Abstract
Side chain amide protons of asparagine and glutamine residues in random-coil peptides are characterized by large chemical shift differences and can be stereospecifically assigned on the basis of their chemical shift values only. The bimodal chemical shift distributions stored in the biological magnetic resonance data bank (BMRB) do not allow such an assignment. However, an analysis of the BMRB shows, that a substantial part of all stored stereospecific assignments is not correct. We show here that in most cases stereospecific assignment can also be done for folded proteins using an unbiased artificial chemical shift data base (UACSB). For a separation of the chemical shifts of the two amide resonance lines with differences ≥0.40 ppm for asparagine and differences ≥0.42 ppm for glutamine, the downfield shifted resonance lines can be assigned to Hδ21 and Hε21, respectively, at a confidence level >95%. A classifier derived from UASCB can also be used to correct the BMRB data. The program tool AssignmentChecker implemented in AUREMOL calculates the Bayesian probability for a given stereospecific assignment and automatically corrects the assignments for a given list of chemical shifts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although methods for accurate stereospecific assignments of resonance lines in folded proteins exist, they are often cumbersome and require additional spectrometer time, which is not always available. These stereospecific assignments are often required for a detailed structural analysis of a given protein or an enzymatic reaction but are also crucial for general NMR based bioinformatics. For biological applications the biological magnetic resonance data bank (BMRB) (Ulrich et al. 2007) is the most widely used data base for chemical shifts, which is often evaluated for stereospecific assignments. An example are the side chain amide protons in asparagine and glutamine residues that provide usually two well separated resonance lines.
The signals of the two amide protons can often be assigned stereospecifically by their typical homonuclear NOE (nuclear overhauser effect) (Wüthrich 1986) or heteronuclear J-coupling patterns (McIntosh et al. 1997). The distributions of the chemical shifts of these protons in Asn and Gln can be retrieved from the BMRB including their supposed stereochemical assignment. The probability density distributions for the given protons have, independent of the fact that they are supposed to be assigned stereochemically, two maxima separated by approximately 0.67 ppm (Asn) and 0.68 ppm (Gln) (Fig. 1). Given that the chemical shift differences are mainly determined by the chemical structure as suggested by the values of 0.68 ppm (Asn) and 0.71 ppm (Gln) obtained in the random-coil model peptides Gly-Gly-Asn-Ala and Gly-Gly-Gln-Ala (Bundi and Wüthrich 1979), respectively, such bimodal probability distributions are difficult to explain by a simple physical model. In Gly-Gly-Asn-Ala-NH2 and Gly-Gly-Gln-Ala-NH2 the resonances of the amide protons were assigned stereospecifically by a combination of molecular dynamics with NOESY-spectroscopy. The upfield shifted amide protons were assigned to Hδ21 and Hε21, respectively (Harsch et al. 2013). It has been suggested by Harsch et al. (2013) that a large part of the entries is most probably wrongly submitted as being stereospecifically assigned into the BMRB data base. The large chemical shift difference observed in model peptides allows a direct stereochemical assignment in random-coil structures from the chemical shifts only. If such a chemical shift difference is also found for folded proteins, a direct stereospecific assignment of these protons from the chemical shifts alone would be possible. However, such a difference cannot be directly deduced from the data deposited BMRB database (see above). In the paper presented here, we will show that a stereochemical assignment is also possible for proteins from the chemical shifts alone and that the reliability of the corresponding assignments can be calculated by a Bayesian ansatz.
