Abstract
4-(2-amino-2-oxoethyl) phenyl 4-methylbenzenesulfonate (APMBS), a novel compound, has been synthesized and characterized by spectro-analytical techniques. X-ray diffraction data reveals it crystallizes in the triclinic crystal system with space group \(P\overline{1}\). The structure is stabilized through strong N–H⋯O, weak C–H⋯O and C–H⋯π hydrogen bond interactions yielding a supramolecular structure. The N1A–H1AA⋯O4A, C4–H4⋯O3, and C14–H14⋯Cg1 (Ring: C8–C13) interactions link the molecules into dimers, while the interaction C9-H9–O2 results in the formation of a molecular chain along the a-axis. The optimized structural geometry, by and large, as calculated using DFT and HF techniques is in conformity with the corresponding X-ray data. The DFT computation further aid in the follow up investigations including frontier molecular orbitals (FMOs), density of states (DOS), atomic charges, and molecular electrostatic potential surface (MEP). For a comprehensive description of various molecular intersections, Hirsheld surface and fingerprint plots analysis has been reported, besides the combined atoms-in-molecules (QTAIM) and reduced density gradient (RDG) analysis. Docking studies have been performed against the target protein Tubulin-Colchicine: Stathmin-Like Domain Complex.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Crystal engineering, an interdisciplinary field of research, plays a vital role in various domains of science and technology, including physics, chemistry, biology, engineering, materials science, pharmaceuticals, etc. It involves the design and assembly of solid-state structures with desirable properties by exploiting an interplay of interactions existing between the molecules. Thus, the role of hydrogen bonding-a fundamental aspect of crystal engineering, becomes important due to the existence of weak non-covalent interactions (NCIs), including weak hydrogen bonds and π⋯π interactions. The study of such weak interactions in the crystal structure influences the crystal packing, thus impact the material’s properties [1,2,3,4,5,6,7,8].
Organo-sulfur compounds are of great significance due to their extensive applications in various scientific disciplines and also for their adaptability in chemical alterations. They are used in the synthesis of sulfur dyes, for their ability to produce vibrant colors and durability. Besides being integral to the production of surfactants which acts as active agents in many detergents [9], they have found a place in the production of ion-exchange resins used in softening and purification of water. Apart from these compounds being used in the agricultural field in the form of fungicides, pesticides, and herbicides to protect various crops [10], their use has also been found in the food industry as preservatives for the maintenance of freshness and color of food products like dried fruits and wines [11]. The aryl tosylates have recently gained interest as valuable electrophilic alternatives to aryl halides in cross-coupling reactions catalyzed by transition metals [12,13,14,15]. Therefore, we report the synthesis of a tosyl ester derivative, 4-(2-amino-2-oxoethyl) phenyl 4-methylbenzenesulfonate (APMBS), using diglyme as the solvent system. A variety of analytical methods have been used to characterize its chemical and crystallographic properties, including NMR, FT-IR, SCXRD, and advanced quantum mechanical calculations using Density Functional Theory (DFT) and Hartree–Fock (HF) methods. A comprehensive analysis of non-covalent interactions (NCIs) has been made by employing Hirshfeld surface, the quantum theory of atoms-in-molecules (QTAIM), and the reduced density gradient (RDG) techniques. Molecular docking of APMBS has been carried out against the target protein Tubulin-Colchicine: Stathmin-Like Domain Complex (PDB ID: 1SA0). Tubulin, consisting of an αβ-heterodimer, assembles into microtubules which are dynamic structures capable enough to polymerize and depolymerize exerting control over various essential cellular processes [16]. It plays a pivotal role in numerous cellular activities, including the maintenance of cell shape and cell division. This vital function of tubulin makes it an attractive and significant target in the development of anti-cancer drugs [17].
Experimental Methods
Synthesis
We used commercially available chemicals without purification. 1 mmol of 4-hydroxyphenyl acetamide and 1.5 mmol of K2CO3 were suspended in 5 mL of diglyme at 0 °C. Then, 1.1 mmol of p-toluene sulfonyl (tosyl) chloride was added in lots. The resulting mixture was stirred continuously for 30 min at 0 °C, followed by stirring for 2 h at 25 °C. After the conclusion of the chemical reaction, 10 mL of methylene dichloride (MDC) and 10 mL of water were added. The separated organic phase was washed twice with 5 mL of saturated NaHCO3 solution and then with brine. After collecting the organic layer and drying it over Na2SO4, solvent was removed through evaporation under vacuum conditions. The purification of the crude was achieved through column chromatography (silica gel = 300–400; ethyl acetate/petroleum ether = 1:3), thus yielding the desired product. Using a methanol solvent and a slow evaporation technique, good-quality single crystals have been produced. TLC (silica gel 60 F-254 plates) has been used to monitor the reaction progress. Ultraviolet light (UV) has been used for the visualisation of products. Scheme-1 shows the synthetic procedure of APMBS. Yield: 275 mg (90%); white solid; mp 130–132 °C.
