Abstract
A novel metal complex was synthesized using freshly prepared 2-Amino-5-nitro-N-[(E)-thiophen-2-yl-methylidene]aniline ligand with Zn (II) sulphate heptahydrate in a 1:1 molar ratio. The ligand and the complex were characterized using different spectroscopic techniques, and the complex was assigned a distorted square pyramidal geometry. Additionally, DNA binding assays and antibacterial activity were used to assess the biological perspectives for the synthesized complex, including the ligand and complex which was further confirmed by molecular docking. Fluorescence Spectroscopy, viscosity measurement, and adsorption measurement were used to investigate the interaction of the Zn (II) complex with CT-DNA. A comparative in vitro antibacterial activity study against Escherichia coli, Klebsiella pneumoniae, Bacillus subtilis, and Staphylococcus aureus strains were studied with free ligand and Zn (II) metal complex. The stable geometry of the complex was additionally established through computational simulation utilizing density functional theory, which was followed by the calculation of several electronic properties. The ADMET characteristics of the complex and ligand were also assessed using ADMET analysis. The in-silico ADMET properties pointed to a significant drug-likeness feature in the synthesized compounds, based on the Lipinski criteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Schiff bases and their transition metal complexes are essential compounds with many exciting properties and extensive medicinal, agricultural, pharmaceutical, and material science applications. The Schiff base and its complexes have been reported for various biological activities such as antioxidant, anti-bacterial, anti-fungal, anti-tumor, etc. [1]. When Schiff base ligands combine with transition metal ions, they show different electronic, geometrical, and biological properties. This is because the ligand moiety is made up of different groups of atoms. Over time, ligands with nitrogen-sulfur donor sites and their derivatives have garnered significant attention because of the possibility of their donor site being available. For metal chelation, these donor atoms are readily accessible [2,3,4,5,6]. Owing to the site-specific binding properties of metal ions, DNA binding with metal complexes provides a new manifesto for developing new chemotherapeutic drugs [7]. The schiff base and its complexes, which are synthesized from a sequence of NS donor atoms, have demonstrated an amazing level of activity against some antifungal and antibacterial agents [8, 9]. Various analogs are found to be capable of binding to DNA, hilting its replication and apoptosis [10, 11].
The study shows that when a ligand is coupled to metal ions present in biological systems (Zn2 + , Cu2 + , Co2 + , Mn2 + , Ni2 + , Fe2 + , VO2 +), its biological activity is significantly enhanced [12]. Zinc is an essential element that plays a very important role in various biological processes. Their complexes bind strongly with DNA and also exhibit predominant anti-tumor activities. Due to its pivotal role in numerous physiological and pathological processes, it is responsible for cell proliferation, differentiation, and viability, including apoptosis [13].
Apart from choosing the transition metals, the design of the ligand framework is regularly crucial as it can modulate the oral or systemic bioavailability of metal ions, secure the selective target DNA or enzyme, and protect and deliver a metal ion at the target site [14, 15]. A slight change in the ligand framework can drastically improve the therapeutic potential of the metal complex via the uptake of pharmacologically active metal ions. In this case, Schiff base ligands work as a platform because they can bind to metals in a flexible way, have biological properties that can't be changed, and can change their structure to get the desired molecule. Therefore, the design of Schiff base-based metal complexes tethered with a heterocyclic moiety shows a multi-fold increase in biological properties [16, 17]. Computational simulation also helps in this regard to understand the stable geometry of the complex with the calculations of different electronic parameters.
Materials, Reagents and Methods
The chemicals thiophene-2-carbaldehyde and 4-nitro-o-phenylenediamine, CT-DNA, and Ethidium Bromide were purchased from Sigma Aldrich, Germany. Zinc(II) sulphate heptahydrate and solvents like methanol, dimethylsulphoxide were purchased from SD Fine Chemicals, India. Solvents were purified prior to their use following standard literature procedures.
Instruments
A Vario Micro Cube elemental analyzer instrument was used for the elemental analysis of ligand and complex. The 1H and 13C NMR spectra were recorded using a Bruker 400 MHz FT-NMR instrument (DMSO-d6 as solvent). A Jasco V-530 UV–Vis spectrophotometer was used to record the UV–Visible spectra of the ligand and its metal complex. A Perkin Elmer RX-1 FT-IR Spectrometer with IR range of 400–4000 cm−1 was used to record the IR spectra using KBr pellets. A Jasco V-530 Spectrophotometer was used to carry out fluorescence emission experiments at room temperature. A Jasco V-530 double beam UV–Vis spectrophotometer was used to study the thermal denaturation of DNA. Viscosity was studied using an Ostwald viscometer maintained at constant room temperature. A Systronics India conductivity meter TDS-308 was used to measure specific conductance of synthesized complex in DMSO solvent at room temperature. Thin-Layer Chromatography was used to affirm the purity of ligand and its complex. Finally, an open-capillary technique was used to determine the melting point of the ligand and its complex.
Synthesis of 2-Amino-5-Nitro-N-[(E)-Thiophen-2-yl-Methylidene]Aniline Schiff Base
4-nitro-o-phenylenediamine (1 mmol, 0.153 g) and thiophene-2-carbadehyde (1 mmol, 0.112 g) were dissolved in ethanol and refluxed for 2 h with constant stirring. The yellow color precipitate was obtained after 2 h of reaction, was filtered using Whatman 41 filter paper, and was washed several times with cold ethanol. The product was kept in vacuum desiccators for 24 h.
