Abstract
NMR isotope shifts occur due to small differences in nuclear shielding when nearby atoms are different isotopes. For molecules dissolved in 1:1 H2O:D2O, the resulting mixture of N-H and N-D isotopes leads to a small splitting of resonances from adjacent nuclei. We used multidimensional NMR to measure isotope shifts for the proteins CUS-3iD and CspA. We observed four-bond 4∆N(ND) isotope shifts in high-resolution 2D 15N-TROSY experiments of the perdeuterated proteins that correlate with the torsional angle psi. Three-bond 3∆C’(ND) isotope shifts detected in H(N)CO spectra correlate with the intraresidue H-O distance, and to a lesser extent with the dihedral angle phi. The conformational dependence of the isotope shifts agree with those previously reported in the literature. Both the 4∆N(ND) and 3∆C’(ND) isotope shifts are sensitive to distances between the atoms giving rise to the isotope shifts and the atoms experiencing the splitting, however, these distances are strongly correlated with backbone dihedral angles making it difficult to resolve distance from stereochemical contributions to the isotope shift. H(NCA)CO spectra were used to measure two-bond 2∆C’(ND) isotope shifts and [D]/[H] fractionation factors. Neither parameter showed significant differences for hydrogen-bonded sites, or changes over a 25° temperature range, suggesting they are not sensitive to hydrogen bonding. Finally, the quartet that arises from the combination of 2∆C’(ND) and 3∆C’(ND) isotope shifts in H(CA)CO spectra was used to measure synchronized hydrogen exchange for the sequence neighbors A315-S316 in the protein CUS-3iD. In many of our experiments we observed minor resonances due to the 10% D2O used for the sample deuterium lock, indicating isotope shifts can be a source of spectral heterogeneity in standard NMR experiments. We suggest that applications of isotope shifts such as conformational analysis and correlated hydrogen exchange could benefit from the larger magnetic fields becoming available.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chemical shifts are exquisitely sensitive to changes in magnetic environment. In staphylococcal nuclease for example, the single site mutation E43S changes the chemical shift for F76, some ~ 30 Å away on the other side of the protein (Wilde et al. 1988). Similarly, subtle differences in loop conformations due to proline cis/trans isomerization can double NMR signals throughout a protein (Alexandrescu et al. 1990; Alexandrescu et al. 1989; Kaplan et al. 2013; Kaplan et al. 2021). One can consider isotope shifts a manifestation of the sensitivity of chemical shifts (Hansen 1988). An isotope shift (specifically, secondary isotope shift) is a small chemical shift change for a nucleus when a nearby atom is replaced by a different isotope (e.g. 1H/2H, 12C/13C, or 14N/15N). For example, protein 15N backbone nuclei measured by NMR in a mixture of 50% H2O/ 50% D2O (henceforth 1:1 H:D) will have slightly different chemical shifts if they are directly bonded by amide protons (H) as opposed to deuterons (D) (Abildgaard et al. 2009). In the most commonly used nomenclature (Hansen 1988), which we employ in this paper, the one-bond isotope shift described above is designated 1∆N(D). The superscript indicates the number of bonds separating the site with alternative isotopes (H or D) from the nucleus (N) affected by the change. Isotope shifts can occur over multiple bonds, with shift size generally decreasing as the number of intervening bonds increases (Hansen 1988; Sun and Tugarinov 2012; Tuchsen and Hansen 1991; Uchida et al. 2005). Nevertheless, isotopes shifts up to 12 bonds away from the site of isotope substitution have been reported for small organic molecules (Berger and Künzer 1983).
Isotope shifts are thought to occur because the lower vibrational energy of the heavier isotope results in a shorter bond and higher nuclear shielding. This causes an upfield (lower frequency) shift for the heavier compared to lighter isotopologue (Hansen 1988). The factors contributing to the sizes of isotope shift are not fully understood but are thought to include bond length, bond order and hybridization, substituents, H-bonding, inter-atomic distances and stereochemical factors (Hansen 1988; Lambert and Greifenstein 1974). The latter dependence on stereochemistry and interatomic distances makes it possible to extract structural information from some of the isotope shifts (Abildgaard et al. 2009; LeMaster et al. 1994; Ottiger and Bax 1997; Sun and Tugarinov 2012; Tuchsen and Hansen 1991; Tugarinov 2013; Zhang and Tugarinov 2013).
Isotope shifts are additionally important because they can influence the appearance of NMR spectra. Uniform replacement of nonexchangeable hydrogens with deuterons by expressing proteins in D2O (perdeuteration) is common for NMR studies of large proteins. Perdeuteration can cause changes for 15N (Mal et al. 1998) and 13C chemical shifts (Sun and Tugarinov 2012) that can be represented as the sums of isotope shifts from all neighboring sites where hydrogen isotopes are replaced with deuterons (Maltsev et al. 2012). These isotope shifts can affect reported chemical shifts, chemical shift-based dihedral restraints in NMR structure calculations, and analyses of residual structure using chemical shift indices. Methods have been proposed to correct for chemical shifts changes arising from perdeuteration (Venters et al. 1996). Isotope effects can also modulate scalar relaxation through H/D isotope exchange (Kateb et al. 2007).
Isotope shifts can be characterized for backbone as well as sidechain nuclei. 2∆C’(ND) isotope shifts have been used for ‘E and Z’ assignments of Asn and Gln -NH2 groups (Liu et al. 2008), while 1∆N(D) isotope shifts can provide information on hydrogen-bonding and sidechain amide rotameric states (Liu et al. 2008). Line-shape analysis on signals from H/D isotopologues has been used to study fast HX in side-chain Thr and Ser hydroxyl, and Cys sulfhydryl groups (Takeda et al. 2014). Using exchange spectroscopy, hydrogen exchange (HX) rates for aromatic hydroxyls in Tyr residues were determined from the 2∆Cζ(OD) isotope splitting that arises depending on whether the phenol group is occupied by H or D (Takeda et al. 2009). Similarly, the SOLEXSY experiment measures fast backbone amide exchange rates through exchange spectroscopy of N(H) and N(D) signals for proteins dissolved in 1:1 HD (Chevelkov et al. 2010).
An application of isotope shifts of particular interest to our lab is to investigate the degree to which HX is correlated between different residues. HX studies have provided insight into protein solvent accessibility, dynamics, and stability (Dempsey 2001). Yet little is known about the structural basis of the “opening” processes that contribute to exchange, in particular the degree to which HX events occurs simultaneously at different labile sites. The most general method to obtain this information is through NOESY experiments that measure correlated exchange through HN-HN NOEs (Roder et al. 1985; Wagner 1980; Wang et al. 1987). In spite of the elegance of this method it may not be applicable to all types of structures, for example sequential HN-HN NOEs becomes vanishingly small in β-sheet structure (Wüthrich 1986). An alternative approach for these cases is to establish connections between HN amide protons across peptide bonds. This can be done through isotope shifts (and in principle through J-couplings). Thus 2∆C’(ND) and 3∆C’(ND) isotope effects, also called β- and γ-shifts, were used to compare the amount of correlated HH to DD exchange compared to single site (HH to HD or DH) exchange in 1D-NMR studies of selectively 13C-labeled subtilisin inhibitor (Uchida et al. 2005).
Here we use 2D and 3D NMR methods to investigate protein backbone isotope shifts. The studies were conducted on two small globular proteins containing mostly antiparallel β-barrel structures. CUS-3iD (115-residues, 11.9 KDa) is the insertion domain from the coat protein of phage CUS-3 (Tripler et al. 2019, 2015). CspA (69-residues, 7.3 KDa) is the mRNA-chaperone cold shock protein A from E. coli (Feng et al. 1998). We used high-resolution 1H-15N TROSY spectra to measure trans-peptide 4∆N(ND) isotope shifts and analyzed their sizes in terms of psi backbone dihedral angles and N(i)-HN(i + 1) distances across peptide bonds. We used HNCO spectra to measure intraresidue 3∆C’(ND) ‘γ’ isotope shifts. The data tested a previously reported correlation between the size of the γ-shift and a proposed 5-atom intra-residue HN to O atom distance (Tuchsen and Hansen 1991), as well as the 4-atom HN to C’ distance. We compared H(CA)CO and H(NCA)CO experiments for measuring 2∆C’(ND) ‘β’ isotope shifts and ϕ = [D]/[H] fractionation factors, and looked at the temperature dependence of the two parameters. We tested the utility of H(CA)CO spectra for measuring correlated hydrogen exchange between neighboring amino acids in the sequence. We discuss how the ubiquitous 10% solvent fraction of D2O commonly used for the NMR deuterium lock (henceforth 9:1 H:D) can be a source of spectral heterogeneity due to isotope shifts. Finally, we comment on how measurements of isotope shifts would benefit from the availability of higher magnetic fields.
Experimental
NMR spectroscopy
2H, 13C, 15N triple-labeled (henceforth 3L) and 13C, 15N double-labeled (henceforth 2L) samples of CUS-3iD (Tripler et al. 2019, 2015) and 3L-CspA (Alexandrescu et al. 2001) were produced as previously described. Samples were dissolved in either 9:1 H:D (standard sample conditions, with 10% (%v/%v) D2O for the deuterium lock), or 1:1 H:D (50% H2O/50% D2O, %v/%v) for measuring isotope shifts. To ensure consistency between solvent fractions, samples were split into two tubes and lyophilized. One aliquot was taken up in 9:1 H:D the other in 1:1 H:D. To achieve isotope equilibration, the CUS-3iD samples were incubated for two weeks at 30 °C, since separate HX experiment monitored by 1H-15N HSQC spectroscopy showed that solvent exchange was mostly complete after 1 week at 25 °C and pH 6.3. CspA exchanges much faster (Jaravine et al. 2000), so samples were equilibrated for one day in 1:1 H:D or 9:1 H:D at pH 5.4 at 25 °C. Final sample conditions were: 2L-CUS-3iD, 0.7 mM protein at pH 6.3; 3L-CUS-3iD, 1.0 mM protein at pH 6.1; 3L-CspA, 1.0 mM protein at pH 5.4. All samples were contained in Shigemi microcells with volumes between 220 and 270 µL.
