 2008;453(7195):657 – 61.
Cohen BC, Shamay A, Argov-Argaman N. Regulation of lipid droplet size in mammary epithelial cells by remodeling of membrane lipid composition-a potential mechanism. PLoS One. 2015;10(3):e0121645.
Couvreur S, Hurtaud C, Marnet PG, Faverdin P, Peyraud JL. Composition of milk fat from cows selected for milk fat globule size and offered either fresh pasture or a corn silage-based diet. J Dairy Sci. 2007;90(1):392–403.
Walstra P. Studies on milk fat dispersion. II. The globule size distribution of cow’s milk. Neth Milk Dairy J. 1969;23:99–110.
King JOL. The association between fat percentage of cow’s milk and the size and number of fat globules. J Dairy Res. 1957;24:198–200.
Thiam AR, Farese RV Jr, Walther TC. The biophysics and cell biology of lipid droplets. Nat Rev Mol Cell Biol. 2013;14(12):775 – 86.
Düzgüneş N, Wilschut J, Fraley R, Papahadjopoulos D. Studies on the mechanism of membrane fusion. Role of head-group composition in calcium- and magnesium-induced fusion of mixed phospholipid vesicles. Biochim Biophys Acta. 1981;642(1):182 – 95.
Haque ME, McIntosh TJ, Lentz BR. Influence of lipid composition on physical properties and peg-mediated fusion of curved and uncurved model membrane vesicles: “nature’s own” fusogenic lipid bilayer. Biochemistry. 2001;40(14):4340–8.
Shi X, Li J, Zou X, Greggain J, Rødkær SV, Færgeman NJ, et al. Regulation of lipid droplet size and phospholipid composition by stearoyl-CoA desaturase. J Lipid Res. 2013;54(9):2504–14.
Walker AK, Jacobs RL, Watts JL, Rottiers V, Jiang K, Finnegan DM, et al. A conserved SREBP-1/phosphatidylcholine feedback circuit regulates lipogenesis in metazoans. Cell. 2011;147(4):840 – 52.
Masedunskas A, Chen Y, Stussman R, Weigert R, Mather IH. Kinetics of milk lipid droplet transport, growth, and secretion revealed by intravital imaging: lipid droplet release is intermittently stimulated by oxytocin. Mol Biol Cell. 2017;28(7):935 – 46.
Monks J, Dzieciatkowska M, Bales ES, Orlicky DJ, Wright RM, McManaman JL. Xanthine oxidoreductase mediates membrane docking of milk-fat droplets but is not essential for apocrine lipid secretion. J Physiol. 2016;594(20):5899 – 921.
Boström P, Andersson L, Rutberg M, Perman J, Lidberg U, Johansson BR, et al. SNARE proteins mediate fusion between cytosolic lipid droplets and are implicated in insulin sensitivity. Nat Cell Biol. 2007;9(11):1286–93.
Vance JE. Phospholipid synthesis and transport in mammalian cells. Traffic. 2015;16(1):1–18.
Leonardi R, Frank MW, Jackson PD, Rock CO, Jackowski S. Elimination of the CDP-ethanolamine pathway disrupts hepatic lipid homeostasis. J Biol Chem. 2009;284(40):27077–89.
Albright CD, da Costa KA, Craciunescu CN, Klem E, Mar MH, Zeisel SH. Regulation of choline deficiency apoptosis by epidermal growth factor in CWSV-1 rat hepatocytes. Cell Physiol Biochem. 2005;15(1–4):59–68.
Niculescu MD, Yamamuro Y, Zeisel SH. Choline availability modulates human neuroblastoma cell proliferation and alters the methylation of the promoter region of the cyclin-dependent kinase inhibitor 3 gene. J Neurochem. 2004;89(5):1252–9.
Voelker DR. Disruption of phosphatidylserine translocation to the mitochondria in baby hamster kidney cells. J Biol Chem. 1985;260(27):14671–6.
Jacobs RL, Lingrell S, Zhao Y, Francis GA, Vance DE. Hepatic CTP:phosphocholine cytidylyltransferase-alpha is a critical predictor of plasma high density lipoprotein and very low density lipoprotein. J Biol Chem. 2008;283(4):2147–55.