Synthesis of [{(2-Amino-5-Nitro-N-[(E)-Thiophen-2-yl-Methylidene]Aniline-κ3N1:N4:S)(Sulphato-κ2O1:O3)}Zinc(II)]
A Zn (II) Sulphate Heptahydrate (0.1 mmol, 0.287 g) was dissolved in ethanol and was added dropwise to the equimolar hot ethanolic solution of the ligand (0.1 mmol, 0.247 g) in a two neck round bottom flask. The reaction was kept under refluxed condition for 10 h at 45–50 °C. The change of the color from yellow to dark yellow was noticeable during the reaction and consequently it resulted in the dark yellow precipitate. The precipitate was filtered using whatman 41 filter paper and was washed thoroughly using cold ethanol three times. The product was collected and kept in vacuum desiccators for 36 h.
Computational Study
Computational simulation using density functional theory (DFT) of the synthesized complex was performed in an aqueous medium with Gaussian 09 software [18, 19]. B3LYP hybrid functional [20] with 6-31 g(d,p) basis set [21] was used in an ethanol medium. The frequency calculations of the optimized geometry of the complex indicated that the geometry lies on the minima to the potential energy surface [22, 23] as no imaginary frequencies were obtained.Different electrochemical parameters like Mulliken Charge Distribution, HOMO–LUMO, and Molecular Electrostatic Potential (ESP) maps were also analysed to understand the characteristics of the synthesized complex [24, 25].
DNA Binding Experiment
Absorption Study
The absorption study is the best technique to study the interaction of the complex with the CT-DNA. As per the literature, the Tris–HCl buffer was prepared and used during the study. The purity of the CT-DNA was confirmed as it showed absorbance peaks at 260 nm, which further indicated that the CT-DNA was free of any contamination [26]. During the absorbance study, the concentration of the complex was fixed at 10 µM, and to prevent the absorption of free CT-DNA, an equal amount of CT-DNA was added to both the reference and solution chamber. Therefore, the spectra obtained were only due to the interaction of CT-DNA with the complex. The Wolfe-Shimer equation was used to calculate the intrinsic binding constant (kb) [27,28,29].
The data obtained was used for plotting the [DNA]/(\(\upvarepsilon\)a-\(\upvarepsilon\)f) versus [DNA] plot and calculating the intrinsic binding constant. From the above equation \(\upvarepsilon\)a, \(\upvarepsilon\)f and \(\upvarepsilon\)b denotes the extinction coefficients observed for charge transfer, complex free in solution, and complex fully bound to DNA, respectively. The intrinsic binding is calculated as per the literature, where kb is obtained as the ratio of slope to the intercept [30].
Fluorescence Emission Spectroscopy
The relative binding of the complex to CT-DNA is investigated using an Ethidium bromide CT-DNA solution. Ethidium bromide is widely used as a fluorescent DNA structural probe for studying the mode of binding of complexes to DNA. A 3 ml solution of 2.0 µM DNA and 1 × 10–5 M EB was titrated by 5–30 × 10–5 M compound (λex = 540 nm, λem = 540 – 680 nm). The obtained spectra were solved using the Stern–Volmer equation, I0/I = 1 + Ksvr, and the quenching constant Ksv was calculated. Here, I and I0 denote the fluorescence intensities in the presence and absence of the complex, respectively and ‘r’ is the concentration of the complex [31].
Thermal Denaturation Study
A thermal denaturation investigation of DNA was studied using a temperature-control programmer spectrometer, where the temperature was set to elevate by 5 °C min−1. The absorbance of DNA in both the absence and presence of complex was observed [28, 30, 31].
Viscosity Measurements
The viscosity measurements of the complex were studied by gradually raising the concentration of complex from 10 to 100 µM keeping the concentration of CT-DNA fixed at 100 µM at a temperature of 25 ± 1 °C. The flow time for each measurement was recorded three times with the help of a stopwatch, and the average time was computed. The values were calculated using the relation ƞ = (t-to)/to, where t and to are the flow times of CT-DNA containing complex and buffer respectively. The data were plotted as (ƞ/ƞo)1/3 verus 1/R, where ƞ and ƞo are the viscosity of DNA in absence and presence of complex and R is equal to the ratio of concentration of DNA to the concentration of complex (R = [DNA]/[complex]) [31,32,33].
DNA Cleavage Study
Agar gel electrophoresis was used to detect the ability of the ligand and its metal complex to cleave supercoiled pBR322 plasmid DNA in the presence and absence of the oxidant H2O2. DNA (20 µM) and H2O2 (200 µM) concentrations were held constant in each reaction mixture, while the concentration of the complex (5–15 µM) was varied [30]. After letting the samples sit for two hours at 37 °C, 1 µL of loading buffer (Bromophenol Blue in H2O) was added to each one. The samples were then carefully loaded onto the 0.8% agarose gel electrophoresis chamber wells, along with a control sample that was just DNA. In a dark chamber for two hours at 50 V, a tris–acetic acid-EDTA buffer with a pH of 8.3 was used for the electrophoresis. The gel was photographed under UV light (280 nm) following electrophoresis [34].
Antimicrobial Activity
Using the disk diffusion technique, the antimicrobial activity of the ligand and its corresponding metal complex was evaluated (in vitro) for antibacterial activity in accordance with a previously published methodology [35]. During the course of the study, two Gram ( +) (Bacillus subtilis and Staphylococcus aureus) and two Gram (-) (Escherichia coli and Klebsiella pneumoniae) bacterial strains were examined. From Hi-media laboratory Pvt Ltd, Mumbai, India, the Nutritional Agar (NA) medium was procured. 100 ml of distilled water were used to suspend 2.8 g of NA, which was then brought to a full boil. Poured into a sterile Petri dish and allowed to set, it was autoclaved for 15 min at 15 lbs of pressure (121 ºC) to sterilize it. Sterility check was done after the agar was solidified [36]. Each bacterium's pure cultures were prepared in nutrient broth and incubated for the entire night at 37 ºC. Following incubation, a non-toxic, sterile cotton swab was dipped into the microbial growth to inoculate each of the necessary plates. By firmly pressing and rotating the swab against the tube's wall above the liquid's level, extra inoculate was eliminated. The swab was equally streaked across the plate's whole surface in three different directions to inoculate the media. The ligand and complex test solutions were prepared in DMSO at concentrations of 5, 7.5, and 10 mg/mL. The ligand and complex test solutions were prepared in DMSO at concentrations of 5, 7.5, and 10 mg/mL. The agar well diffusion method was used to conduct the assay for antibacterial activity. After the test organisms were inoculated into sterile Petri dishes, sterile nutritional agar (NA) was added. Each Petri plate was thoroughly prepared (9 mm) using a sterile cork borer. Using a sterile pipette, about 200 μl of a synthetic chemical (10 mg/ml) was added to each well (microorganism-inoculated plates), and the mixture was then incubated for 24 h at 37 ºC. Following incubation, the widths of the inhibitory zones were measured and all tested plates were inspected. Every experiment was run in three duplicates. There was a control, DMSO. The standard deviation of the mean (SD) is used to express the zone of inhibitions as mean values [37].
