Abstract
Two new mononuclear mixed ligand copper(II) complexes [Cu(PBO)(Gly-gly)(H2O)]·ClO4·1.5H2O (1) and [Cu(PBO)(Gly-L-leu)(H2O)]·ClO4 (2) (PBO is 2-(2′-pyridyl)benzoxazole, Gly-gly and Gly-L-leu are Glycyl-glycine anion and Glycyl-L-leucine anion, respectively), have been prepared and characterized by various analytical and spectral techniques. The interactions of the complexes with DNA were investigated using multi-spectroscopic methods (absorption, emission, circular dichroism), viscometry and electrochemical titration as well as molecular docking technique. The results indicated that 1 and 2 are bound to calf thymus DNA (CT-DNA) through an intercalative mode. The thermodynamic analyses revealed that the reactions between the Cu(II) complexes with DNA are spontaneous with negative Gibbs free energy (ΔG). The positive changes of enthalpy (ΔH) and entropy (ΔS) suggested that the binding processes are dominated by hydrophobic interaction accompanying with endothermic. Also, the complexes exhibited efficient oxidative cleavage of pBR322 plasmid DNA in the presence of ascorbic acid, probably induced by •OH as reactive oxygen species. In addition, 1 and 2 displayed excellent antioxidant activities with the IC50 values of 0.112 and 0.191 μM, respectively, using the mean of nitroblue tetrazolium (NBT) photochemical reduction under a nonenzymatic condition. Moreover, the complexes were screened for their in vitro cytotoxicity against three human carcinoma cell lines (HeLa, PC-3 and A549), in which 2 owns higher cytotoxicity, which was consistent with DNA binding and cleavage ability order of the complexes. This results showed the in vitro biochemical potentials of the Cu(II)-dipeptide complexes with aromatic heterocyclic, viz. effective metallopeptide-nucleases, SOD mimics and non-platinum chemotherapeutic metallopharmaceuticals and their structure-activity relationship, which may contribute to the rational molecular design of new metallopeptide based chemotherapeutic agents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Recent years, cancer has shown an increased uptrend in its incidence and become one of the primary health killers [1, 2]. Meanwhile, considerable research suggested that DNA is the major target molecule for most antiviral and antitumor therapies, although there are other biological targets such as RNA, topoisomerase, or some protein [3, 4]. Additionally, DNA plays a crucial role in the life process, since it contains all genetic information required for cellular function [5, 6]. Therefore, the binding and cleavage studies of small molecules with DNA are extremely useful for illuminating the action mechanism of some DNA-target drugs. Furthermore, there have been considerable research focusing on the binding of compounds with DNA, but only limited thermodynamic analyses about the interaction of compounds to DNA have been performed. It is also important to study the thermodynamics of DNA-compound interaction so as to thoroughly understand the driving forces behind the binding of compounds with DNA.
Over the past few decades, the growing interest has been focused on transition metal complexes owing to their applications as therapeutic agents and their photochemical properties that make them potential probes of DNA structure [7]. Copper as an essential micronutrient for human has attracted numerous inorganic chemists to study its complexes [8]. Moreover, copper is a vital co-factor of several enzymes involved in oxidative metabolism, superoxide dismutase, tyrosinase, cytochromoxidase, ascorbic acid oxidase and ceruloplasmin etc. [9], and could also participate in many redox-related transformations as a redox-active metal [10]. Copper(II) complexes have been found to exhibit a wide range of biochemical activity in a variety of application areas [11–14]. Meanwhile, benzoxazoles and their derivatives are important fragments in medicinal chemistry because of their wide range of biological activities such as antibacterial, anticancer, anti-measles virus and anti-HIV activities [15–17]. Besides, dipeptides as the auxiliary ligand of the complexes could improve biological compatibility and recognition in the physiological system due to their structure similar to protein. In consideration of the possible synergistic effect, it is of great interest and significance to explore the photochemical properties and biochemical activities of the benzoxazoles derivatives based copper(II)-dipeptide complexes.
By considering the above facts in our mind, herein two new copper(II)-dipeptide complexes of 2-(2′-pyridyl)benzoxazole: [Cu(PBO)(Gly-gly)(H2O)]·ClO4·1.5H2O (1) and [Cu(PBO)(Gly-L-leu)(H2O)]·ClO4 (2) were synthesized and characterized using various spectroscopic methods. The DNA binding and cleavage properties were explored by multiple spectroscopic techniques (UV-vis, fluorescence, CD), viscometer titration, cyclic voltammetry (CV) and electrophoresis measurements as well as molecular docking study. The thermodynamic relationship of DNA-complex interaction was revealed employing fluorescence quenching measurements. Moreover, the antioxidant activities of the complexes were investigated using the photochemical reduction of nitroblue tetrazolium assays, and their cytotoxicity were evaluated by MTT assays.
Experimental
Materials and Instruments
All reagents were commercial quality and used without further purification. Gly-gly (Aladdin), Glycyl-L-leucine (Aladdin), pyridine-2-carboxylic acid (Aladdin), 2-aminophenol (Aladdin) and ethidium bromide (EthBr) (Sigma) were used as received. Calf thymus (CT)-DNA (Sigma) was stored at 4 °C and used as received. pBR322 DNA was obtained from MBI Fermentas (Lithuania). Nitroblue tetrazolium (NBT), riboflavin (VB2), N,N,N′,N′- tetramethylethyl-enediamine (TEMED) and 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT), were purchased from Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China). Human cancer cell lines of HeLa (cervical), A549 (pulmonary) and PC-3 (prostatic) were obtained from the Laboratory Animal Center of Sun Yat-Sen University. Deionized water was used throughout the experiments.
Microanalyses (C, H, and N) were carried out with a Vario EL elemental analyzer. UV-Vis absorption spectra were carried out by using a Shimadzu 2550 spectrophotometer using cuvettes of 1 cm path length. IR spectra were recorded, using KBr pellets, with a Bruker Vertex 70 FT-IR spectrophotometer in the range 4000–400 cm−1. The circular dichroism (CD) spectra were measured on a Chirascan CD spectropolarimeter and fluorescence spectra on a Hitachi F-4500 spectrophotometer. Electrospray ionization mass spectra were obtained on an API4000 triple quadrupole mass spectrometer. Electron-spin-resonance (ESR) spectra for the complexes were obtained on a Bruker EMX A300 spectrometer. Cyclic voltammetry measurements were performed on a CHI 660A Electrochemical Workstation. Molar conductance was measured on a DDS-11A digital conductometer.
