Abstract
A new theoretical approach was recently addressed to predict cyclodextrin host–guest binding constants with the GFN2-xTB semiempirical quantum method. Within such a strategy, a set of starting supramolecular arrangements is automatically obtained through the UD-APARM software, and many optimized host–guest systems are used to obtain each binding constant. In the present work, within the scope of the multi-equilibrium treatment, we carried out a theoretical study concerning the host–guest systems formed with paraoxon (PRX), methyl-parathion (MPTN), and parathion (PTN) into α-cyclodextrin (α-CD), for which experimental data were addressed. Those guests correspond to pesticides in use, and their inclusion plays a role in remediation technology. The procedure for estimating binding constants for the host–guest system is discussed in terms of the ranges for the supramolecular parameters employed in exploring the GFN2-xTB Potential Energy Surface (PES). As a result, by investigating an unprecedented number of starting systems (3,076), we identified that proper exploration of the GFN2-xTB PES gives a reliable prediction of the binding constant in solution. Furthermore, with the study of different starting associations, for PTN/α-CD, we found an excellent linear correlation (R2 = 0.987) between GFN2-xTB data and experimental information, which, as in our previous study, supports the discussed methodology for application in predicting binding constants for CD-based host–guest systems.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cyclodextrins (CDs) [1, 2] comprise a family of cyclic oligosaccharides produced by an enzymatic process. They can accommodate guest molecules in their cavities to form inclusion compounds [3,4,5]. These complexes are also called host–guest systems. The formation of such a supramolecular system is central to many practical applications of CDs [6]. For example, the interaction of CD with a contaminant is responsible for changes in the degradation rate [7, 8]. Specifically, in terms of remediation, in which CD found many applications [8,9,10,11,12], host–guest systems present different solubilities when compared to isolated pesticides, in which complexation can be a strategy for remotion [13]. CDs can also be employed to mitigate the hazard posed by chemical warfare agents. The inclusion of organophosphorus such as GB (sarin) and GD (soman) and simulants [14] by cyclodextrins (CDs) [14] provide, for instance, a test case for establishing supramolecular agent–simulant correlations.
In the present work, the inclusion compounds formed with α-CD and the pesticides paraoxon (PRX) [15], parathion (PTN) [16, 17], and methyl-parathion (MPTN) [18] for which experimental data were addressed were investigated theoretically. The motivation of the present work is related to the test of a recently developed theoretical methodology in which hundreds of starting supramolecular systems were investigated in a multi-equilibrium approach through a semiempirical quantum method [19].
The experimental characterization of an inclusion compound is well-addressed in the literature. For instance, a NOESY experiment can attest to the formation of such a supramolecular system in a solution through the correlation between the protons of the CD and guest [20,21,22,23]. Isothermal Titration Calorimetry (ITC) is the most common experimental approach for determining thermodynamic parameters. The association constants are distinct for a series of homologous compounds and may vary with temperature, solvent, and pH [23,24,25,26,27]. Throughout ITC, one can determine the association constants named binding constants [28]. However, the scenario is distinct concerning the theoretical approach applied to study CD-based inclusion systems. Cyclodextrins have been the subject of numerous theoretical investigations [29,30,31,32,33,34]. Due to the development of modern computers, the processing capacity has increased considerably, allowing the application of sophisticated quantum methods to CDs. However, not only the size of the CDs poses a challenge to the theoretical investigation but also the flexibility of the host–guest system [35]. In this respect, parameters to account for many possible spatial associations of host and guest can aid the theoretical investigation.
Recently a new approach to characterize supramolecular systems with a focus on the axes of inertia was developed [36], which naturally applies to CD-based systems in differentiation and to obtain cartesian coordinates of starting systems to evaluate stability. The relative position and rotation of the host and guest are evaluated based on a set of association parameters. Specifically for CD-based inclusion compounds, one Cartesian system is constructed with the axis of the inertia of the CD. The positive orientation of such a Cartesian system is determined by evaluating the center of mass of the oxygen atoms, as shown in Fig. 1. Since the number of oxygen atoms for CD differs for each rim from the developed characterization scheme [36], the head and tail identification are promptly identified for any CD orientation in space, as illustrated in Fig. 1. In addition, the head and tail identifications may be relevant in studying CD-based inclusion compounds [37].
Axes of inertia for individual molecules in a supramolecular system formed with a CD (host) and a guest to evaluate the relative position and rotation [36]. As shown, the red arrow in the center of mass of the CD (Z axis) determines the orientation of inclusion through the head or tail rim of the cavity.
The axes of inertia approach allow the study of inclusion on a systematic basis because one can obtain the supramolecular parameters for a given geometry, or with the parameters, one can obtain the geometry. The software to obtain geometry, named UD-APARM, is available for free use (https://github.com/anconi-lab). The keyword SCAN is the central application of such software in the theoretical investigations concerning supramolecular systems. Using the SCAN keyword allows for systematically obtaining hundreds or thousands of starting associations of two non-bonded systems. Through the UD-APARM software, the guest molecule can pass over the cavity of the CD with subsequent rotation along with the inclusion axis. Therefore, the procedure implemented in UD-APARM automatically generates a set of starting supramolecular systems to be investigated. One of the fundamental features of the axes of inertia implementation is related to the possibility of reproducing at any time the set of supramolecular systems studied by a particular contribution. Only the isolated starting Cartesian coordination of the molecules employed is required to obtain an entire supramolecular set of starting arrangements.
Within the study of CD-based host–guest systems, more than one type of inclusion compound can coexist in equilibrium [38,39,40,41], a fundamental aspect related to a theoretical multi-equilibrium approach. However, due to the significant number of CD-based systems to be investigated, the computational cost is a bottleneck to the theoretical investigation in a multi-equilibrium approach. A semiempirical method is well known for its small computational cost, but some implementations are not recommended for studying CDs [42]. In such a context, Grimme and co-workers [43, 44] recently developed a methodology based on tight binding that is an excellent choice for treating CD-based systems and was applied to study such types of supramolecular systems [26, 45].
