Abstract
A liposome containing the β-cyclodextrin (β-CD) complex of norfloxacin (NFLX) was prepared and its ternary structure characterized. Structural information about the NFLX/β-CD complex was obtained by infrared spectroscopy and powder X-ray diffraction. Nuclear magnetic resonance (NMR) ROESY experimental results confirmed the inclusion structure, with the results well matching those of molecular docking studies. The stoichiometry and the association constant of the complex were also determined. Free multilamellar vesicles composed of soybean phospholipids and cholesterol as membrane materials were prepared and loaded with NFLX, β-CD, and the NFLX/β-CD complex. Diffusion coefficients measured by 19F NMR spectroscopy were compared. In addition, saturation transfer difference NMR experiments were performed to elucidate the structural differences of these liposomes. The results confirmed the formation of the ternary inclusion system comprising the liposome and the NFLX/β-CD complex. Transmission electron micrographs showed the morphological features and particle size differences of these liposomes.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Chemical compounds studied in this article
Norfloxacin (PubChem CID 4539), β-cyclodextrin (PubChem CID 444041)
Cyclodextrins (CDs), cyclic oligosaccharides consisting of six–eight glucopyranose units linked by α-(1, 4) bonds, have a truncated cone shaped structure with a hydrophobic cavity. The secondary and primary hydroxyl groups of the anhydroglucopyranose moieties are arranged on the exterior of CD molecules rendering their external surfaces polar while the internal surface is apolar. The CD cavity can accommodate small or medium-sized molecules to form inclusion complexes. These complexes are of particular interest in the pharmaceutical industry because incorporation of drug molecules into the cavity of CDs significantly changes the properties of the drug, enhancing, for example, its chemical stability, bioavailability, and solubility [1–3].
A liposome is an artificial spherical vesicle composed of a lamellar phase lipid bilayer. Liposomes can be used as vehicles to administer nutrients and pharmaceutical drugs. While hydrophilic drugs can be encapsulated within aqueous compartments of liposomes, lipophilic drugs are usually entrapped within the lipid bilayer. The concept of “drugs-in-cyclodextrin-in-liposome” systems combines the advantages of both CDs (e.g., increasing drugs solubility) and liposomes (e.g., targeting of drugs) into a single system thus circumventing problems associated with each of them. This concept, first introduced by McCormack and Gregoriadis in 1994, has been successfully applied to several drugs and different CD derivatives [4–7].
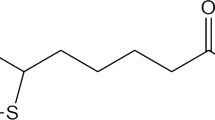
Norfloxacin (NFLX) is one of the second-generation fluoroquinolone drugs known to cause phototoxicity in vivo. Liposomes containing the riboflavin/γ-CD complex reportedly have a highly protective effect on the photosensitive riboflavin molecule [8]. It is therefore likely that incorporation of the CD complex of NFLX into a liposome may also improve the photostability of this fluoroquinolone. The molecular structures of NFLX and β-CD are shown in Fig. 1.
This paper reports the preparation and characterization of the ternary inclusion structure of the NFLX/β-CD complex in a liposome. The NFLX/β-CD complex was characterized by powder X-ray diffraction, infrared (IR) spectroscopy, and nuclear magnetic resonance (NMR) spectroscopy. Liposomes containing NFLX, β-CD, and the NFLX/β-CD complex were prepared independently from free multilamellar vesicles (MLVs). By comparing the diffusion behavior of NFLX in the drug containing systems through 19F NMR spectroscopy and investigating the saturation transfer difference (STD) NMR spectra of these liposomes, the structure of the ternary system could be characterized. This study is the first to investigate the structure of complex between a fluoroquinolone drug and β-CD and a liposome containing this complex by NMR spectroscopy.