Synthesis
Synthesis of [Cu(PBO)(Gly-gly)(H2O)]·ClO4·1.5H2O (1)
The ligand 2-(2′-pyridyl)benzoxazole (PBO) was prepared by the condensation of pyridine-2-carboxylic acid and 2-aminophenol as reported early [18]. 1 was achieved by the process: Gly-gly (0.066 g, 0.5 mmol) dissolved in water (5 ml), which was deprotonated by using a solution of NaOH (0.020 g, 0.5 mmol), was added to a solution of Cu(ClO4)2·6H2O (0.185 g, 0.5 mmol) with stirring. A solution of PBO (0.098 g, 0.5 mmol) in MeOH (20 ml), was added dropwise to the above solution and then stirring was continued at 50 °C for 2 h. The resulting solution was filtered and allowed to evaporate at room temperature until blue microcrystalline precipitates formed. It was then further purified by recrystallization from 80% MeOH. Yield: 0.193 g (72%). IR bands (KBr pellet, cm−1): 3432 ν(−OH); 3289 ν(−NH); 3096 ν as(−NH2); 2922 ν s(−NH2); 1622 ν as(−COO−); 1552 ν(C = N); 1473 ν(C = C); 1381 ν s(−COO−); 1103 ν(ClO4 −); 629 ν(Cu–O); 472 ν(Cu–N). UV-vis (MeOH) [λmax/nm (ε/M−1 cm−1)]: 231 (14969), 303 (28102), 642 (62.76). Molar conductance, Λ M (MeOH, 299 K): 106.7 S−1 cm2 mol−1. ESI-MS (MeOH) displays a peak at m/z 389.9 [Cu(PBO)(Gly-gly)]+. ESR (MeOH, 90 K): g|| = 2.2817, g⊥ = 2.0656. Elemental Anal. Calcd (%) for C16H20N4O10.5ClCu (1): C, 35.90; H, 3.77; N, 10.47; Found (%): C, 36.17; H, 3.98; N, 10.19.
Synthesis of [Cu(PBO)(Gly-L-leu)(H2O)]·ClO4 (2)
2 was prepared by a similar procedure as for 1, using Gly-L-leu instead of Gly-gly. Yield: 0.121 g (43%). IR bands (KBr pellet, cm−1): 3415 ν(−OH); 3297 ν(−NH); 3103 ν as(−NH2); 2963 ν s(−NH2); 1634 ν as(−COO−); 1553 ν(C = N); 1482 ν(C = C); 1387 ν s(−COO−); 1120 ν(−ClO4 −); 621 ν(Cu–O); 495 ν(Cu–N). UV-vis (MeOH) [λmax/nm (ε/M−1 cm−1)]: 230 (15811), 304 (29815), 639 (58.41). Molar conductance, Λ M (MeOH, 299 K): 104.3 S−1 cm2 mol−1. ESI-MS (MeOH) displays a peak at m/z 446.0 [Cu(PBO)(Gly-L-leu)]+. ESR (MeOH, 90 K): g|| = 2.2762, g⊥ = 2.0603. Elemental Anal. Calcd (%) for C19H23N4O8ClSCu (2): C, 42.56; H, 4.47; N, 9.93; Found (%): C, 42.76; H, 4.28; N, 9.90.
DNA Binding Experiments
Absorption Spectra Measurements
The UV absorbance at 260 nm and 280 nm of the CT-DNA solution in 50 mM NaCl/5 mM Tris–HCl buffer (pH = 7.2) gives a ratio of 1.8–1.9, indicating that the DNA was sufficiently free of protein [19]. The concentration of CT-DNA per nucleotide was determined from its absorption intensity at 260 nm, taking the molar absorption coefficient of 6600 M−1 cm−1 [20].
Electronic absorption titration were performed by keeping the fixed concentration of the complexes (50 μM) and gradually increasing the concentration of CT-DNA in the 50 mM NaCl/5 mM Tris–HCl buffer solution (pH = 7.2). The absorption data were analyzed to assess the binding constant K b, which was determined using the Wolfe–Shimmer Eq. [21]:
where ε a, ε f and ε b correspond to the apparent extinction coefficient (A obsd/[compound]), the extinction coefficient for free compound and the extinction coefficient for compound in the fully bound form, respectively. A plot of [DNA]/(ε a – ε f) vs. [DNA], where [DNA] is the concentration of DNA, gives K b as the ratio of the slope to the intercept.
Fluorescence Spectra Experiments
Fluorescence titration quenching experiments were carried out by titrating the complexes into EthBr–DNA solution containing 8 μM EthBr and 10 μM DNA at three different temperature (300, 305, 310 K). The fluorescence spectra were recorded with excitation wavelength at 525 nm and emission wavelength in the range of 540 to 680 nm. For each addition, the solutions were allowed to keep for 10 min to equilibrate.
Viscosity Measurements
Viscosity measurements were performed using Ostwald’s viscometer at 29.0 ± 0.1 °C in a thermostatic water-bath. Data were presented as (η/η 0)1/3 vs. the ration of the concentration of the complexes and CT-DNA, where η and η 0 are the viscosity of CT-DNA (200 μM) in the presence and absence of the complexes, respectively. Viscosity values were calculated according to the relation η = (t − t 0)/t 0, where t and t 0 are the observed flow time in the presence and absence of the samples, respectively. Flow time was measured with a digital stopwatch, and each sample was measured three times.
Circular Dichroism (CD) Spectra Measurements
The CD spectra of CT-DNA (100 μM) were conducted in the Tris–HCl buffer solution (pH = 7.2) at ambient temperature by increasing [Complex]/[CT-DNA] ratios (r = 0.0, 0.2, 0.4, 0.6, 0.8, 1.0). The test system was in protection of nitrogen atmosphere during the experiment. Spectra were collected after averaging over three scans from 220 to 320 nm and subtracting the buffer background.
Electrochemical Measurements
All the electrochemical measurements were carried out in a single compartment cell at 298 K by using 10 mM Tris HCl/50 mM NaCl buffer solution (pH = 7.2) as the supporting electrolyte. The experiments were performed in 0.3 mM complexes solution in the absence and presence of CT-DNA. Cyclic voltammetry measurements were performed with a three-electrode system (a glass carbon electrode, a platinum electrode and a saturated calomel electrode (SCE) were used as working, auxiliary, and reference electrodes, respectively). All solutions were deoxygenated by purging with nitrogen prior to measurements.