With the axes of inertia, recently, the GFN2-xTB semiempirical quantum method in a multi-equilibrium scope was applied to compute binding constants in an aqueous solution for CD-based host guest systems [19]. According to our findings, GFN2-xTB binding constants were obtained with an excellent linear correlation to experimental data. However, as the theoretical methodology was recently developed, we are still refining the approach focusing on the experimental binding energies.
In the present work, our motivation consists of testing the methodology recently developed in reproducing the experimental trend addressed for the host–guest systems focused. The GFN2-xTB semiempirical method and the multi-equilibrium approach [19] were used to estimate binding constants. The theoretical results were compared to experimental information [46], in which the discussion focuses on the scan definition, a fundamental aspect of obtaining reliable theoretical outcomes.
Computational details
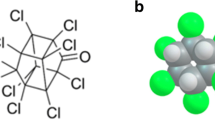
The phosphorothioates studied herein are shown in Fig. 2. Their starting geometries were obtained from PubChem [47] with the following identifiers: CID 9395 (PRX), CID 991 (PTN), and CID 4130 (MPTN). The starting PubChem structures of PRX, PTN, and MPTN were submitted to the CREST software [48] to obtain the most favorable conformers. Next, X-ray crystallography data were used to obtain the starting structures for α-CD [49], in which the explicit water molecules were removed. The starting α-CD structure was optimized with the GFN2-xTB level of theory. Finally, the UD-APARM software was employed to construct hundreds of starting supramolecular associations. All starting structures were included in Supplementary Information.
Structures of the guests studied herein: paraoxon (PRX), parathion (PTN), and methyl-parathion (MPTN). The differences between the pesticides PRX and MPTN concerning PTN are in red, which forms the most stable inclusion compound with α-CD [46].
According to the recently developed approach [36], the UD-APARM software automatically constructed a set of starting non-bonded arrangements with distinct distances of center of mass (r), the position of the guest relative to the CD cavity (θ), rotation of the guest along its axis (α Euler angle), and distinct relative position of the guest along inclusion (β Euler angle). Each set of parameters creates a unique supramolecular system formed with a CD and one guest. In the present work, 792 arrangements were obtained with UD-APARM as the starting set for each supramolecular system investigated (PRX/α-CD, MPTN/α-CD, and PTN/α-CD), totaling 2,376 starting associations. The system in Fig. 1 corresponds to one of the 792 starting arrangements investigated herein. Such systems were submitted to unconstrained optimization at the GFN2-xTB level of theory in a vacuum and aqueous solution through the ALPB [50] continuum solvation model. The multi-equilibrium approach was employed [19]. The starting arrangements studied in the present work were obtained with the ranges included in Table 1, named herein as Starting Input (SI).
The starting systems formed with α-CD and the guests PRX, PTN, and MPTN were generated by UD-APARM according to Table 1. Initially, 792 points were investigated for each host/guest pair. To obtain such starting set, α Euler angle that accounts for guest rotation and inclusion was changed from 0 to 330 degrees in 11 steps. The distance of the center of mass of the host and guest was changed from 0.0 to 8.0 Å within 16 steps, and the guest orientation along the inclusion was changed to 180 degrees (β Euler angle). The guest passes through the CD cavity (change in the θ polar angle). It is worth noting that the data in Table 1, required to prepare the UD-PARAM input and the cartesian coordinates of the starting isolated molecules (available as Supplementary Information), allow one to reproduce the entire set of arrangements investigated in the present work. The parameters in Table 1 were modified to explore the GFN2-xTB PES. Such modifications will be discussed in the results and discussion section. The optimized supramolecular systems were characterized in an aqueous solution using the software APARM (https://github.com/anconi-lab). The APARM software also checked the supramolecular arrangements. Within this study, the root-mean-square deviation (RMSD) was also used in the analysis (http://github.com/charnley/rmsd) to help the discussion concerning the different optimized arrangements obtained. After the initial analysis, a new set of supramolecular starting systems were obtained. The modifications in the range for the UD-APARM input parameters were included as Supplementary Information (Tables S1-S4). Much additional information, along with xTB keywords and data from python3 routines, are described in the Supplementary Information along with the starting geometries of the isolated molecules to allow to obtain the entire set of supramolecular systems investigated herein, to reproduce the total contribution or to test other methods form the starting set of supramolecular systems investigated herein.
Results and discussion
According to the experiment [46], the binding constants correspond to 1.92, 2.28, and 2.77 for including PRX, MPTN, and PTN into α-CD, respectively. The experimental information attests that PTN@α-CD corresponds to the most favorable host–guest system among the systems studied herein. Analyzing the structures in Fig. 2 sheds light on exciting aspects concerning the inclusion of such pesticides into α-CD.
It is noticeable that replacing the sulfur atom attached to the phosphorus in PTN (Fig. 2) for the oxygen atom, which corresponds to the difference between PTN and PRX, decreases the binding constant (log K) from 2.77 to 1.92. The difference corresponds to a change in the Gibbs energy of formation from –3.78 to − 2.62 kcal.mol−1 (difference in K from 5.9 × 102 to 83.3 M−1 [46]). It is also noticeable that replacing the ethyl group from PTN with a methyl group (as in MPTN) changes the Gibbs energy of formation from –3.78 to –3.11 kcal.mol−1. Such modification corresponds to the decrease in the binding constant from 5.9 × 102 to 1.9 × 102 M−1 [46]. Therefore, for the inclusion of PRX, PTN, and MPTN into α-CD, we see that sulfur instead of oxygen attached to phosphorus and ethyl instead of methyl attached to oxygen bonded to phosphorus give rise to the most favorable inclusion compound with α-CD, the PTN@α-CD.
Concerning the GFN2-xTB multi-equilibrium approach, the main objective of the present work corresponds to predicting such an experimental tendency for the inclusion of the pesticides studied herein into α-CD. Using the ranges included in Table 1, we obtained the values of binding constant equal to 4.20, 4.49, and 3.55 for PRX@α-CD, MPTN@α-CD, and PTN@α-CD, respectively. As identified previously [19], GFN2-xTB multi-equilibrium approach gives overestimated Gibbs energies and overestimated binding constants. However, using the range in Table 1 in the scope of the multi-equilibrium approach with GFN2-xTB does not predict the experimental trend. Hence, the theoretical data computed for the PTN@α-CD should be higher than those evaluated.
