Abstract
Common variable immunodeficiency (CVID) is the most common adult-onset primary antibody deficiency disease due to various causative genes. Several genes, which are known to be the cause of different diseases, have recently been reported as the cause of CVID in patients by performing whole exome sequencing (WES) analysis. Here, we found FANC gene mutations as a cause of adult-onset CVID in two patients. B cells were absent and CD4+ T cells were skewed toward CD45RO+ memory T cells. T-cell receptor excision circles (TRECs) and signal joint kappa-deleting recombination excision circles (sjKRECs) were undetectable in both patients. Both patients had no anemia, neutropenia, or thrombocytopenia. Using WES, we identified compound heterozygous mutations of FANCE in one patient and homozygous mutation of FANCA in another patient. The impaired function of FANC protein complex was confirmed by a monoubiquitination assay and by chromosome fragility test. We then performed several immunological evaluations including quantitative lymphocyte analysis and TRECs/sjKRECs analysis for 32 individuals with Fanconi anemia (FA). In total, 22 FA patients (68.8%) were found to have immunological abnormalities, suggesting that such immunological findings may be common in FA patients. These data indicate that FANC mutations are involved in impaired lymphogenesis probably by the accumulation of DNA replication stress, leading to CVID. It is important to diagnose FA because it drastically changes clinical management. We propose that FANC mutations can cause isolated immunodeficiency in addition to bone marrow failure and malignancy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common variable immunodeficiency (CVID) is primarily caused by defective development and maturation of B cells and plasmacytes, which results in antibody deficiencies [1, 2]. T-cell abnormalities may exist and include decreases in the cell number and/or function [3]. Although CVID is the most common primary immunodeficiency disease (PID) associated with hypogammaglobulinemia in adults and more than 13 causative genes were found to date, most genetic mechanisms leading to CVID remain unknown [4]. Several causative genes of CVID have been recently identified by whole exome sequencing (WES), and a part of CVID has been found to be a non-classical phenotype of other diseases [5,6,7].
Fanconi anemia (FA) is a congenital bone marrow failure syndrome that exhibits malformations and malignancies due to the defect of 19 genes [8]. FANC products are involved in DNA repair [9]. Although FA has a wide range of symptoms, hematological symptoms are the most common, and 80% of patients progress to bone marrow failure by 10 years of age [10]. Several reports are available about the immunological status in FA patients. It has been reported that some FA patients have increased infection susceptibility, which is not explained by neutropenia alone [11].
Total absolute lymphocytes, B cells, and NK cells were reduced and serum IgG and IgM levels were significantly lower in FA patients than in controls [12]. Serological response in FA mice after human papillomavirus vaccination were significantly decreased, suggesting that a primary immune dysfunction independently occurs from bone marrow failure [13]. However, isolated immunodeficiency without hematological anomaly in FA has not been reported to date.
Other DNA repair disorders such as Nijmegen breakage syndrome, Cernunnos syndrome, Bloom syndrome, ataxia telangiectasia, ICF (Immunodeficiency, Centromeric instability, and Facial anomalies) syndrome, and PMS2 deficiency are reported to have CVID-like phenotypes or immunological aberrations related to the impairment of V(D)J rearrangement, class switch recombination (CSR), or somatic hypermutations (SHMs) [14, 15]. Here, we report the isolated immunodeficiency caused by FANC mutations.
We identified two cases of adult CVID as a result of FANC mutations using WES and confirmed the diagnosis of FA by functional analysis. We subsequently examined immunological aspects of 32 FA patients by quantitative lymphocyte and T-cell receptor excision circles (TRECs)/signal joint kappa-deleting recombination excision circles (sjKRECs) analyses. This report is one of the largest immunological studies for FA and the first report that identified that FANC may be the responsible gene for CVID.
Methods
Study Design
These studies were approved by the institutional review boards of National Defense Medical College, Tokyo Medical and Dental University, and Tokai University School of Medicine. Informed consent was obtained according to the Declaration of Helsinki.