Spectral Analysis
A Perkin Elmer FT/IR-4600typeA Spectrophotometer has been employed to record the FT-IR (KBr) spectra. An Agilent Technologies 1H NMR (at 400 MHz) and 13C NMR (at 100 MHz) spectrometer has been employed to record the NMR spectra. Using solvent resonance as the internal standard in dimethyl sulfoxide (DMSO) solution, the chemical shifts have been measured in parts per million (ppm) with reference to tetramethylsilane (TMS).
FT-IR (KBr, ν, cm−1): 3669.8, 2986.2, 1662.3 (C = O), 1503.2, 1376.9, 1151.2, 1091.5, 872.6, 545.7. 1H NMR (DMSO, 400 MHz) δ 7.60 (m, 2H, J = 8 Hz), 7.27 (d, 2H, J = 8 Hz), 7.16 (d, 2H, J = 8 Hz), 7.05 (brd, NH), 6.79 (d, 2H, J = 8 Hz), 6.32 (brd, NH), 3.36 (s, CH2); 2.36 (s, CH3); 13C NMR (DMSO, 100 MHz): δ 173.1 (C = O), 148.2, 145.5, 134.8, 132.0, 130.5, 129.9, 128.3, and 122.2 (Ar–C), 41.9 (CH2), 21.7 (CH3). HRMS (EI): m/z (M+) calcd for C15H15NO4S: 305.3419.
X-ray Intensity Data Collection and Structure Solution
Bruker Apex-II CCD diffractometer was employed to collect the intensity datasets with MoKα radiation (λ = 0.71073 Å). The data were collected using ω/φ scans at room temperature (290 K) with diffraction angles ranging from 4.1° to 50.1°. The data reduction and integration were performed by SAINT [18] program while absorption corrections using SADABS [19]. The SHELXT-2018 [20, 21] program was used for the structure solution (via intrinsic phasing), and the electron density maps (E-map) were used to determine the position of all non-H atoms. The SHELXL-2018 [22] program was used for the full-matrix least-squares on F2 to perform anisotropic refinement on all non-H atoms. During the refinement process, the H-atom positions were determined using difference Fourier maps, with an exception of those on C10, C14, and nitrogen atoms (fixed geometrically at the ideal coordinates), and all H-atoms were constrained to ride on their parent atoms. A total of 249 parameters were refined using 2524 unique reflections, which converged the final R-factor to 0.0526 [weighted R-factor (wR) = 0.145]; the goodness of fit in the last refinement cycle being S = 1.08. The atomic scattering coefficients were derived from the data provided in the International Tables for X-ray Crystallography [23]. The crystal structure and its geometrical features were obtained using PLATON [24], PARST [25], and MERCURY [26] software. The precise crystallographic data for APMBS are presented in Table 1.
Computational Methodology
The structural optimization of the system has been achieved by using Density Functional Theory (DFT) and Hartree–Fock (HF) approaches in its ground spin state in the gas phase, employing 6–311 + + G (d, p) basis set and B3LYP functional using the Gaussian 09W software [27, 28]. The HF calculations provided crucial insights into the optimal structural geometry while the DFT calculations help derive various electronic parameters, including EHOMO, ELUMO (Energy of Highest Occupied and Lowest Unoccupied Molecular Orbitals), the ∆Egap (energy gap of Frontier molecular orbitals), related quantum chemical parameters, DOS, atomic charges, and MEP. The frequency calculations using DFT at the same level of theory have been performed to correlate the theoretical predictions with the experimental results. The CrystalExplorer 17.5 software was used to calculate the crystal voids, Hirshfeld surfaces (dnorm, shape index, and curvedness plots), and the fingerprint plots [29]. The combined QTAIM/RDG studies were performed with the help of Multiwfn software [30] by using the.chk files outputted from the Gaussian single point energy calculations. These calculations provide valuable insights into the nature of NCIs within the molecular system and the presence of void spaces in the crystal lattice. The theoretical calculations were performed using atomic co-ordinates as retrieved from the CIF file. AutoDock Vina software [31] helped yield in silico molecular docking. The target protein Tubulin-Colchicine: Stathmin-Like Domain Complex (PDB ID: 1SA0) file was downloaded from the PDB database [32]. The.pdbqt [Protein Data Bank, Partial Charge (Q), & Atom Type (T)] files for both the protein and ligand molecule (APMBS) were prepared, and the grid box configuration was finalized using AutoDock Tools, with the binding site of the target identified by the grid centre coordinates set for docking simulations at X = 123.10, Y = 97.70, and Z = 5.99 dimensions. Discovery Studio 4.1 Visualizer software [33] has been used to visualise the complete ligand–protein interactions.
Results and Discussion
Synthesis and Spectroscopy
4-(2-amino-2-oxoethyl) phenyl 4-methylbenzenesulfonate was successfully synthesized in a single-step reaction using 4-hydroxyphenyl acetamide and tosyl chloride as starting materials. The reaction has been facilitated by the use of K2CO3 as a reagent at room temperature in diglyme solvent, employing a nucleophilic displacement mechanism. The selection of diglyme as the solvent has been driven by its environmentally friendly properties, including reduced volatility and lower toxicity compared to other conventional organic solvents commonly used in laboratory settings. The synthesis has been verified subsequently through various spectroanalytical techniques (Scheme 1).