Experiments were done on a Bruker Avance 600 MHz solution NMR instrument equipped with a TCI cryogenic probe, that is part of the Francis Bitter Magnet Lab of MIT. Topspin 2.1 software was used for NMR data acquisition and processing. Alternatively NMR spectra were processed in iNMR 6.3 (Mestrelab Research, Santiago de Compostela, Spain) and analyzed with CcpNmr Analysis version 2.5.2 (Vranken et al. 2005). Published assignments were used to analyze the NMR data for Cus3-ID (BMRB 25263) and CspA (BMRB 4296). Unless otherwise noted, all experiments were done with sequences from the Bruker pulse program catalogue.
4∆N(ND) isotope shifts
To characterize 4∆N(ND) isotope shifts, 1H-15N TROSY data on 3L samples of CUS3-iD and CspA in either 1:1 H:D or 9:1 H:D were obtained using the pulse sequence trosyetf3gpsi. 13C decoupling was achieved with the Bruker Topspin “-Dlabel_CN” flag in ZGOPTNS. The flag refocuses 13C-15N J-coupling in the center of the N evolution period using a smooth linear 180° Chirp shaped pulse on 13C (Crp60,0.5,20.1). The d1 interscan delay was 1 s. Acquisition times were 1HN 114 ms and 15N 913 ms, for FID resolutions of 1HN 8.8 Hz and 15N 1.1 Hz. Total experiment times for each 1H-15N TROSY experiment were 22 h. The data were processed using Topsin 2.1 with Gaussian window functions of GB = 0.05, LB = − 1 for 1HN, and GB = 0.24, LB = − 3 for 15N, to digital resolutions of 2.0 Hz/pt for 1HN, and 0.2 Hz/pt for 15N. The TDeff parameter of the Bruker Topspin software was used to test how many points were needed to resolve the isotope shifts in the 15N dimension. With TD1 = 1024 (310 ms acquisition time) all but the strongest isotope shifts were unresolved; most could not be separated with less than TD1 = 2048 (620 ms acquisition time), with TD1 = 3000 (913 ms acquisition time) being optimal for the experiments.
The temperature for the CUS-3iD experiments was 303 K while that for CspA was 298 K. Uncertainties in 4∆N(ND) isotope shifts were estimated by repeating the 1H-15N TROSY experiment on a separate 0.8 mM 3L-sample of CUS-3iD in 1:1 H:D. Additional data for CUS-3iD to compare spectra in 1:1 H:D, 9:1 H:D, and 99.96% D2O were obtained at 288 K, with the lower temperature chosen to reduce the extent of exchange in D2O. The partially exchanged spectrum was obtained 2.5 days after dissolving 3L CUS-3iD in D2O.
We also examined if 4∆N(ND) isotope shifts could be measured using 15N-HSQC experiments using the pulse sequence hsqcetf3gp, rather than 15N-TROSY. The HSQC data were inferior to TROSY both in terms of sensitivity and line-shape. Although CUS-3iD is only an 11.9 KDa protein, the very long 15N acquisition times needed to resolve the 4∆N(ND) isotope shifts benefit from the optimal 15N transverse relaxation achievable with 1H-15N TROSY compared with 1H-15N HSQC experiments. We therefore concluded that the optimal detection of 4∆N(ND) isotope shifts requires 15N-TROSY experiments in conjunction with perdeuterated 3L protein samples.
3∆C’(ND) isotope shifts
3∆C’(ND) isotope shifts were determined from 3L samples of CUS3-iD and CspA with 2D H(N)CO spectra obtained with the pulse program hncogp3d_HJdec_CO. This is a modified version of the Bruker hncogp3d sequence that includes a 180° 1H pulse with constant phase 0 in the center of the C’ evolution d0 delays, to ensure no long-range 1H J-couplings affected C’ lineshapes. Initially studies were done on 2L samples of CUS3-iD, however, we switched to 3L samples to improve sensitivity and reduce sources of long-range 1H J-couplings. With the aliphatic protons removed in 3L samples, the 2D H(N)CO experiments showed no difference when the 180° 1H pulse was included or excluded, thus any residual splitting is due to isotope shifts or unresolved 13C-13C J-couplings. Spectra were acquired for samples in both 1:1 H:D and 9:1 H:D to identify the isotope shifts. The interscan delay d1 was 1 s. The data for CUS-3iD were obtained at 288 oK, those for CspA at 298 oK. Acquisition times were 1HN 143 ms and 13C’ 377 ms, for FID resolutions of 1HN 3.5 Hz and 13C’ 1.3 Hz. Total acquisition times were 9 h. The data were processed using iNMR with SB = 90 for 1HN and SSB = 50° for 13C’, with zero-filling to digital resolutions of 3.8 Hz/pt for 1HN and 0.16 Hz/pt for 13C’. Uncertainties in 3∆C’(ND) isotope shifts were estimated by repeating the H(N)CO experiment on a separate 0.8 mM 3L-sample of CUS-3iD.
Temperature dependence of 2∆C’(ND) isotope shifts and ϕ fractionation factors
2D H(NCA)CO spectra obtained with the Bruker pulse program hncacogp3d were used to determine 2∆C’(ND) isotope shifts and ϕ fractionation factors for 3L samples of CUS3-iD. The interscan delay d1 was 1 s, and acquisition times were 1HN 132 ms and 13C’ 110 ms, for FID resolutions of 1HN 3.8 Hz and 13C’ 4.5 Hz. Total experiment times for each spectrum were 3.5 h. To look at the temperature dependence of the parameters, data were obtained at six temperatures: 288 oK, 293 oK, 298 oK, 303 oK, 308 oK, and 313 oK. Spectra were processed using iNMR with Gaussian functions of GB = 0.63, LB = 15 for 1HN and GB = 0.63, LB = 1 for 13C’, to digital resolutions of 3.8 Hz/pt for 1HN and 0.66 Hz/pt for 13C’. Uncertainties in 3∆C’(ND) isotope shifts and ϕ values were estimated two ways. In the first, H(NCA)CO spectra were collected on a separate 0.8 mM 3L-sample of CUS-3iD. In the second, an H(NCA)CO spectrum was obtained on the same sample used for the temperature titration, since the entire temperature series was collected on one sample. Errors from the two approaches were similar. The 2∆C’(ND) isotope shifts were calculated from the chemical shift difference between the upfield (D) and downfield (H) components of 13C doublets obtained for 3L CUS-3iD in 1:1 H:D. The fractionation factor ϕ was calculated from the ratio of uncorrected peak heights ID/IH using the program CcpNmr Analysis.
H(CA)CO spectroscopy to investigate correlated exchange
To test the feasibility of obtaining data on correlated HX using multidimensional NMR, a 0.7 mM 2L CUS-3iD sample in 99.96% D2O at a temperature of 288 K, was investigated with 2D H(CA)CO spectra using the Bruker pulse sequence hcacogp3d. The 1H offset was set to 6.5 ppm to avoid suppression of signals in the 1Hα region, and the 1H spectral window was set to 9 ppm. Acquisition times were 1HN 95 ms and 13C’ 377 ms, for FID resolutions of 1HN 5.3 Hz and 13C’ 1.3 Hz. Total experiment times, depending on the number of transients averaged, were 3 h for the earliest time point following dissolution in D2O, switched to 6 h for later time points between 9 and 117 h, and 24 h for the last three time points between days 5 and 8. Data were processed with the iNMR program using Gaussian functions of GB = 0.02, LB = 9 for 1Hα and GB = 0.17, LB = 0.5 for 13C’, and zero-filled to digital resolutions of 5.3 Hz/pt for 1HN and 0.17 Hz/pt for 13C’. The correlated exchange parameter Xβγ for residues A315-S316 was calculated by line-fitting the resolved quartet components of the A315 signal in the spectrum at 9 h, using the integration approach previously described (Uchida et al. 2005).
Structure analysis
To correlate isotope shifts with structural parameters we used the CUS3-iD segment (Chain A of the asymmetric unit, residues 223–337) from the 2.8 Å-resolution cryo-EM structure of phage CUS-3 (deposited under PDB accession code 8SKG, Richard Whitehead and C.M.T., in preparation), and the 2.0 Å-resolution X-ray diffraction structure of CspA (PDB 1MJC). Backbone dihedrals were obtained with the “Torsion Angles” function of the What If (Vriend 1990) server (https://swift.cmbi.umcn.nl/servers/html/chiang.html). Interatomic distances were calculated with the “get_distance” command of PyMol.