Boström P, Rutberg M, Ericsson J, Holmdahl P, Andersson L, Frohman MA, et al. Cytosolic lipid droplets increase in size by microtubule-dependent complex formation. Arterioscler Thromb Vasc Biol. 2005;25(9):1945–51.
Murphy S, Martin S, Parton RG. Quantitative analysis of lipid droplet fusion: inefficient steady state fusion but rapid stimulation by chemical fusogens. PLoS One. 2010;5(12):e15030.
Barber MC, Clegg RA, Travers MT, Vernon RG. Lipid metabolism in the lactating mammary gland. Biochim Biophys Acta. 1997;1347(2–3):101 – 26.
Schanche JS, Schanche T, Ueland PM. Inhibition of phospholipid methylation in isolated rat hepatocytes by analogues of adenosine and S-adenosylhomocysteine. Biochim Biophys Acta. 1982;721(4):399–407.
Zivkovic AM, Bruce German J, Esfandiari F, Halsted CH. Quantitative lipid metabolomic changes in alcoholic micropigs with fatty liver disease. Alcohol Clin Exp Res. 2009;33(4):751–8.
Vance DE, Ridgway ND. The methylation of phosphatidylethanolamine. Prog Lipid Res. 1988;27(1):61–79.
Hörl G, Wagner A, Cole LK, Malli R, Reicher H, Kotzbeck P, et al. Sequential synthesis and methylation of phosphatidylethanolamine promote lipid droplet biosynthesis and stability in tissue culture and in vivo. J Biol Chem. 2011;286(19):17338–50.
Farber SA, Slack BE, Blusztajn JK. Acceleration of phosphatidylcholine synthesis and breakdown by inhibitors of mitochondrial function in neuronal cells: a model of the membrane defect of Alzheimer’s disease. FASEB J. 2000;14(14):2198 – 206.
Rector RS, Thyfault JP, Morris RT, Laye MJ, Borengasser SJ, Booth FW, et al. Daily exercise increases hepatic fatty acid oxidation and prevents steatosis in Otsuka Long-Evans Tokushima Fatty rats. Am J Physiol Gastrointest Liver Physiol. 2008;294(3):G619–G26.
Gao G, Chen FJ, Zhou L, Su L, Xu D, Xu L, et al. Control of lipid droplet fusion and growth by CIDE family proteins. Biochim Biophys Acta. 2017;1862(10 Pt B):1197 – 204.
Wang W, Lv N, Zhang S, Shui G, Qian H, Zhang J, et al. Cidea is an essential transcriptional coactivator regulating mammary gland secretion of milk lipids. Nat Med. 2012;18(2):235 – 43.
Mudd AT, Alexander LS, Berding K, Waworuntu RV, Berg BM, Donovan SM, et al. Dietary prebiotics, milk fat globule membrane, and lactoferrin affects structural neurodevelopment in the young piglet. Front Pediatr. 2016;4:4.
Timby N, Domellöf E, Hernell O, Lönnerdal B, Domellöf M. Neurodevelopment, nutrition, and growth until 12 mo of age in infants fed a low-energy, low-protein formula supplemented with bovine milk fat globule membranes: a randomized controlled trial. Am J Clin Nutr. 2014;99(4):860–8.
Timby N, Hernell O, Vaarala O, Melin M, Lönnerdal B, Domellöf M. Infections in infants fed formula supplemented with bovine milk fat globule membranes. J Pediatr Gastroenterol Nutr. 2015;60(3):384–9.
Folch J, Lees M, Sloane Stanley GH. A simple method for the isolation and purification of total lipides from animal tissues. J Biol Chem. 1957;226(1):497–509.
Bionaz M, Loor JJ. Identification of reference genes for quantitative real-time PCR in the bovine mammary gland during the lactation cycle. Physiol Genomics. 2007;29(3):312–9.
Harvatine KJ, Bauman DE. SREBP1 and thyroid hormone responsive spot 14 (S14) are involved in the regulation of bovine mammary lipid synthesis during diet-induced milk fat depression and treatment with CLA. J Nutr. 2006;136(10):2468–74.