DNA Cleavage Experiments
The pBR322 DNA (250 ng) cleavage experiments were performed by agarose gel electrophoresis. The samples were incubated for 1 h at 37 °C in the dark, and a loading buffer was added. Then the samples were electrophoresed for 40 min at 100 V on 0.8% agarose gel using Tris–Boric acid–EDTA buffer (pH = 8.3). Control experiments were carried out using 50 μM ascorbic acid, 20 μM Cu(ClO4)2·6H2O and 20 μM ligands. Additionally, to identify the reactive oxygen species (ROS) involved in the cleavage reaction, the ROS scavengers DMSO, tert-butyl alcohol, and ethanol (hydroxyl radical), sodium azide (NaN3) (singlet oxygen) and superoxide dismutase (SOD) (superoxide anion radical) were used. After electrophoresis, bands were visualized by UV light and photographed with a Bio-Rad Laboratories-Segrate Gel Imaging System.
Molecular Docking Procedure
The molecular docking studies were performed using the DOCK6 [22] program to investigate the interaction of the complexes and DNA. The structures of the complexes were sketched by Gaussian viewer. And the full geometry optimizations were executed with the Gaussian09 program package using the hybrid density functional theory (B3LYP) method and 6-31G(D) basis set. The crystal structure of the DNA containing iodinated 8-mer oligonucleotides d(5′-G-dIU-TGCAAC-3′) (PDB ID: 454D) [23, 24] was downloaded from the protein data bank. This DNA fragment included a cavity between GC/GC consecutive base pairs. Meanwhile, the water molecules and the substrate were deleted. The receptor active site was defined and selected, and a 5 Å extra margin grid box were constructed to enclose the whole DNA molecule. Then the two complexes were docked to the DNA with grid-based scoring. Other parameters were default settings. All calculations were performed on a Dell T7500 server with dual XEON 5660 cores, RedHat linux operating system. Visualization of the docked pose has been carried out by using PyMol molecular graphics program.
Superoxide Dismutase Activity
The superoxide radicals (O2 −•) were generated by the system of TEMED-VB2-NBT and determined by the modified nitroblue tetrazolium (NBT) photochemical reduction method [25]. The solution contained the tested complexes (0.05–0.80 μM), TEMED (100 μM), NBT (93.20 μM), and VB2 (6.80 μM) in phosphate buffer (0.05 M, pH = 7.8). The reactions were monitored by detecting the concentration of the reduced NBT (blue formazan) at 560 nm. The suppression percentage (I) of NBT reduction was calculated using the following equation: I (%) = (1 – A i/A 0) × 100, where A i and A 0 present the slopes of the straight line of absorbance values as a function of time in the presence and absence of the complexes, respectively. The IC50 is defined as the necessary concentration of the complexes to scavenge 50% reduction of NBT. Each sample was measured in three parallel times.
In Vitro Cytotoxicity Assays (MTT)
The MTT cytotoxicity assay was performed as previously described by Mosmann [26]. Three tumour cells of HeLa (cervical), A549 (pulmonary) and PC-3 (prostatic) were seeded into 96-well microtiter plates (1 × 104 cells per well) and incubated overnight in an atmosphere with 5% CO2 at 37 °C. The tested compounds were dissolved in DMSO and added to the wells. DMSO was used as the vehicle control. The plates were incubated in a 5% CO2 incubator at 37 °C for 48 h. Then 20 μL of MTT solution (5 mg/mL) was added to each well after 4 h incubation. The formed formazan crystals were solubilized in 150 μL DMSO, and the optical density of the purple formazan colour formed was measured spectrophotometrically at 595 nm. The cytotoxicity of each sample was expressed as IC50 value, which is the concentration of the complex that caused 50% inhibition of cell growth. All the samples were assayed in triplicate.
Results and Discussion
Synthesis and Characterization
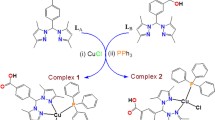
The titled complexes were synthesized by the reaction of copper(II) perchlorate hexahydrate with Glycyl-glycine/Glycyl-L-leucine and PBO in equimolar quantities using 80% MeOH as solvent (Scheme 1). The complexes were characterized by molar conductivity, elemental analyses, ESI-MS, UV-visible, IR and ESR data. The molar conductivity values of the two complexes in MeOH suggest that the complexes are 1:1 type electrolytes [27]. The results of the elemental analyses were found to be in good agreement with the molecular formula of the complexes. Besides, the ESI-MS of the complex 1 displayed a peak at m/z = 389.9 in MeOH matching exactly with [Cu(PBO)(Gly-gly)]+ (m/z = 446.0 for 2, [Cu(PBO)(Gly-L-leu)]+).
In the IR region, the broad peak near 3420 cm−1 was the stretching vibration ν(−OH) of the water molecules. The band near at 3289 cm−1 for 1 (3297 cm−1 for 2) was the stretching vibration ν(−NH) of the secondary amide in dipeptide. The two bands near 3096 and 2922 cm−1 for 1 (3103 and 2963 cm−1 for 2) were ascribed, respectively, to asymmetric ν as(−NH2) and symmetric ν s(−NH2) stretching vibrations. And the two peaks at 1622 and 1381 cm−1 for 1 (1634 and 1387 cm−1 for 2) were attributed to asymmetric ν as(−COO−) and symmetric ν s(−COO−) stretching frequencies, respectively. The frequency separation Δν [= ν as(−COO−) − ν s(−COO−)] > 200 cm−1 suggested the unidentate nature of the carboxylato group [28]. Besides, the bands at 1552 cm−1 and 1473 cm−1 for 1 (1553 cm−1 and 1482 cm−1 for 2) can be assigned to the ring stretching frequencies ν(C = N) and ν(C = C) of ligand PBO, indicating that PBO was coordinated to the central Cu2+. Moreover, the complexes showed very strong bands at 1103 cm−1 for 1 and 1120 cm−1 for 2, which can be attributed to ν(Cl − O) of ClO4 −. The bands at 472 and 629 cm−1 for 1 (495 and 621 cm−1 for 2) most likely belong to the ν(Cu–N) and ν(Cu–O) stretching vibrations, respectively.
The UV–visible emission spectra of the complexes were recorded in methanol solution at ambient temperature. The bands observed at 231 nm and 303 nm for 1 (230 nm and 304 nm for 2) were assigned as π → π* transitions of ligand PBO. Complexes 1 and 2 exhibited a broad and weak bands in the lower energy regions at 642 and 639 nm, respectively, which could be attributed to the d → d transitions of Cu(II) in a distorted square pyramidal environment [29].