Molecular Docking
Auto Dock Vina programe (version 1.1.2) [38] was used to perform the molecular docking of complex with the selected protein PDB ID (https://www.rcsb.org). The receptor structure was defined as rigid and the grid dimensions varied from being 44, 50 and 80 for X,Y and Z dimensions for protein having PDB ID’s 1BNA with 1 Å spacing. Similarly, the dimensions for X, Y and Z for 5ZH8 was 62, 58 and 60, 2GCX was 40, 44 and 40, 2BH0 was 56, 54 and 40, 1XFF was 40, 46 and 48, respectively with 1 Å spacing.
Results and Discussion
The ligand and Zn (II) complex are characterized by different analytical and spectroscopic techniques (FT-IR, 1H and 13C NMR, UV–Vis, Molar conductance, SC-XRD, PXRD, and Mass Spectrometry) and their formation is depicted in scheme 1.
The analytical data of compounds (both ligand and complex) and physical properties are listed in Table 1. Both compounds are soluble in Methanol and DMSO. All the analytical data are consistent with the calculated value and refer to forming a mononuclear Zn (II) complex with a 1:1 molar ratio [30]. The molar conductance of the complex was measured, and the value indicates the non-electrolytic nature (Table 1) [39].
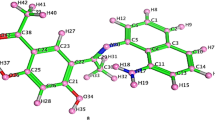
It was possible to obtain the Ligand's Single Crystal appropriate for Single Crystal XRD. The CCDC/Deposit no. assigned to the CIF file upon deposit was 2280507. Regretfully, the crystal results agreed with Geiger et al.'s earlier research [40] which they have only published with the crystal structure. Thus, this work has covered every detail, explanation, and application pertaining to the ligand and its Zn (II) complex.
Spectroscopic Characterization
Infrared Spectroscopy
The ligand and complex's IR spectra were acquired using the KBr pellet method in the 4000–400 cm−1 spectral region, as shown in Fig. S1. This information gave structural details necessary for the complex's formation.
The schiff base ligand's azomethine v(C = N) group has a distinctively strong band at 1621 cm−1 [41]. The band shifts to 1603 cm−1 in the case of the Zn (II) complex. This suggested that azomethine N had a role in the formation of complex [39]. Furthermore, the ligand displayed a distinctive band of N–H stretching (-NH2 group) at 3453 cm−1 and 3339 cm−1 [42], which are present in the schiff base ligand. The presence of the sulfato group in the metal complex is what causes the emergence of a new medium band at 1141–1096 cm–1 [43]. The M–N bond is responsible for the 471 cm−1 in the ML complex, which is absent from the spectra of the free ligand. The new M-S bond in the synthesized complex ML may be responsible for a second band at 630 cm−1.
ESI-mass Spectrometry
The mass spectra of the ligand and complex are shown in Fig. S2a and b respectively, where the peak [M +] at m/z = 247.30 precisely matches the predicted ligand's computed value, which has the chemical formula C11H9N3O2S. The mass spectrum of the complex is also shown in Fig. 2, where the molecular weight of the Zn (II) complex C11H9N3O6S2Zn is confirmed by the peak [M +] at m/z = 408.45. The other peaks, which correspond to the other segments—C11H11N3O4SZn, C11H9N3O2SZn, and C11H9N3O2S—are located at m/z = 342.33, 312.46, and 247.30, respectively. Therefore, we may infer that the expected ligand and complex have been synthesized based on the total mass spectrum of both the ligand and the complex.
1H and 13C NMR Spectral Analysis
The 1H and 13C NMR of the ligand and complex are shown in figs. S3a, b and S4a , b respectively. The Schiff base ligand's azomethine proton (-CH = N-) displayed a peak at 8.938 ppm [44, 45]. However, this azomethine proton peak is displaced to 8.792 ppm in the case of the Zn (II) complex. This could suggest that the azomethine N and Zn (II) ion are coordinated. The ligand also had a peak at 6.526 ppm, which might be the result of the ligand's –NH2 proton. But in the synthesized Zn (II) complex, this peak shifted to 5.971 ppm, suggesting that the -NH2 group was involved in complexation [46]. In the 13C NMR Spectra of the ligand, we observed a characteristic peak at 154.168 ppm and 151.164 ppm for the C atom of the azomethine group (-CH = N-) and –NH2 group respectively. While the complex shows a clear shift in the spectra (149.778 ppm and 148.561 ppm).
Electronic Absorption Study
The UV–Vis absorption spectra of the ligand and its associated complex are shown in Fig. S5. It was measured in DMSO solvent (1 × 10–5 M) at ambient temperature. Two distinct bands were visible in the ligand's electronic absorption spectra at 296 nm and 354 nm. The phenyl ring's π-π* transitions are responsible for the higher energy band at 296 nm, while the Schiff base's azomethine (-CH = N) group's n-π* transitions are responsible for the other lower energy band at 354 nm [46, 47]. The complex's spectra similarly exhibits same absorption bands, however they experience a blue shift (hypochromic shift). This change in the spectra is explained by the complex that forms with the Zn (II) ion. The spectra lack the d-d transition due to its d10 configuration. Consequently, it suggests that the compound has a diamagnetic nature.