Results
4∆N(ND) isotope shifts depend on the psi-angle and N(i) to H(i + 1) distance
15N-TROSY spectra obtained with high resolution in the indirect dimension show splitting of 15N signals for 3L-CUS-3iD samples in 1:1 H:D. The splitting is not detected in equivalent samples in H2O, indicating it is due to a H/D isotope shift (Fig. 1A). Possible mechanisms for the isotope shift are shown in Fig. 1B. In this and subsequent schematics, the green star indicates the nucleus for which the isotope splitting is observed, and the green arrow points to the proton detected in ‘inverse’ heteronuclear NMR experiments (Cavanagh et al. 2006). Importantly, isotope shifts cannot be seen for the atoms bound to protons used for NMR detection, thus we can rule out a one-bond 1∆N(D) isotope effect. The amide nitrogen of residue i could be occupied by H or D, however, only H will be detected in 1H-15N TROSY. The fraction of molecules with D on the nitrogen i will result in a proportional loss in signal. The remaining possibility for the isotope effect is a 4∆N(ND) four-bond shift that could involve one of the three mechanisms outlined in Fig. 1B. The through H-bond pathway can be ruled out because amide nitrogens involved in H-bonds in the structure of CUS-3iD, as well as those that are not H-bonded show the 4∆N(ND) isotope shift splitting. Of the remaining pathways two observations indicate that the subsequent residue, i + 1, dominates the 4∆N(ND) isotope shift splitting for the nitrogen of residue i (red in Fig. 1B). The first is that none of the residues preceding one of the seven prolines in the CUS-3iD sequence show the isotope effect. Because proline does not have an amide hydrogen it cannot undergo an H/D equilibrium. By contrast five of the seven residues following a proline have an isotope shift suggesting the effect is due to residue i + 1. The second observation is the pattern of isotope shifts when CUS-3iD is partially exchanged in D2O (Fig. 1C). The pink contours in Fig. 1C are due to amide protons in CUS-3iD that survive exchange with D2O. The superpositions of spectra is shown “as is”, without any offset. The pink spectrum for the sample in D2O has 1H signals showing a slight downfield shift of ~ 0.015 ppm compared to the green spectrum in 1:1 H:D. This 1H shift could itself be due to an isotope effect from the protein or solvent (Hansen 1988) but was not analyzed further. For the 15N isotope shift, some of the HX-protected TROSY correlations overlay on the downfield component of the 1:1 H:D doublet (H signal), some are split and show both upfield and downfield signals, and some overlay on the upfield component (D signal). Figure 1D shows the HX protection pattern for the CUS-3iD sequence. Residues shown in red represent clusters of HX protected residues that are contiguous in the sequence. The residues shown in blue are those that are protected from HX but are followed by an unprotected residue in the sequence. All of the HX protected residues in the D2O spectrum that overlay with the upfield component of the 15N isotope shift doublet are followed by an unprotected residue in the CUS-3iD sequence (blue in Fig. 1D). Conversely, residues with a downfield or split 15N signal, are those followed by an HX-protected residue (red in Fig. 1D). A few of the residues give split signals in D2O, with sufficient resolution to see the downfield (H) signal decrease at the expense of the increasing upfield (D) signal, as the amide proton from the subsequent residue exchanges with D2O. We therefore conclude that the 4∆N(ND) isotope shift of residue i is dominated by the H/D isotopomer state of residue i + 1, although we do not understand why residue i-1 does not contribute. Consistent with our interpretation, measurements of 4∆N(ND) isotope shifts with an isotope-filtered method showed that the 4∆N(Ni+iD) isotope shifts were much larger than 4∆N(Ni-iD) (Tugarinov 2013).
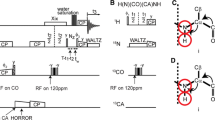
The 4∆N(ND) isotope shift. A Superposition of 15N-TROSY spectra for 3L-CUS-3iD in 9:1 H:D (H2O, black contours) and 1:1 H:D (green contours). The data were obtained at a temperature of 303 oK. Spectra obtained in 9:1 H:D are offset by + 0.05 ppm in the 1H dimension to better illustrate the splitting in 1:1 H:D, denoted by vertical red lines. For comparison, the same spectra without the offset are shown in Fig. S1A. For the sample in 9:1 H:D (black) small peaks from D isotopologues can sometimes also be seen at low contours (black arrows) due to the 10% D2O used for the deuterium lock. B Schematic showing three possible origins for a 4∆N(ND) isotope shift: (1) the subsequent residue in the sequence (i + 1, red), (2) the preceding residue (i-1, purple), (3) a four-bond connection mediated through an H-bond (orange). As described in the main text, our observations suggest the subsequent residue makes the dominant contribution to the isotope shift (red pathway). C Superposition of 15N-TROSY spectra for 3L-CUS-3iD in 1:1 H:D (green) and partially exchanged in 99.96% D2O (purple). The data were collected at 288 oK to minimize HX. Surviving protected amide protons in D2O retain either the upfield or downfield component of the doublet seen in the 1:1 H:D solvent. D Sequence of CUS-3iD showing amide protons protected from exchange in D2O and whether the shift of the protected amide proton corresponds to the upfield (blue) or downfield (red) component of the doublet observed in 1:1 H:D. The protection pattern shows that the upfield component appears when the subsequent residue is not protected, consistent with a D atom at the subsequent nitrogen in the sequence
As with other types of isotope shifts (Abildgaard et al. 2009; Hansen 1988; Mal et al. 1998; Maltsev et al. 2012) the downfield and upfield components of 4∆N(ND) correspond to H and D isotopes, respectively (Fig. 1A). Of 89 residues in CUS-3iD that could be analyzed 14 show no detectable splitting. Most of these were in β-sheet secondary structure. The remaining resolvable isotope shifts range from 3 to 66 ppb (Supplementary Table S1), with a mean value and standard deviation (s.d.) of 32 ± 15. Based on duplicate measures on CUS-3iD samples, we estimate that the error in the in the 4∆N(ND) isotope shifts was about ± 3 ppb, or 10% of the average 32 ppb size of non-zero 4∆N(ND) isotope shifts. The largest 4∆N(ND) isotope shifts were seen for glycines, the residues with the highest propensity to be found outside of regular secondary structure.
We examined possible relationships between 4∆N(ND) isotope shifts and secondary structure, and found a correlation with the backbone dihedral angle psi (Ψ). The data can be fit to a Karplus-type equation as shown in Fig. 2A. It is notable that the backbone dihedral angle psi is defined by the four atoms N(i), Cα(i), C’(i), N(i + 1), the same atoms spanned by the 4∆N(ND) isotope shift (red in Fig. 1B). As an additional test, we looked at the correlation between 4∆N(ND) and the distance from the site of isotope exchange H(i + 1) to the site experiencing the isotope shift N(i), henceforth dNH(i,i + 1). A similar relationship between 4∆N(ND) and the distance dNH(i,i + 1) was reported previously (Tugarinov 2013). The correlation between 4∆N(ND) and the dNH(i,i + 1) distance is comparable to that with the dihedral angle psi (Fig. 2B), since psi and dNH(i,i + 1) are correlated in protein structures (Fig. S2A, B). Alpha helical conformations have psi angles near zero that bring the N and H atoms in the five-atom sequence N-Cα-C’-N-H close together, whereas the extended β-strand conformation results in larger distances between the two atoms.
Correlations between 4∆N(ND) isotope shifts and structure parameters. A Relationship between 4∆N(ND) and backbone psi values. The blue curve shows a least square fit of the data to a Karplus equation with the fitting parameters 4∆N(ND)/ppb = (− 1.4 ± 4.1)cos2Ψ + (20 ± 2)cosΨ + (32 ± 3), R-value = 0.75. The Karplus coefficients were within experimental error when the data for CUS-3iD and CspA were fit individually, with R-values of 0.78 and 0.74, respectively. The error bars for the CUS-3iD are from duplicate measurements. B Correlation between 4∆N(ND) and the distance from the site of the isotope effect to the split NMR signal dNH(i,i + 1). Line-fit parameters are 4∆N(ND) = − 32 ± 3*dNH(i,i + 1) + 151 ± 10, R-value = 0.72. C Ramachandran map for the CUS-3iD cryo-EM structure with residues color-coded according to sizes of the 4∆N(ND) isotope shifts. The smallest 4∆N(ND) values are for residue in β-sheet regions (green), the largest for psi angles near zero (red). D Ramachandran map for the CspA X-ray structure. Approximate probability potentials for the Ramachandran map are shown with solid and dashed lines for favorable and allowed regions, respectively
The 4∆N(ND) isotope shift shows clear secondary structure discrimination (Fig. 2C). The smallest 4∆N(ND) isotope shifts below 25 ppb (or for those residues where an isotope shift cannot be resolved) occur for residues in extended β-sheet structure. The largest 4∆N(ND) isotope shifts, greater than 50 ppb, occur for residues near the left and right-handed α-helical regions, with psi angles near 0. A previous study reported that scalar 3JNN couplings depend on backbone psi angles (Theis et al. 1997). The sizes of protein 3JNN backbone couplings are in the range of 0.1 to 0.3 Hz, an order of magnitude smaller than the sizes of 4∆N(ND) isotope shifts at 60 MHz. The dependence of 3JNN backbone couplings on the psi dihedral angle is opposite to that of the 4∆N(ND) isotope shifts, with the largest couplings occurring for β-sheet conformations and the smallest for psi angles near 0 degrees.
To further test the relationship between 4∆N(ND) and structural parameters, we examined a second protein CspA (Table S2) for which there is a 2.0 Å resolution X-ray structure available (PDB code 1MJC, (Schindelin et al. 1994)). The results for CspA and CUS-3iD were similar for dNH(i,i + 1) distances and psi dihedral angles (Fig. 2A, B). The discrimination of 4∆N(ND) isotope shifts for α-helix and β-sheet is also evident for CspA (Fig. 2D).
Intraresidue 3∆C’(ND) isotope shifts depend on the distances H-N-Cα-C’-O and H-N-Cα-C’
Protein backbone carbonyl signal splitting due to three-bond 3∆C’(ND) (Fig. 3A) and two-bond 2∆C’(ND) isotope shifts (Fig. 4A) were first described some 40 years ago. The isotope shifts were used to study hydrogen exchange (Kainosho and Tsuji 1982), as well as to obtain some of the first site-specific NMR resonance assignments for proteins (Kainosho et al. 1985; Tuchsen and Hansen 1988). The larger inter-residue isotope shift 2∆C’(ND), is sometimes called the β-shift. The smaller intra-residue isotope shift 3∆C’(ND), is also called the γ-shift (Fig. 4A).