As mentioned, when CD-based host–guest systems are theoretically analyzed by the GFN2-xTB method, the Gibbs energy of formation obtained is overestimated. Despite that, a strong correlation with the experimental data can be achieved [19]. Assuming that the GFN2-xTB multi-equilibrium approach gives reliable trends based on Gibbs energy of formation for the CD-based host–guest system, the issue identified is related to the initial range for the UD-APARM software (values in Table 1). The range indicated in Table 1 probably did not allow the location of the most representative supramolecular systems in an aqueous solution (ALPB), at least for the PTN@α-CD compound. In principle, the most significant deviation occurred for including the PTN into α-CD.
With a focus on the ranges in Table 1, the goal corresponds to identifying the supramolecular parameters in which the most stable inclusions occur at the GFN2-xTB PES. It is well established that CD-based host–guest compounds are flexible [35]. It is noteworthy that the set of starting supramolecular parameters is not identical to those identified after optimizations, as evaluated by the APARM software. Therefore, the Starting Input (SI) was modified to locate new stable supramolecular host–guest systems. The new study employed the SI data as references in setting new inputs. The analysis was conducted for each pesticide studied herein. After optimization, the APARM values are analyzed to determine a new range for the UD-APARM software. With a focus on the PRX/α-CD associations, some representative starting and optimized parameters were included in Table 2. For PRX pesticide, the most stable inclusions obtained at the GFN2-xTB level of theory had initial values of the center of mass between host and guest (r parameter) between 1 and 2 Å. Furthermore, the Euler angle α, which accounts for the rotation of the guest inside the CD cavity, varies from 120 to 180, in which the other parameters are kept fixed (θ, φ, Euler β, and Euler γ). The most favorable PRX@α-CD compounds optimized at GFN2-xTB (ALPB) level of theory are depicted in Fig. 3.
More stable PRX@α-CD systems derived from different starting points (different supramolecular parameters: A and B in Table 2).
We see from Fig. 3 that the host–guest systems identified here as PRX@α-CD (1) and PRX@α-CD (2) are quite similar, being the main difference the guest rotation inside the α-CD cavity (α Euler parameter). We also see from Table 2 that distinct starting sets of supramolecular parameters (A and B) give rise to different stable inclusion compounds, PRX@α-CD (1) and PRX@α-CD (2). Both systems contribute to the binding constant, coherent with the discussion about the existence of more than one host–guest system at equilibrium in solution.
As shown in Table 2, the inclusion of PRX pesticide occurs more favorably at a distance r of 1.5 Å with a close polar angle (θ) and at different Euler angles (α, β, and γ). Analysis of the APARM parameters and the RMSD plays a role in the investigation. A similar optimized host–guest system was identified from distinct starting parameters (data not shown). After the data analysis, a new UD-APARM input was defined, resulting in a new scan (Table S1). The new UD-APARM input (PRX/SI2) generated 77 starting arrangements.
After GFN2-xTB optimizations of the new 77 starting points (Table S1), eight arrangements presented negative values for Gibbs free energy (ΔG) within a 3.0 kcal.mol−1 window. After optimizations, APARM data (the optimized values for r, θ, φ, α, β, and γ) and RSMD were evaluated. The resulting optimized arrangements obtained with the new scan (PRX/SI2, with 77 points) had already been identified. Therefore, the starting set of supramolecular arrangements obtained by the original scan (Table 1) was sufficient to locate the most representative structures for the PRX@α-CD host–guest system.
For the MPTN pesticide, a new input was also defined based on the SI structural parameters (Table 1) and the resulting optimized values. Therefore, in Table 3, some SI parameters and the corresponding optimized data were included. The focus is on the most stable optimized host–guest system identified for such pesticide as MPTN@α-CD (1) because it was verified that more than one input gives rise to such a stable inclusion compound.
According to Table 3, the inclusion of MPTN pesticide occurs more favorably at a distance r of approximately 1.2 Å, and, as for PRX, the nitro group and the aromatic portion of the pesticide are included in the α-CD cavity. Furthermore, the ΔG value of the inclusion is more negative than the reported value for the inclusion of PRX, in agreement with the experimental information related to binding constant [46].
From the initial UD-APARM data, the most favorable arrangements for the inclusion of MPTN into α-CD occurs between 0 and 2 Å with the angle θ varying between 0 and 180, with the θ = 00 identified for the small starting r value (r = 0 Å). For higher distances, between 0.5 and 2 Å, the starting θ = 180 produces the most favorable inclusion compound focused. According to Table 3, the starting values for φ, Euler α, Euler β, and Euler γ angles corresponding to 0, 120, 180, and 0, respectively, give rise to the MPTN@α-CD (1) arrangement. Noticeably, different inputs give rise to the same host–guest system, the optimized host–guest compounds in Fig. 4. With the analysis of Table 3, a new UD-APARM input (MPTN/SI2) was defined for MPTN/α-CD associations (Table S2), generating 41 new starting arrangements.
After optimization, all arrangements showed negative values for the Gibbs free energy (ΔG) of inclusion in an aqueous solution (with ALPB). Subsequently, the APARM data and the RSMD were evaluated to compare the optimized host–guest systems. The analysis of the optimized systems attested that the arrangements obtained from the new SI for MPTN/α-CD (Table S2), within a negligible Gibbs energy difference (0.02 kcal mol−1), have already been identified. For this reason, for MPTN/α-CD, the application of SI was sufficient to locate the most representative inclusion compounds for such a pesticide.
The same procedure was carried out for the PTN@α-CD host–guest system. However, this system presented a greater variety in the structural data of the more stable arrangements. For a better definition of the new input, the most stable UD-APARM structures located by the SI were analyzed, and the data was collected in Table 4. From such a table, we see that distinct starting associations give rise to the most stable host–guest system: PTN@α-CD (1). We also calculated a Gibbs energy of inclusion (binding energy) equal to − 3.99 kcal.mol−1, a value less negative when compared to PRX@α-CD (1) and MPTN@α-CD (1), which corresponds to − 5.21 and − 5.99 kcal.mol−1, respectively. According to experimental information [46], the binding energy for PTN@α-CD (1) should be more negative (because the log K for such a compound is higher).