Powder X-Ray Diffraction
The phase purity of the ligand and complex is confirmed using powder X-ray diffraction technique. PXRD of the ligand and complex was recorded at 2θ = 5–70° range (at wavelength 1.54 Å), as shown in Fig. S6. The sharp peaks in the case of ligand indicate the crystalline nature which was missing in the case of complex.
Study of the Electronic Parameters from Computational Simulation
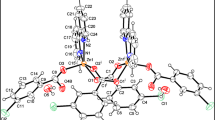
Figure 1 represents the optimized structure of the synthesized complex in the ethanol medium. Bond distance measurements clearly represents that Zn is stabilized by the two ‘O’ and two ‘N’ atoms. Interaction of ‘S’ with Zn (3.5434 Å) is comparatively weaker. DFT study clearly indicated that the complex is stable enough.
Mulliken Charges Analysis
Mulliken charge distribution analysis helps to understand the charge accumulation on each and every atom during complex formation. Charge distribution in the synthesized complex is represented by Fig. 2(a) where the red end indicates high negative charge accumulation and the light green end indicates high positive charge accumulation. Mulliken charge analysis indicates that the positive charge accumulated on the Zn metal and it is well attached with negative charge accumulated O. Charge distribution reflects major interaction present in between ‘Zn’ and ‘O’ atom and ‘S’ atom did not contribute much more here, also reflected from bond distance values of Fig. 1.HOMO–LUMO energy gap [Fig. 2(b)] of the complex is quite small (3.36 eV) [21]. ESP map [Fig. 2(c)] indicated the surface charge of the complex. The combination of negative surface (reddish) and positive surface (bluish) well supports the Mulliken charge analysis study.
HOMO–LUMO and ESP Analysis
HOMO–LUMO energy gap of the complex is represented by Fig. 2(b). As per the calculation, HOMO–LUMO energy gap of the complex is quite small (3.36 eV) [21]. A low band gap of the complex can increase its application in the field of light sensitivity, catalysis, and optoelectronics. ESP map [Fig. 2(c)] indicates the surface charge of the complex. Sulphate ends reflects the negative surface (reddish) which combined with the Positive surface (bluish) of Zn. Combination of surface charges indicates the stability of the complex and this observation well supports the Mulliken charge analysis study.
DNA Binding study
Absorption Study
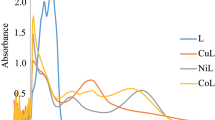
All metal complexes with DNA have their binding properties determined using electronic absorption spectroscopy. Variations in wavelength and absorbance are frequently used to study how DNA binds to metal complexes. When an incremental concentration of CT-DNA solution was added to a fixed concentration of metal complex solutions, a decrease in absorption intensity (hypochromism) at 354 nm was observed. Hypochromism, which denotes a strong contact between the electronic states of the chromophore and the DNA bases, is commonly observed when a complex binds to DNA by intercalation because of a strong bond between an aromatic chromophore and a base pair of DNA [48, 49]. The Wolfe-Shimer equation was used to calculate the intrinsic binding constant Kb, and the value of Kb was 2.7 × 104 M−1 for complex (Fig. 3).
EB-Competitive Study with Fluorescence Emission Spectroscopy
The Zn (II) complex's DNA binding mechanism with CT-DNA was investigated using ethidium bromide (EB) displacement. The use of the planar cationic dye EB as a susceptible fluorescence molecule for inherent DNA is well known [7]. The planar phenanthridine ring of EB intercalates between neighboring DNA base pairs, resulting in the EB-DNA conjugate having a prominent emission band at 559 nm [30]. DNA intercalating probes are compounds that change the EB-DNA system's fluorescence intensity. Regardless of the binding modes, they provide a precise tool for assessing the complex's affinity for binding DNA [7]. The synthesized Zn (II) complex does not show an emission band in solution, CT-DNA, or EB when stimulated at 540 nm. Therefore, it is essential to examine the observed changes in the emission spectra of the EB-DNA solution with the introduction of complex in order to determine the EB displacement capability of the synthesized complex [49]. Figure 4 shows the anticipated emission band at 559 nm upon the addition of increasing quantities of complex to CT-DNA that had previously been treated with EB solution. The calculated quenching constant (Ksv) value for the synthesized compound is 4.5(± 0.2) × 103 M−1 (Fig. 4).
Thermal Denaturation Study
Thermal denaturation studies of DNA provide additional insight into the mode of binding and binding strength of complexes into the DNA helix by providing information on the conformational alteration as the temperature is increased [50]. The thermal denaturation profile of the CT-DNA solution with and without the complex is shown in Fig. 5. The melting temperature (Tm) of the CT-DNA solution was 80 °C; however, the Tm value rose sharply to 85 °C upon the addition of complex. When complex is present, CT-DNA melts at a higher temperature (Tm = 5 °C), which supports complex intercalative binding to DNA [51].
Viscosity Measurement
Viscosity measurements were used to investigate the complex's interaction with DNA. The spacing between the DNA base pairs existing at the intercalation site to host the binding molecule is frequently increased when DNA is intercalated with metal complexes. Consequently, the DNA helix lengthens, raising the viscosity of the DNA [52, 53]. The DNA helix lengthened and became more viscous as base pairs were split apart to create space for the binding complex in a classical interaction mode. Conversely, a partial or unconventional intercalation might bind to the DNA helix and shorten it, reducing its effective length and viscosity [54, 55]. Figure 6 illustrates how the relative viscosity of the DNA solution continuously increases as the complex concentration in the CT-DNA solution rises. The UV–Vis and fluorescence spectrum data are consistent with the increased viscosity of the complex solution, suggesting that the synthesized complex may bind to DNA via intercalation mode.