The 3∆C’(ND) isotope shift. A Illustration of the measurement of 3∆C’(ND) isotope shifts from an H(N)CO spectrum. In the diagram the green arrow indicates the proton used for NMR detection, the green”*” indicates the signal split by the H/D isotope shift, and the red arrow shows the origin of the 3∆C’(ND) isotope shift. B Superposition of H(N)CO spectra for 3L-CUS-3iD in 9:1 H:D (black contours) and 1:1 H:D (green contours). The red vertical lines denote the 3∆C’(ND) isotope shift splits. The spectra obtained in 9:1 H:D are offset by + 0.05 ppm in the 1H dimension to better illustrate the splitting in 1:1 H:D. For comparison, the same spectra without the offset are shown in Fig. S1B. C Correlation between the sizes of 3∆C’(ND) isotope shifts and the H to O distance, dHO in the intra-residue 5-atom ring H-N-Cα-C’-O. Data from the proteins CUS-3iD and CspA are shown by blue circles and brown squares, respectively. The linear fit of all data gave 3∆C’(ND) = − 18 ± 1*dHO + 80 ± 4, R-value = 0.88. The individual proteins CUS-3iD and CspA gave fitting parameters within experimental errors of those obtained for all the data, and R-values of 0.87 and 0.88, respectively. D Correlation between the sizes of 3∆C’(ND) isotope shifts and the distance dHC’ in the 4-atom sequence H-N–Cα-C’ (one atom less than in panel C). The linear fit of all data gave 3∆C’(ND) = − 47 ± 5*dHC’ + 160 ± 14, R-value = 0.73. Error bars in C and D are from duplicate experiments for CUS-3iD
2∆C’(ND) isotope shifts and ϕ fractionation factors from H(NCA)CO experiments. A Illustration of the measurement of 2∆C’(ND) “β” isotope shifts (solid red) from H(CA)CO spectra. B Schematic showing 2∆C’(ND) “β” isotope shift in H(NCA)CO experiments. Additional 3∆C’(ND) “γ” isotope shifts (dotted red lines) could also be obtained from both H(CA)CO or H(NCA)CO experiments, if they are acquired with sufficient resolution in 13C (about 2 × as much as for the β-shifts). C H(NCA)CO spectrum recorded on a 2L-sample of CUS-3iD at 303 K. D H(NCA)CO spectrum collected on a 3L-sample of CUS-3iD,with the exact same parameters as in C. Note the improvement in sensitivity due to perdeuteration. E Expansion of the H(NCA)CO spectrum of 3L-CUS-3iD at 303 K, corresponding to the blue rectangle in D. Green contours are from a sample in 1:1 H:D. Black contours are from a sample in 9:1 H:D. For clarity, the spectra in 9:1 H:D were shifted downfield by 0.05 ppm. Spectra without this offset are shown in Fig. S1C. At sufficiently low contour levels, isotope shifts from the 10% deuterated population can be readily seen in the black spectrum, as indicated by the horizontal arrows. Most amino acids in the H(NCA)CO spectrum show the additional expected weaker sequential HN(i)-C’(i-1) crosspeaks, as exemplified by the A259-S258. The 13C resolution (9 Hz, 110 ms acquisition time) was insufficient to detect the additional potential γ splitting for this sequential C’(i-1) signal (dotted red arrow in B). (F) Same expansion as in E showing the effects of temperature. The 2∆C’(ND) values were obtained from the isotope shift splitting in the 13C dimension, the ϕ-values from the ratio of intensities of the D (upfield) and H (downfield) components
Because the 3D HNCO experiment has the highest sensitivity amongst triple-resonance experiments, we anticipated it would lend itself to measurements of 3∆C’(ND) isotope shifts as outlined in the scheme of Fig. 3A. For a protein sample in 1:1 H:D, the HNCO spectrum is expected to give the 3∆C’(ND) isotope shift of the previous residue in the sequence but not 2∆C’(ND). As described previously, isotope shifts arising from the amide proton used for detection in the HNCO experiment (green arrow in Fig. 3A) cannot be measured. The H(N)CO experiment is therefore selective for only the 3∆C’(ND) isotope shift of the previous residue. A portion of the 2D H(N)CO spectrum is shown Fig. 3B. The black contours are from a 2D H(N)CO spectrum obtained for CUS-3iD in H2O. The green contours are from a spectrum for the protein in for 1:1 H:D and show 3∆C’(ND) splittings in the 13C-dimension indicated by the red vertical lines. A complication for the measurement of 3∆C’(ND) isotope shifts with uniformly 13C-labeled samples, is that additional splittings are often observed. We think these are likely due to unidentified long-range two and three-bond 13C-13C scalar couplings that are comparable in size to the 3∆C’(ND) isotope shifts. Even though we used perdeuterated samples to avoid 1H-13C scalar couplings, the sizes of 2JCβC’ (0.4 Hz), 2JCαC’ (0.6 Hz), 3JC’C’ (2 Hz), and 3JCβC’ (2 Hz) 13C-13C couplings (Lohr and Ruterjans 1999; Schmidt et al. 2010) all approach the average 3∆C’(ND) isotope shift of 4 Hz at the 150 MHz 13C-frequency of the spectrometer used in this work. Interference from 13C-13C scalar couplings would not have been a problem with the earlier experiments that used natural 13C-abundance, or selectively 13C-labeled protein samples. Our approach to circumventing interference from 13C-13C scalar couplings was to carefully compare the splitting pattern obtained in H2O and 1:1 H:D samples. The J-couplings should be equivalent in both samples but the 3∆C’(ND) isotope shifts are unique to the 1:1 H:D sample. Nevertheless, potential interference from 13C-13C scalar couplings makes the analysis of 3∆C’(ND) isotope shifts more subjective and error prone than for the 4∆N(ND) isotope shifts described in the previous section. A way to circumvent interference from 13C-13C J-couplings would be to use a 13C constant-time version of the HNCO experiment, however, this approach is unlikely to work with the large 13C-dimension acquisition times of 0.38 s needed to resolve the small 3∆C’(ND) isotope shifts. An alternative method to measure 3∆C’(ND) isotope shifts using isotope-filtering has been previously described (Zhang and Tugarinov 2013).
Using the 2D H(N)CO experiment we obtained 46 and 44 3∆C’(ND) isotope shifts for CUS-3iD (Table S3) and CspA (Table S4), respectively. Resolvable 3∆C’(ND) isotope shifts in this work ranged from 5 to 50 ppb, with a median value of 28 ppb. The sizes of 3∆C’(ND) isotope shifts obtained here are in good agreement with those of 25–30 ppb, previously reported in the literature (Tuchsen and Hansen 1988; Uchida et al. 2005; Zhang and Tugarinov 2013). Based on replicate experiments on two different CUS-3iD samples we estimate the error in the 3∆C’(ND) isotope shifts was about ± 4 ppb, or ~ 15% of the average 28 ppb size of the non-zero 3∆C’(ND) isotope shifts.
The intra-residue 3∆C’(ND) isotope shift was proposed to depend on the H to O distance in the intra-residue 5-atom ring H-N-Cα-C’-O (Tuchsen and Hansen 1991; Zhang and Tugarinov 2013), henceforth dHO. We were able to confirm this correlation (Fig. 3C). Since the previously described 4∆N(ND) isotope shift depends on the distance between the atom originating the shift and the atom experiencing the splitting, we wanted to see if a similar relationship held for 3∆C’(ND). To this end, we looked for a possible relationships between 3∆C’(ND) and the 4-atom chain distance H-N-Cα-C’, henceforth dHC’. We observed a correlation between the 3∆C’(ND) isotope shift and dHC’ (Fig. 3D) that was slightly poorer than with the dHO distance (Fig. 3C). The dHC’ distance is highly correlated (Fig. S2C, D) to the torsional angle phi, defined by the atoms C’(i-1), N(i), Cα (i) and C’(i). Consequently, the 3∆C’(ND) isotope shift is also correlated with the dihedral angle phi (R-value = 0.68).
2∆C’(ND) isotope shifts and ϕ fractionation factors are invariant with temperature
The large ~ 70 ppb inter-residue 2∆C’(ND) isotope shifts in ubiquitin were previously obtained from isotope splittings in H(CA)CO experiments (LiWang and Bax 1996), and through isotope-filtered experiments (Zhang and Tugarinov 2013). The intensity ratios of C’(i-1) resonances arising from N-D (i) and N–H(i) isotopomers in H(CA)CO experiments were used to calculate ϕ fractionation factors (LiWang and Bax 1996). Earlier efforts to measure ϕ-values used a series of 1H-15N HSQC correlation experiments over a range of H2O:D2O ratios (Bowers and Klevit 1996, 2000; Khare et al. 1999; Loh and Markley 1994). It has been pointed out that measurements of ϕ-values at different solvent ratios can be affected by the ensuant differences in viscosity, as well as intramolecular NOEs for exchangeable protons (LiWang and Bax 1996). However, subsequent work on the same ubiquitin protein found no evidence for such effects (Bowers and Klevit 2000).
The fractionation factor measures the equilibrium constant ϕ = [D]/[H] for exchangeable sites, in mixed H:D solvents (Khare et al. 1999; LiWang and Bax 1996). While fractionation factors have an established role for identifying low-barrier H-bonds in small molecules or enzyme active sites (Cao and Bowie 2014; Shan et al. 1996) their use to assess H-bond strength in proteins has been controversial. (Bowers and Klevit 1996; Khare et al. 1999; Loh and Markley 1994)(Bowers and Klevit 1996; Khare et al. 1999; LiWang and Bax 1996; Loh and Markley 1994) We thought if protein fractionation factors were sensitive to H-bonding, they might change with temperature. 3hJNC’ through-hydrogen couplings suggest that most protein H-bonds lengthen and weaken with increasing temperature (Cordier and Grzesiek 2002; Hong et al. 2013). The size of the 2∆C’(ND) isotope shift has been attributed to H-bond strength, specifically H-bond enthalpy (Tuchsen and Hansen 1991).