Patients
Patients 1 and 2 were diagnosed with CVID. In patient 1, TRECs and sjKRECs had been found to be undetectable in a previous study [16]. Patient 2 who was involved in another cohort of patients with hypogammaglobulinemia was referred to the Department of Pediatrics in Tokyo Medical and Dental University. Both patients fulfilled the revised clinical criteria for CVID by the European Society for Immunodeficiency [17].
We also included 32 clinically diagnosed FA patients confirmed by FANC mutation analyses, chromosome breakage tests, and monoubiquitination assays as reported previously [18].
Diagnosis of refractory cytopenia with multilineage dysplasia (RCMD), refractory anemia with excess blasts (RAEB), and acute myeloid leukemia (AML) was based on World Health Organization (WHO) classifications. Aplastic anemia (AA) was defined as hypoplastic marrow with two of the following: neutrophil count <1.5 × 109/L, platelet count (Plt) <100 × 109/L, and hemoglobin level (Hb) <10 g/dL. Pancytopenia was defined as the following: total white cell count <4.0 × 109/L, Plt <150 × 109/L, and Hb <13.5 g/dL (male) or 12.0 g/dL (female).
Lymphopenia and a decrease in B cells, T cells, and NK cells were defined as below 5th percentile of reference values of lymphocyte subsets in age-appropriate norms for healthy children [19].
Whole Exome Sequencing
For the exome sequencing of patient 1, genomic DNA was enriched for protein-coding sequences with a SureSelect Human All Exon V5 kit 2 (Agilent Technologies, Santa Clara, CA, USA). This was followed by massively parallel sequencing with the HiSeq 2000 platform with 100 bp paired-end reads (Illumina, San Diego, CA, USA).
For the exome sequencing of patient 2, genomic DNA was enriched for protein-coding sequences with a TruSeq Exome Enrichment Kit (Illumina, San Diego, CA, USA). This was followed by massively parallel sequencing with the Genome Analyzer IIx platform with 100 bp paired-end reads (Illumina, San Diego, CA, USA).
Candidate germline variants were detected through our in-house pipeline for exome-sequencing analysis with minor modifications for the detection of germline variants. The obtained sequences were aligned to the human reference genome (hg19) with the Burrows–Wheeler Aligner. After removal of duplicate artifacts caused by PCR, the single nucleotide variants with a detected variation frequency >0.25 and insertion–deletions with a detected variation frequency >0.1 were called. All candidate pathogenic variants in FA genes were verified by PCR and Sanger sequencing.
Flow Cytometry Analysis
The mononuclear cells of patients’ peripheral blood were analyzed using a FACS Calibur (Becton Dickinson, USA). We used monoclonal antibodies as follows: anti-CD3-FITC (clone SK7), anti-CD4-FITC/PerCP (clone SK3), anti-CD8-PE/PerCP (clone SK1), anti-CD16-PE (clone B73.1), anti-CD31-PE (clone WM59), anti-CD45RA-FITC (clone L48), and anti-CD45RO-APC (clone UCHL1) obtained from Becton Dickinson, USA; and anti-CD3-APC (clone UCHT1), anti-CD19-FITC (clone J4)/APC (clone 119), and anti-CD56-PE-Cy5 (clone N901) obtained from Becton Coulter, USA.
Measurement of TRECs and sjKRECs Copy Numbers
TRECs and sjKRECs were analyzed using real-time PCR on genomic DNA from whole blood samples as previously reported [20, 21].
FANCD2 Monoubiquitination Assay and Chromosome Fragility Tests in Peripheral Blood Lymphocytes
FANCD2 monoubiquitination assays were conducted using peripheral blood lymphocytes or fibroblasts cultured for 24 h and treated with mitomycin C 0.1 μg/mL [22].
Chromosome fragility tests of lymphocytes [22] were conducted using peripheral blood lymphocytes cultured for 72 h and treated with mitomycin C 0.02 μg/mL and diepoxybutane 0.1 μg/mL. Chromosome fragility tests of fibroblasts [23] were conducted using fibroblasts cultured for 24 h and treated with mitomycin C 0.02 μg/mL.