DNA Cleavage Study
DNA endonucleolytic cleavage processes that are initiated by metal ions are of great and increasing significance [56, 57]. The cleavage reaction on plasmid DNA (pBR322 DNA) can be seen via agarose gel electrophoresis. DNA can be broken down by the complex into its linear and nicked forms, as well as back into its super-coiled form. The intact supercoiled form of circular plasmid DNA will migrate quite quickly when electrophoresed. If one strand is scissile (nicking), the super coil will relax creating an open circular form that moves more slowly. If both strands split, a linear form that moves between Forms 1 and 2 will be produced [58] as shown in Fig. 7. The cleavage effect upon irradiation of the plasmid pBR322 DNA in the presence of different concentration of complex. In figure Lane 1 DNA alone, Lane 2 DNA + H2O2, Lane 3 DNA + ligand + H2O2 and Lane 4–6 DNA + Zn(II) complex + H2O2 with increasing concentration of complex. As the complex's concentration rises, Form 1 progressively declines and Form 2 steadily increases. The results point to concentration-dependent single-strand cleavage of the super-coiled Form1 to the nicked Form 2, according to the study. The varied binding affininty of the complex to DNA may be the cause of the varying cleaving efficacy.
Antibacterial Activity
Using two gram positive (Staphylococcus aureus, Bacillus subtilis) and gram negative (Escherichia coli, Klebsiella pneumonia) bacteria, the antibacterial activity of the ligand and complex was evaluated. It was highly anticipated that the ligand and complex would exhibit good action against bacteria, as seen in Fig. 8 and Table 2, given the presence of biologically active donor sites (N and S). It was discovered that the metal complex and ligand were very effective against Escherichia coli, with high inhibition zone diameters of 14 mm and 11 mm, respectively. Regarding other bacterial strains, Klebsiella pneumonia, Bacillus subtilis, and Staphylococcus aureus exhibit descending patterns in the sequence of activity of the ligand and complex. A prior article examined the effectiveness of standard drugs against several strains of bacteria. The findings indicated that minor modifications in ligand and complex could potentially improve the antibacterial activity [59, 60].
Molecular Docking
Using the Auto Dock Vina program, the complex was tested for interaction with various bacteria and DNA receptors (Fig. 9 and Table 3). Our earlier statements are further confirmed by the complex's binding affinity of -7.7 kcal/mol with protein (1BNA), which suggests that the complex interacts with DNA. Further molecular docking of the complex was done using the receptor proteins of Escherichia coli (1XFF), Bacillus subtilis (2BH0), Klebsiella pneumoniae (2GCX), and Staphylococcus aureus (5ZH8). The binding affinities were reported to be -8.0, -6.3, -6.7, and -7.6 correspondingly. Thus, the total finding suggests that the complex exhibits robust interaction with many bacterial protein receptors, claiming the previously mentioned antibacterial activity.
Pharmacokinetic Properties
An in-silico theoretical ADME prediction study has been conducted on a Schiff base and its complex. For this, the web application Swiss ADME was utilized. The bioavailability score was calculated by estimating several factors using Swiss ADME software. Important pharmacokinetic factors for any chemical include its molecular weight, hydrogen donors, hydrogen acceptors, rotatable bonds, lipophilicity (Log PO/W), gastrointestinal absorption (GI), water soluble capacity (Log S), CYP1A2 inhibitor, and Blood–Brain Barrier (BBB) [61]. From Table 4, it is evident that the compounds (Ligand and complex) with bioavailability 55% have consensus liphophilicity (Log PO/W) value 1.79 and 1.24 for ligand andcomplex respectively. It is evident that the Ligand and the complex meet the requirements for druglikness and do not violate the Lipinski rule, making them viable candidates for use as drugs.
Conclusion
This study reports the synthesis and characterization of a Zn (II) complex with Schiff base ligand obtained from 4-nitro-o-phenylenediamine and thiophene-2-carbadehyde. The physical measurements' result shows that the Zn (II) metal ion is coordinated by two oxygen of the sulphate ion, one azo methine nitrogen, and one sulfur atom of the thiophene group; as a result, it may adopts a distorted square pyramidal geometry. The synthesized complex has no electrolytic properties and is stable. The experiment's outcome shows that the complex exhibits remarkable interactions with CT-DNA and validates an intercalative method of binding. DFT studies clearly reflected the stabile geometry of the complex from the calculations of different electronic parameters. The pharmacokinetics study has revealed that the Ligand and Complex could act as potential drug candidate. The antibacterial activity and molecular docking reports reveals that the complex has high activity over selected bacteria.
Data Availability
All data generated or analyzed during this study are included in the manuscript.