Experimental
Materials
NFLX (HPLC purity ≥99.8 %) and cholesterol (HPLC purity ≥99 %) were purchased from Aladdin Reagents, Shanghai, China. β-CD (content ≥99 %) was purchased from Tianjin Chemical Reagents Co., Tianjin, China. Soybean phospholipids (PC content >90 %) were purchased from Shanghai Taiwei Pharmaceutical Co., Ltd., Shanghai, China. D2O was purchased from Cambridge Isotope Laboratories, Inc., Andover, USA. All chemicals used were analytical grade. All of the materials were directly used without further purification.
Preparation of NFLX/β-CD complex
Solid NFLX (0.319 g, 100 mmol) and β-CD (1.135 g, 100 mmol anhydrous) were dissolved in 25 mL of 100 mM NaOH solution. This solution was magnetically stirred for 24 h at room temperature before it was freeze-dried. The solid product was then collected and vacuum-dried at 60 °C to obtain NFLX/β-CD complex.
Characterization of NFLX/β-CD complex via IR spectroscopy and powder X-ray diffraction
IR spectroscopy spectra were obtained by using a Nicolet 6700 Fourier-transform IR (FTIR) spectroscopy with KBr pelleting. The spectra ranged from 400 to 4000 cm−1 at 4 cm−1 resolution. Powder X-ray diffraction was measured in a Rigaku diffractometer using Cu Kα (λ = 1.5406 Å) with 40 mA, 40 kV, and 3°/min scanning rate.
Stoichiometry of NFLX/β-CD complex through the continuous variation method (Job’s plot)
β-CD and NFLX (both 20 mM, in 100 mM NaOH–D2O solution) were mixed to a standard volume in various molar ratios (e.g., 75:525 µL and 150:450 µL), while maintaining the total concentration constant. Eight samples were prepared. All the samples were maintained at room temperature overnight until equilibrium was reached. The 1H NMR spectra were acquired at 298 K using a presaturation pulse sequence to suppress the residual water signal. The chemical shifts were calibrated using sodium 2,2-dimethyl-2-silapentane-5-sulfonate (DSS). The signals of H2, H5, and H8 of NFLX were compared to obtain the Job plots.
Molecular modeling (geometry optimization and molecular docking)
The initial molecular structure of NFLX was created using the GaussView program. The β-CD structure was extracted from a 3CGT protein crystal structure file [9]. Both of the initial structure files were converted to Gaussian input files, and the geometries of the files were optimized using the semi-empirical PM3 method with the Gaussian 03 program package [10]. The output files (NFLX and β-CD) were set to be the ligand and macromolecule of the AutoDock program, respectively. Molecular docking was performed with a Lamarckian genetic algorithm using the AutoDock 4.2 program [11]. The AutoDockTools program was used to set the docking parameters and analyze the docking files.
Preparation of the NFLX, β-CD, and NFLX/β-CD complex liposomes
Soybean phospholipids (100 mg) and cholesterol (25 mg) were dissolved in 10 mL of CHCl3 in a 50 mL flask. Subsequently, NaOH in D2O (100 mM, 5 mL) was added. The organic solvent was removed via rotary evaporation under vacuum at 38 °C, leaving a thin layer of lipids on the wall of the flask. This thin layer was removed from the flask via ultrasonic oscillation, and the lipids were diluted with D2O (5 mL) to compensate for the loss during rotary evaporation. Through ultrasonic oscillation of the opaque liquid, uniform MLVs were generated. Subsequently, 1 mM of 100 mM NaOH in D2O was added to 1 mL of the MLV. The sample was dispersed via ultrasonic probe (300 W, 5 s on and 15 s off) for 10 min to obtain a reference liposome for the STD NMR experiments. By replacing the aqueous phase with 1 mL NFLX, 1 mL β-CD, and 1 mL NFLX/β-CD complex (each 20 mM in 100 mM of NaOH in D2O), the liposomes of the NFLX, β-CD, and NFLX/β-CD complex were prepared. The free liposome and liposome of NFLX were filtered using a 0.22 µm filter membranes (aqueous phase). The liposomes of β-CD and NFLX/β-CD complex could not be filtered through 0.22 µm filter membranes because particle sizes were larger than the filter pores. Therefore, these liposomes were filtered using 0.45 µm membranes instead. All the liposomes were colorless colloidal solutions. Except for the reference sample, all of the other liposome samples had the same concentration of encapsulated NFLX, β-CD, or NFLX/β-CD complex (10 mM).