We found that the previously described approach of using H(CA)CO experiments to measure 2∆C’(ND) and ϕ-values (LiWang and Bax 1996) posed some disadvantages for our CUS-3iD protein. These included a limited dispersion of 1Hα and 13Cα resonances in CUS-3iD, that 1Hα resonances near the solvent signal were difficult to observe, and that the spectrum had a large range of peak intensities due to residues with different 1Hα -13Cα relaxation properties. As an alternative to the H(CA)CO experiment (Fig. 4A) we measured the 2∆C’(ND) isotope shifts and ϕ-values using H(NCA)CO experiments (Fig. 4B). The H(NCA)CO experiment offers the advantages of (1) better 1HN chemical shift dispersion, (2) more uniform 1HN signal intensities and (3) most importantly the ability to use perdeuterated 3L protein samples. Perdeuteration results in a large increase in sensitivity with the long 13C acquisition times needed to resolve the 2∆C’(ND) isotope shift splittings (Fig. 4 C, D). The H(NCA)CO spectra (Fig. 4E) allowed us to calculate ϕ-values from D/H peak height intensity ratios (Table S5) and 2∆C’(ND) values from the chemical shift difference between H and D resonances (Table S6). Data over a series of temperatures were used to look at the temperature-dependence of the parameters (Fig. 4F).
The 2∆C’(ND) isotope shifts for CUS-3iD at 288 K range from 48 to 121 ppb, with a median value of 76 ppb (corresponding to 11 Hz at 13C 150 MHz). The 2∆C’(ND) isotope shifts are about twice as large as 4∆N(ND) and 3∆C’(ND), and are in excellent agreement with previously published data (LiWang and Bax 1996; Tuchsen and Hansen 1991; Uchida et al. 2005; Zhang and Tugarinov 2013). To assess the accuracy of the 2∆C’(ND) isotope shifts we collected replicate data in two ways. In the first, we collected data for a duplicate sample of CUS-3iD, and obtained an average 2∆C’(ND) error of ± 7 ppb. In the second, we obtained on the same sample used for the temperature titration, giving an average 2∆C’(ND) error of ± 5 ppb. The second approach for error estimation probably more relevant to the temperature titration, since all the data were obtained on the same sample. The 2∆C’(ND) data for CUS-3iD at 288 K together with experimental uncertainties are shown in Fig. 5A. No differences were observed between the 2∆C’(ND) values of residues forming H-bonds and those that do not. The insensitivity of 2∆C’(ND) isotope shifts to secondary structure was also noted in a previous study (Zhang and Tugarinov 2013).
Temperature dependence of 2∆C’(ND) isotope shifts and ϕ fractionation factors in CUS-3iD. A Sizes of 2∆C’(ND) isotope shifts from H(NCA)CO spectra of 3L CUS-3iD at a temperature of 288 K. B Difference in slopes between the upper (D) and lower (H) component of the 2∆C’(ND) doublet as a function of temperature. C Fractionation factors (ϕ-values) of 3L CUS-3iD in 1:1 H:D at a temperature of 288 K. D Slopes of the change in ϕ fractionation factors as a function of temperature. For panels A and C error bars are from duplicate measurements on the same sample at a temperature of 288 K. For panels B and D error bars are the SEM obtained from linear least squares fits of the respective data as a function of temperature. In each panel, black symbols indicate residues that form H-bonds in the CUS-3iD structure. The secondary structure of the protein is shown at the top of the graph
To characterize the temperature dependence of the 2∆C’(ND) shifts we carried out linear least squares fits of 13C’ chemical shifts as a function of temperature. This gives slopes for the upfield (H) and downfield (D) component of each doublet (Fig. 4F). Figure 5B shows the difference in the slopes between the H and D signal, together with SEM values. Nearly all residues give slopes for the H and D components that are the same within experimental uncertainty (Fig. 5B). Stated another way, the 2∆C’(ND) isotope shifts are invariant with temperature. We note that CUS-3iD is a moderately stable protein, which limited our studies to a temperatures between 288 and 313 oK (15 °C to 40 °C). For very stable proteins that remain folded up to 80 °C or 90 °C, differences in 2∆C’(ND) values might be detectable over a larger temperature range. Our results suggest if there is a change in 2∆C’(ND) with temperature, as might be expected if the parameter is related to H-bonds, the effect is very small.
We next looked at ϕ-value fractionation factors obtained from uncorrected D/H peak height ratios in the same H(NCA)CO dataset used to extract 2∆C’(ND) isotope shifts. For the CUS-3iD protein at a temperature of 288 K, the ϕ-values spanned a range of 0.77 to 1.44 with an average value of 1.07. Our average value is very similar to that published for ubiquitin determined either from three-dimensional H(CA)CO experiments (LiWang and Bax 1996), or from a series of 1H-15N HSQC at different H2O:D2O solvent ratios (Bowers and Klevit 2000). In contrast to the average, the ranges of values in the two ubiquitin studies are markedly different. It is unlikely that different NMR experiments could account for differences in the range of ϕ-values. Rather the ubiquitin study by H(CA)CO spectroscopy is unique in that it used corrections to account for differences in the T2 values of 1Hα, 13Cα, 13C’ nuclei, when the adjacent N atom is bonded to H or D (LiWang and Bax 1996). These corrections require an X-ray structure of the protein, as well as additional information on parameters such as protein correlation time, amongst others. In practice, the corrections applied to ubiquitin reduce the range of ϕ-values in half for the corrected (1.01 < ϕ < 1.16) compared to the uncorrected (1.03 < ϕ < 1.33) data.
Our ϕ-value data for the CUS-3iD protein shows no correlation to H-bonds or secondary structure (Fig. 5C). Most ϕ-values are within experimental uncertainty of unity—that is there is no preference for H or D for most residues in the protein. Duplicate sets of measurements at 288 oK, were used to estimate an average uncertainty in ϕ of ± 0.1. When we looked for changes with temperature we saw apparently random variations of ~ 20% of each ϕ-value. We conclude that there is no systematic change in ϕ-values with temperature, within an experimental uncertainty of ~ 15–20% (Fig. 5D). The precision of the ϕ-values measured in this work is low but not without precedent, as sets of ϕ-values for ubiquitin from two different groups found no correlation between individual residues (Bowers and Klevit 2000; LiWang and Bax 1996). We conclude that error in protein ϕ fractionation factors is large compared to the range of ϕ-values spanned by different residues. Alternatively, accurate use of NMR ϕ-values require corrections that include correlation time and X-ray structure data (LiWang and Bax 1996) that would make accurate studies of ϕ-values over a range of conditions such as different temperatures impractical.
Use of 2∆C’(ND) and 3∆C’(ND) isotope shifts to assess the cooperativity of HX across peptide bonds
The combination of β and γ isotope shifts for the C’ resonance of a given residue (Fig. 4A), should give a quartet pattern of signals at sufficiently high resolution (Tuchsen and Hansen 1988; Uchida et al. 2005). The intensities of the four quartet components from downfield to upfield corresponds to the combinations of isotope populations H(i)H(i + 1), D(i)H(i + 1), H(i)D(i + 1), and D(i)D(i + 1). Because the isotope shifts link information on amide proton occupancy across peptide bonds, they can be used to determine the extent to which HX is correlated. Such an approach was used to interrogate correlated HX using 1D NMR of the protein SSI selectively 13C-labeled at leucine and valine sites (Uchida et al. 2005). To examine if this approach is amenable to multi-dimensional NMR of uniformly 13C-labeled samples, we obtained H(CA)CO data on a 2L-sample of CUS3-iD as a function of incubation time in 99.96% D2O. Note that 1Hα detection is necessary for H(CA)CO spectra, precluding the use of 3L-samples.
Figure 6A shows the full H(CA)CO spectrum collected after 54 h of exchange in D2O. The spectrum illustrates the wide distribution of crosspeak intensities. The center of the spectrum shows intense signals due to dynamic residues in the protein. In contrast, signals from structured regions of the protein, downfield of 5 ppm and upfield of 4 ppm in the 1Hα dimension, are considerably weaker. Figure 6B shows expansions of the region containing crosspeaks from the C-terminus (shown for reference) and residue A315, after various exchange times in D2O. To resolve the splitting from the smaller 3∆C’(ND) isotope shifts, the data were collected with 13C acquisition times of 377 ms corresponding to an FID resolution of 1.3 Hz. The 13C resolution was thus about 3 times larger than that used to measure 2∆C’(ND) isotope shifts from the same 2D H(CA)CO spectra (LiWang and Bax 1996).