The FTIR spectra of APMBS shows a sharp peak at 1662 cm−1 due to the C=O stretching of the unconjugated amide group. The N–H stretching peak is exhibited at 3669.8 cm−1. The characteristic peaks have been observed at 1376–1091 cm−1 range due to the presence of SO2 group. This confirms the product formation and also signifies the stability of the tosyloxy group. The theoretical IR spectra for APMBS is consonance with the corresponding experimental spectra (Fig. S1).
The 1H– and 13C–NMR spectra of the compound APMBS (Figs. S2 and S3) were acquired in DMSO-d6, and the signals were calibrated with respect to tetramethylsilane (TMS), and the chemical shifts were assigned to the respective protons and carbon atoms based on their chemical environment. The 1H NMR spectra of the APMBS shows two singlets at δ 3.36 and 2.36 ppm, integrating for two protons in the CH2 and three protons in the CH3 groups, respectively. Also, the ortho coupled aromatic protons having coupling constant J = 8.0 Hz exhibit four doublets in the range of 7.60–6.79 ppm, integrating eight protons of CH=CH. The NH protons were observed as a downfield singlet at 6.32 ppm. The 13C NMR spectra reveals eleven signals corresponding to the total number of carbon atoms. Two signals were observed in the aliphatic region, one at 41.9 ppm, which was ascribed to the carbon atom of the –CH2 group, and the other at 21.7 ppm, which was linked to the benzylic carbon or –CH3 group carbon atom. The signals for the six aromatic carbons of the phenyl rings were observed between 102 and 148 ppm. The carbonyl of the amide group was observed at 173.1 ppm. The proton and carbon spectra confirms the structure of APMBS.
X-ray and Optimized Structure Analysis
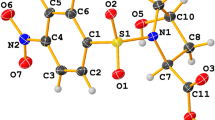
The compound crystallizes in the triclinic system (space group \(P\overline{1}\)) with Z = 2. The intramolecular hydrogen bond C6–H6⋯O2 forms S(5) graph set motif (Fig. 1). The amide group shows rotational disorder, occupying two positions with site occupancies of 0.50 for both parts. The dihedral angle between the two phenyl rings is 72.55° and the crystal structure is stabilized through C–H⋯O, C–H⋯π, and N–H⋯O hydrogen bonds (Table 2). The intermolecular weak H–bond C9–H9⋯O2 links the molecules along the a-axis and forms a C(6) chain (Fig. S4), while the intermolecular hydrogen bonds N1A–H1AA⋯O4A and C4–H4⋯O3 results in the formation of dimers having \({R}_{2 }^{2 }\left(8\right)\) and \({R}_{2 }^{2 }\left(10\right)\) ring set motifs (Fig. 2a and b). Beside this, a centrosymmetric inverted dimer has also been observed (Fig. 2c) which results in the formation of a supramolecular architecture (Fig. 3). The experimentally observed bond lengths and bond angles are in agreement with some analogous structures [34,35,36,37,38]. The structural optimization has been achieved using DFT and HF methods. The plot for variation in selected bond lengths and bond angles (Tables S1 and S2) are shown in Figs. S5 and S6, respectively. From the bond lengths plot, it is seen that the HF values are in more agreement with the experimental values while the DFT values are slightly higher than the experimental value but these deviations are within the acceptable range of experimental errors. However, from the bond angles plot, it is seen that the theoretical values of calculated bond angles are in agreement with the experimentally calculated angles with slight deviations except for the angles C8–O3⋯S1 and C11–C14⋯C15 which shows large deviation. This deviation of bond angles from the experimental structure might be attributed to the existence of rotational disorder within the amide group atoms.
Molecular Orbital Analysis
Frontier molecular orbitals (FMO) provide better insights about the biological potential and chemical reactivity of a molecule [39]. In the instant case, the HOMO–LUMO energies as EHOMO = − 6.906 eV and ELUMO = − 1.833 eV, respectively. The HOMO is localized over the whole molecular moiety, except for the methylbenzene part, while the LUMO too is localized, except for the acetamide group (Fig. 4). The energy gap (∆ELUMO-HOMO = 5.073 eV) and the global reactivity parameters as computed by using Koopman’s theorem [40] are presented in Table 3. A large value of the energy gap is indicative of reduced chemical reactivity and high kinetic stability, thus making the molecule chemically hard, with effective electron donating capacity.
DOS Analysis and Atomic Charge Distribution
The Density of States (DOS) is a representation of the number of states per unit energy in a material, as a function of energy. It acts as a useful tool when the discrete energy levels are artificially stretched into continuous curves. The Total DOS (TDOS) provides a comprehensive view of the system's overall orbitals, while the Partial DOS (PDOS) breaks down the total density of states into contributions from specific atomic or molecular orbitals within the system [41,42,43]. Figure 5 shows the TDOS and PDOS plots for the APMBS.
The position of HOMO molecular orbital is indicated by the dotted line at -0.25381 au in the vertical direction. The black and grey lines at the bottom show the position of various occupied and unoccupied orbitals, respectively. To analyse the contribution of s-, p- and d-orbitals to the whole system orbital, the PDOS spectra clearly indicates that the contribution of s- orbitals is more than that of the p-orbitals in the energy range − 1.25 to − 0.68 au (approx.) and the contribution of the p-orbitals increases as it moves towards the HOMO orbital. The contribution of d-orbitals in comparison to s- and p-orbitals to the occupied molecular orbitals is minimal. Also, the Frontier molecular orbitals do not have degenerate energy levels and energies are solely contributed by single HOMO and LUMO orbitals.