Chemical shift distributions of side chain amide protons in asparagines and glutamine residues. Data were taken from the BMRB database. a Asn Hδ21, b Gln Hε21, c Asn Hδ22, d Gln Hε22. The distributions were created by binning with 0.05 ppm. (Black) all entries of the BMRB where two chemical shift values were given, (red) only amide groups with ambiguity code 1 (claimed as stereospecifically assigned). f(δ) is the frequency of the chemical shifts δ when the areas of the gray curves are normalized to 1
Materials and methods
Calculation of side chain amide chemical shift distributions
Experimental amide proton chemical shifts were taken from the BMRB (state: 12.01.2017). When only one chemical shift value is given in the datatbase for one of the two atoms of the amide group, both chemical shifts of the sidechain protons were omitted. 417 entries of proton chemical shifts were therefore removed for Asn and 308 entries for Gln. At the end, the database for Asn was reduced to 20,220 sidechain groups and the database of Gln was reduced to 18,474 sidechain groups respectively. The unbiased artificial chemical shift data base (UACSB) was recalculated from the unbiased protein structural data base NH3D (Sillitoe et al. 2013). The structures with added protons were energy minimized in explicit water using the molecular dynamics program GROMACS (version 4.6.1) (Berenden et al. 1995; Hess et al. 2008). The final energy minimized structure of each model was then used for a molecular dynamics simulation in explicit water. After equilibration, 400 ps trajectories have been calculated. Eleven equidistant structures were extracted from these trajectories for creation of the UACSB database. Since stereospecifically defined atom names may change after energy minimization (bond rotation), it was checked again and if necessary corrected after minimization with the AUREMOL (Gronwald and Kalbitzer 2004) tool IUPACify. Chemical shifts were calculated using SHIFTS (version 4.3) (Osapay and Case 1991) or SHIFTX2 (version 1.04) (Neal et al. 2003). Since the stereospecific assignments of the random coil values of the side chain amide protons were incorrect for Asn in Gln in SHIFTS and incorrect for Asn in SHIFTX1, the stereospecific assignments of the random coil basis was corrected for UACSB by us (Harsch et al. 2013). The program REDUCE (Word et al. 1999), which is recommended by SHIFTS (Osapay and Case 1991) for adding protons to X-ray structures, also uses a definition of the names of two protons that is not in agreement with the stereochemical IUPAC definition. Details about UACSB will be published elsewhere.
Assessment of stereospecific assignment
The side chain amide groups contain a pair of protons called Hδ21 and Hδ22 in Asn and Hε21 and Hε22 in Gln with a stereochemistry, which is defined in the IUPAC/IUB recommendations (Markley et al. 1998). In the following, the corresponding chemical shifts are called δ 1 and δ 2. The observed chemical shifts δ can always be separated into three contributions, a contribution from the intrinsic chemical structure δ rc, a contribution from the three-dimensional structure δ 3D, and a term δ cor containing contributions (e.g. exchange processes), that cannot be separated completely in δ rc and δ 3D. A good approximation is to define δ rc by values taken from random coil peptides. Unfortunately, only for a few non-equivalent resonances stereospecific assignments in random-coil models are published. For the side chain amide groups they are given by Harsch et al. (2013). The chemical shift δ is then given by
For the chemical shift difference Δδ = δ 2 − δ 1 one obtains
If C 1 is the class of correct stereospecific assignments and C 2 the class of incorrect stereospecific assignments, then the conditional probability P(C 1| Δδ) can be calculated according to Bayes that for a given Δδ a stereochemical assignment is correct by
When a correct assignment is not known a priori, then P(C 1) = P(C 2) = 0.5 can be assumed. However, if other information such as information from NOESY-spectroscopy becomes available, P(C 1) and P(C 2) would have to be adapted.
By using difference distributions the problem is reduced to one dimension but information about the individual chemical shifts is lost. Equation 3 can be rewritten for pairs of chemical shifts by
For becoming more realistic the probability distributions have to be smoothened. Here we use a kernel density estimator with Gaussian kernel k(δ)
The bandwidth h of the kernel was either determined by Silvermans rule (Silverman 1998)
or according to Nasser (2006) using a dynamic bandwith h i (Eq. 7). In Eq. (6) the overall bandwidth h is determined by the standard deviation σ of the underlying chemical shift dataset and the number n of data points in the set. The dynamic bandwith h i replaces h in the kernel of Eq. 5 with
h i is the bandwidth of the predicted chemical shift value δ i and is calculated over a sliding window of the size 2m + 1 data points in the sorted set of chemical shift values. The sliding window is centered around the i-th value. This kind of smoothing was used for Fig. 2.