Use of H(CA)CO spectra to investigate correlated hydrogen exchange. A H(CA)CO spectrum of 2L CUS-3iD obtained 54 h after dissolving the protein in 99.96% D2O. The spectrum in A was processed to enhance sensitivity (GB = 0.03, LB = 11 for 1Hα and GB = 0.03, LB = 7 for 13C’). Crosspeaks for the C-terminal carboxyl signal, and the backbone carbonyl of residue A315 are indicated in the spectrum. Note the wide range of crosspeak intensities in the spectrum and the relatively poor chemical shift dispersion. B Selected HX time points monitoring changes in the H(CA)CO spectra following dissolution in D2O. The spectra were processed for resolution enhancement with GB = 0.02, LB = 9 for 1Hα and GB = 0.17, LB = 0.5 for 13C’. The quartet of signals for residue A315 due to different isotope combinations, is indicated in the figure
Using this approach we were able to investigate correlated exchange for the pair of residues A315-S316, as shown in Fig. 6B. The β-shift obtained from the H(NCA)CO for residue A315 is 77 ppb. From separate H(N)CO data, the smaller γ-shift for A315 is 10 ppb. The values are close to the averages found in the CUS3-iD protein, so A315 should be a representative example. The four components of the quartet that change in intensity as a function of incubation time in D2O are labeled according to their different isotope compositions. Using separate conventional 1H-15N HSQC spectra HX experiments of CUS3-iD in D2O, we determined that the amide proton of A315 is fully exchanged in about 21 h. The S316 amide proton exchanges more slowly, persisting up to 34 h. All Hα-C’ correlations in the H(CA)CO spectrum are observed throughout the course of the experiment following dissolution in D2O, however, the positions of the crosspeaks change as the 13C’ resonances experience different isotope shifts, depending on whether adjacent N atoms are bonded to H or D. At the earliest 3 h timepoint we observe two downfield signals with the 3∆C’(ND) γ-splitting of 10 ppb for the C’ of A315. This is because the amide proton of this residue starts to exchange before the amide proton of S316. As S316 also begins to exchange by 9 h, we see the complete quartet of signals reflecting the onset of partial exchange at both sites. By 15 h most of the A315 amide protons have exchanged and the C’ signal of A315 appears upfield due to the β-shift arising from deuteration of the amide of S316. Some residual γ-splitting due to incomplete exchange of the amide proton of A315 is still observed in the 15 h spectrum. By 54 h both sites have fully exchanged, and only one signal is observed for the C’ of A315 that experiences the maximum β and γ shifts, as both the A315 and S316 amides are fully deuterated.
At the 9 h time point we see all the components of the quartet (Fig. 6B). We can use a previously published integration approach (Uchida et al. 2005) to calculate the cooperativity factor Xβγ of 0.20 ± 0.02 for the A315-S316 pair. The parameter Xβγ ranges from 1 for fully correlated sites, to 0 for uncorrelated HX (Uchida et al. 2005). Thus HX at A315 is largely independent of that at S316. We examined the H(CA)CO spectrum to see how many other residues are amenable to this type of analysis and found only 5–10 residues have sufficiently resolved quartet components to assess HX cooperativity.
Discussion
In this paper we obtained 4∆N(ND), 3∆C’(ND) and 2∆C’(ND) isotope shifts from the splittings between ND and NH signals of protein samples dissolved 1:1 H:D. The experiments used to measure the isotope shifts are some of the most common in the arsenal of biomolecular NMR, including 1H-15N TROSY, HNCO, HCACO, and HN(CA)CO. A previously published method used 2D sub-spectra of an intra-HA[CA/CO/N/CB] pulse sequence to obtain six isotope shifts 1∆N(NiD), 4∆N(Ni+1D), 2∆Cα(NiD), 3∆Cα(Ni+1D), 2∆C’(Ni+1D), and 3∆C’(NiD) arising from deuteration of amide protons in 1:1 H:D samples (Zhang and Tugarinov 2013). The 2D sub-spectra use BIRD filtering elements to select correlations arising from one of the four possible isotope species (NiH,Ni+1H), (NiH,Ni+1D), (NiD,Ni+1H), (NiD,Ni+1D) in a dipeptide fragment. Thus for a 4∆N(D) isotope shift for example, the N(H) and N(D) signals are measured separately in two sub-spectra, rather than simultaneously as in our 15N-TROSY approach. An advantage of the isotope-filtering method is that all isotope shifts are measured as sub-spectra from a single intra-HA[CA/CO/N/CB] experiment (Zhang and Tugarinov 2013). The intra-HA[CA/CO/N/CB] experiment uses Hα protons for detection precluding the use of perdeuterated proteins. This could be a disadvantage for proteins with poor Cα chemical shift dispersion or relaxation properties. For the four-bond nitrogen isotope shift, however, isotope-filtered 2D HN(COCAN[H/D]) and intra-HN(CACON[H/D]) experiments were developed to measure 4∆N(Ni-1D) and 4∆N(Ni+1D) effects on perdeuterated proteins (Tugarinov 2013). Overall the results from the sub-spectra and isotope shift splitting approaches are similar, since both measure the same chemical shift differences.
We concluded based on the lack of 4∆N(ND) isotope shifts for residues preceding prolines, and isotope shift patterns when the residue following a protected amide proton is exchanged in D2O (Fig. 1B), that the 4∆N(ND) isotope shift of residue i is primarily due to the H/D isotopomer state of the amide nitrogen of residue i + 1. In isotope-filtered experiments where 4∆N(Ni+1D) and 4∆N(Ni-1D) isotope shifts were individually measured the former were noticeably larger (Tugarinov 2013). Whereas the 4∆N(Ni+1D) were exclusively positive and had mean and s.d. values of 36 ± 14 ppb similar to the ones obtained here (32 ± 15 ppb), the 4∆N(Ni-1D) isotope shifts were positive and negative with a mean and s.d. of -2 ± 9 ppb (Tugarinov 2013), consistent with our conclusion. We found that for the two proteins CUS3-iD and CspA that the size of the 4∆N(ND) isotope shift is correlated with the backbone dihedral angle psi of residue i. The 4∆N(ND) isotope shifts show clear discrimination between α-helical and β-sheet secondary structure (Fig. 2B, D), and could be used for psi angle dihedral restraints. The 4∆N(ND) isotope shift also correlates with the d(NiHi+1) distance between the site of isotope substitution and the nucleus experiencing the isotope shift, because this distance is correlated with the dihedral angle psi (Fig. S2A,B). A similar correlation between 4∆N(Ni+1D) and d(NiHi+1) was found for ubiquitin (Tugarinov 2013). The data were modeled to a Morse equation since that function was used to analyze isotope shifts in small molecules (Tugarinov 2013). For our data (Fig. 2B) an F-test on the limiting ratio of reduced χ2 ratios (Garland et al. 2009), showed inclusion of a third fitted variable in the Morse function does not result in a significant fit improvement compared to a linear fit.
The next largest isotope shift we looked at is the three bond 3∆C’(ND), or γ-shift, with average values of ~ 30 ppb. The γ-shifts were previously reported to correlate with the intraresidue H–O distance in the 5-atom ring H-N–Cα-C-O (Tuchsen and Hansen 1991; Zhang and Tugarinov 2013). We were able to confirm this correlation using a data set of 90 values obtained using 2D H(N)CO spectroscopy on 3L samples of the protein CUS-3iD and CspA. The correlation between 3∆C’(ND) isotope shifts and intraresidue H–O distances was previously attributed to intra-residue H-bonding (Tuchsen and Hansen 1991). We think such a mechanism is unlikely since these types of putative H-bonds would require energetically unfavorable N–H-O angles (Adalsteinsson et al. 1996), and because the correlation between γ-shift and the H–O distance persists for distances between 2.5 and 4.5 Å that are too long for H-bonds (Fig. 3C). It has been proposed that the relationship between 3∆C’(ND) and intra-residue H–O distances could be used to provide structural constraints for NMR structure calculations (Tuchsen and Hansen 1991).
A feature shared by the 3∆C’(ND) and 4∆N(ND) isotope shifts is that the largest values occur when the atom responsible for the isotope shift and the atom experiencing it are at their closest distances. Conversely, the isotope shifts tend to their minimum values or are unresolved when the corresponding distances dNH(i,i + 1) and dHC’ tend to a maximum. The distances dNH(i,i + 1) and dHC’ are highly correlated with the backbone dihedral angles psi and phi, respectively, making it difficult to establish if the size of isotope effect is due to a stereochemical or distance effect. A distance-dependence could signify that nuclear shielding, thought to be responsible for the isotope shifts (Hansen 1988), becomes more effective at shorter distances. Support for this hypothesis comes from the observation that the sizes of isotope shifts generally increase as the number of intervening bonds decreases between the site of isotope substitution and isotope shift (Berger and Künzer 1983). Additional indirect support for a distance-dependence could come from the large size and relative uniformity of the 2∆C’(ND) isotope shifts (LiWang and Bax 1996) who’s corresponding distance dC’H(i,i + 1) is fixed to a narrow range of ~ 0.05 Å by the planarity of the peptide bond (Fig. S3). At the same time the isotope shift cannot depend exclusively on distances, since non-bonded N–H and H–O distances in the 3D structures of folded proteins can be comparable to the distances spanned by the N–Cα-C’-N–H and H-N–Cα-C-O atom chains corresponding to the 4∆N(ND) and 3∆C’(ND) isotope shifts.
We used 2D H(NCA)CO spectra to characterize the temperature dependence of 2∆C’(ND) isotope shifts and ϕ = [D]/[H] fractionation factors in the protein CUS3-iD. We find that neither parameter discriminates sites involved in H-bonding. Moreover, neither parameter changes with temperature within the experimental uncertainty of ~ 6% for the isotope shifts, and ~ 20% for the ϕ-values. Since H-bonds in proteins have been reported to typically decrease with increasing temperature (Cordier and Grzesiek 2002; Hong et al. 2013), this suggests that neither of the two parameters is sensitive to protein H-bonding.