The X-band ESR signals of the complexes were recorded in MeOH (90 K) at frequency of 9.46 GHz under the magnetic field strength 3000 ± 1000 gauss using tetracyanoethylene (TCNE) as field marker. The ESR spectra revealed an anisotropic signal with g|| = 2.2817, g⊥ = 2.0656, and A|| = 163.6 G for 1 and g|| = 2.2762, g⊥ = 2.0603, and A|| = 166.3 G for 2. The trend g|| > g⊥ > ge (2.0023) revealed that the unpaired electron was located in the dx2–y2 orbital of the Cu(II) ions characteristic of local symmetry, indicating an approximate square pyramidal geometry for the complexes [30].
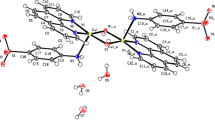
Therefore, from the above results, the structure of the two Cu(II) complexes were considered an approximate square-pyramidal geometry where four equatorial positions were occupied by PBO (N, N) and dipeptide (N, O) and the axial position was occupied by a H2O molecule.
DNA-Binding Studies
Electronic Absorption Spectra
Electronic absorption spectroscopy is an efficient and widely employed method to study the binding mode of metal complexes to DNA. Due to the stacking interaction between aromatic chromophore of complex and DNA base pairs, complex bound to DNA through intercalation usually results in hypochromism and bathochromism. The absorption spectra of the two complexes in the range of 200–400 nm are depicted in Fig. 1. As increasing the concentration of CT-DNA, the hypochromism (1: 10.29%, 2: 12.21%) of the LMCT absorption peak in 303 nm was observed, which might be attributed to the heteroaromatic rings of the complexes intercalating into DNA base pairs. The binding constants (K b) are calculated as 1.421 × 104 M−1 (1) and 2.681 × 104 M−1 (2), respectively. The K b values followed the trend of 2 > 1, indicating that the DNA binding strength of 2 is stronger.
Although the binding mode and affinity of the two complexes with DNA were determined with the electronic absorption spectra, it is necessary to carry out other experiments to confirm further (cf. below).
Fluorescence Spectra
Fluorescence spectroscopy was applied to study the interaction characteristics between chromophore and other compounds. The Stern–Volmer constant (K SV) was used to evaluate the fluorescence quenching efficiency according to the classical Stern–Volmer Eq. [31]:
where F 0 and F are the fluorescence intensities of EthBr-DNA in the absence and presence of the complexes, respectively, and [Q] is the concentration of the complexes. Sterm–Volmer equation was employed to determine K SV by the linear regression of F 0/F against [Q]. As shown in Fig. 2, incremental addition of the complexes concentration caused the decrease in fluorescence intensity, which suggesting that the complexes could partly displace EthBr from the EthBr–DNA system, characteristic for the intercalative binding of the complexes to DNA. Additionally, the obtained K SV values in 300 K for complexes 1 and 2 were found to be 3.774 × 103 M−1 and 4.977 × 103 M−1, respectively. Obviously, 2 has a higher DNA binding affinity than 1.
The Mechanism of Quenching
The mechanisms of quenching are usually classified as two types: static and dynamic quenching. Static quenching refers to fluorophore–quencher complex formation, while dynamic quenching refers to a process in which the fluorophore and the quencher come into contact during the transient existence of the exited state [32]. We can distinguish the two mechanisms according to the relationship of temperature dependence of the K SV. Generally, dynamic quenching depends on diffusion, higher temperatures lead to larger diffusion coefficients, which cause more collision probability, and thus the K SV can be increased by rising the temperature. In contrast, the temperature increment may decrease compounds stability, and thus lower values of the K SV were obtained in the static quenching [33]. The Stern–Volmer plots for quenching of EthBr-DNA fluorescence by the complexes at three different temperatures were shown in Fig. 3 and relevant data were summarized in Table 1. The Stern–Volmer plots were linear which showed that only one type of quenching process occurs, either dynamic or static quenching. Apparently, the fluorescence quenching process is dynamic quenching because the K SV increased with the temperature.
Thermodynamic Studies
In order to have a better understanding of the driving forces behind the binding of metal complexes to DNA, the thermodynamic parameters such as enthalpy (ΔH), entropy (ΔS) and Gibbs free energy (ΔG) with three different temperatures were obtained and analyzed. From the thermodynamic standpoint, if ΔH > 0 and ΔS > 0, the main force is hydrophobic interaction; if ΔH < 0 and ΔS < 0, the major force is the Van der Waals force or hydrogen bond; if ΔH ≈ 0 and ΔS > 0, electrostatic force plays major role in the reaction [34]. The ΔH, ΔS and ΔG for a binding reaction were calculated from the vant’t Hoff equation and Gibbs–Helmholtz Eq. [33]:
where K is the Stern–Volmer quenching constant at the corresponding temperature and R is the gas constant. The ΔH can be regarded as a constant if the temperature does not vary significantly. The thermodynamic data were listed in Table 1. The positive ΔH and ΔS values suggest that the binding of the complexes to DNA is an endothermic and entropy increasing process. And the hydrophobic interaction plays a major role in this process. Moreover, the binding process of the complexes with CT-DNA is entropy-driven due to |ΔH| < |TΔS|. And the negative value of ΔG reveals that the process is spontaneous.
Viscosity Measurements
The viscosity measurements of CT-DNA is regarded as the least ambiguous and the most critical tests of a DNA binding mode in solution in the absence of crystallographic or NMR structural data. Intercalation leads to an increase in the DNA viscosity because the DNA base pairs are separated to accommodate the bound ligand. The partial or non-classical mode of binding could bend or kink the DNA helix, which reduces its effective length and thereby its viscosity [35]. As seen in Fig. 4, the relative viscosity of DNA steadily increases upon increasing the concentration of the complexes. This may be explained by the fact that the complexes can insert the DNA adjacent base pairs, leading to an increase in the separation of base pairs at insertion sites, as a consequence, an increase in overall DNA contour length. The increased degree of viscosity supported the order of the binding strength of the complexes to DNA: 2 > 1.