Distribution of the pairwise sidechain amide proton chemical shift differences. The chemical shifts differences Δδ of the proton resonances of the sidechain amide groups δ(Hδ22) − δ(Hδ21) of Asn and δ(Hε22) − δ(Hε21) of Gln were calculated from the BMRB data. The chemical shifts difference distributions were smoothed by the application of a kernel density estimator with a Gaussian kernel (Silverman 1998). a Asn, b Gln. f(∆δ) is the frequency of the chemical shift differences ∆δ when the area of the gray curves is normalized to 1
In addition to the Bayesian analysis, the data were also analyzed with a support vector machine from scikit-learn (Pedregosa et al. 2011).
Software
The programs GROMACS, SHIFTS, SHIFTX2, and scikit-learn 0.14.1 can be downloaded from http://www.gromacs.org, casegroup.rutgers.edu, http://www.shiftx2.ca, and scikit-learn.org respectively. The program AUREMOL as well as the chemical shift data base are available at http://www.auremol.de.
Results and discussion
Chemical shift distribution of side chain amide protons in the BMRB data base and in the chemical shift data base UACSB
If the hypothesis, that the bimodal chemical shift distribution is the consequence of errors in the stereospecific assignments in the BMRB only, is correct, the chemical shift difference of a given pair of amide protons should be bimodal and the two maxima of the distribution should, as it is indeed observed (Fig. 2), be located symmetrically about 0 ppm. The obtained distributions of the pairwise differences (Fig. 2) is surprisingly similar for Asn and Gln, in contrast to the rather different frequency distributions obtained for the individual chemical shifts. Again this is plausible, since the distributions of chemical shifts in the bimodal distribution of Fig. 1 depend on the distribution of correct and incorrect stereochemical assignments of the data base as well as the non-correlated absolute structure dependent shifts δ 3D of the resonances that may be quite different for Asn and Gln. However, the differences of the pairwise chemical shifts Δδ are mainly determined by the random-coil chemical shift difference Δδ rc,, that is in most cases larger than Δδ 3D, and the distribution of correct and incorrect stereochemical assignments.
Experimentally determined chemical shift distributions are, especially with regard to stereospecific assignments, prone to errors, but chemical shift data bases recalculated from three-dimensional structures have the advantage that assignment errors cannot occur. Usually, chemical shifts are calculated on the basis of random-coil chemical shifts modified by tertiary structure effects. Although the accuracy of the predictions depends on the chemical shift prediction program used, general trends caused by tertiary structure effects can be predicted rather well. For this work, we used the unbiased chemical shift data base UACSB which is based on a structurally unbiased set NH3D of X-ray structures (Sillitoe et al. 2013).
Figure 3 compares the chemical shift distributions obtained from UACSB with the distributions obtained from the BMRB data base. It shows an almost perfect agreement with a subset of the chemical shift distributions obtained from the BMRB indicating that the bimodal distributions probably are assignment artifacts. As suggested, the chemical shift distributions of the two protons are well separated. As a rule, the Hδ21 resonance in Asn is shifted downfield relative to the Hδ22 resonance and the Hε21 resonance in Gln is shifted downfield relative to the Hε22 resonance. In UACSB one can select which of the two programs SHIFTS and SHIFTX2 is applied to the structures refined in explicit water. Note that we had to introduce a correction for the stereospecific assignment (see Materials and Methods). The two distributions shown here were calculated with SHIFTS, which creates somewhat better results for the amide protons. A detailed analysis of the general differences between the two programs will be presented elsewhere since they are not important in the actual context of stereochemical assignment.