The atomic charge distribution plays an important role which affects the dipole moment and atomic polarizability of a molecular system [44]. The atomic charges for APMBS have been computed for Mulliken analysis by using HF/DFT techniques (Table S3, Fig. S7). The charge on nitrogen and oxygen atoms is negative while it is positive on the sulphur and hydrogen atoms: The magnitude of the charge varies as we move from HF to DFT, except for H1A and H1C.
MEP Analysis
The MEP surface generally predicts a molecule's reactivity by visual inspection of various coloured patches that correlate to various electrostatic potential (ESP) values. Maximum negative ESP (related with electrophilic reactivity) is indicated by the red colour zone, while the blue colour zone indicates maximum positive ESP (associated with nucleophilic reactivity) and the green represents the region of neutral ESP. Thus, the ESP increases in the order of red, orange, yellow, green, and blue across various coloured regions. In the MEP surface and contour map (Fig. 6a, b) of the molecule, as generated using the DFT, the area around the oxygen atoms bonded to the sulphur and carbon atom of the amide group exhibits negative ESP (red); thus making these sites prone to an electrophilic attack. On the contrary, the region around the hydrogens bonded to the nitrogen of the amide group and other carbon atoms exhibits positive ESP (blue) and makes these sites more prone to the nucleophilic attack. The presence of these reactive sites provides valuable insights into the molecule's biological activity and hence serves as confirmation of its good electrical properties [45, 46].
Hirshfeld Surface (HS) and Crystal Voids Analysis
To characterize various non-covalent intermolecular contacts in the crystal structure, HS and 2D fingerprint plots (2D-FPs) have been used. The dnorm surface has been used to describe the contacts shorter, longer and comparable to the van der Waals (vdW) separation using red, blue, and white colour on the dnorm surface [47]. The prominent deep red colour spots on the dnorm surface are due to the presence of shorter N–H⋯O and H–H intermolecular contacts while the tiny light red colour spots are due to the presence of shorter C–H⋯O intermolecular contacts (Fig. 7a). The shape index plot, being highly sensitive to even minor changes in the surface structure, helps identify the complementary bump convex and hollow concave regions located around the donor and acceptor atoms. In Fig. 7b, the presence of a red hollow region on the phenyl ring indicates the occurrence of a C–H⋯π interaction [48]. The lack of complementary red-blue regions on the shape index surface suggests that there is no π⋯π stacking interactions in the structure and this fact is further ascertained by the non-existence of flat green region (Fig. 7c).
To evaluate the impact of different molecular interactions within the crystal structure, fingerprint plots have been used which reveal complementary regions where one molecule serves as an acceptor (de < di) while the other acts as a donor (de > di) [49]. Figure 8 illustrates the plots of different interaction types, each contributing relatively to the overall pattern, while forming a complete fingerprint plot. The O⋯H/H⋯O intermolecular contacts, appearing in the form of two sharp symmetrical spikes in the FPs, contribute 27.8% to the total Hirshfeld surface and are associated with strong N–H⋯O and weak C–H⋯O hydrogen bond interactions. The highest contribution to the 2D FPs is from the H–H contacts which contributes 46.5% to the total HS, thus indicating the significance of weak non-covalent interactions in the crystal structure. The remaining 0.1% contribution which is not reflected in the plots, is from the S–O/O–S and the S–H/H–S contacts.
While assessing the mechanical stability of a crystal, a crucial factor demanding thorough investigation is the comprehensive analysis of crystal voids and their corresponding volumes [50]. The crystal voids occupy a volume of 43.23 Å3 and a surface area of 203.95 Å2, thus occupying 6.04% volume of the unit cell (714.64 Å3) (Fig. S8). Such a relatively low percentage of void space suggests that the molecules are tightly packed within the crystal structure, which in turn signifies a high degree of mechanical stability.
Combined QTAIM/RDG Theoretical Studies
Combining Quantum Theory of Atoms in Molecules (QTAIM) and Reduced Density Gradient (RDG) analysis provides a potent method for exploring non-covalent interactions (NCIs), viz. strong as well as weak interactions in the molecular systems. QTAIM assesses electron density features at bond critical points (BCPs), such as charge density (ρ), Laplacian of ρ (∇2ρ), and total energy density (H(r)). Positive H(r) signals non-covalent dominance, while negative H(r) indicates partial covalent character. Furthermore, positive H(r) and ∇2ρ signify weak interactions, whereas H(r) < 0 and ∇2ρ > 0 denotes relatively stronger interactions [51]. RDG analysis, utilizing color-mapped isosurfaces and scatter plots, correlates RDG with sign (λ2)ρ (λ2 is the Hessian eigen value). The value of sign (λ2)ρ < 0 indicates attractive interactions, sign(λ2)ρ > 0 signifies repulsive interactions, and sign(λ2)ρ ≈ 0 suggests vdW interactions while the ρ serves as a fundamental parameter to determine the strength of these interactions. Color intensity (blue for strong attractions, green for weak interactions, and red for repulsions) on 2D scatter plots and isosurfaces corresponds to interaction strength [52,53,54].