Acknowledgements
The authors would like to acknowledge Dr. Sergei Grigoryan for his assistance in confocal microscopy imaging. This research was partially supported by the Nutrigenomics Center of the Hebrew University of Jerusalem, Israel, and by the Israeli Dairy Board grant #8200327.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Lipid droplet fusion. Representative fusion event of two lipid droplets, observed in MEC treated with 100 μM palmitic acid + 10 μM DZA and stained for their lipid droplets with Nile red. Optical sections were recorded with time-lapse imaging and reconstructed as 3D images from deconvolved images. Arrows indicate a pair of fusing lipid droplets. (AVI 78 KB)
10911_2017_9386_MOESM2_ESM.tif
DZA addition has no effect on live cell number compared to palmitic acid alone. Number of live cells determined by Trypan blue staining was similar in MEC treated with 10 μM DZA+100 μM palmitic acid relative to palmitic acid alone, both for 24 h. Data are presented as mean ± SD. (TIF 29 KB)
10911_2017_9386_MOESM3_ESM.tif
Time and dose response for NaN3 + NaF and oleic acid treatments. a Addition of 5 mM NaN3 + 20 mM NaF to medium containing 100 μM oleic acid eliminated MEC presenting large lipid droplets (>2.5 μm) from culture. Distribution of MEC phenotype, distinguished by the presence or absence of large lipid droplets after 24 h of treatment, was determined by chi-square test (P ≤ 0.05). b Phospholipid composition (weight %) of MEC treated as in (a) shows lower PE and higher sphingomyelin percent in oleic acid + NaN3 + NaF treatment relative to oleic acid alone. Data are presented as mean ± SD (*P ≤ 0.05). c, d Effects of NaN3 + NaF + oleic acid concentration and duration of treatment on live cell number. c 5 mM NaN3 + 20 mM NaF in the presence of 100 μM oleic acid for 24 h decreased the number of cells relative to treatment with oleic acid alone. d 2.5 mM NaN3 + 10 mM NaF in the presence of 360 μM oleic acid did not change live cell number relative to oleic acid alone after 2 and 4 h of treatment. Number of live cells was determined by Trypan blue staining. e MEC treated with 360 μM oleic acid + 2.5 mM NaN3 + 10 mM NaF presented smaller droplets relative to oleic acid alone after 4 h of treatment. Neutral lipids were stained with Nile red (red) and nuclei were stained with DAPI (blue). Scale bars, 20 μm. Data are presented as mean ± SD (*P ≤ 0.05). (TIF 912 KB)
Relatively slow movement of lipid droplets in MEC treated with oleic acid + NaN3 + NaF. Time-lapse imaging of Nile red-stained lipid droplets in MEC treated with 360 μM oleic acid + 2.5 mM NaN3 + 10 mM NaF reveals slower movement relative to oleic acid alone (see Online Resource <link rid="Sec22">5</link>). (AVI 199 KB)
Lipid droplet movement in MEC treated with oleic acid. Time-lapse imaging of Nile red-stained lipid droplets in MEC treated with 360 μM oleic acid reveals rapid movement and noticeable fusion between lipid droplets. (AVI 286 KB)
10911_2017_9386_MOESM6_ESM.tif
Differential responsiveness of MEC cultures to similar oleic acid concentration. Different batches of MEC primary culture were treated with 100 μM free oleic acid for 24h. Culture #1 showed a much greater number of large lipid droplets than culture #2. The largest droplet measured in each culture (2 replicates, 30 cells per replicate) was 5.0 μm and 3.7 1 μm in culture #1 and culture #2 respectively. The average diameter of the 3 largest lipid droplets was 3.4±0.7 μm and 2.8±0.3 in culture #1 and culture #2 respectively. Neutral lipids were stained with Nile red (red) and nuclei were stained with DAPI (blue). Scale bars, 20 μm. (TIF 1919 KB)
10911_2017_9386_MOESM8_ESM.pdf
PSD mRNA sequence and sequence amplified by real-time PCR. mRNA coding sequence of bovine PSD (NCBI reference sequence: NM_001024475.1, gi|66792777:137-1387) with forward and reverse primers marked in bold and underlined, and PCR product marked with light gray. PCR product sequence was verified by sequencing. (PDF 32 KB)
Rights and permissions
About this article
Cite this article
Cohen, BC., Raz, C., Shamay, A. et al. Lipid Droplet Fusion in Mammary Epithelial Cells is Regulated by Phosphatidylethanolamine Metabolism. J Mammary Gland Biol Neoplasia 22, 235–249 (2017). https://doi.org/10.1007/s10911-017-9386-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10911-017-9386-7