Chemical shift distribution of side chain amide obtained from UACSB. The distribution of chemical shifts a δ (Hδ21) and c δ(Hδ22) of Asn and b δ(Hε21) and d δ (Hε22) of Gln. (Gray) distributions obtained from the BMRB, (red) distributions obtained from UACSB. The distributions were smoothed by the application of a kernel density estimator with a Gaussian kernel. The bandwidth of the kernel was derived by a sliding window. The bandwidth is the standard deviation σ of the window which contained 15% of the whole dataset and was centered on the current data point. The random-coil values of the UACSB distribution were corrected to achieve the best match between the experimental and calculated distributions. The correction values were −0.025 ppm for Asn Hδ21, −0.078 ppm for Asn Hδ22, 0.021 ppm for Gln Hε21 and −0.046 ppm for Gln Hε22. f(δ) is the frequency of the chemical shifts δ when the area of the gray curves is normalized to 1
Deconvolution of the experimental side chain amide chemical shift distribution
The question arises if we can deconvolute the BMRB chemical shift distribution for obtaining a “true”, corrected distribution. For such a deconvolution a classifier is required that determines if a pair of chemical shift values has been correctly assigned or if the assignment has to be corrected. The simplest classifier has been proposed by Harsch et al. (2013) with the assumption that P corr (Δδ) = 1 for Δδ < 0, P corr (Δδ) = 0.5 for Δδ = 0, and P corr (Δδ) = 0 for Δδ > 0. Since it represents the probability that a stereochemical assignment is correct, a more accurate definition would acknowledge that the assignment is always correct when both chemical shifts are identical (Δδ = 0). This leads to the improved definition P corr (Δδ) = 1 for Δδ ∈ (−∞, 0] and P corr (Δδ) = 0 for Δδ ∈ (0, ∞]. A second classifier could be a Bayesian classifier derived from the theoretical Δδ distribution (Fig. 4). Using these classifiers, one could reorganize the BMRB data for finding the most probable distribution (Fig. 5) of the experimental data. If we assume a confidence level of 95%, approximately 55% of the stereochemical assignments in the BMRB would be correct, approximately 32% are recognized as exchanged, and for approximately 13% of the assignment we could not make decision for this confidence level (Table 1).
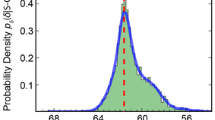
Theoretical amide chemical shift differences and Bayesian probabilities. The ∆δ distributions were calculated from UACSB (grey). The distributions were smoothed by the application of a kernel density estimator with a Gaussian kernel (Silverman 1998). In addition, the probability P(∆δ) of the corresponding Bayesian classifier (green, y2-axis) is shown (see “Materials and methods” section). A Gausssian fit of the ∆δ distributions of UACSB is plotted in red and the probability curve of the corresponding Bayesian classifier is plotted in blue (y2-axis). a Asn, b Gln. f(∆δ) is the frequency of the chemical shift differences ∆δ when the area of the gray curves is normalized to 1
Reconstructed experimental chemical shifts of side chain amide protons. Corrected chemical shift distributions were calculated from the BMRB distributions by using the Bayesian classifier (see Fig. 4). a (Hδ21) and c (Hδ22) of Asn and b (Hδ21) and d (Hδ22) of Gln. A confidence level of 95% was used for the correction of the order of the sidechain NH2-proton resonances of Asn and Gln. f(δ) is the frequency of the chemical shifts δ when the area of the gray curves is normalized to 1.
A program for determining the stereochemical assignment from chemical shifts only
When a pair of amide protons has been assigned, from the chemical difference the probability for a given stereochemical assignment can be calculated by applying the Bayes theorem (Eq. 3) to the chemical shift difference distribution. The corresponding software tool AssignmentChecker is implemented in the program AUREMOL (menu point: Assignment—Check stereospecific assignment of sidechain NH2-groups). Here, either a pair of chemical shifts can be entered manually or a list of chemical shifts can be read in and the stereochemical assignment will be corrected. Independent of the software presented, at a confidence level >95% a stereochemical assignment is correct when the chemical shift difference δ(Ηδ21) − δ(Ηδ22) is greater than 0.4 ppm for Asn or δ(Ηε21) − δ(Ηε22) is greater than 0.42 ppm for Gln, respectively.