After optimization using the semiempirical GFN2-xTB method, it was verified that the inclusion of the PTN pesticide occurs more favorably at a distance r of approximately 1.9 Å, a system identified as PTN@α-CD (1). The PTN@α-CD obtained from (E) input corresponds to the PTN@α-CD (1) with 0.1 Å of difference. The binding energy contradicts the experimental data, as the inclusion of PTN in α-CD is expected to be more favorable than other pesticides. We concluded that SI (Table 1) did not allow the location of the most representative supramolecular systems for the inclusion of the PTN pesticide.
The most stable supramolecular system identified by the SI (Table 1), according to Table 4, occurs from input with initial values between 0.5 and 2.5 Å for the distance of the center of mass and with the angle θ varying between 0 and 180 (mainly equal to 180). The Euler angle α varies between 90 and 150, and the angles φ, Euler β, and Euler γ are 0, 180, and 0, respectively. This information defined a new input (Table S3) that generated 294 new arrangements. The number of steps for r and α Euler (Table S3) accounts for the huge number of new points to investigate for this new UD-APARM input (PTN/SI2). After GFN2-xTB (ALPB) optimizations, the data was collected in Table 5.
Unlike the data found for PRX and MPTN, two new unique and distinguishable arrangements were identified (Table 5) by applying the PTN/SI2 (Table S3). Gibbs energies for formation are more negative than those identified for the host–guest system PTN@α-CD (1). However, the values are not sufficient to account for the experimental trend. The strong dependence between the guest position inside the CD cavity and the binding energy is noticeable. For instance, The PTN@α-CD (new 1) arrangement showed a similar RMSD value to the PTN@α-CD-1, but after optimization, a 6° difference was found in the Euler angle α. This difference stabilized the PTN@α-CD (new 1) host–guest system in 1.16 kcal.mol−1. The analysis of Table 5 suggests that α Euler angle plays a role in determining new stable arrangements. Therefore, a new set of starting parameters (PTN/SI3, Table S4) was defined in which α Euler rotation angle varies from 0 to 300 in 11 steps. With the variation of the distance of the center of mass (from 0.8 to 2.0 in 11 steps), 288 PTN/α-CD supramolecular arrangements were obtained. Data are included in Table 6. Furthermore, it was verified that the starting arrangements obtained with the UD-APARM software with the PTN/SI3 give rise to two new GFN2-xTB optimized stable arrangements with more negative ΔG of inclusion than previously identified (Table 6).
The Gibbs energy of inclusion for the PTN@α-CD (new 3) is more negative than other PTN@α-CD compounds identified that comprise the inclusion of PRX and MPTN. Therefore, we carried out new investigations concerning variations of the UD-APARM range of parameters for PTN/α-CD beyond the PTN/SI3 set of parameters. With new investigations (data not shown), no stable host–guest PTN/α-CD systems were identified with Gibbs energy of inclusion more negative than computed for PTN@α-CD (new 3). Therefore, a limit seems to be reached to include PTN in α-CD through the theoretical approach adopted.
According to data in Table 6, the PTN@α-CD (new 3) and (new 4) host–guest systems are very similar. The distances of the center of mass are very close (1.8 and 1.7 for new 3 and new 4, respectively). Furthermore, the polar angle (θ) and the relative rotational parameters α and β Euler angles differ by 2 degrees. In terms of visualization, it is noticeable that it is challenging to distinguish the new PTN@α-CD arrangements (new 3 and new 4) and PTN@α-CD (1) by simple analysis, as illustrated in Fig. 5. Despite the similarities, the new 3-inclusion compound is the most stable one. The stabilization of the PTN@α-CD (new 3) can be attributed to the formation of a favorable intermolecular interaction between a CD primary hydroxyl group and the PTN sulfur atom (Fig. 5). Such a type of interaction is well known to contribute to the stabilization of other supramolecular systems comprising sulfur atoms [51, 52]. In the present study, such interactions are responsible for a difference in 2.85 kcal.mol−1.
To compute the new theoretical log K for PTN@α-CD, the host–guest systems (new 1 to 4) were considered. Table 7 presents the values of the binding constants obtained by the semiempirical method GFN2-xTB (log KxTB) in a multi-equilibrium scope. The data was obtained using the UD-APARM to construct the starting arrangements and APARM to check and identify the host–guest systems to be included in the binding constant calculations.
A good linear correlation (R2 = 0.987) was obtained between experimental and GFN2-xTB (ALPB) data in Table 7 in a multi-equilibrium scope for CD-based host–guest systems (Fig. 6). It is also noticeable that many systems contribute to each binding constant (Table 7). To compute the binding constants 16, 10, and 6, geometries and theoretical data were used for PRX@α-CD, MPTN@α-CD, and PTN@α-CD, respectively. In the study of CD-based systems, many systems at equilibrium in aqueous media are expected. As previously carried out [19], using the UD-APARM to explore the GFN2-xTB PES gives reliable theoretical information. It was demonstrated that the enlargement of the scan for specific points was essential to obtain a good correlation since new stable arrangements were identified, especially for the PTN@CD system. Table 7 also includes the errors computed with the adjusted Log KxTB, for which a 3% or less error was computed when experimental and adjusted data were compared. The small error associated with adjusted data attests to the robustness of the GFN2-xTB implementation [43, 53] in association with the ALPB continuum solvent model [50] that gives valuable information if the searching protocol identified the representative supramolecular arrangements in a multi-equilibrium scope [19] for CD-based systems.
Linear correlation between experimental and theoretical binding constants for including PRX, MPTN, and PTN pesticides with α-CD. Theoretical data were obtained using the multi-equilibrium approach at the GNF2-xTB theory level (ALPB) with the aid of the UD-APARM software to explore the PES and APARM to identify and check the host-guest systems to be considered in calculations.
At this point of the present study, analyzing the available theoretical method to study CD-based host–guest systems can enhance the potential of the discussed GFN2-xTB (ALPB) multi-equilibrium approach.