To describe various interactions existing in the crystal structure, we have used four dimers (Fig. 9a–d) along with colored mapped isosurfaces and QTAIM distributed BCPs and bond paths (BPs). The corresponding 2D scatter plots are shown in Fig. S9. Figure 9a describes the existence of C9–H9⋯O2 hydrogen bond characterized by the BCP and BP along with the green coloured RDG isosurface between H9 and O2 atom confirming its weak attractive nature as indicated by the dotted rectangle. Figure 9b describes the existence of N–H⋯O interactions [\({R}_{2 }^{2 }\left(8\right)\) ring set motif] characterized by the BCPs and BPs along with the little bluish green isosurfaces establishing its attractive nature. The bluish green isosurfaces indicates that these interactions are stronger relative to all other interactions in the crystal structure supported by the bluish green peak in the scatter plot (Fig. S9b) around ρ = 0.016 a.u. (approx.). Similary, the existence of \({R}_{2 }^{2 }\left(10\right)\) ring set motif formed by the C4–H4⋯O3 interactions (Fig. 9c) is characterized by the presence of BPs and BCPs (shown by dotted rectangles) and its attractive nature is supported by the green RDG isosurfaces between the H4 and O3 atoms. The greenish RDG isosurface localized over a large region in the two molecules (Fig. 9d) shows the existence of the of C–H⋯π interactions in the crystal structure and supported well by QTAIM BCPs and BPs between the H14B and C13 atom of the benzene ring (C8–C13). The topological parameters at BCPs for various interactions derived from the QTAIM analysis are given in Table S4. The C–H⋯O weak intramolecular hydrogen bond as mentioned in the crystal structure description is not evident in the QTAIM analysis of BCPs and BPs, suggesting the contact is extremely weak or it does not exist. However, the RDG analysis provides a unique perspective, revealing the presence of a green–red RDG isosurface between the interacting atoms which signifies an electron density distortion caused by the proximity of nearby atom [55]. This distortion is not be evident in the QTAIM analysis but is highlighted by the RDG analysis, emphasizing the electron density's role in characterizing various interactions in the QTAIM analysis. The red isosurfaces in the benzene rings are due to steric effect and sharp spikes appear in the 2D scatter plots around 0.024 a.u. corresponding to it. In addition, QTAIM analysis also predicts the existence of the weak C–H⋯N hydrogen bond interaction but this interaction is not observed geometrically.
Combined QATIM/RDG analysis showing a C–H⋯O hydrogen bond that form chains. b N–H⋯O hydrogen bond that forms \({R}_{2 }^{2 }\left(8\right)\) ring set motif. c C–H⋯O hydrogen bond that forms \({R}_{2 }^{2 }\left(10\right)\) ring set motif d Centrosymmetric inverted dimer linked through C–H⋯π interactions
Molecular Docking Analysis
The binding position of the ligand at the active site of the 1SA0 target protein is shown in Fig. 10a. The 2D binding interactions of the ligand at the binding site of target protein Tubulin (PDB code: 1SA0) are shown in Fig. 10b. Table 4 contains information about the binding energy, type of interaction, and binding distance of the ligand with the target protein Tubulin. The resulting ligand–protein complex is stabilized by three hydrogen bond (HBs), one electrostatic, and three hydrophobic bond interactions. The ARG158 residue engages in two conventional hydrogen bonds (HBs) with the ligand molecule. Specifically, the two hydrogen atoms of ARG158 form interactions with the ligand's oxygen atom at distances of 2.78 Å and 2.58 Å, respectively. Additionally, another conventional hydrogen bond occurs between the ligand's oxygen atom and the ARG112 residue, with a bond distance of 2.65 Å. The electrostatic interactions (Pi-Anion) occur between the six-membered ring of the ligand and oxygen atom of the active amino acid ASP199 at a distance of 3.85 Å. The hydrophobic interaction (Amide-Pi Stacking) takes place between the six-membered ring of the APMBS ligand and the nitrogen and carbon atoms of the corresponding residues, ARG264 and PRO263. This interaction is observed at a distance of 4.32 Å. The alkyl hydrophobic bonding interactions occur between the carbon atom of the ligand and the residues ARG264 and PRO263, with distances of 4.13 Å and 5.32 Å, respectively. The binding energy score of -7.9 (Kcal/mol) for ligand-1SA0 complex shows that APMBS has the potential to act as a potent anti-microtubule agent and hence can be useful in the development of anti-cancer drug.