Somewhat more significant results can be expected when both chemical shifts and not only the chemical shift difference of the sidechain amide group resonances are used as input. We tested the two-dimensional Bayesian ansatz (Eq. 4) as well as a support vector machine (Schölkopf and Smola 2002), which worked with the chemical shift values as input. For both methods, the results improved only marginally compared to the one-dimensional Bayesian ansatz (Eq. 3), which had the chemical shift difference Δδ of the resonance lines of the sidechain NH2-groups as input. Therefore, we implemented only this type in AUREMOL.
Structural implications
The theoretical chemical shift distributions calculated from the unbiased structural data set (Fig. 4) as well as the corrected experimental distributions from the BMRB (Fig. 5) are characterized by a central Gaussian like distribution but also by significant frequency densities in the overlap region of the Hδ21/ε21 and Hδ22/ε22 distributions. The simulated data show that such an overlap should exist. However, in the corrected data from the BMRB the intensity in this range is significantly larger than in the data derived from the UACSB. This is the result of the fact that the values of the BMRB in the overlap region could not be redistributed at a confidence level >95% and thus the original stereospecific assignment was retained. An additional effect that would lead to identical chemical shift values of the two protons is the fast 180° flip around the C–N bond that would lead to identical experimental values. For asparagine and glutamine in the random-coil Ac-GGXA-NH2 the flip rates are 1.3 and 0.4 s−1 at 293 K, respectively (Harsch et al. 2013). The large reorientaional correlation times imply that fast exchange averaging may only occur in special cases in proteins. The main factor leading to structure dependent deviations from the random coil chemical shifts of amide protons are electric field effects and ring current effects. In addition, paramagnetic shifts may occur when paramagnetic centers are present. The BMRB data base contains a small number of structures with paramagnetic ligands that can lead to extreme paramagnetic shifts up to 111 ppm but are statistically irrelevant and are not contained in the theoretical data base.
The calculated chemical shifts from the unbiased structural data base show that sometimes upfield chemical shifts of individual Hδ21/ε21 or downfield chemical shifts of individual Hδ22/ε22 protons occur that are larger than the corresponding random-coil chemical shift differences. In fact, the largest upfield shifts of Hδ21, Hδ22, Hε21, and Hε22 are 2.39, 2.25, 3.32, and 1.68 ppm, respectively. Correspondingly, the largest downfield shifts values for Hδ21, Hδ22, Hε21, and Hε22 are 8.97, 8.27, 8.86, and 8.23 ppm, respectively. Since in the BMRB even more extreme experimental values are found for diamagnetic proteins this is clearly not an artifact of the chemical shift calculations by SHIFTS. Inspection of Fig. 3 also shows that large downfield shifts for amide protons are more likely than upfield shifts.
Correlated effects in chemical shifts were not in the focus of this paper but promise new, interesting insights. A simple example are the distribution of chemical shift differences. Large differences indicate that the two protons of an amide group experience very different shifts although they are located rather close to each other. For the differences δ(Hδ22) − δ(Hδ21) and δ(Hε22) − δ(Hε21) values between −4.37 and 3.17 ppm and −4.66 and 2.89 ppm, respectively, are calculated. In the BMRB even more extreme values are reported.
Conclusion
We have shown here that in many cases side chain amide protons can be stereochemically assigned from their chemical shift difference only. However, one has to have in mind that this is a probability statement with a predefined error probability. There may be rare cases where the assignment may be wrong. The stereochemical assignment can be important when the amide group is involved in a specific biological interaction and it will increase the precision of distance limits. However, it is well-known that the precision of the distance restraints only determines the accuracy of the 3D structures when the total number of restraints is in the low to intermediate range (<10 restraints per residue). In cases of ambiguity, other methods such as homonuclear NOE (nuclear Overhauser effect) (Wüthrich 1986) or heteronuclear J-coupling patterns should be used. Conversely, chemical shift information can be added if the latter methods cannot accomplish a unique solution. In addition, we have shown that simulated chemical shift data bases have clear advantages when the experimental data base itself is not flawless or limited in numbers as it is expected for stereochemical assignments.