Circular Dichroic Spectra
Circular dichroic spectral techniques can provide some useful information on how the conformation of DNA is influenced by the binding of small molecules. The B form DNA conformation shows two conservative CD bands in the UV region: a positive band at 276 nm due to the π–π stacking of base pairs and a negative band at 246 nm due to right-hand helicity [36]. As shown in Fig. 5, the CD spectra of DNA with the addition of the complexes shown a decrease in both the positive and negative peaks, with the former being affected slightly more than the latter. Meanwhile, red shifts were observed for the positive (3–5 nm) and negative (1–2 nm) bands, indicating that the binding disturbed the right handed helicity and base stacking of DNA, and induced certain conformational changes of B-DNA, such as the conversion from B → C like structure [37]. The larger decrease of positive bands suggested the complexes intercalated into the DNA base stacking and thereby unwinding the double helix [38, 39]. Hydrophobic interaction played a major role in this process. Furthermore, the variation of intensity follows the trend 2 > 1, which is consistent with the results obtained by the above experiments.
Electrochemical Titration
The cyclic voltammetric technique was applied to further investigate the interaction mode between the metal complexes and DNA. The typical cyclic voltammograms and related parameters of the copper(II) complexes with and without DNA were shown in Fig. 6 and Table 2, respectively. A quasi-reversible redox response was observed along with the ratio of the oxidation peak current to reduction peak current (I pa/I pc) ≈ 1. In general, if the interaction mode is intercalation binding, formal potential shifts to a more positive value, while electrostatic interaction results in a negative shift in the formal potential [40, 41]. Apparently, the anodic peak potential (E pa) and formal potential (E 1/2) of the complexes showed positive shifts, indicating an intercalative binding mode of the complexes with DNA. This kind of interaction was principally due to the hydrophobic forces [42]. Additionally, the I pa and I pc of 1 and 2 experienced an apparent decline in the presence of CT-DNA. According to Kelly et al., the drop of peak currents is due to the reason that the complex bound to DNA and the product was non-electroactive, decreasing the concentration of electroactive species in solution and resulting in the decline of the peak currents [43]. The observed drop of the voltammetric current can be ascribed to slow diffusion of the complex-DNA compound. The more prominent decrease of the peak currents and shift of the peak potentials observed for 2 over 1 upon addition of DNA suggest that the interaction force of the complexes to DNA follow the order of 2 > 1, which is consistent with the above results obtained by viscosity measurements and spectral methods.
From reversible redox reactions of the free and bound species, the corresponding equilibrium constants for the binding of the reduced and oxidized forms to DNA can be obtained from the Nernst Eq. [37].
where E b 0’and E f 0’ are the formal potentials of the Cu(I)/Cu(II) couple in the bound and free forms, respectively, and K + and K 2+ the corresponding binding constants for the binding of the Cu(I) and Cu(II) species to DNA, respectively (Scheme 2). The values of K +/K 2+ were 1.22 and 2.87 for 1 and 2, respectively, indicating that the reduced form of the Cu(II) complexes interact with DNA more strongly than the oxidized form.
DNA Cleavage Activity
The Cleavage of pBR322 DNA
The DNA cleavage by the complexes have been studied using supercoiled pBR322 plasmid DNA as a substrate, and the cleavage reactions were monitored by agarose gel electrophoresis. When plasmid DNA is subjected to electrophoresis, the fastest migration will be observed for the intact supercoiled DNA (Form I). If one strand is cleaved, the supercoiled DNA will relax to generate a slower-moving open circular form (Form II). If both strands are cleaved, a linear form (Form III) will be generated which migrates in between Form I and Form II [44]. As shown in Fig. 7, complexes 1 and 2 exhibited corresponding abilities to cleave the DNA. Meanwhile, the control experiments demonstrated that the ascorbic acid, metal salts Cu(ClO4)2·6H2O, ligands or complexes didn’t show prominent cleavage of DNA. Concentration dependent cleavage in the presence of 50 μM ascorbic acid (lanes 7–10) was observed for the complexes. 2 had higher cleavage activity, which may be associated with its higher DNA binding affinity and Cu(II)/Cu(I) redox potential (cf. above).
Cleavage of plasmid pBR322 DNA (250 ng) by 1 (a) and 2 (b) after 1 h incubation at 37 °C. Lane 1, DNA control; lane 2, DNA + ascorbic acid (50 μM); lane 3, DNA + Cu(ClO4)2·6H2O (20 μM); lane 4, DNA + PBO (20 μM); lane 5, DNA + Gly-gly (a)/Gly-L-leu (b) (20 μM); lane 6, DNA + 1 or 2 (20 μM); lanes 7–10, DNA + ascorbic acid (50 μM) + 1 or 2 (5, 10, 15 and 20 μM, respectively)
Mechanistic Investigation of the DNA Cleavage
In order to explore the active oxygen species responsible for DNA damage, DNA cleavage in presence of hydroxyl radical (•OH) scavengers (DMSO, tert-butyl alcohol, and ethanol), singlet oxygen (1O2) quencher (NaN3) and superoxide anion radical (O2 −•) scavenger (SOD) were investigated (Fig. 8). The experimental result showed that the cleavage was inhibited significantly in the presence of •OH scavengers (lanes 3–5), which indicated that the cleavage reaction could involve •OH. The presence of 1O2 quencher didn’t significantly reduce the efficiency of DNA cleavage, ruling out the possibility of involvement of diffusible 1O2 or singlet oxygen-like entities in the cleavage. Furthermore, DNA cleavage was promoted with addition of superoxide dismutase (O2 −• scavenger) to the reaction mixture (lane 8), and free SOD exhibited no any DNA cleavage (lane 2), which validates that O2 −• could be involved in the DNA strand scission in a round-about way, in which the complexes can catalyze the dismutation O2 −• to generate H2O2 and O2. Moreover, we have known that the Cu(I) species binds to DNA with affinity higher than Cu(II) species from the above electrochemical test. And thus DNA is made more accessible for the ROS (•OH) produced by Fenton type reaction [45], resulting in higher DNA cleavage ability.