The use of Molecular Mechanics (MM) and Molecular Dynamics (MD) for CDs cause controversy when applied to the study of CD-based systems. We have, for instance, early contributions of Dodziuk in which the length dependence of the MD average energy is discussed concerning the experimental trend for complexes of enantiomeric α-pinenes with α-cyclodextrin [54]. In addition, concerning the MM, the test of AMBER, CVFF, CFF91, and MMX force fields and five values of dielectric constants for which no preference was addressed for the selective complexation of decalins by β-cyclodextrin was the basis for concluding that MM is not suitable for reliable analysis of chiral recognition by CDs [55]. The use of MD for CD was also addressed in other selected contributions in which simple MD at vacuum, MM/PBSA, MM/GBSA, and Free Energy Perturbation (FEP) methodologies were employed to provide good qualitative results [56, 57]. The theoretical data for CD-based host–guest systems of such contributions are in qualitative accordance with the experimental trend, mainly concerning the average values computed. It is noticeable that MM/PBSA or MM/GBSA methodology implies the extraction of geometries from the MD simulations carried out with explicit solvent molecules to treat the snapshots evaluated along the production within a continuum approach. In our group, we also have experience in the use of MD and faced the issue related to the fluctuations along a productive MD length. Such fluctuations usually precludes the evaluation of a preference host–guest system due to the small interaction energy compared to the standard deviation computed, even with the use of the MM-PBSA/MM-GBSA methodology [58].
Regarding quantum mechanics, we also contributed to the study of CD-based systems. Usually, the computational cost precludes the investigation of many systems required to model CD-base systems. Notably, low computational cost semiempirical methods such as PM3 must be avoided due to the formation of non-physical artifacts in the study of CD-based systems [59]. An improvement in the implementation corresponds to PM6 and PM6-DH [14] with significant corrections to treat non-bonded systems.
According to the experience of our group, in early studies, the use of a more sophisticated method such as DFT can impose a computational cost that may allow the analysis of only interaction energies [60], even with a scan performed for rigid associations with complete optimization of representative associations [61]. By some point, with DFT, we started to employ the statistical thermodynamics and continuum model to obtain Gibbs energies of inclusion in solution [58, 62, 63]. But a limited number of systems were investigated due to the computational costs despite evaluating Gibbs energies.
In summary, concerning the application of the theoretical method to CD chemistry, it can be stated that MM uses is debatable; MD can give qualitative information with the average values, but the fluctuations usually with the order of magnitude of the differences under investigation, DFT can be prohibitive to accounts for many starting associations and some semiempirical methods such as PM3 cannot be employed.
Within this context, the development of the tight-binding method with the dispersion correction by Grimme and co-workers in 2017 [43] and the development of the GFN2-xTB in 2019, a semiempirical low-cost robust method with dispersion contribution [53] paved the way for fast investigation of CD-based systems through quantum mechanics and statistical thermodynamics. In 2020 a new scheme to characterize supramolecular systems and systematically obtain hundreds of supramolecular associations with the axis of inertia within a reproducible basis through the UD-APARM software was developed by Anconi [36]. The combination of GFN2-xTB and the UD-APARM allows estimating binding constants that comprise many CD-based host–guest systems in equilibrium in aqueous media, the essence of the multi-equilibrium approach. In such methodology, hundreds of starting systems were fully optimized with frequency evaluation to apply the statistical thermodynamics and use Gibbs energies of complexation. Since the individual Gibbs energies of inclusion for CD are overestimated due to the GFN2-xTB implementation, the binding constants are greater than the experimental values. Still, a good linear correlation allows one to adjust the values for a more realistic comparison. Moreover, the data can include a host–guest system not yet synthesized.
With the xTB/UD-APARM multi-equilibrium approach, the experimentalist will have access to a representative set of supramolecular systems that can aid the interpretation of experimental information. Furthermore, predictions can be made regarding the formation of novel inclusion compounds because the approach gives true binding constants (estimated with Gibbs energies of inclusion). Finally, the computation cost allows the study of hundreds or thousands of starting systems that can be promptly obtained to reproduce the entire theoretical contributions using the UD-APARM software. The starting set of supramolecular systems can also be employed for test and comparison with other theoretical methodologies. It is worth noting that investigating many systems accounts for the flexible nature of CD-based host–guest systems.
Due to its recent development [19], the method is under test, which motivates us to reproduce data previously addressed in literature in a series of contributions. Furthermore, developing the procedure to account for additional exploration in the PES of the GFN2-xTB level of theory will allow the prediction of the binding constant for systems not yet obtained experimentally. Our group is already engaged in such theoretical development.
Conclusion
The present work computed through the GFN2-xTB semiempirical quantum method, in a multi-equilibrium scope, the stability constants for the host–guest systems formed by α-CD and the phosphorothioates: paraoxon (PRX), parathion (PTN) and methyl parathion (MPTN). Within the present work, unprecedented 3,076 supramolecular systems systematically obtained through the UD-APARM software were optimized at GFN2-xTB level of theory at the gas phase and in solution through the continuum ALPB model. Based on the results, we concluded that:
-
The multi-equilibrium approach applied to the GFN2-xTB (ALPB) level of theory with the massive use of the UD-APARM software to explore the PES provides overestimated stability constants with a good correlation with experimental data, which allows for obtaining through linear correlation reliable adjusted values.
-
Enlarging the scan for specific points is essential to obtain a good correlation between experimental and theoretical data since the original UD-APARM starting set of arrangements may not include the most stable systems, as identified for PTN@α-CD.
-
For the CD-based host–guest systems investigated (PRX@α-CD, MPTN@α-CD, and PTN@α-CD), a good correlation (R2 = 0.987) was obtained with error less or equal to 3% for adjusted binding constants estimated with the GFN2-xTB (ALPB) level of theory in a multi-equilibrium scope.