Results
Patients
Patient 1 is a 40-year-old female who had no history of susceptibility to infection until 39 years of age. There was no history of immunodeficiency and consanguineous marriage in the family. She had a short stature (−2.2 SD) and three café au lait spots. She was incidentally found to have hypogammaglobulinemia (IgG = 4.98 g/L) at 32 years of age, but immunoglobulin replacement therapy had not been started. She had recurrent pneumonia at 39 years of age and was referred to our hospital for further study. She had hypogammaglobulinemia (IgG, 3.65 g/L; IgA, 0.46 g/L; IgM, <0.02 g/L) without pancytopenia (white blood cells, 5000/μL; neutrophils, 2130/μL; hemoglobin, 11.8 g/dL; platelets, 19.2 × 104/μL). Her bone marrow evaluation revealed decreased cellularity without dysplasia (Table 1). While lymphocytes numbers were within normal range, FACS analysis revealed that CD19+ B cells were severely reduced to 0.02% of lymphocytes. The CD4+/CD8+ ratio was 1.15, but 90.1% of CD4+ T cells were skewed toward CD45RO+ memory T cells (Fig. 1a). All antibodies against measles, rubella, and mumps were lower than the protective levels despite vaccination. The copy number of TRECs, a marker for T-cell neogenesis, was 660 copies/μg DNA at 39 years of age, and sjKRECs, a marker for B-cell neogenesis, were undetectable (<102 copies/μg DNA). The copy numbers of TRECs gradually decreased and became undetectable at 40 years of age. The patient was tentatively diagnosed with adult onset CVID and periodic immunoglobulin replacement therapy was started. Development of cytopenia has not been observed and no additional symptoms have occurred.
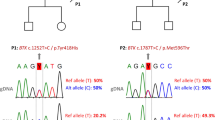
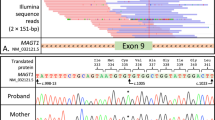
Evaluation of patients 1 and 2. a Flow cytometric analysis of mononuclear cells. B cells are absent and CD4+ T cells are skewed toward CD45RO+ memory T cells in both patients. b Sequencing results of FANCE (patient 1) and FANCA (patient 2). c Monoubiquitination assay of lymphocytes from patient 1 and fibroblasts from patient 2. FANCD2-L (ubiquitinated FANCD2) was absent in both patients even after 24-h stimulation with or without 100 ng/mL mitomycin C (MMC). d Chromosome fragility test of fibroblasts in patient 2. Chromosome breakage was increased by 24-h stimulation of fibroblasts derived from FA patients and patient 2 with MMC 0.02 μg/mL. e Chromosome fragility test of lymphocytes in patient 1. Chromosome breakage was analyzed by 72-h culture of lymphocytes with MMC 0.02 μg/mL and diepoxybutane 0.1 μg/mL. The chromosome fragility of patient 1 was similar to that of typical FA patients
Patient 2 is a 27-year-old female who was found to have hypogammaglobulinemia when she suffered from sinusitis at 19 years of age. Before 19 years, susceptibility to infection was not evident. There was no history of immunodeficiency and consanguineous marriage in the family. She had a short stature (−2.9 SD). She was diagnosed with CVID, and immunoglobulin replacement therapy was started. Epstein–Barr virus (EBV) was persistently detected at 21 years of age (EBV = 3.4 × 103 copies/μg DNA). She was referred to our hospital at 23 years of age. Her complete blood count was normal (white blood cells, 3500/μL; neutrophils, 2240/μL; hemoglobin, 12.5 g/dL; platelets, 12.2 × 104/μL). Her marrow evaluation revealed decreased cellularity without dysplasia. CD19+ B cells were reduced to 0.38% of lymphocytes and CD20+CD27+IgD− switched memory B cells were reduced to 6% of CD20+CD27+ memory B cells. The CD4+/CD8+ ratio was 0.47, and 91.5% of CD4+ T cells were skewed toward CD45RO+ memory T cells (Fig. 1a). Copy numbers of TRECs and sjKRECs were undetectable (<102 copies/μg DNA). At 23 years of age, because of chronic active Epstein–Barr virus infection of NK cell type (EBV, 8.6 × 104 copies/μg DNA in NK cells), she underwent human leukocyte antigen (HLA) matched unrelated donor bone marrow transplantation after reduced-intensity conditioning with fludarabine 75 mg/m2, melphalan 100 mg/m2, and anti-thymocyte globulin 2.5 mg/kg [24, 25]. Common Terminology Criteria for Adverse Events (CTCAE) grade III gastrointestinal disorders emerged during the conditioning regimen. Grade III acute graft-versus-host disease (gut stage 3) with severe diarrhea occurred and improved after the administration of prednisolone. The patient is now 4 years after bone marrow transplantation, alive, and well with complete donor chimerism.