Conclusions
In the present paper, a novel tosyl ester derivative, 4-(2-amino-2-oxoethyl) phenyl 4-methylbenzenesulfonate (APMBS), has been synthesized and characterized by 1H NMR, 13C NMR, FT-IR spectroscopic techniques. Crystallographic studies reveal that the compound APMBS crystallizes in the triclinic system with space group \(P\overline{1}\). The molecular packing is stabilized through a unique combination of weak intra- and intermolecular C–H⋯O and strong intermolecular N–H⋯O hydrogen bond interactions, besides the intermolecular C–H⋯π hydrogen bond interactions. FMOs analysis shows a large energy gap of 5.073 eV, thus predicting high stability and low reactivity for the molecule. DOS analysis shows the contribution of s-, p-, and d-orbitals to the overall system orbitals and predicts that d-orbitals contribute more to the occupied orbitals as compared to s- and p-orbitals. The MEP surface and atomic charge distribution indicates the presence of reactive sites prone to the nucleophilic and electrophilic attack in the molecule. The HS analysis confirms that the strong N–H⋯O, weak C–H⋯O and C–H⋯π interactions play a crucial role in the crystal structure stabilization. FPs analysis shows that H–H contacts contribute 46.5% to the total HS, thus indicating the significance of weak non-covalent interactions in the crystal structure. The combined AIM and RDG studies provide strong support for the HSA and 2D-FPs' assertion that the non-covalent interactions are crucial for the stabilization of the crystal structure. Molecular docking analysis shows that the molecule has the potential to be useful in the development of anti-cancer drug.
Data Availability
No datasets were generated or analysed during the current study.
References
Nangia AK, Desiraju GR (2019) Crystal engineering: an outlook for the future. Angewandte Chem Int Ed 58:4100–4107. https://doi.org/10.1002/anie.201811313
Desiraju GR (2013) Crystal engineering: from molecule to crystal. J Am Chem Soc 135:9952–9967. https://doi.org/10.1021/ja403264c
Biradha K, Su C-Y, Vittal JJ (2011) Recent developments in crystal engineering. Cryst Growth Des 11:875–886. https://doi.org/10.1021/cg101241x
Desiraju GR, Vittal JJ, Ramanan A (2011) Crystal engineering: a textbook. World Scientific, Singapore
Chopra D, Dey D (2014) Computational approaches towards crystal engineering in molecular crystals. Acta Crystallogr A Found Adv 70:C642–C642. https://doi.org/10.1107/S2053273314093577
Shukla R, Saeed A, Simpson J, Chopra D (2017) Quantitative investigation of C–H π and other intermolecular interactions in a series of crystalline N-(substituted phenyl)-2-naphthamide derivatives. CrystEngComm 19:5473–5491. https://doi.org/10.1039/C7CE01310H
Yao Z-F, Wang J-Y, Pei J (2018) Control of π–π stacking via crystal engineering in organic conjugated small molecule crystals. Cryst Growth Des 18:7–15. https://doi.org/10.1021/acs.cgd.7b01385
Garg U, Azim Y, Alam M (2021) In acid-aminopyrimidine continuum: experimental and computational studies of furan tetracarboxylate-2-aminopyrimidinium salt. RSC Adv 11:21463–21474. https://doi.org/10.1039/D1RA01714D
Weil ED (1991) Recent industrial organosulfur chemistry. Phosphorus Sulfur Silicon Relat Elem 59:31–46. https://doi.org/10.1080/10426509108045699
Lamberth C, Walter H, Kessabi FM et al (2015) The significance of organosulfur compounds in crop protection: current examples from fungicide research. Phosphorus Sulfur Silicon Relat Elem 190:1225–1235. https://doi.org/10.1080/10426507.2014.984033
Ariza JJ, García-López JD, Aguinaga-Casañas MA et al (2024) Enhancing food preservation and safety: synergistic effects of Allium-derived organosulfur compounds and outer membrane permeabilization peptide L-11. Food Control 165:110691. https://doi.org/10.1016/j.foodcont.2024.110691
Cremlyn RJ (1996) An introduction to organosulfur chemistry. John Wiley & Sons, Chichester
Patrick GL (2001) An introduction to medicinal chemistry, 2nd edn. Oxford University Press, Oxford
LisbetKværnø WM, Hauser H, Carreira EM (2005) Carbohydrate sulfonyl chlorides for simple, convenient access to glycoconjugates. Org Lett 7:1145–1148. https://doi.org/10.1021/ol0502127
Park NH, Teverovskiy G, Buchwald SL (2014) Development of an air-stable nickel precatalyst for the amination of aryl chlorides, sulfamates, mesylates, and triflates. Org Lett 16:220–223. https://doi.org/10.1021/ol403209k
Perez EA (2009) Microtubule inhibitors: differentiating tubulin-inhibiting agents based on mechanisms of action, clinical activity, and resistance. Mol Cancer Ther 8:2086–2095. https://doi.org/10.1158/1535-7163.MCT-09-0366