References
Ashraf T, Ali B, Qayyum H, Haroone MS, Shabbir G (2023) Pharmacological aspects of schiff base metal complexes: A critical review. Inorg Chem Commun 150:110449
Akbar Ali M, Huq Mirza A, Nazimuddin M, Rahman H, Butcher RJ (2002) The preparation and characterization of mono- and bis-chelated cadmium(II) complexes of the di-2-pyridylketone Schiff base of S-methyldithiocarbazate (Hdpksme) and the X-ray crystal structure of the |Cd(dpksme)2|· 0.5MeOH complex. Transit Met Chem 27:268–273
Zangrando E, Islam MT, Islam MAAAA, Sheikh MC, Tarafder MTH, Miyatake R, Zahan R, Hossain MA (2015) Synthesis, characterization and bio-activity of nickel(II) and copper(II) complexes of a bidentate NS Schiff base of S-benzyl dithiocarbazate. Inorganica Chim Acta 427:278–284
Pereira GA, Massabni AC, Castellano EE, Costa LAS, Leite CQF, Pavan FR, Cuin A (2012) A broad study of two new promising antimycobacterial drugs: Ag(I) and Au(I) complexes with 2-(2-thienyl)benzothiazole. Polyhedron 38:291–296
Ylldlz M, Tan E, Demir N, Ylldlrlm N, Ünver H, Kiraz A, Mestav B (2015) Synthesis and spectral, antimicrobial, anion sensing, and DNA binding properties of Schiff base podands and their metal complexes. Russ J Gen Chem 85:2149–2162
Latif MA, Ahmed T, Hossain MS, Chaki BM, Abdou A, Kudrat-E-Zahan M (2023) Synthesis, Spectroscopic Characterization, DFT Calculations, Antibacterial Activity, and Molecular Docking Analysis of Ni(II), Zn(II), Sb(III), and U(VI) Metal Complexes Derived from a Nitrogen-Sulfur Schiff Base. Russ J Gen Chem 93:389–397
Aazam ES, Zaki M (2020) Synthesis and Characterization of Ni(II)/Zn(II) Metal Complexes Derived from Schiff Base and Ortho-Phenylenediamine: In vitro DNA Binding, Molecular Modeling and RBC Hemolysis. ChemistrySelect 5:610–618
Kudrat-E-Zahan M, Islam MS (2015) Synthesis, characterization, and antimicrobial activity of complexes of Cu(II), Ni(II), Zn(II), Pb(II), Co(II), Mn(II), and U(VI) containing bidentate Schiff base of [S-methyl-3-(4-methoxybenzylidine)dithiocarbazate]. Russ J Gen Chem 85:979–983
Hossain MS, Khushy KA, Latif MA, Hossen MF, Asraf MA, Kudrat-E-Zahan M, Abdou A (2022) Co(II), Ni(II), and Cu(II) Complexes Containing Isatin-Based Schiff Base Ligand: Synthesis, Physicochemical Characterization, DFT Calculations, Antibacterial Activity, and Molecular Docking Analysis. Russ J Gen Chem 92:2723–2733
Rosenberg B, Vancamp L, Trosko JE, Mansour VH (1969) Platinum compounds: a new class of potent antitumour agents. Nature 222:385–386
Rodríguez MR, Lavecchia MJ, Parajón-Costa BS, González-Baró AC, González-Baró MR, Cattáneo ER (2021) DNA cleavage mechanism by metal complexes of Cu(II), Zn(II) and VO(IV) with a schiff-base ligand. Biochimie 186:43–50
Omer MAS, Liu JC, Deng WT, Jin NZ (2014) Syntheses, crystal structures and antioxidant properties of four complexes derived from a new Schiff base ligand (N1E, N2E)-N 1, N2-bis(1-(pyrazin-2-yl)ethylidene)ethane-1,2 diamine. Polyhedron 69:10–14
Ozturk H, Niazi P, Mansoor M, Monib AW, Alikhail M, Azizi A (2023) The Function of Zinc in Animal, Plant, and Human Nutrition. J Res Appl Sci Biotechnol 2:35–43
Boyle KM, Barton JK (2016) Targeting DNA mismatches with rhodium metalloinsertors. Inorganica Chim Acta 452:3–11
Thompson KH, Orvig C (2003) Boon and bane of metal ions in medicine. Science 300:936–939
Kaushik S, Paliwal SK, Iyer MR, Patil VM (2023) Promising Schiff bases in antiviral drug design and discovery. Med Chem Res 1–14
Kumar B, Devi J, Manuja A (2023) Synthesis, structure elucidation, antioxidant, antimicrobial, anti-inflammatory and molecular docking studies of transition metal(II) complexes derived from heterocyclic Schiff base ligands. Res Chem Intermed 49:2455–2493
Sieffert N, Wipff G (2015) Uranyl extraction by N, N-dialkylamide ligands studied using static and dynamic DFT simulations. Dalt Trans 44:2623–2638
Riaz S, Jaffar K, Perveen M, Riaz A, Nazir S, Iqbal J (2021) Computational study of therapeutic potential of phosphorene as a nano-carrier for drug delivery of nebivolol for the prohibition of cardiovascular diseases: a DFT study. J Mol Model 27:1–15. https://doi.org/10.1007/s00894-021-04907-w
Becke AD (1992) Densityâ-functional thermochemistry. I. The effect of the exchangeâ-only gradient correction. J Chem Phys 96:2155–2160
Ghosh K, Mridha NK, Khan AA, Baildya N, Dutta T, Biswas K, Ghosh NN (2022) CO2 activation on transition metal decorated graphene quantum dots: An insight from first principles. Phys E Low-dimensional Syst Nanostruct 135:114993
Baildya N, Mazumdar S, Mridha NK, Chattopadhyay AP, Khan AA, Dutta T, Mandal M, Chowdhury SK, Reza R, Ghosh NN (2023) Comparative study of the efficiency of silicon carbide, boron nitride and carbon nanotube to deliver cancerous drug, azacitidine: A DFT study. Comput Biol Med. https://doi.org/10.1016/j.compbiomed.2023.106593
Dutta T, Chowdhury SK, Ghosh NN, Chattopadhyay AP, Das M, Mandal V (2022) Green synthesis of antimicrobial silver nanoparticles using fruit extract of Glycosmis pentaphylla and its theoretical explanations. J Mol Struct. 1247:131361. https://doi.org/10.1016/j.molstruc.2021.131361