Diagnosis of FA in Patients 1 and 2
We identified compound heterozygous mutations of FANCE c.419T > C (p.L140P) and c.648delC (p.V216fs) in patient 1 and homozygous mutations of FANCA c.190_191insT (p.Glu65ArgfsX5) in patient 2 using WES. No other mutations in known causative genes for bone marrow failure syndromes including FA, and for PIDs were found in both patients. FANCE (patient 1) and FANCA (patient 2) mutations in the peripheral blood, fibroblasts, and neutrophils were confirmed by Sanger sequencing (Fig. 1b).
Karyotype analysis by G-banding with phytohemagglutinin stimulation showed abnormal chromosome breakage, indicating the chromosomal instability (Table 1). We performed FANCD2 monoubiquitination assays [26] and chromosome fragility tests [22] for peripheral blood lymphocytes in patient 1 and for fibroblasts in patient 2. Reduced FANCD2 monoubiquitination and increased chromosome fragility were confirmed in both patients (Fig. 1c–e). Based on these results, we diagnosed both with FA. Our results indicated that FANC mutations can cause isolated immunodeficiency.
Immunological Analysis of 32 FA Patients
Along with the 2 indicated cases, we analyzed immunological aspects of 32 typical FA patients confirmed by FANC mutation analyses, chromosome breakage tests, and monoubiquitination assays.
Mean age ± SD at examination of FA was 9.6 ± 8.7 years (range, 0.3–29 years). The causative genes were as follows: FANCA (n = 16), FANCG (n = 11), FANCC (n = 1) FANCI (n = 1), FANCL (n = 1), FANCN (n = 1), and FANCP (n = 1). The hematological status at examination were as follows: aplastic anemia (AA; n = 17), refractory cytopenia with multilineage dysplasia (RCMD; n = 7), refractory anemia with excess of blasts (RAEB; n = 4), mild anemia and mild thrombocytopenia (n = 2), acute myeloid leukemia (AML; n = 1), and normal peripheral blood with solid tumor (n = 1).
Total numbers of lymphocytes, B, T, and NK cells and the copy numbers of TRECs and sjKRECs are summarized in Table 2. Lymphopenia was observed in five patients (34.4%). CD19+ B cells were decreased in 10 patients (55.0%), CD3+ T cells were decreased in 4 patients (20.0%), the CD4+/CD8+ ratio was inverted in 4 patients (20.0%), and CD3−CD16+CD56+ NK cells were reduced in 10 patients (55.0%) out of 20 patients examined with flow cytometric analyses.
TRECs and sjKRECs were measured in 28 FA patients. TRECs were detectable in all patients. sjKRECs were undetectable in five patients (17.9%), including the patient who had a normal percentage of B cells (P13). sjKRECs of P13 were normal at 4 months of age, but were undetectable at 1.1 years of age.
Discussion
We identified FANC mutations in two adult-onset CVID patients. This is the first report to describe that FANC mutations may cause isolated immunodeficiency without cytopenia. We found that 22 of 32 typical FA patients (68.8%) had abnormal lymphopoiesis. Because FANC plays important roles in hematopoietic stem cell differentiation [27], it is plausible that both hematopoiesis (other than lymphopoiesis) and lymphopoiesis are abnormal in FA.