NMR spectroscopy
All of the NMR experiments were performed at 298 K by using a Bruker Advance II 500 MHz spectrometer with a band observed probe head, operating at the frequency of 500.13 MHz (1H NMR). Tetramethylsilane was used as an internal standard. The 1H NMR details were as follows: spectral width 5000 Hz, scan number 16, relaxation delay 2 s, and data points 64 K. The 1H NMR chemical shifts were calibrated using DSS. The NFLX concentration to measure the chemical shifts and diffusion coefficients was 20 mM in 100 mM NaOH/D2O, and the concentration of the NFLX/β-CD complex for the NMR ROESY experiments was 50 mM.
To perform 19F NMR DOSY experiments, the resonance frequency was set to 470 MHz, and a combination of bipolar gradient pulses and longitudinal eddy delay pulse sequence was used. A series of spectra was acquired with a gradient strength ranging from 2 to 95 % of the maximum gradient amplitude output with 2.5 % steps. The data obtained were processed using the TopSpin T1/T2 package.
For the STD NMR experiments, a pseudo-2D pulse program with a spin-lock filter and water suppression via gradient-tailored excitation (WATERGATE) were used. The total saturation time was set to 2 s. The saturation frequency for the selective pulse was applied with on-resonance at 0 ppm and off-resonance at −30 ppm. The spectra were recorded internally via phase cycling. The spin-lock pulse length was set to 150 ms. The relaxation delay was set to 2 s. Free induction decay was acquired in 32 K complex data points, which covered a spectral width of 8000 Hz. All the STD NMR spectra were acquired after 512 scans. For processing, an exponential window function of 0.3 Hz and two times of zero filling were used prior to Fourier transformation.
Transmission electron micrograph
The free liposome, NFLX liposome, and β-CD and NFLX/β-CD complex samples (20 µL) were transferred separately into copper grids (200 mesh) using filter paper. Ten minutes later, 1 % phosphotungstic acid solution (10 µL for each copper grid) was added for dyeing at room temperature. The copper grids were transferred into a transmission electron microscope (JEOL, JEM-2100). The particulate appearances were observed and photographed.
Results and discussion
FTIR spectroscopy and powder X-ray diffraction
Comparison of the FTIR spectra, and the powder X-ray diffractions of β-CD, NFLX, and the NFLX/β-CD complex indicated the formation of the inclusion structure (Fig. 2).
The powder X-ray diffraction patterns of NFLX and β-CD revealed the existence of crystalline material, while the sample obtained for the NFLX/β-CD complex from freeze-drying possessed an amorphous form. In the IR spectrum of the NFLX/β-CD complex characteristic signals of both components were observed, but the β-CD bands were dominant because of the molar excess of anhydroglucose moieties in the sample with respect to NFLX. A clear indication for the presence of NFLX in the complex is the peak at 1630 cm−1.
1H NMR spectroscopy
The 1H NMR spectra of β-CD, NFLX, and the NFLX/β-CD complex in 100 mM NaOH/D2O are shown in Fig. 3.
Comparison of the spectrum of the complex with the ones of the free components revealed that complex formation causes small, but visible shifts of the signals of NFLX and β-CD. Downfield shifts were observed for the following signals: H5 (δ7.71, 1H, d), H2 (δ8.27, 1H, s), H1, H7α (δ3.05, 4H, m), H7β (δ2.90, 4H, m), and H10 (δ4.15, 2H, t) of NFLX; and H2 (δ3.50, 1H, d), H4 (δ3.44, 1H, t), and H1 (δ4.94, 1H, d) of β-CD, while the following signals shifted upfield: H8 (δ6.86, 1H, d) of NFLX; and H3 (δ3.86, 1H, t), H5, and H6 (δ3.82–3.74, 3H, m) of β-CD. The large shielding effect on H3 (δ3.86–3.80), H5, and H6 (δ3.82–3.74 to δ3.78–3.70) of the β-CD molecule indicate the incorporation of NFLX in the β-CD cavity.