Using 2D H(CA)CO experiments with a sufficiently long acquisition time of 377 ms in the 13C’ dimension we were able to resolve both β and γ-isotope shifts of residue A315 in the protein CUS3-iD, and thus determine the extent of synchronized amide proton hydrogen exchange between residues A315-S316 (Fig. 6). These types of experiments have previously only been done with 1D NMR methods (Uchida et al. 2005). The main hurdles with this approach are the limited 1Hα and 13Cα chemical shift dispersion in the 2D H(CA)CO spectrum, the wide range of signal intensities, and the high sensitivity demands of the experiment. The 2D H(NCA)CO experiments gives better sensitivity for 2∆C’(ND) isotope shifts, mainly because it lends itself to experiments on perdeuterated protein samples. In principle high-resolution 2D H(NCA)CO experiments could resolve both β and γ isotope shifts. Note that in contrast to H(CA)CO which reports on the γ-shift of residue i (Fig. 4A), the γ-shift in the H(NCA)CO experiment would come from residue i-1 (Fig. 4B). Therefore the H(NCA)CO experiment could provide unique information on correlated HX between residues separated by two positions in the sequence, whereas H(CA)CO monitors correlated HX between neighboring residues. Preliminary data on the feasibility of using the H(NCA)CO experiment to monitor correlated HX, indicates that for this approach to work the central residue in a tripeptide segment needs to exchange slower than the neighboring two residues. Since the detection of isotope shifts occurs through the amide proton of the central residue (Fig. 4B), once this amide proton has exchanged the experiment provides no information on the HX status of the neighboring residues. There are only about five triplet segments in the CUS-3iD protein protected from HX (Fig. 1D), and even less that meet the requirement that the central residue is the slowest exchanger. Alternatively, information on correlated HX could be investigated using 4∆N(ND) isotope shifts or NOESY spectroscopy (Roder et al. 1985; Wagner 1980; Wang et al. 1987).
Throughout this work we pointed out that the isotope shifts that we optimally measured for protein samples in 1:1 H:D, can also be detected in 9:1 H:D samples at sufficient sensitivity. The 9:1 H:D solvent fraction is ubiquitously used in solution NMR for the deuterium lock. With the benefit of clearly resolvable isotope shifts for samples in 1:1 H:D, we can see minor isotopologue species due to the 10% D2O in experiments that include 1H-15N TROSY (Fig. 1A), H(CA)CO, and H(NCA)CO (Fig. 4E). These minor species are a source of spectral heterogeneity that could potentially complicate NMR assignments, or confound experiments that seek to detect small 5–10% populations of species in studies of conformational equilibria or ligand binding. Similar, but more severe problems associated with the 10% fraction of D2O for the deuterium lock have been described for NMR studies of Lys and Arg sidechains, where isotopomer combinations such as -NHD, -NH2D or -NHD2 severely complicate spectra and can lead to N/D exchange broadening (Nguyen et al. 2019). These problems were circumvented with coaxial NMR tubes where the D2O for the deuterium lock is kept separate from the protein solution (Nguyen et al. 2019). A drawback of coaxial NMR tubes is that they can lower sensitivity and make shimming more difficult.
A primary limitation in the application of isotope shifts to conformational analysis and HX cooperativity, is the loss of sensitivity in the indirect dimension that occurs due to the long acquisition times needed to resolve small chemical shift differences. A second complicating factor is that small long-range scalar J-couplings can interfere with analyses if these have comparable sizes to the isotope shifts. Both issues are expected to improve at higher magnetic fields. The size of the isotope shift will scale with increasing magnetic field. In contrast, J-couplings are invariant to magnetic field, and would effectively become smaller compared to isotope shifts on a larger magnet. The experiments in this work were done on a 600 MHz instrument. On a 1.2 GHz spectrometer half the number of points in the indirect dimension would achieve comparable resolution, reducing experiment times by half.
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
Abbreviations
- ϕ:
-
[D]/[H] fractionation factor
- 1D, 2D, 3D:
-
One-, two, three-dimensional
- 2L:
-
13C,15N double-labeled sample
- 3L:
-
2H,13C,15N triple-labeled sample
- CspA:
-
Cold shock protein A from E. coli
- CUS-3iD:
-
Insertion domain from the coat protein of phage CUS-3
- D:
-
2H isotope of hydrogen
- FID:
-
Free induction decay
- H:
-
1H isotope of hydrogen
- HSQC:
-
Heteronuclear single-quantum coherence
- 9:1 H:D:
-
solvent fraction containing 90% H2O and 10% D2O (%v/%v)
- 1:1 H:D:
-
solvent fraction containing 50% H2O and 50% D2O (%v/%v)
- HX:
-
Hydrogen exchange
- H-bond:
-
Hydrogen bond
- NOE:
-
Nuclear Overhauser effect
- NOESY:
-
NOE spectroscopy
- ppb:
-
Parts per billion
- ppm:
-
Parts per million
- s.d.:
-
Standard deviation
- SEM:
-
Standard error of the mean
- β-shift:
-
2∆C’(ND) isotope shift
- γ-shift:
-
3∆C’(ND) isotope shift
- S/N:
-
Signal to noise ratio
- TROSY:
-
Transverse relaxation optimized spectroscopy
References
Abildgaard J, Hansen PE, Manalo MN, LiWang A (2009) Deuterium isotope effects on 15N backbone chemical shifts in proteins. J Biomol NMR 44:119–126. https://doi.org/10.1007/s10858-009-9316-0
Adalsteinsson H, Maulitz AH, Bruice TC (1996) Calculation of the potential energy surface for intermolecular amide hydrogen bonds using semiempirical and ab initio methods. J Am Chem Soc 118:7689–7693
Alexandrescu AT, Ulrich EL, Markley JL (1989) Hydrogen-1 NMR evidence for three interconverting forms of staphylococcal nuclease: effects of mutations and solution conditions on their distribution. Biochemistry 28:204–211. https://doi.org/10.1021/bi00427a028
Alexandrescu AT, Hinck AP, Markley JL (1990) Coupling between local structure and global stability of a protein: mutants of staphylococcal nuclease. Biochemistry 29:4516–4525. https://doi.org/10.1021/bi00471a003
Alexandrescu AT, Snyder DR, Abildgaard F (2001) NMR of hydrogen bonding in cold-shock protein A and an analysis of the influence of crystallographic resolution on comparisons of hydrogen bond lengths. Protein Sci 10:1856–1868. https://doi.org/10.1110/ps.14301
Berger S, Künzer H (1983) Extremely long-range 2H isotope effects on the chemical shifts in the 13C-NMR spectra of compounds with conjugated double bonds. Angew Chem Int Ed Engl 22:321–322
Bowers PM, Klevit RE (1996) Hydrogen bonding and equilibrium isotope enrichment in histidine-containing proteins. Nat Struct Biol 3:522–531. https://doi.org/10.1038/nsb0696-522
Bowers PM, Klevit RE (2000) Hydrogen bond geometry and 2H/1H fractionation in proteins. J Am Chem Soc 122:1030–1033
Cao Z, Bowie JU (2014) An energetic scale for equilibrium H/D fractionation factors illuminates hydrogen bond free energies in proteins. Protein Sci 23:566–575. https://doi.org/10.1002/pro.2435
Cavanagh J, Fairbrother WJ, Palmer AG III, Skelton NJ (2006) Protein NMR Spectroscopy Principles and Practice. Protein NMR Spectroscopy, 2nd edn. Elsevier Inc., Amsterdam
Chevelkov V, Xue Y, Rao DK, Forman-Kay JD, Skrynnikov NR (2010) 15N H/D-SOLEXSY experiment for accurate measurement of amide solvent exchange rates: application to denatured drkN SH3. J Biomol NMR 46:227–244. https://doi.org/10.1007/s10858-010-9398-8
Cordier F, Grzesiek S (2002) Temperature-dependence of protein hydrogen bond properties as studied by high-resolution NMR. J Mol Biol 317:739–752. https://doi.org/10.1006/jmbi.2002.5446
Dempsey CE (2001) Hydrogen exchange in peptides and proteins using NMR spectroscopy. Prog Nucl Magn Reson Spectrosc 39:135–170
Feng W, Tejero R, Zimmerman DE, Inouye M, Montelione GT (1998) Solution NMR structure and backbone dynamics of the major cold-shock protein (CspA) from Escherichia coli: evidence for conformational dynamics in the single-stranded RNA-binding site. Biochemistry 37:10881–10896. https://doi.org/10.1021/bi980269j
Garland CW, Nibler JW, Shoemaker DP (2009) Experiments in Physical Chemistry, 8th edn. McGraw-Hill, New York
Hansen PE (1988) Isotope effects in nuclear shielding. Prog in NMR Spectrosc 20:207–255
Hong J, Jing Q, Yao L (2013) The protein amide (1)H(N) chemical shift temperature coefficient reflects thermal expansion of the N-H O=C hydrogen bond. J Biomol NMR 55:71–78. https://doi.org/10.1007/s10858-012-9689-3
Jaravine VA, Rathgeb-Szabo K, Alexandrescu AT (2000) Microscopic stability of cold shock protein A examined by NMR native state hydrogen exchange as a function of urea and trimethylamine N-oxide. Protein Sci 9:290–301. https://doi.org/10.1110/ps.9.2.290
Kainosho M, Tsuji T (1982) Assignment of the three methionyl carbonyl carbon resonances in Streptomyces subtilisin inhibitor by a carbon-13 and nitrogen-15 double-labeling technique A new strategy for structural studies of proteins in solution. Biochemistry 21:6273–6279. https://doi.org/10.1021/bi00267a036
Kainosho M, Nagao H, Imamura Y, Uchida K, Tomonaga N, Nakamura Y, Tsuji T (1985) Structural studies of a protein using the assigned back-bone carbonyl carbon-13 NMR resonances. J Mol Struct 126:549–562