It is currently unclear why the two patients do not have cytopenia. However, there are at least three possible explanations for this phenomenon. First, FANC mutations in these patients were only present in lymphoid progenitor cells (LPCs) through somatic mutations and not in other lineages. However, these mutations were also found in the fibroblasts and the neutrophils from the patient 1, indicating that these diseases were not caused by somatic mosaicism of LPCs nor reversion of mutation in neutrophils but by germline mutations. Second, a genotype–phenotype correlation may exist; however, the previously reported nonsense mutations in exon 2 of FANCA [28, 29], resembling the mutation in patient 2, is associated with typical FA, excluding this possibility for the patient. For patient 1, as this is the first reported patient with hypomorphic FANCE mutation in one allele, this possibility can be considered. Third, the defect in the lymphoid system and in other hematopoietic systems may progress over different time courses. It is known that abnormal hematopoiesis (neutropenia, anemia, and thrombocytopenia) progresses with age in FA patients [10]. Similarly, abnormal lymphopoiesis may progress with age. In the two indicated cases, it is feasible to assume that abnormal lymphopoiesis preceded abnormal hematopoiesis. To clarify these possibilities, we examined the hematologic defects of patient 1, but patient 1 did not show anemia, neutropenia, or thrombocytopenia until the time of this writing. Patient 2 could not be evaluated because she was treated with stem cell transplantation. Additional studies are underway using induced pluripotent stem cells derived from the patient.
Our results on the FACS analysis of 32 typical FA patients are consistent with those of previous studies that reported reduced B and NK cells and an increased percentage of cytotoxic T cells in some FA patients [12, 30]. sjKRECs were undetectable in five patients, including the patient who had a normal percentage of B cells (P13). The patient was sjKRECs positive at 0.3 years of age but became sjKRECs negative at 1.1 years of age. Neogenesis of B cells may be progressively impaired; however, the patient was RCMD, and the effect of malignancy could not be denied. sjKRECs are associated with B-cell neogenesis [16]. FA patients with normal CD19+ B-cell counts but with undetectable sjKRECs could progress to a deficiency of B cells with age. These findings indicate that FANC genes may be involved in lymphopoiesis.
In our patient cohort, 88% had abnormal lymphocyte subsets and 15% had undetectable sjKRECs among the AA patients, 67% had abnormal lymphocyte subsets and 40% had undetectable sjKRECs among the RCMD patients, and 33% had abnormal lymphocyte subsets and none had undetectable sjKRECs among the RAEB patients. Immunological defects in typical FA patients do not seem to progress with malignant transformations. In addition, in the two cases of CVID phenotype FA, no malignant findings were observed in the bone marrow. Thus, a decrease of TRECs and sjKRECs was not associated with malignancy.
In patient 2, CTCAE grade III gastrointestinal disorders emerged during the conditioning regimen, and grade III acute graft-versus-host disease of the gastrointestinal tract occurred [31]. High dose of alkylating agents, such as melphalan, cause severe damage to FA patients [32, 33]. We believe melphalan in the conditioning regimen was the main reason for the gastrointestinal intestinal disorders. High dose of alkylating agents may be avoided in CVID patients with decreased TRECs or sjKRECs without known causative genes because there are possibilities of DNA repair disorders such as FA.
It is suggested that individuals with FA have immunological disorders, which manifest over time, and that one should consider the diagnosis of FA in older individuals with a CVID phenotype, especially with diminished B cells. This is important as this diagnosis results in dramatic changes in clinical management regarding malignancy screening regimens as well as potential transplant approaches.
CVID is a clinically heterogeneous group. Although about 13 causative genes have been reported, the causative genes have been identified in only around 30% of the patients [34]. There is no report so far that FANC mutations cause CVID. Whole exome sequencing (WES) is an effective tool for identifying the genetic cause for PIDs. It has been recently revealed using WES that part of CVID has new causative genes [7, 35]. We identified the cause of sporadic CVID as a new phenotype of a known disease using WES and revealed the usefulness of WES for CVID once again.