Cleavage of plasmid pBR322 DNA (250 ng) by 1 (a) and 2 (b) in the presence of different typical reactive oxygen species scavengers after 1 h incubation at 37 °C. Lane 1, DNA control; lane 2, DNA + ascorbic acid (50 μM) + SOD (15 units); lane 3, DNA + ascorbic acid (50 μM) + 1 or 2 (10 μM); lanes 4–8, DNA + ascorbic acid (50 μM) + 1 or 2 (10 μM) + [DMSO (0.2 M), tert-butyl alcohol (0.2 M), EtOH (0.2 M), NaN3 (0.2 M) and SOD (15 units), respectively]
Molecular Docking Analysis
Molecular docking technique is a well-documented computational tool to understand the interaction of a small molecule (ligand) and a macromolecule (receptor). It can contribute to the rational drug design and the mechanistic study target DNA mainly in a non-covalent mode [46]. As shown in Fig. 9, the resulting model showed that 1 and 2 could intercalate into the cavity between GC/GC consecutive base pairs through heteroaromatic rings and stabilized by hydrophobic interactions and hydrogen bonds. There are certain hydrogen-bonding interactions between the complexes and DNA, the complex 1:H24…454D:DC-13:O2 (2.4 Å), while the complex 2:H40…454D:DA-6:O4’ (2.2 Å) and H25…454D:DA-6:O4’ (2.8 Å). The resulting Grid Score of docked 1 and 2 with DNA were found to be −41.674 and −41.973, respectively. The more negative the Grid Score, the more potent the binding between complex and DNA. The results indicated that the binding strength of 2 to DNA is stronger than 1, which is correlated well with the above experimental DNA binding results.
Antioxidant Activity
The superoxide anion radical (O2 −•) as a kind of clinically important reactive oxygen species in the human body involved in various physiological and pathophysiological processes such aging and ascarcinogenesis etc. [47]. A few diseases have been proved to involve overproduction of the O2 −•, which attracts much attention in the search for excellent SOD mimics [48]. As shown in Fig. 10, the inhibitory effect of the tested complexes was concentration related, and 1 and 2 exhibited excellent antioxidant activities with IC50 values of 0.112 and 0.191 μM, respectively. This could be related to the flexible geometry around the center copper(II) contributing to O2 −• approach, with fast exchange of axial weakly coordinated H2O [25]. Moreover, the electrons of nitrogen heteroatomic ligands, such as pyridine and oxazole, can facilitate stability of complex–O2 −• interaction in favor of high SOD activity. And the IC50 values of suppression ratio (%) for O2 −• were listed in Table 3 (including native SOD and other excellent SOD mimics) [49–52]. The complexes are potent SOD mimics due to their much lower molecular weight compared with that of the native Cu,Zn-SOD (MW 32000 Da), although the activities of the complexes are slightly lower than the SOD. The SOD-like activity of 1 was better than 2, which could be due to the larger steric hindrance induced by the branched group of Gly-L-leu in comparison to Gly-gly, making the approach of the O2 −• to the active centre of copper ions more difficult. The mechanism for the complexes scavenging O2 −• was shown in Scheme 3. A six–coordinate octahedral adduct initially formed, which was quite unstable due to the influence of the Jahn–Teller effect [53], then the rapid internal transformation of Cu(II) and Cu(I) was made possible via electron transfer between copper and reactive oxygen radical anion, and eventually the reaction of catalyzing the dismutation O2 −• was realized. The excellent SOD-like activity of the complexes would be helpful to develop new potential antioxidants and new therapeutic reagents for some diseases [54].
Cytotoxic Activity
The in vitro cytotoxic activities of the complexes against cell lines of HeLa (cervical), PC-3 (prostatic) and A549 (pulmonary) were evaluated by MTT assay and presented in Table 4. Cisplatin was used as a positive control. The cell viability decreased with increasing complexes’ concentrations from 3.125 to 200 μM indicating that the cytotoxicity of the complexes was concentration-dependent. In addition, 2 possessed higher cytotoxicity than 1, which could be due to the increasing lipophilicity of 2 with the chain length of dipeptide, making the drug penetrate the cell membrane more easily. And the stronger ability of 2 to bind and cleave DNA could relate this. Meanwhile, the adherence of the positively charged metal complexes to the plasma membrane by electrostatic attraction is facilitated by the different concentration gradient of the complexes before its transport across the membrane. The cytotoxicity of complexes 1 and 2 against A549 were superior to HeLa and PC-3, which indicated that the complexes tend to inhibit A549 cell lines. The in vitro cytotoxicity of the complexes was lower than cisplatin, but their cytotoxicity toward HeLa was higher than 5-Fluorouracil [55], which is widely applied to the treatment of cancer as clinical drug, indicating that the complexes still have the potential to act as effective chemotherapeutic drugs.
Conclusions
Two new mononuclear mixed ligand copper(II) complexes [Cu(PBO)(Gly-gly)(H2O)]·ClO4·1.5H2O (1) and [Cu(PBO)(Gly-L-leu)(H2O)]·ClO4 (2) have been prepared and characterized by various analytical and spectroscopic techniques. The interactions of the complexes with DNA were investigated using a lot of physico-chemical methods. The results indicated that the complexes were bound to DNA with an intercalative mode. And the processes were dominated by hydrophobic interaction accompanying with endothermic. Also, the complexes exhibited efficient activities of oxidative cleavage DNA in the presence of ascorbic acid, probably induced by •OH. Moreover, the complexes all showed cytotoxicity of some extent and excellent SOD-like activities, which is related to the redox property and hydrophobicity of the complexes. This work would be helpful to rationally design new metallopeptide based chemotherapeutic agents and probes of DNA structure.