Kaplan AR, Maciejewski M, Olson R, Alexandrescu AT (2013) NMR assignments for the cis and trans forms of the hemolysin II C-terminal domain. Biomol NMR Assign 8:419–423
Kaplan AR, Olson R, Alexandrescu AT (2021) Protein yoga: conformational versatility of the Hemolysin II C-terminal domain detailed by NMR structures for multiple states. Protein Sci 30:990–1005. https://doi.org/10.1002/pro.4066
Kateb F, Pelupessy P, Bodenhausen G (2007) Measuring fast hydrogen exchange rates by NMR spectroscopy. J Magn Reson 184:108–113. https://doi.org/10.1016/j.jmr.2006.09.022
Khare D, Alexander P, Orban J (1999) Hydrogen bonding and equilibrium protium-deuterium fractionation factors in the immunoglobulin G binding domain of protein G. Biochemistry 38:3918–3925. https://doi.org/10.1021/bi9827114
Lambert JB, Greifenstein LG (1974) Origin of the chemical-shift isotope effect stereochemical evidence. J Amer Chem Soc 96:5120–5124
LeMaster DM, LaIuppa JC, Kushlan DM (1994) Differential deuterium isotope shifts and one-bond 1H–13C scalar couplings in the conformational analysis of protein glycine residues. J Biomol NMR 4:863–870. https://doi.org/10.1007/BF00398415
Liu A, Wang J, Lu Z, Yao L, Li Y, Yan H (2008) Hydrogen-bond detection, configuration assignment and rotamer correction of side-chain amides in large proteins by NMR spectroscopy through protium/deuterium isotope effects. Chembiochem 9:2860–2871. https://doi.org/10.1002/cbic.200800467
LiWang AC, Bax A (1996) Equilibrium protium/deuterium dractionation of backbone amides in U-13C/15N labeled human ubiquitin by triple resonance NMR. J Am Chem Soc 118:12864–12865
Loh SN, Markley JL (1994) Hydrogen bonding in proteins as studied by amide hydrogen D/H fractionation factors: application to staphylococcal nuclease. Biochemistry 33:1029–1036. https://doi.org/10.1021/bi00170a023
Lohr F, Ruterjans H (1999) Alternative E.COSY techniques for the measurement of 3J(C (i) (') (-1), C (i) (beta) ) and (3) J(H (i) (N), C (i) (beta) ) coupling constants in proteins. J Biomol NMR 13:263–274. https://doi.org/10.1023/A:1008378719908
Mal TK, Matthews SJ, Kovacs H, Campbell ID, Boyd J (1998) Some NMR experiments and a structure determination employing a [15N,2H] enriched protein. J Biomol NMR 12:259–276. https://doi.org/10.1023/a:1008238009056
Maltsev AS, Ying J, Bax A (2012) Deuterium isotope shifts for backbone (1)H, (1)(5)N and (1)(3)C nuclei in intrinsically disordered protein alpha-synuclein. J Biomol NMR 54:181–191. https://doi.org/10.1007/s10858-012-9666-x
Nguyen D, Chen C, Pettitt BM, Iwahara J (2019) NMR methods for characterizing the basic side chains of proteins: electrostatic interactions hydrogen bonds, and conformational dynamics. Methods Enzymol 615:285–332. https://doi.org/10.1016/bs.mie.2018.08.017
Ottiger M, Bax A (1997) An empirical correlation between amide deuterium isotope effects on 13Calpha chemical shifts and protein backbone conformation. J Am Chem Soc 119:8070–8075
Roder H, Wagner G, Wuthrich K (1985) Amide proton exchange in proteins by EX1 kinetics: studies of the basic pancreatic trypsin inhibitor at variable p2H and temperature. Biochemistry 24:7396–7407. https://doi.org/10.1021/bi00346a055
Schindelin H, Jiang W, Inouye M, Heinemann U (1994) Crystal structure of CspA, the major cold shock protein of Escherichia coli. Proc Natl Acad Sci USA 91:5119–5123. https://doi.org/10.1073/pnas.91.11.5119
Schmidt JM, Hua Y, Lohr F (2010) Correlation of (2)J couplings with protein secondary structure. Proteins 78:1544–1562. https://doi.org/10.1002/prot.22672
Shan SO, Loh S, Herschlag D (1996) The energetics of hydrogen bonds in model systems: implications for enzymatic catalysis. Science 272:97–101. https://doi.org/10.1126/science.272.5258.97
Sun H, Tugarinov V (2012) Precision measurements of deuterium isotope effects on the chemical shifts of backbone nuclei in proteins: correlations with secondary structure. J Phys Chem B 116:7436–7448. https://doi.org/10.1021/jp304300n
Takeda M, Jee J, Ono AM, Terauchi T, Kainosho M (2009) Hydrogen exchange rate of tyrosine hydroxyl groups in proteins as studied by the deuterium isotope effect on C(zeta) chemical shifts. J Am Chem Soc 131:18556–18562. https://doi.org/10.1021/ja907911y
Takeda M, Miyanoiri Y, Terauchi T, Yang CJ, Kainosho M (2014) Use of H/D isotope effects to gather information about hydrogen bonding and hydrogen exchange rates. J Magn Reson 241:148–154. https://doi.org/10.1016/j.jmr.2013.10.001
Theis K, Dingley AJ, Hoffmann A, Omichinski JG, Grzesiek S (1997) Determination of backbone nitrogen-nitrogen J correlations in proteins. J Biomol NMR 10:403–408. https://doi.org/10.1023/A:1018373601391
Tripler TN, Maciejewski MW, Teschke CM, Alexandrescu AT (2015) NMR assignments for the insertion domain of bacteriophage CUS-3 coat protein. Biomol NMR Assign. https://doi.org/10.1007/s12104-015-9604-4
Tripler TN, Kaplan AR, Alexandrescu AT, Teschke CM (2019) Conservation and divergence of the I-domain inserted into the ubiquitous HK97 coat protein fold in P22-like bacteriophages. J Virol. https://doi.org/10.1128/JVI.00007-19
Tuchsen E, Hansen PE (1988) Carbonyl 13C NMR spectrum of basic pancreatic trypsin inhibitor: resonance assignments by selective amide hydrogen isotope labeling and detection of isotope effects on 13C nuclear shielding. Biochemistry 27:8568–8576. https://doi.org/10.1021/bi00423a010
Tuchsen E, Hansen PE (1991) Hydrogen bonding monitored by deuterium isotope effects on carbonyl 13C chemical shift in BPTI: intra-residue hydrogen bonds in antiparallel beta-sheet. Int J Biol Macromol 13:2–8. https://doi.org/10.1016/0141-8130(91)90002-c
Tugarinov V (2013) Four-bond deuterium isotope effects on the chemical shifts of amide nitrogens in proteins. Magn Reson Chem 51:722–728. https://doi.org/10.1002/mrc.4007
Uchida K, Markley JL, Kainosho M (2005) Carbon-13 NMR method for the detection of correlated hydrogen exchange at adjacent backbone peptide amides and its application to hydrogen exchange in five antiparallel beta strands within the hydrophobic core of Streptomyces subtilisin inhibitor (SSI). Biochemistry 44:11811–11820. https://doi.org/10.1021/bi050467s
Venters RA, Farmer BT 2nd, Fierke CA, Spicer LD (1996) Characterizing the use of perdeuteration in NMR studies of large proteins: 13C, 15N and 1H assignments of human carbonic anhydrase II. J Mol Biol 264:1101–1116. https://doi.org/10.1006/jmbi.1996.0699
Vranken WF et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696. https://doi.org/10.1002/prot.20449
Vriend G (1990) WHAT IF: a molecular modeling and drug design program. J Mol Graph 8(52–56):29. https://doi.org/10.1016/0263-7855(90)80070-v
Wagner G (1980) A novel application of nuclear Overhauser enhancement (NOE) in proteins: analysis of correlated events in the exchange of internal labile protons. Biochem Biophys Res Commun 97:614–620. https://doi.org/10.1016/0006-291x(80)90308-3
Wang QW, Kline AD, Wuthrich K (1987) Amide proton exchange in the alpha-amylase polypeptide inhibitor Tendamistat studied by two-dimensional 1H nuclear magnetic resonance. Biochemistry 26:6488–6493. https://doi.org/10.1021/bi00394a030
Wilde JA, Bolton PH, Dell’Acqua M, Hibler DW, Pourmotabbed T, Gerlt JA (1988) Identification of residues involved in a conformational change accompanying substitutions for glutamate-43 in staphylococcal nuclease. Biochemistry 27:4127–4132. https://doi.org/10.1021/bi00411a033
Wüthrich K (1986) NMR of proteins and nucleic acids. John Wiley & Sons, New York
Zhang D, Tugarinov V (2013) Accurate measurements of the effects of deuteration at backbone amide positions on the chemical shifts of (1)(5)N, (1)(3)Calpha, (1)(3)Cbeta, (1)(3)CO and (1)Halpha nuclei in proteins. J Biomol NMR 56:169–182. https://doi.org/10.1007/s10858-013-9733-y
Acknowledgements
We thank Richard Whitehead III for the coordinates of the cryo-EM CUS-3 phage structure, and Morgan Wagner for help with the analysis of the temperature-dependence of 2∆C’(ND) isotope shifts and ϕ fractionation factors. We thank anonymous reviewers for bringing to our attention papers on isotope shifts that we missed in our original draft. A.T.A. thanks Prof. Mei Hong and the Hong lab for their hospitality during his sabbatical visit at MIT and for use of their solution 600 MHz instrument. The NMR experiments used equipment at the MIT-Harvard Center for Magnetic Resonance, which is supported by the P41 grant GM132079. M.H. is partially supported by NIH grant AG059661. A.J.D. was partially supported by an NIH fellowship F31AG069418. A.T.A. and C.M.T. were supported by NIH grant GM076661.
Author information
Authors and Affiliations
Contributions
ATA conceived the study and experiments. ATA collected the data with help from AJD. ATA analyzed the data. CMT contributed samples and structural data. CMT and ATA acquired funding. ATA wrote the paper with help from AJD. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Alexandrescu, A.T., Dregni, A.J. & Teschke, C.M. NMR detection and conformational dependence of two, three, and four-bond isotope shifts due to deuteration of backbone amides. J Biomol NMR 77, 93–109 (2023). https://doi.org/10.1007/s10858-023-00414-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-023-00414-7