References
Szejtli, J.: Introduction and general overview of cyclodextrin chemistry. Chem. Rev. 98(5), 1743–1753 (1998). https://doi.org/10.1021/cr970022c
Rekharsky, M.V., Inoue, Y.: Complexation thermodynamics of cyclodextrins. Chem. Rev. 98(5), 1875–1917 (1998). https://doi.org/10.1021/cr970015o
Jansook, P., Ogawa, N., Loftsson, T.: Cyclodextrins: structure, physicochemical properties and pharmaceutical applications. Int. J. Pharm. 535(1–2), 272–284 (2018). https://doi.org/10.1016/j.ijpharm.2017.11.018
Dhiman, P., Bhatia, M.: Pharmaceutical applications of cyclodextrins and their derivatives. J. Incl. Phenom. Macrocycl. Chem. 98(3–4), 171–186 (2020). https://doi.org/10.1007/s10847-020-01029-3
Del Valle, E.M.M.: Cyclodextrins and their uses: a review. Process Biochem. 39(9), 1033–1046 (2004). https://doi.org/10.1016/S0032-9592(03)00258-9
Crini, G.: Review: a history of cyclodextrins. Chem. Rev. 114(21), 10940–10975 (2014). https://doi.org/10.1021/cr500081p
Kim, D.H., Lee, S.E., Pyo, Y.C., Tran, P., Park, J.S.: Solubility enhancement and application of cyclodextrins in local drug delivery. J. Pharm. Investig. 50(1), 17–27 (2020). https://doi.org/10.1007/s40005-019-00434-2
Landy, D., Mallard, I., Ponchel, A., Monflier, E., Fourmentin, S.: Remediation technologies using cyclodextrins: an overview. Environ. Chem. Lett. 10(3), 225–237 (2012). https://doi.org/10.1007/s10311-011-0351-1
Wang, J., Chen, B.: Adsorption and coadsorption of organic pollutants and a heavy metal by graphene oxide and reduced graphene materials. Chem. Eng. J. 281, 379–388 (2015). https://doi.org/10.1016/j.cej.2015.06.102
Yadav, M., Thakore, S., Jadeja, R.: A review on remediation technologies using functionalized cyclodextrin. Environ. Sci. Pollut. Res. 29(1), 236–250 (2022). https://doi.org/10.1007/s11356-021-15887-y
Sikder, M.T., Rahman, M.M., Jakariya, M., Hosokawa, T., Kurasaki, M., Saito, T.: Remediation of water pollution with native cyclodextrins and modified cyclodextrins: a comparative overview and perspectives. Chem. Eng. J. 355, 920–941 (2019). https://doi.org/10.1016/j.cej.2018.08.218
Waris, K.H., Lee, V.S., Mohamad, S.: Pesticide remediation with cyclodextrins: a review. Environ. Sci. Pollut. Res. 28(35), 47785–47799 (2021). https://doi.org/10.1007/s11356-021-15434-9
Flaherty, R.J., Nshime, B., DeLaMarre, M., DeJong, S., Scott, P., Lantz, A.W.: Cyclodextrins as complexation and extraction agents for pesticides from contaminated soil. Chemosphere 91(7), 912–920 (2013). https://doi.org/10.1016/j.chemosphere.2013.02.005
Sambrook, M.R., Vincent, J.C., Ede, J.A., Gass, A., Cragg, P.J.: Experimental and computational study of the inclusion complexes of β-cyclodextrin with the chemical warfare agent soman (GD) and commonly used simulants. RSC Adv. 7, 38069–38076 (2017). https://doi.org/10.1039/c7ra03328a
Lorke, D.E., Nurulain, S.M., Hasan, M.Y., Kuča, K., Petroianu, G.A.: Combined pre- and posttreatment of paraoxon exposure. Molecules (2020). https://doi.org/10.3390/molecules25071521
Rani, M., Shanker, U.: Degradation of traditional and new emerging pesticides in water by nanomaterials: recent trends and future recommendations. Int. J. Environ. Sci. Technol. 15(6), 1347–1380 (2018). https://doi.org/10.1007/s13762-017-1512-y
Sharma, A., et al.: Worldwide pesticide usage and its impacts on ecosystem. SN Appl. Sci. 1(11), 1–16 (2019). https://doi.org/10.1007/s42452-019-1485-1
Garcia, S.J., Abu-Qare, A.W., Meeker-O’Connell, W.A., Borton, A.J., Abou-Donia, M.B.: Methyl parathion: a review of health effects. J. Toxicol. Environ. Heal. —Part B Crit. Rev. 6(2), 185–210 (2003). https://doi.org/10.1080/10937400306471
Anconi, C.P.A., Souza, L.C.A.: Multi-equilibrium approach to study cyclodextrins host–guest systems with GFN2-xTB quantum method: a case study of phosphorothioates included in β-cyclodextrin. Comput. Theor. Chem. 1217, 113916 (2022)
Fernandes, C.M., Carvalho, R.A., Pereira da Costa, S., Veiga, F.J.B.: Multimodal molecular encapsulation of nicardipine hydrochloride by β-cyclodextrin, hydroxypropyl-β-cyclodextrin and triacetyl-β-cyclodextrin in solution. Structural studies by 1H NMR and ROESY experiments. Eur. J. Pharm. Sci. 18(5), 285–296 (2003). https://doi.org/10.1016/S0928-0987(03)00025-3
Mura, P.: Analytical techniques for characterization of cyclodextrin complexes in aqueous solution: a review. J. Pharm. Biomed. Anal. 101, 238–250 (2014). https://doi.org/10.1016/j.jpba.2014.02.022
Saha, S., Roy, A., Roy, M.N.: Mechanistic investigation of inclusion complexes of a sulfa drug with α- and β-cyclodextrins. Ind. Eng. Chem. Res. 56(41), 11672–11683 (2017). https://doi.org/10.1021/acs.iecr.7b02619
Saha, S., Roy, A., Roy, K., Roy, M.N.: Study to explore the mechanism to form inclusion complexes of β-cyclodextrin with vitamin molecules. Sci. Rep. 6, 1–12 (2016). https://doi.org/10.1038/srep35764
Samuelsen, L., Holm, R., Schönbeck, C.: Cyclodextrin binding constants as a function of pH for compounds with multiple pKa values. Int. J. Pharm. 585, 119493 (2020). https://doi.org/10.1016/j.ijpharm.2020.119493
Tafazzoli, M., Ghiasi, M.: Structure and conformation of α-, β-and γ-cyclodextrin in solution: theoretical approaches and experimental validation. Carbohydr. Polym. 78(1), 10–15 (2009)