Other bone marrow failure syndromes, such as dyskeratosis congenita (DC), have immunological abnormalities. Patients with DC frequently show lymphopenia, low B-cell and NK-cell number, and decreased T-cell function before other characteristic symptoms of DC develop [36]. This resembles our cases of FA.
The immune system receives various genetically damaging stresses that occur during its maturation and immune responses such as the V(D)J recombination, the immunoglobulin CSR, and the generation of SHMs. Many DNA repair disorders such as Cernunnos syndrome, Bloom syndrome Nijmegen Breakage syndrome, ataxia telangiectasia, ICF, and PMS2 exhibit immunodeficiency by the impairment of V(D)J rearrangement, CSR, or SHMs [14, 15]. It has been observed that the FA cell lines showed a several fold increase in the frequency of aberrant rearrangements associated with V(D)J coding joint formation [37] and that the loss of Fancc impairs antibody-secreting cell differentiation in mice [38]. The impaired V(D)J recombination and/or CSR may account at least in part for the immunodeficiency observed in FA patients.
Most DNA repair disorders that cause CVID are non-homologous end-joining disorders [14, 15]; however, we found CVID phenotype in FA, which impairs in homologous recombination (HR) [39]. Our findings might suggest a new role of FANC complex and HR in lymphopoiesis.
This is the first report that describes FANC as a causative gene for CVID. We propose here that FA should be considered in the differential diagnosis of CVID. Treatment and management associated with FA must be considered for these patients. In addition, our findings expand the clinical spectrum of FA.
References
Durandy A, Kracker S, Fischer A. Primary antibody deficiencies. Nat Rev Immunol. 2013;13:519–33.
Wehr C, Kivioja T, Schmitt C, et al. The EUROclass trial: defining subgroups in common variable immunodeficiency. Blood. 2008;111:77–85.
Cunningham-Rundles C, Bodian C. Common variable immunodeficiency: clinical and immunological features of 248 patients. Clin Immunol. 1999;92:34–48.
Salzer U, Warnatz K, Peter HH. Common variable immunodeficiency—an update. Arthritis Res Ther. 2012;14:223.
Parvaneh N, Casanova J-L, Notarangelo LD, Conley ME. Primary immunodeficiencies: a rapidly evolving story. J Allergy Clin Immunol. 2013;131:314–23.
Al-Herz W, Bousfiha A, Casanova J-L, et al. Primary immunodeficiency diseases: an update on the classification from the International Union of Immunological Societies Expert Committee for Primary Immunodeficiency. Front Immunol. 2014;5:162.
Fliegauf ML, Bryant V, Frede N, et al. Haploinsufficiency of the NF-κB1 subunit p50 in common variable immunodeficiency. Am J Hum Genet. 2015;97:389–403.
Dong H, Nebert DW, Bruford EA, Thompson DC, Joenje H, Vasiliou V. Update of the human and mouse Fanconi anemia genes. Hum Genomics. 2015;9:32.
Duxin JP, Walter JC. What is the DNA repair defect underlying Fanconi anemia? Curr Opin Cell Biol. 2015;37:49–60.
Butturini A, Gale RP, Verlander PC, Adler-Brecher B, Gillio AP, Auerbach AD. Hematologic abnormalities in Fanconi anemia: an International Fanconi Anemia Registry study. Blood. 1994;84:1650–5.
Fagerlie SR, Bagby GC. Immune defects in Fanconi anemia. Crit Rev Immunol. 2006;26:81–96.
Korthof ET, Svahn J, Peffault de Latour R, et al. Immunological profile of Fanconi anemia: a multicentric retrospective analysis of 61 patients. Am J Hematol. 2013;88:472–6.