References
Siegel R, Ma J, Zou Z, Jemal A (2014) Cancer statistics, 2014. CA-Cancer J Clin 64:9–29
Tabassum S, Al-Asbahy WM, Afzal M, Shamsi M, Arjmand F (2012) DNA binding and cleavage studies of new sulfasalazine-derived dipeptide Zn(II) complex: validation for specific recognition with 5′-TMP. J Lumin 132:3058–3065
Arun T, Subramanian R, Raman N (2016) Novel bio-essential metal based complexes linked by heterocyclic ligand: synthesis, structural elucidation, biological investigation and docking analysis. J Photoch Photobio B 154:67–76
Huang HY, Zhang PY, BL Y, Chen Y, Wang JQ, Ji LN, Chao H (2014) Targeting nucleus DNA with a cyclometalated dipyridophenazineruthenium(II) complex. J Med Chem 57:8971–8983
Kumar RS, Arunachalam S (2009) DNA binding and antimicrobial studies of polymer–copper(II) complexes containing 1,10-phenanthroline and l-phenylalanine ligands. Eur J Med Chem 44:1878–1883
Singh R, Afzal M, Zaki M, Ahmad M, Tabassum S, Bharadwaj PK (2014) Synthesis, structure elucidation and DFT studies of a new coumarin-derived Zn(II) complex: in vitro DNA/HSA binding profile and pBR322 cleavage pathway. RSC Adv 4:43504–43515
Metcalfe C, Thomas JA (2003) Kinetically inert transition metal complexes that reversibly bind to DNA. Chem Soc Rev 32:215–224
Rad FV, Housaindokht MR, Jalal R, Hosseini HE, Doghaei AV, Goghari SS (2014) Spectroscopic and molecular modeling based approaches to study on the binding behavior of DNA with a copper(II) complex. J Fluoresc 24:1225–1234
Sarkar S, Mukherjee T, Sen S, Zangrando E, Chattopadhyay P (2010) Copper(II) complex of in situ formed 5-(2-pyridyl)-1,3,4-triazole through C-S bond cleavage in 1,2-bis(2-pyridylmethylthio)-bis-ethylsulphide: synthesis, structural characterization and DNA binding study. J Mol Struct 980:117–123
Pravin N, Devaraji V, Raman N (2015) Targeting protein kinase and DNA molecules by diimine-phthalate complexes in antiproliferative activity. Int J Biol Macromol 79:837–855
Nagaraj K, Sakthinathan S, Arunachalam S (2014) Synthesis, CMC determination, antimicrobial activity and nucleic acid binding of a surfactant copper(II) complex containing phenanthroline and alanine schiff-base. J Fluoresc 24:589–598
Pulimamidi RR, Ravula C, Battu S (2016) New bio-based Cu(II) complexes and study of their anti-cancer activities. J Fluoresc 26:1183–1197
He FH, Tao L, Li XW, Li YT, Wu ZY, Yan CW (2012) Syntheses and structures of new dicopper(II) complexes bridged by N-(2-hydroxyphenyl)-N′-(3-aminopropyl)oxamide: DNA-binding properties and cytotoxic activities. New J Chem 36:2078–2087
Mendu P, Gyana Kumari C, Ragi C (2015) Synthesis, characterization, DNA binding, DNA cleavage and antimicrobial studies of schiff base ligand and its metal complexes. J Fluoresc 25:369–378
Sun A, Prussia A, Zhan W, Murray EE, Doyle J, Cheng L, Yoon J, Radchenko EV, Palyulin VA, Compans RW, Liotta DC, Plemper RK, Snyder JP (2006) Nonpeptide inhibitors of measles virus entry. J Med Chem 49:5080–5092
Seenaiah D, Reddy PR, Reddy GM, Padmaja A, Padmavathi V, Krishna NS (2014) Synthesis, antimicrobial and cytotoxic activities of pyrimidinyl benzoxazole, benzothiazole and benzimidazole. Eur J Med Chem 77:1–7
Alper-Hayta S, Arisoy M, Temiz-Arpaci Ö, Yildiz I, Aki E, Özkan S, Kaynak F (2008) Synthesis, antimicrobial activity, pharmacophore analysis of some new 2-(substitutedphenyl/benzyl)-5-[(2-benzofuryl) carboxamido]benzoxazoles. Eur J Med Chem 43:2568–2578
Czerwieniec R, Kapturkiewicz A, Lipkowski J, Nowacki J (2005) Re(I)(tricarbonyl)+ complexes with the 2-(2-pyridyl)-N-methyl-benzimidazole, 2-(2-pyridyl)benzoxazole and 2-(2-pyridyl)benzothiazole ligands- syntheses, structures, electrochemical and spectroscopic studies. Inorg Chim Acta 358:2701–2710
Marmur JA (1961) Procedure for the isolation of deoxyribonucleic acid from micro-organisms. J Mol Biol 3:201–208
Vamsikrishna N, Kumar MP, Tejaswi S, Rambabu A, Shivaraj (2016) DNA binding, cleavage and antibacterial activity of mononuclear Cu(II), Ni(II) and Co(II) complexes derived from novel benzothiazole schiff bases. J Fluoresc 26:1317–1329
Wolfe A, Shimer GHJ, Meehan T (1987) Polycyclic aromatic hydrocarbons physically intercalate into duplex regions of denatured DNA. Biochemistry 26:6392–6396
Lang PT, Brozell SR, Mukherjee S, Pettersen EF, Meng EC, Thomas V, Rizzo RC, Case DA, James TL, Kuntz ID (2009) DOCK 6: combining techniques to model RNA-small molecule complexes. RNA 15:1219–1230
Hu W, Deng SW, Huang JY, Lu YM, Le XY, Zheng WX (2013) Intercalative interaction of asymmetric copper(II) complex with DNA: experimental, molecular docking, molecular dynamics and TDDFT studies. J Inorg Biochem 127:90–98
Kielkopf CL, Erkkila KE, Hudson BP, Barton JK, Rees DC (2000) Structure of a photoactive rhodium complex intercalated into DNA. Nat Struct Mol Biol 7:117–121
Le XY, Liao SR, Liu XP, Feng XL (2006) Synthesis, structure and SOD-like activity of a ternary Cu(II) complex with 1,10-phenanthroline and L-valinate. J Coord Chem 59:985–995
Mosmann T (1983) Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods 65:55–63
Geary WJ (1971) The use of conductivity measurements in organic solvents for the characterisation of coordination compound. Coordin Chem Rev 7:81–122
Nakamoto K (2009) Infrared and raman spectra of inorganic and coordination compounds (part B), 6th edn. John Wiley and Sons, New York, p. 65
Reddy PR, Raju N, Satyanarayana B (2011) Synthesis, characterization, and DNA binding and cleavage properties of copper(II)-tryptophanphenylalanine-1,10-phenanthroline/2,2′-bipyridine complexes. Chem Biodivers 8:131–144
Hathaway BJ (1972) The correlation of the electronic properties and stereochemistry of mononuclear {CuN4-6} chromophores. J Chem Soc Dalton Trans:1196–1199. doi:10.1039/DT9720001196
Lakowicz JR, Weber G (1973) Quenching of fluorescence by oxygen. A probe for structural fluctuations in macromolecules. Biochemistry 12:4161–4170