Usacheva, T.R., et al.: Complexation of cyclodextrins with benzoic acid in water-organic solvents: a solvation-thermodynamic approach. Molecules (2021). https://doi.org/10.3390/molecules26154408
Sadrerafi, K., Moore, E.E., Lee, M.W.: Association constant of β-cyclodextrin with carboranes, adamantane, and their derivatives using displacement binding technique. J. Incl. Phenom. Macrocycl. Chem. 83(1–2), 159–166 (2015). https://doi.org/10.1007/s10847-015-0552-5
Bouchemal, K., Mazzaferro, S.: How to conduct and interpret ITC experiments accurately for cyclodextrin–guest interactions. Drug Discov. Today 17(11–12), 623–629 (2012)
Lipkowitz, K.B.: Applications of computational chemistry to the study of cyclodextrins. Chem. Rev. 98(5), 1829–1873 (1998). https://doi.org/10.1021/cr9700179
Castro, E.A., Barbiric, D.A.J.: Current theoretical methods applied to study cyclodextrins and their complexes. An. des la Asoc. Quim. Argent. 90(4–6), 1–44 (2002). https://doi.org/10.1002/chin.200551276
Nagaraju, M., Sastry, G.N.: Theoretical studies on inclusion complexes of cyclodextrins. J. Phys. Chem. A 113(34), 9533–9542 (2009)
Anconi, C.P.A., Nascimento, C.S., Fedoce-Lopes, J., Dos Santos, H.F., De Almeida, W.B.: Ab initio calculations on low-energy conformers of α-cyclodextrin. J. Phys. Chem. A 111(48), 12127–12135 (2007). https://doi.org/10.1021/jp0762424
Ding, B., Yu, Y., Geng, S., Liu, B., Hao, Y., Liang, G.: Computational methods for the interaction between cyclodextrins and natural compounds: technology, benefits, limitations, and trends. J. Agric. Food Chem. 70(8), 2466–2482 (2022). https://doi.org/10.1021/acs.jafc.1c07018
Dinar, K.: Theoretical and experimental study on inclusion complexation between guests and cyclodextrins: a critical review. J. Maroc. Chim. Hétérocyclique 19(4), 77–84 (2020). https://doi.org/10.48369/IMIST.PRSM/jmch-v19i4.24288
Dodziuk, H.: Rigidity versus flexibility. A review of experimental and theoretical studies pertaining to the cyclodextrin nonrigidity. J. Mol. Struct. 614(1–3), 33–45 (2002)
Anconi, C.P.A.: Relative position and relative rotation in supramolecular systems through the analysis of the principal axes of inertia: ferrocene/cucurbit[7]uril and Ferrocenyl azide/β-cyclodextrin case studies. ACS Omega 5(10), 5013–5025 (2020). https://doi.org/10.1021/acsomega.9b03914
Anconi, C.P.A., Nascimento, C.S., De Almeida, W.B., Dos Santos, H.F.: The role played by head-tail configuration on the molecular weight distribution of α-cyclodextrin tubes. J. Incl. Phenom. Macrocycl. Chem. 60(1–2), 25–33 (2008). https://doi.org/10.1007/s10847-007-9348-6
Faucci, M.T., Melani, F., Mura, P.: 1H-NMR and molecular modelling techniques for the investigation of the inclusion complex of econazole with α-cyclodextrin in the presence of malic acid. J. Pharm. Biomed. Anal. 23(1), 25–31 (2000). https://doi.org/10.1016/S0731-7085(00)00260-0
Dahlheim, C.E., Dali, M.M., Naringrekar, V.H., Miller, S.A., Shukla, R.B.: Multidisciplinary investigation of atypical inclusion complexes of β-cyclodextrin and a phospholipase-A2 inhibitor. J. Pharm. Sci. 94(2), 409–422 (2005). https://doi.org/10.1002/jps.20245
Floare, C.G., Pirnau, A., Bogdan, M.: 1H NMR spectroscopic characterization of inclusion complexes of tolfenamic and flufenamic acids with β-cyclodextrin. J. Mol. Struct. 1044, 72–78 (2013). https://doi.org/10.1016/j.molstruc.2012.11.021
Jug, M., Mennini, N., Kövér, K.E., Mura, P.: Comparative analysis of binary and ternary cyclodextrin complexes with econazole nitrate in solution and in solid state. J. Pharm. Biomed. Anal. 91, 81–91 (2014). https://doi.org/10.1016/j.jpba.2013.12.029
Casadesús, R., et al.: Testing electronic structure methods for describing intermolecular H ⋯ H interactions in supramolecular chemistry. J. Comput. Chem. 25(1), 99–105 (2004). https://doi.org/10.1002/jcc.10371
Grimme, S., Bannwarth, C., Shushkov, P.: A robust and accurate tight-binding quantum chemical method for structures, vibrational frequencies, and noncovalent interactions of large molecular systems parametrized for all spd-block elements (Z = 1–86). J. Chem. Theory Comput. 13(5), 1989–2009 (2017). https://doi.org/10.1021/acs.jctc.7b00118
Bannwarth, C., et al.: Extended tight-binding quantum chemistry methods. Wiley Interdiscip. Rev. Comput. Mol. Sci. 11(2), 1–49 (2021). https://doi.org/10.1002/wcms.1493
Ferrero, R., et al.: On the interactions of melatonin/β-cyclodextrin inclusion complex: a novel approach combining efficient semiempirical extended tight-binding (xTB) results with ab initio methods. Molecules 26(19), 5881 (2021). https://doi.org/10.3390/molecules26195881
Kamiya, M., Mitsuhashi, S., Makino, M.: Catalytic properties of cyclodextrins on the hydrolysis of parathion and paraoxon in aquatic medium containing humic acids. Chemosphere 25(12), 1783–1796 (1992). https://doi.org/10.1016/0045-6535(92)90019-N
Kim, S., et al.: PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Res. 49(D1), D1388–D1395 (2021). https://doi.org/10.1093/nar/gkaa971
Pracht, P., Bohle, F., Grimme, S.: Automated exploration of the low-energy chemical space with fast quantum chemical methods. Phys. Chem. Chem. Phys. 22(14), 7169–7192 (2020). https://doi.org/10.1039/c9cp06869d