Contreras-García J, Johnson ER, Keinan S, Chaudret R, Piquemal JP, Beratan DN, Yang W (2011) NCIPLOT: A program for plotting noncovalent interaction regions. J Chem Theory Comput 7:625–632
Boto RA, Peccati F, Laplaza R, Quan C, Carbone A, Piquemal JP, Maday Y, Contreras-Garcĺa J (2020) NCIPLOT4: Fast, Robust, and Quantitative Analysis of Noncovalent Interactions. J Chem Theory Comput 16:4150–4158
Kallur M, Chandraprabha MN, Rajan HK, Khosla A, Manjunatha CJET (2022) Synthesis, Characterization of Cerium Oxide Nanoparticles and Evaluation of DNA Binding Interactions. ECS Trans 107:15935–15943
Wolfe A, Shimer GH, Meehan T (1987) Polycyclic Aromatic Hydrocarbons Physically Intercalate into Duplex Regions of Denatured DNA. Biochemistry 26:6392–6396
Singha UK, Pradhan S, Mishra DK, Gurung P, Chettri A, Sinha B (2023) Synthesis, physicochemical characterisation and DNA binding study of a novel azo Schiff base Ni(II) complex. Eur J Chem 14:280–286
Sabolová D, Sovová S, Janovec L, Timko E, Jager D, Tóthová J (2023) Synthesis, characterization and DNA binding properties of 3,6-bis[(alkylamino)ureido]acridines. Chem Pap 77:3155–3162
Mishra DK, Singha K, Das A, Dutta S, Kar P, Chakraborty A, Sen A, Sinha B, Kumar Mishra D, Kumar Singha U (2018) Journal of Coordination Chemistry DNA Binding, amelioration of oxidative stress, and molecular docking study of Zn(II) metal complex of a new Schiff base ligand DNA Binding, amelioration of oxidative stress, and molecular docking study of Zn(II) metal complex of a new Schiff base ligand. J Coord Chem 71:2165–2182
Sunita M, Anupama B, Ushaiah B, Gyana Kumari C (2017) Synthesis, characterization, DNA binding and cleavage studies of mixed-ligand copper (II) complexes. Arab J Chem 10:S3367–S3374
Chaires JB, Dattagupta N, Crothers DM (1982) Studies on Interaction of Anthracycline Antibiotics and Deoxyribonucleic Acid: Equilibrium Binding Studies on Interaction of Daunomycin with Deoxyribonucleic Acid. Biochemistry 21:3933–3940
Shahabadi N, Razlansari M (2022) Insight into the binding mechanism of macrolide antibiotic; erythromycin to calf thymus DNA by multispectroscopic and computational approaches. J Biomol Struct Dyn 40:6171–6182
Richter S, Singh S, Draca D, Kate A, Kumbhar A, Kumbhar AS, Maksimovic-Ivanic D, Mijatovic S, Lönnecke P, Hey-Hawkins E (2016) Antiproliferative activity of ruthenium(II) arene complexes with mono- and bidentate pyridine-based ligands. Dalt Trans 45:13114–13125
Sumrra SH, Atif AH, Zafar MN, Khalid M, Tahir MN, Nazar MF, Nadeem MA, Braga AAC (2018) Synthesis, crystal structure, spectral and DFT studies of potent isatin derived metal complexes. J Mol Struct 1166:110–120
Hassan AU, Sumrra SH, Zafar MN, Nazar MF, Mughal EU, Zafar MN, Iqbal M (2022) New organosulfur metallic compounds as potent drugs: synthesis, molecular modeling, spectral, antimicrobial, drug likeness and DFT analysis. Mol Divers 26:51–72
Hassan AU, Sumrra SH, Imran M, Chohan ZH (2022) New 3d multifunctional metal chelates of sulfonamide: Spectral, vibrational, molecular modeling, DFT, medicinal and in silico studies. J Mol Struct. https://doi.org/10.1016/j.molstruc.2021.132305
Trott O, Olson AJ (2010) AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Abdallah SM, Mohamed GG, Zayed MA, El-Ela MSA (2009) Spectroscopic study of molecular structures of novel Schiff base derived from o-phthaldehyde and 2-aminophenol and its coordination compounds together with their biological activity. Spectrochim Acta Part A Mol Biomol Spectrosc 73:833–840
Geiger DK, Cristina Geiger H, Donohoe JS (2012) 2-Amino-5-nitro-N-[(E)-thiophen-2-yl-methylidene]aniline. Acta Crystallogr Sect E Struct Rep Online 68(10):o2867–o2867. https://doi.org/10.1107/S1600536812037464
Khalil EAM, Mohamed GG (2022) Preparation, spectroscopic characterization and antitumor-antimicrobial studies of some Schiff base transition and inner transition mixed ligand complexes. J Mol Struct. 1249:131612. https://doi.org/10.1016/j.molstruc.2021.131612
Sayed FN, Mohamed GG, Deghadi RG (2023) Structural characterization and molecular docking studies of biologically active platinum(II) and palladium(II) complexes of ferrocenyl Schiff bases. J Mol Struct. https://doi.org/10.1016/j.molstruc.2023.134904
Sellmann D (1971) K. Nakamoto: Infrared Spectra of Inorganic and Coordination Compounds . Second Edition, John Wiley & Sons. New York, London, Sydney, Toronto, 1970. 338 Seiten, zahlreiche Abbildungen und Tabellen. Preis: 140s . Berichte der Bunsengesellschaft für Phys Chemie 75:603–604
Neelakantan MA, Rusalraj F, Dharmaraja J, Johnsonraja S, Jeyakumar T, Sankaranarayana Pillai M (2008) Spectral characterization, cyclic voltammetry, morphology, biological activities and DNA cleaving studies of amino acid Schiff base metal(II) complexes. Spectrochim Acta Part A Mol Biomol Spectrosc 71:1599–1609
Abd El-Lateef HM, Mohamad ADM, Shehata MR, Abu-Dief AM (2022) Targeted synthesis of two iron(III) tetradentate dibasic chelating Schiff base complexes toward inhibition of acidic induced steel corrosion: Empirical and DFT insights. Appl Organomet Chem. 36(7):e6718. https://doi.org/10.1002/aoc.6718