Jordan M, Kamath K (2007) How do microtubule-targeted drugs work? an overview. Curr Cancer Drug Targets 7:730–742. https://doi.org/10.2174/156800907783220417
Bruker (2017) SAINT data reduction and correction program v.8.38A. Bruker AXS Inc., Madison
Sheldrick GM (2016) SADABS (2016/2) Bruker/Siemens area detector absorption correction program. Bruker AXS, Madison
Sheldrick GM (2015) SHELXT—integrated space-group and crystal-structure determination. Acta CrystallogrA Found Adv 71:3–8. https://doi.org/10.1107/S2053273314026370
Dolomanov OV, Bourhis LJ, Gildea RJ et al (2009) OLEX2: a complete structure solution, refinement and analysis program. J Appl Crystallogr 42:339–341. https://doi.org/10.1107/S0021889808042726
Sheldrick GM (2015) Crystal structure refinement with SHELXL. Acta Crystallogr C Struct Chem 71:3–8. https://doi.org/10.1107/S2053229614024218
(1995) International Tables for Crystallography. Volume C. Corrigenda and Addenda to the First Edition. ActaCrystallogr A 51:441–444. https://doi.org/10.1107/S0108767395099958
Spek AL (2009) Structure validation in chemical crystallography. Acta Crystallogr D Biol Crystallogr 65:148–155. https://doi.org/10.1107/S090744490804362X
Nardelli M (1995) PARST 95—an update to PARST: a system of Fortran routines for calculating molecular structure parameters from the results of crystal structure analyses. J Appl Crystallogr 28:659–659. https://doi.org/10.1107/S0021889895007138
Macrae CF, Bruno IJ, Chisholm JA et al (2008) Mercury CSD 2.0—new features for the visualization and investigation of crystal structures. J ApplCrystallogr 41:466–470. https://doi.org/10.1107/S0021889807067908
Becke AD (1993) Density-functional thermochemistry. III. The role of exact exchange. J ChemPhys 98:5648–5652. https://doi.org/10.1063/1.464913
Frisch MJ, Trucks GW, Schlegel HB, et al (2016) Gaussian 09, Revision D.01. Gaussian Inc., Wallingford CT
Spackman PR, Turner MJ, McKinnon JJ et al (2021) CrystalExplorer: a program for Hirshfeld surface analysis, visualization and quantitative analysis of molecular crystals. J ApplCrystallogr 54:1006–1011. https://doi.org/10.1107/S1600576721002910
Lu T, Chen F (2012) Multiwfn: a multifunctional wavefunctionanalyzer. J Comput Chem 33:580–592. https://doi.org/10.1002/jcc.22885
Morris GM, Huey R, Lindstrom W et al (2009) Autodock4 and autodocktools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Biovia DS (2017) Discovery studio visualizer. Accelrys Inc., San Diego
Vembu N, Nallu M, Garrison J, Youngs WJ (2003) 4-Chlorophenyl 4-toluenesulfonate: supramolecular aggregation through C–H–O, C–H–Cl and C–H π interactions. Acta Crystallogr Sect E Struct Rep Online 59:o936–o938. https://doi.org/10.1107/S1600536803012066
Singh D, Sharma R, Nikam A et al (2024) Spectral, quantum chemical, X-ray crystallographic, Hirshfeld surface, energy framework, and molecular docking investigations of 4-acetylphenyl 4-methylbenzenesulfonate (APMBS). J Mol Struct 1295:136612. https://doi.org/10.1016/j.molstruc.2023.136612
Manivannan V, Vembu N, Nallu M et al (2005) 4-Nitrophenyl 4-toluenesulfonate: supramolecular aggregation through weak C–H–O, C–H π and van der Waals interactions. Acta Crystallogr Sect E Struct Rep Online 61:o75–o78. https://doi.org/10.1107/S1600536804032015
Vembu N, Nallu M, Spencer EC, Howard JAK (2003) Methyl 4-tosyloxybenzoate: supramolecular aggregation through C–H–O, C–H π and π–π interactions. Acta Crystallogr Sect E Struct Rep Online 59:o1009–o1011. https://doi.org/10.1107/S1600536803013333
Chen X, Yu M (2006) 4-Formyl-2-methoxyphenyl 4-methylbenzenesulfonate. Acta Crystallogr Sect E Struct Rep Online 62:o5775–o5776. https://doi.org/10.1107/S1600536806049221
Uzun S, Esen Z, Koç E et al (2019) Experimental and density functional theory (MEP, FMO, NLO, Fukui functions) and antibacterial activity studies on 2-amino-4-(4-nitrophenyl)-5,6-dihydrobenzo [h] quinoline-3-carbonitrile. J Mol Struct 1178:450–457. https://doi.org/10.1016/j.molstruc.2018.10.001
Manne R, Åberg T (1970) Koopmans’ theorem for inner-shell ionization. Chem Phys Lett 7:282–284. https://doi.org/10.1016/0009-2614(70)80309-8
Bougherara H, Kadri R, Kadri M et al (2021) Complex of 4-(2-aminophenyl)−1,2,3-thiadiazole with 2,3-dichloro- 5,6-dicyano-1,4-benzoquinone: Experimental study and investigation at different exchange-correlation functionals. DOS, NBO, QTAIM and RDG analyses. J Mol Struct 1223:128855. https://doi.org/10.1016/j.molstruc.2020.128855
Banerjee A, Das D, Ray PP et al (2021) Phenoxo-bridged dinuclear mixed valence cobalt( <scp>iii</scp> / <scp>ii</scp> ) complexes with reduced Schiff base ligands: synthesis, characterization, band gap measurements and fabrication of Schottky barrier diodes. Dalton Trans 50:1721–1732. https://doi.org/10.1039/D0DT03707A