Puliti, R., Mattia, C.A., Paduano, L.: Crystal structure of a new α-cyclodextrin hydrate form. Molecular geometry and packing features: Disordered solvent contribution. Carbohydr. Res. 310(1–2), 1–8 (1998). https://doi.org/10.1016/S0008-6215(98)00150-5
Ehlert, S., Stahn, M., Spicher, S., Grimme, S.: Robust and efficient implicit solvation model for fast semiempirical methods. J. Chem. Theory Comput. 17(7), 4250–4261 (2021). https://doi.org/10.1021/acs.jctc.1c00471
Krepps, M.K., Parkin, S., Atwood, D.A.: Hydrogen Bonding with Sulfur 2001. Cryst. Growth Des. (2001). https://doi.org/10.1021/cg015505v
Platts, J.A., et al.: Directionality of hydrogen bonds to sulfur and oxygen. J. Am. Chem. Soc. 118, 2726–2733 (1996)
Bannwarth, C., Ehlert, S., Grimme, S.: GFN2-xTB—an accurate and broadly parametrized self-consistent tight-binding quantum chemical method with multipole electrostatics and density-dependent dispersion contributions. J. Chem. Theory Comput. 15(3), 1652–1671 (2019). https://doi.org/10.1021/acs.jctc.8b01176
Dodziuk, H., Lukin, O.: Dependence of the average energy between the 1: 2 complexes of enantiomeric a -pinenes with a -cyclodextrin on the length of dynamic simulation. Chem. Phys. Lett. 327, 18–22 (2000)
Bonnet, P., Jaime, C., Morin-allory, L.: α -, β, and γ -cyclodextrin dimers. Molecular modeling studies by molecular mechanics and molecular dynamics simulations. J. Org. Chem. 66, 689–692 (2001)
I. Bea, M. Gotsev, P. Ivanov et al., Chelate Effect in Cyclodextrin Dimers: A Computational (MD, MM/PPSA, and MM/GBSA) study, 71, 5, 2056–2063 (2006)
Bea, I., Jaime, C., Kollman, P., Bea, I.: Molecular recognition by β-cyclodextrin derivatives : molecular dynamics, free-energy perturbation and molecular mechanics/poisson—boltzmann surface area goals and problems. Theor. Chem. Acc. 108, 286–292 (2002). https://doi.org/10.1007/s00214-002-0384-4
Pereira, R.A., da Silva Borges, W.M., Peraro, C.R., Anconi, C.P.A.: Theoretical inclusion of deprotonated 2,4-D and dicamba pesticides in ß-cyclodextrin. J. Incl. Phenom. Macrocycl. Chem. 86(3–4), 343–349 (2016). https://doi.org/10.1007/s10847-016-0665-5
Casadesús, R., Moreno, M., González-Lafont, À., Lluch, J.M., Repasky, M.P.: Testing electronic structure methods for describing intermolecular H··· H interactions in supramolecular chemistry. J. Comput. Chem. 25(1), 99–105 (2004). https://doi.org/10.1002/jcc.10371
Anconi, C.P.A., et al.: Inclusion complexes of a -cyclodextrin and the cisplatin analogues oxaliplatin, carboplatin and nedaplatin : a theoretical approach. Chem. Phys. Lett. 515(1–3), 127–131 (2011). https://doi.org/10.1016/j.cplett.2011.09.005
Teixeira, M.G., et al.: Theoretical and Experimental study of inclusion complexes formed by isoniazid and modified β—cyclodextrins: 1 H NMR structural determination and antibacterial activity evaluation. J. Phys. Chem. B 12, 5 (2014). https://doi.org/10.1021/jp409579m
Pereira, R.A., Anconi, C.P.A., Nascimento, C.S., De Almeida, W.B., Dos Santos, H.F.: Stability and spatial arrangement of the 2,4-dichlorophenoxyaceticacid and β-cyclodextrin inclusion compound: a theoretical study. Chem. Phys. Lett. 633(1), 158–162 (2015). https://doi.org/10.1016/j.cplett.2015.05.044
Anconi, C.P.A., Santos, T.M.R., Souza, A.C.: Host—guest intermolecular hydrogen bonds and stability in aqueous media : the benzaldehyde/β-CD case study. J. Incl. Phenom. Macrocycl. Chem. 89(1), 137–142 (2017). https://doi.org/10.1007/s10847-017-0734-4
Acknowledgements
The authors thank Professor Hélio Ferreira dos Santos for the access to the NEQC (Núcleo de Estudos em Química Computacional, UFJF, Brazil) computer facility and also thanks the Laboratório Central de Computação Científica (LCC) da UFLA for providing additional computing resources. Furthermore, professor Cleber Paulo Andrada Anconi also thanks the Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG) for supporting the Laboratório de Química Fundamental (LQF), where the research was conducted.
Funding
Fundação de Amparo à Pesquisa do Estado de Minas Gerais
Author information
Authors and Affiliations
Contributions
JCM made the first calculations and provided the critical analysis of the data and the expansion of the scan to explore the quantum Potential Energy Surface (PES). Furthermore, JCM wrote the first version of the results and discussion section of the manuscript. CLCR, GT, and EFL obtained outcomes from quantum calculations and, in conjunction with JCM, worked with the vast amount of data to estimate the binding constants in aqueous media through the use of in-house routines (python 3) written by Cleber Anconi. Finally, CPAA supervised the research and wrote the manuscript along with inputs from the authors that revised the results and the first versions of the contribution.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Marcelino, J.C., Ribeiro, C.L.C., Teixeira, G. et al. Inclusion of paraoxon, parathion, and methyl parathion into α-cyclodextrin: a GFN2-xTB multi-equilibrium quantum study. J Incl Phenom Macrocycl Chem 103, 263–276 (2023). https://doi.org/10.1007/s10847-023-01192-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10847-023-01192-3