Paul MK, Singh YD, Singh NB, Sarkar U (2015) Emissive bis-salicylaldiminato Schiff base ligands and their zinc(II) complexes: Synthesis, photophysical properties, mesomorphism and DFT studies. J Mol Struct 1081:316–328
Singh A, Maiti SK, Gogoi HP, Barman P (2023) Purine-based Schiff base Co(II), Cu(II), and Zn(II) complexes: Synthesis, characterization, DFT calculations, DNA binding study, and molecular docking. Polyhedron 230:116244. https://doi.org/10.1016/j.poly.2022.116244
Wu JZ, Yuan L, Wu JF (2005) Synthesis and DNA binding of μ-[2,9-bis(2-imidazo[4,5-f][1,10]phenanthroline)-1,10-phenanthroline]bis[1,10-phenanthrolinecopper(II)]. J Inorg Biochem 99:2211–2216
Chen LM, Liu J, Chen JC, Tan CP, Shi S, Zheng KC, Ji LN (2008) Synthesis, characterization, DNA-binding and spectral properties of complexes [Ru(L)4(dppz)]2+ (L = Im and MeIm). J Inorg Biochem 102:330–341
Indumathy R, Weyhermüller T, Nair BU (2010) Biimidazole containing cobalt(iii) mixed ligand complexes: Crystal structure and photonuclease activity. Dalt Trans 39:2087–2097
Abdel-Rahman LH, El-Khatib RM, Nassr LAE, Abu-Dief AM (2013) Synthesis, physicochemical studies, embryos toxicity and DNA interaction of some new Iron(II) Schiff base amino acid complexes. J Mol Struct 1040:9–18
Tarui M, Doi M, Ishida T, Inoue M, Nakaike S, Kitamura K (1994) DNA-binding characterization of a novel anti-tumour benzo[a]phenazine derivative NC-182: Spectroscopic and viscometric studies. Biochem J 304:271–279
Ramdas K, Reddy CVR, Sireesha B (2022) Synthesis, Characterization, DNA Binding, Cleavage, Antibacterial, In vitro Anticancer and Molecular Docking Studies of Ni(II), Cu(II) and Zn(II) Complexes of 3,4,5-Trimethoxy-N-(3-Hydroxy-5-(Hydroxymethyl)-2-Methylpyridin-4-yl)methylene)Benzohydrazide. Arab J Sci Eng 47:407–418
Satyanarayana S, Dabrowiak JC, Chaires JB (1993) Tris(phenanthroline)ruthenium(II) Enantiomer Interactions with DNA: Mode and Specificity of Binding. Biochemistry 32:2573–2584
Nie Y, Dai Z, Fozia ZG, Jiang J, Xu X, Ying M, Wang Y, Hu Z, Xu H (2022) Comparative Studies on DNA-Binding Mechanisms between Enantiomers of a Polypyridyl Ruthenium(II) Complex. J Phys Chem B 126:4787–4798
Johnston DH, Glasgow KC, Thorp HH (1995) Electrochemical Measurement of the Solvent Accessibility of Nucleobases Using Electron Transfer between DNA and Metal Complexes. J Am Chem Soc 117:8933–8938
Hidalgo M, Rodriguez G, Kuhn JG, Brown T, Weiss G, MacGovren JP, Von Hoff DD, Rowinsky EK (1998) A phase I and pharmacological study of the glutamine antagonist acivicin with the amino acid solution aminosyn in patients with advanced solid malignancies. Clin Cancer Res 4:2763–2770
Barton JK, Raphael AL (1984) Photoactivated Stereospecific Cleavage of Double-Helical DNA by Cobalt(III) Complexes. J Am Chem Soc 106:2466–2468
Hassan AU, Sumrra SH (2022) Exploring the Bioactive Sites of New Sulfonamide Metal Chelates for Multi-Drug Resistance: An Experimental Versus Theoretical Design. J Inorg Organomet Polym Mater 32:513–535
Behrami A (2014) Antibacterial activity of coumarine derivatives synthesized from 4-Chloro-chromen-2-one. The comparison with standard drug. Orient J Chem 30:1747–1752
Amoa Onguéné P, Ntie-Kang F, Lifongo LL, Ndom JC, Sippl W, Mbaze LMA (2013) The potential of anti-malarial compounds derived from African medicinal plants, part I: A pharmacological evaluation of alkaloids and terpenoids. Malar J 12:1–26. https://doi.org/10.1186/1475-2875-12-449
Acknowledgements
The authors would like to acknowledge Departmental Special Assistance Scheme under the University Grants Commission, New Delhi (SAP-DRS-III, No. 540/12/DRS/2013) & University of North Bengal, Govt. of West Bengal for financial and instrumental support.
Funding
This declaration is “not applicable”.
Author information
Authors and Affiliations
Contributions
Sudarshan Pradhan: Design and Synthesis, Analytical and Spectroscopic Data Analysis, Writing – original draft. Pritika Gurung: Analytical and Spectroscopic Data Analysis. Anmol Chettri: Analytical and Spectroscopic Data Analysis. Uttam Kumar Singha: Pharmacokinetic properties. Prajal Chhetri: Design and Synthesis. Tanmoy Dutta: DFT Study. Biswajit Sinha: Design and Synthesis, Analytical and Spectroscopic Data Analysis, Writing – final draft, Editing and Communication.
Corresponding author
Ethics declarations
Ethics Approval
This is an observational study. The University of North Bengal Ethics Committee has confirmed that no ethical approval is required.
Consent to Participate
This declaration is “not applicable”.
Competing Interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pradhan, S., Gurung, P., Chettri, A. et al. Synthesis of Novel [{(2-Amino-5-Nitro-N-[(E)-Thiophen-2-yl-Methylidene]Aniline-κ3N1:N4:S)(Sulphato-κ2O1:O3)}Zinc(II)] Complex with Physico-Chemical and Biological Perspective Exploration: A Combined Experimental and Computational Studies. J Fluoresc (2024). https://doi.org/10.1007/s10895-024-03612-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10895-024-03612-0