Abbreviations
- NOE:
-
Nuclear Overhauser effect
- NOESY:
-
Nuclear Overhauser enhancement spectroscopy
- UACSB:
-
Unbiased artificial chemical shift database
References
Berenden HJC, van der Spoerl D, van Dunen R (1995) GROMACS: a message-passing parallel dynamics impmementation. Comp Phys Comm 91:43–56
Bundi A, Wuthrich K (1979) 1H-nmr parameters of the common amino acid residues measured in aqueous solutions of the linear tetrapeptides H-Gly-Gly-X-L-Ala-OH. Biopolymers 18:285–297
Gronwald W, Kalbitzer HR (2004) Automated structure determination of proteins by NMR spectroscopy. Prog Nucl Magn Reson Spectrosc 44:33–96
Harsch T, Dasch C, Donaubauer H, Baskaran K, Kremer W, Kalbitzer HR (2013) Stereospecific assignment of the asparagine and glutamine side chain amide protons in random-coil peptides by combination of molecular dynamic simulations with relaxation matrix calculations. Appl Magn Reson 44(1–2):319–331
Hess B, Kutzner C, van der Spoel D, Lindahl E (2008) GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 4(3):435–447
Markley JL, Bax A, Arata Y, Hilbers CW, Kaptein R, Sykes BD, Wright PE, Wüthrich K (1998) Recommendations for the presentation of nmr structures of proteins and nucleic acids. Pure Appl Chem 70:117–142
McIntosh LP, Brun E, Kay LE (1997) Stereospecific assignment of the NH2 resonances from the primary amides of asparagine and glutamine side chains in isotopically labeled proteins. J Biomol NMR 9:306–312
Nasser, A. (2006) Optimierung der Zuordnung mehrdeutiger NOESY-NMR-Signale unter Anwendung einer Datenbank nichtredundanter Proteinstrukturen. Doctoral thesis, University of Regensburg.
Neal S, Nip AM, Zhang H, Wishart DS (2003) Rapid and accurate calculation of protein 1H, 13C and 15N chemical shifts. J Biomol NMR 26:215–240
Osapay K, Case DA (1991) A new analysis of proton chemical shifts in proteins. J Am Chem Soc 113(25):9436–9444
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O et al (2011) Scikit-learn: machine learning in Python. J Machine Learn Res 12:2825–2830
Schölkopf B, Smola AJ (2002) Learning with kernels support vector machines, regularization, optimization, and beyond. Adaptive computation and machine learning. MIT Press, Cambridge
Sillitoe, I., Cuff, Alison L. (2013) New functional families (FunFams) in CATH to improve the mapping of conserved functional sites to 3D structures. Nucleic Acids Res 41:D490–D498
Silverman BW (1998) Density estimation for statistics and data analysis. Chapman & Hall, London
Ulrich EL, Akutsu H, Doreleijers JF, Harano Y, Ioannidi YE, Lin J, Livny M, Mading S, Maziuk D, Miller Z, Nakatani E, Schulte CF, Tolmie DE, Wenger RK, Yao H, Markley JL (2007) BioMagResBank. Nucl Acids Res 36:D402–D408
Word JM, Lovell SC, Richardson JS, Richardson DC (1999) Asparagine and glutamine: using hydrogen atom contacts in the choice of side-chain amide orientation. J Mol Biol 285(4):1735–1747
Wüthrich K (1986) NMR of Proteins and Nucleic Acids. Wiley, New York
Acknowledgements
This work has been supported by the DFG (FOR1979 and KA 647), the Humboldt Society, the Fonds of the Chemical Industry (VCI), and the Human Frontier Science Program Organization (HFSPO).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Harsch, T., Schneider, P., Kieninger, B. et al. Stereospecific assignment of the asparagine and glutamine sidechain amide protons in proteins from chemical shift analysis. J Biomol NMR 67, 157–164 (2017). https://doi.org/10.1007/s10858-017-0093-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-017-0093-x