Sagaama A, Issaoui N, Bardak F et al (2022) Non covalent interactions analysis and spectroscopic characterization combined with molecular docking study of N′-(4-Methoxybenzylidene)-5-phenyl-1H-pyrazole-3-carbohydrazide. J King Saud UnivSci 34:101778. https://doi.org/10.1016/j.jksus.2021.101778
Priya MK, Revathi BK, Renuka V et al (2019) Molecular structure, spectroscopic (FT-IR, FT-Raman, 13C and 1H NMR) analysis, HOMO-LUMO energies, Mulliken, MEP and thermal properties of new chalcone derivative by DFT calculation. Mater Today Proc 8:37–46. https://doi.org/10.1016/j.matpr.2019.02.078
Raja M, Raj Muhamed R, Muthu S, Suresh M (2017) Synthesis, spectroscopic (FT-IR, FT-Raman, NMR, UV–Visible), NLO, NBO, HOMO-LUMO, Fukui function and molecular docking study of (E)-1-(5-bromo-2-hydroxybenzylidene)semicarbazide. J MolStruct 1141:284–298. https://doi.org/10.1016/j.molstruc.2017.03.117
Ahamed JI, Narendran K, Ambika VR et al (2022) Synthesis, spectral characterization, DFT-computational analyses on novel 4-nitrobenzenaminium benzenesulfonate (4NBASA) crystal. J MolStruct 1266:133548. https://doi.org/10.1016/j.molstruc.2022.133548
Manna P, Seth SK, Das A et al (2012) Anion induced formation of supramolecular associations involving lone pair−π and anion−π interactions in Co(II) malonate complexes: experimental observations, Hirshfeld surface analyses and DFT studies. Inorg Chem 51:3557–3571. https://doi.org/10.1021/ic202317f
Seth SK (2013) Tuning the formation of MOFs by pH influence: X-ray structural variations and Hirshfeld surface analyses of 2-amino-5-nitropyridine with cadmium chloride. Cryst Eng Comm 15:1772. https://doi.org/10.1039/c2ce26682b
Tarahhomi A, van der Lee A, Dumitrescu DG (2019) Evaluation of N–H–O hydrogen bond interactions in two new phosphoric triamides with a P(O)[NHCH(CH 3) 2 ] 2 segment by means of topological (AIM) calculations, Hirshfeld surface analysis and 3D energy framework approach. Z Kristallogr Cryst Mater 234:401–413. https://doi.org/10.1515/zkri-2019-0005
Irrou E, Elmachkouri YA, Oubella A et al (2022) Crystal structure determination, Hirshfeld surface, crystal void, intermolecular interaction energy analyses, as well as DFT and energy framework calculations of 2-(4-oxo-4,5-dihydro-1 H-pyrazolo[3,4-d ]pyrimidin-1-yl)acetic acid. Acta Crystallogr E CrystallogrCommun 78:953–960. https://doi.org/10.1107/S2056989022008489
Kumar P, Frontera A, Pandey SK (2021) Coordination versusspodium bonds in dinuclearZn( <scp>ii</scp> ) and Cd( <scp>ii</scp> ) complexes with a dithiophosphate ligand. New J Chem 45:19402–19415. https://doi.org/10.1039/D1NJ03165A
Basak T, Das D, Ray PP et al (2020) Synthesis, characterization, self-assembly and non-ohmicSchottky barrier diode behaviors of two iron( <scp>iii</scp> ) based semiconductors with theoretical insight. CrystEngComm 22:5170–5181. https://doi.org/10.1039/D0CE00223B
BurudeghattaSundaramurthy C, KesthurNataraju CP, Krishnappagowda LN (2022) Design, synthesis, structural analysis and quantum chemical insight into the molecular structure of coumarin derivatives. Mol Syst Des Eng 7:132–157. https://doi.org/10.1039/D1ME00113B
Jenkins S, Morrison I (2000) The chemical character of the intermolecular bonds of seven phases of ice as revealed by ab initio calculation of electron densities. Chem Phys Lett 317:97–102. https://doi.org/10.1016/S0009-2614(99)01306-8
Km C, Al-Ostoot FH, Shehata EE et al (2021) Synthesis, crystal structure, Hirshfeld surface analysis, DFT calculations, 3D energy frameworks studies of Schiff base derivative 2,2′-((1Z,1′Z)-(1,2-phenylene bis(azanylylidene)) bis(methanylylidene)) diphenol. J Mol Struct 1244:130910. https://doi.org/10.1016/j.molstruc.2021.130910
Funding
No specific grants or funding organizations from the public, commercial, or non-profit sectors provided financial support for this study.
Author information
Authors and Affiliations
Contributions
D.S and R.S. carried out some experimental work and wrote the draft paper. A.N., J.J., S.S. conducted the synthesis and carried out chemical analysis. S.M. helped in molecular docking. R.K. designed the experiment, allowed the laboratory facilities and prepared the final manuscript for submission.
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest between the authors of this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Singh, D., Sharma, R., Nikam, A. et al. Supramolecular X-ray and Quantum Chemical Analysis of 4-(2-amino-2-oxoethyl) phenyl 4-methylbenzenesulfonate: A Tosyl Ester Derivative. J Chem Crystallogr (2024). https://doi.org/10.1007/s10870-024-01027-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10870-024-01027-0