Holmgren SC, Goren EM, Wood BL, Becker PS, Taylor JA. Immune defects in a mouse model of Fanconi anaemia. Br J Haematol. 2012;159:246–50.
Picard C, Al-Herz W, Bousfiha A, et al. Primary immunodeficiency diseases: an update on the classification from the International Union of Immunological Societies Expert Committee for Primary Immunodeficiency 2015. J Clin Immunol. 2015;35:696–726.
Gennery AR. Primary immunodeficiency syndromes associated with defective DNA double-strand break repair. Br Med Bull. 2006;77–78:71–85.
Kamae C, Nakagawa N, Sato H, et al. Common variable immunodeficiency classification by quantifying T-cell receptor and immunoglobulin κ-deleting recombination excision circles. J Allergy Clin Immunol. 2013;131:1437–40.e5.
Ameratunga R, Brewerton M, Slade C, Jordan A, Gillis D, Steele R, Koopmans W, Woon S-T. Comparison of diagnostic criteria for common variable immunodeficiency disorder. Front Immunol. 2014;5:415.
Hira A, Yabe H, Yoshida K, et al. Variant ALDH2 is associated with accelerated progression of bone marrow failure in Japanese Fanconi anemia patients. Blood. 2013;122:3206–9.
Comans-Bitter WM, de Groot R, van den Beemd R, Neijens HJ, Hop WCJ, Groeneveld K, Hooijkaas H, van Dongen JJM. Immunophenotyping of blood lymphocytes in childhood reference values for lymphocyte subpopulations. J Pediatr. 1997;130:388–93.
Nakagawa N, Imai K, Kanegane H, et al. Quantification of κ-deleting recombination excision circles in Guthrie cards for the identification of early B-cell maturation defects. J Allergy Clin Immunol. 2011;128:223–225.e2.
Morinishi Y, Imai K, Nakagawa N, et al. Identification of severe combined immunodeficiency by T-cell receptor excision circles quantification using neonatal Guthrie cards. J Pediatr. 2009;155:829–33.
Yabe M, Yabe H, Hamanoue S, et al. In vitro effect of fludarabine, cyclophosphamide, and cytosine arabinoside on chromosome breakage in Fanconi anemia patients: relevance to stem cell transplantation. Int J Hematol. 2007;85:354–61.
Pinto FO, Leblanc T, Chamousset D, et al. Diagnosis of Fanconi anemia in patients with bone marrow failure. Haematologica. 2009;94:487–95.
Arai A, Imadome K-I, Watanabe Y, Yoshimori M, Koyama T, Kawaguchi T, Nakaseko C, Fujiwara S, Miura O. Clinical features of adult-onset chronic active Epstein–Barr virus infection: a retrospective analysis. Int J Hematol. 2011;93:602–9.
Kawa K, Sawada A, Sato M, et al. Excellent outcome of allogeneic hematopoietic SCT with reduced-intensity conditioning for the treatment of chronic active EBV infection. Bone Marrow Transplant. 2011;46:77–83.
Tomida J, Itaya A, Shigechi T, et al. A novel interplay between the Fanconi anemia core complex and ATR-ATRIP kinase during DNA cross-link repair. Nucleic Acids Res. 2013;41:6930–41.
Ceccaldi R, Parmar K, Mouly E, et al. Bone marrow failure in Fanconi anemia is triggered by an exacerbated p53/p21 DNA damage response that impairs hematopoietic stem and progenitor cells. Cell Stem Cell. 2012;11:36–49.
Yagasaki H, Hamanoue S, Oda T, Nakahata T, Asano S, Yamashita T. Identification and characterization of novel mutations of the major Fanconi anemia gene FANCA in the Japanese population. Hum Mutat. 2004;24:481–90.
Bouchlaka C, Abdelhak S, Amouri A, et al. Fanconi anemia in Tunisia: high prevalence of group a and identification of new FANCA mutations. J Hum Genet. 2003;48:352–61.
Myers KC, Bleesing JJ, Davies SM, et al. Impaired immune function in children with Fanconi anaemia. Br J Haematol. 2011;154:234–40.