Fei BL, Xu WS, Tao HW, Li W, Zhang Y, Long JY, Liu QB, Xia B, Sun WY (2014) Effects of copper ions on DNA binding and cytotoxic activity of a chiral salicylidene Schiff base. J Photoch Photobio B 132:36–44
Gan XJ, Liu SP, Liu ZF, Hu XL (2012) Determination of tetracaine hydrochloride by fluorescence quenching method with some aromatic amino acids as probes. J Fluoresc 22:129–135
Azab HA, Mogahed EM, Awad FK, El Aal RMA, Kamel RM (2012) Fluorescence and electrochemical recognition of nucleosides and DNA by a novel luminescent bioprobe Eu(III)-TNB. J Fluoresc 22:971–992
Satyanarayana S, Dabrowiak JC, Chaires JB (1993) Tris(phenanthroline)ruthenium(II) enantiomer interactions with DNA: mode and specificity of binding. Biochemistry 32:2573–2584
Ivanov VI, Minchenkova LE, Schyolkina AK, Poletayev AI (1973) Different conformations of double- stranded nucleic acid in solution as revealed by circular dichroism. Biopolymers 12:89–110
Mahadevan S, Palaniandavar M (1998) Spectroscopic and voltammetric studies on copper complexes of 2,9-dimethyl-1,10-phenanthrolines bound to calf thymus DNA. Inorg Chem 37:693–700
Xu XY, Wang DD, Sun XJ, Zeng SY, Li LW, Sun DZ (2009) Thermodynamic and spectrographic studies on the interactions of ct-DNA with 5-fluorouracil and tegafur. Thermochim Acta 493:30–36
Li GY, Du KJ, Wang JQ, Liang JW, Kou JF, Hou XJ, Ji LN, Chao H (2013) Synthesis, crystal structure, DNA interaction and anticancer activity of tridentate copper(II) complexes. J Inorg Biochem 119:43–53
Carter MT, Rodriguez M, Bard AJ (1989) Voltammetric studies of the interaction of metal chelates with DNA. 2. Tris-chelated complexes of cobalt(III) and iron(II) with 1,10-phenanthroline and 2,2′-bipyridine. J Am Chem Soc 111:8901–8911
Li X, Li Y, Wu Z, Zheng Y, Yan C (2012) Synthesis, structure, DNA-binding properties and cytotoxic activities of a new one-dimensional polymeric copper(II) complex with N-benzoate-N′-[3-(2-hydroxyl- ethylammino)propyl]oxamide as ligand. Inorg Chim Acta 385:150–157
Nagaraj K, Murugan KS, Thangamuniyandi P, Sakthinathan S (2014) Nucleic acid binding study of surfactant copper(II) complex containing dipyrido[3,2-a:2′-3′-c]phenazine ligand as an intercalator: in vitro antitumor activity of complex in human liver carcinoma (HepG2) cancer cells. RSC Adv 4:56084–56094
Kelly JM, Lyons EG, Putten JMV, SmythM RE (1986) Analytical chemistry, symposium series, electrochemistry, sensors and analysis. Elsevier, Amsterdam, p. 205
Barton JK, Raphael AL (1984) Photoactivated stereospecific cleavage of double-helical DNA by cobalt(III) complexes. J Am Chem Soc 106:2466–2468
Gulumian M, Van Wyk JA (1987) Hydroxyl radical production in the presence of fibres by a Fenton-type reaction. Chem Biol Interact 62:89–97
Rohs R (2005) Molecular flexibility in ab initio drug docking to DNA: binding-site and binding-mode transitions in all-atom Monte Carlo simulations. Nucleic Acids Res 33:7048–7057
Potapov AS, Nudnova EA, Domina GA, Kirpotina LN, Quinn MT, Khlebnikov AI, Schepetkin IA (2009) Synthesis, characterization and potent superoxide dismutase-like activity of novel bis(pyrazole)-2,2 ‘-bipyridyl mixed ligand copper(II) complexes. Dalton Trans 23:4488–4498
Kiningham KK, Clair DKS (1997) Overexpression of manganese superoxide dismutase selectively modulates the activity of Jun-associated transcription factors in fibrosarcoma cells. Cancer Res 57:5265–5271
Fu H, Zhou YH, Chen WL, Deqing ZG, Tong ML, Ji LN, Mao ZW (2006) Complexation, structure, and superoxide dismutase activity of the imidazolate-bridged dinuclear copper moiety with beta-cyclodextrin and its guanidinium-containing derivative. J Am Chem Soc 128:4924–4925
Wu HL, Wang H, Wang XL, Pan GL, Shi FR, Zhang YH, Bai YC, Kong J (2014) V-shaped ligand bis(2-benzimidazolylmethyl)amine containing three copper(II) ternary complexes: synthesis, structure, DNA-binding properties and antioxidant activity. New J Chem 38:1052–1061
Zhou YH, Fu H, Zhao WX, Chen WL, Su CY, Sun HZ, Ji LN, Mao ZW (2007) Synthesis, structure, and activity of supramolecular mimics for the active site and Arg141 residue of copper, zinc-superoxide dismutase. Inorg Chem 46:734–739
Agotegaray MA, Dennehy M, Boeris MA, Grela MA, Burrow RA, Quinzani OV (2012) Therapeutic properties, SOD and catecholase mimetic activities of novel ternary copper(II) complexes of the anti-inflammatory drug Fenoprofen with imidazole and caffeine. Polyhedron 34:74–83
Ramadan AM, El-Naggar MM (1996) Synthesis, characterization and demonstration of superoxide dismutase-like activity of copper(II) chloride, bromide, nitrate, thiocyanate, sulphate, and perchlorate complexes with 2-methyl-amino pyridine. J Inorg Biochem 63:143–153
Wang XZ, Jiang GB, Xie YY, Liu YJ (2014) Synthesis, molecular structure, DNA interaction and antioxidant activity of novel naphthoxazole compound. Spectrochim Acta A 118:448–453
Fu XB, Liu DD, Lin Y, Hu W, Mao ZW, Le XY (2014) Water-soluble DNA minor groove binders as potential chemotherapeutic agents: synthesis, characterization, DNA binding and cleavage, antioxidation, cytotoxicity and HSA interactions. Dalton Trans 43:8721–8737
Acknowledgments
The authors gratefully acknowledge the financial support from the Program of Natural Science Foundation of Guangdong Province (2015A030313423) and Guangdong Province College Students’ Innovation and Entrepreneurship Training Program (No. 1056413042).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOCX 5791 kb)
Rights and permissions
About this article
Cite this article
Gan, Q., Qi, Y., Xiong, Y. et al. Two New Mononuclear Copper(II)-Dipeptide Complexes of 2-(2′-Pyridyl)Benzoxazole: DNA Interaction, Antioxidation and in Vitro Cytotoxicity Studies. J Fluoresc 27, 701–714 (2017). https://doi.org/10.1007/s10895-016-1999-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10895-016-1999-5