Trotti A, Colevas AD, Setser A, et al. CTCAE v3.0: development of a comprehensive grading system for the adverse effects of cancer treatment. Semin Radiat Oncol. 2003;13:176–81.
Latt SA, Stetten G, Juergens LA, Buchanan GR, Gerald PS. Induction by alkylating agents of sister chromatid exchanges and chromatid breaks in Fanconi’s anemia. Proc Natl Acad Sci U S A. 1975;72:4066–70.
Taniguchi T, D’Andrea AD. Molecular pathogenesis of Fanconi anemia: recent progress. Blood. 2006;107:4223–33.
Maffucci P, Filion CA, Boisson B, Itan Y, Shang L, Casanova J-L, Cunningham-Rundles C. Genetic diagnosis using whole exome sequencing in common variable immunodeficiency. Front Immunol. 2016;7:220.
Kuehn HS, Boisson B, Cunningham-Rundles C, et al. Loss of B cells in patients with heterozygous mutations in IKAROS. N Engl J Med. 2016;374:1032–43.
Allenspach EJ, Bellodi C, Jeong D, Kopmar N, Nakamura T, Ochs HD, Ruggero D, Skoda-Smith S, Shimamura A, Torgerson TR. Common variable immunodeficiency as the initial presentation of dyskeratosis congenita. J Allergy Clin Immunol. 2013; doi:10.1016/j.jaci.2012.11.052.
Smith J, Andrau J, Kallenbach S, Laquerbe A, Doyen N, Papadopoulo D. Abnormal rearrangements associated with V(D)J recombination in fanconi anemia. J Mol Biol. 1998;281:815–25.
Sertorio M, Amarachintha S, Wilson A, Pang Q. Loss of Fancc impairs antibody-secreting cell differentiation in mice through deregulating the Wnt signaling pathway. J Immunol. 2016;196:2986–94.
Michl J, Zimmer J, Tarsounas M. Interplay between Fanconi anemia and homologous recombination pathways in genome integrity. EMBO J. 2016;35:909–23.
Acknowledgments
This work was supported by grants from the Ministry of Defense; the Ministry of Education, Culture, Sports, Science, and Technology (nos. 26293250 and 15 K15396); the Ministry of Health, Labor, and Welfare (no. 14427260); the Kawano Masanori Memorial Public Interest Incorporated Foundation for Promotion of Pediatrics; the Practical Research for Rare/Intractable Diseases from Japan Agency for Medical Research and Development, AMED; The Japan Foundation for Pediatric Research; and Drs. M. and W. Hirose, Dr. H. Matsuda, and Dr. H. Seto. We thank Dr. Yuki Tsujita, Dr. Yumiko Ogura, and Dr. Hirokazu Kawaguchi for medical support. We would also like to thank Ms. Kaori Tomita, Ms. Kimiko Gasa, and Dr. Naomi Terada for their skillful technical assistance.
Author information
Authors and Affiliations
Contributions
Contributions: Y.S., N.M., K.I., A.A., M.Y., and H.Y. treated the patients and designed the clinical laboratory tests; K.S. and N.M. performed flow cytometric analysis; K.H. performed TRECs and sjKRECs analyses; M. Takagi established skin fibroblast of patient 2; M.Y. performed the chromosome fragility tests; A.H. and M. Takata performed monoubiquitination assay and validation of FANC mutations; K.Y., Y.O., Y.S., K.C., H.T., S.M., H.M., S.K., and S.O. performed WES of patient 1; O.O. performed WES of patient 2; K.I., T.M., and S.N. designed the overall study, supervised the experiments, and performed analyses and interpretation of the data.
Corresponding author
Ethics declarations
Disclosure of Conflicts of Interest
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Sekinaka, Y., Mitsuiki, N., Imai, K. et al. Common Variable Immunodeficiency Caused by FANC Mutations. J Clin Immunol 37, 434–444 (2017). https://doi.org/10.1007/s10875-017-0396-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-017-0396-4