Stoichiometry of the NFLX/β-CD complex
According to Job’s method of continuous variation, the concentration of a complex in a set of samples with different molar fractions of the individual components reaches a maximum when the molar ratio of the components corresponds to the complex stoichiometry [12]. Here, we used NMR spectroscopy to quantify complex concentration in a set of samples containing NFLX and β-CD. Accordingly, samples with different molar ratios of NFLX to β-CD were prepared. By following the signal shifts of the aromatic H2, H5, and H8 protons of NFLX, the Job plots shown in Fig. 4 was obtained. The maxima of the curves were observed at a molar fraction of 0.5, which indicates that the NFLX/β-CD complex has a 1:1 stoichiometry.
Association constant of the NFLX/β-CD complex in 100 mM NaOH/D2O
The diffusion coefficient (D) of a substance is a physical parameter associated with its Brownian molecular motion in solution. The Stokes–Einstein equation relates the diffusion coefficient to factors such as molecular size and shape, temperature, and viscosity. Diffusion coefficients can be measured by DOSY NMR experiments [13].
In the case of equilibria that involve the formation of a complex between two components, which are fast on the NMR time scale, the diffusion coefficient observed in the DOSY NMR experiment represents the weighted average (Dobs) of the diffusion coefficients of the bound molecule (Dbound) and the free guest (Dfree).
where χ is the molar fraction of the bound guest.
Since the molecular size of NFLX is smaller than that of β-CD, the diffusion coefficient of the NFLX/β-CD complex can be assumed to be similar to that of free β-CD. By measuring the diffusion coefficients of NFLX (Dobs) and β-CD (Dbound) and of NFLX in the presence of a known amount of β-CD (Dfree), the association constant K a can be calculated by using the following equation [14, 15]:
where χ is the molar fraction of the bound guest, [H]0 is the total concentration of the host, and [G]0 is the total concentration of the guest. On this basis, the D-values of NFLX (10 mM), β-CD (10 mM), and NFLX/β-CD complex (10 mM) were measured and averaged over three independent measurements. From the obtained results (Dfree = 4.32 × 10−10 m/s, Dobs = 4.17 × 10−10 m/s, and Dbound = 2.67 × 10−10 m/s) the stability constant K a of the NFLX/β-CD complex was calculated to amount to 11 M−1.
NOE measurements of the NFLX/β-CD complex
2D ROESY NMR spectroscopy is an important tool to obtain information about the structure of non-covalently stabilized complexes. The presence of NOE correlation peaks between the signals of guest and host protons indicates that they are arranged at a maximum distance of 0.5 nm in space. In addition, the relative intensities of the NOE peaks correlate with the number of absorbing protons and their distance. In the case of CD complexes, crosspeaks between guest signals with the signals of the H3 and H5 protons of the β-CD molecule are particularly informative because these protons are located inside the β-CD cavity. Partial contour plots of the ROESY NMR spectrum of the NFLX/β-CD complex are shown in Fig. 5. Correlation peaks of H2, H5, and H8 of NFLX with H3 and H6 of β-CD were observed. The signal intensity of the crosspeak to H5 was the strongest, whereas that to H2 was the weakest. Correlation peaks between the signals of H7α and H7β of NFLX with H6 and H5 of β-CD were also observed. However, no correlations were found between H11 and H12 of NFLX with any of the β-CD signals. These results suggest that upon formation of the inclusion complex between NFLX and β-CD the piperazine and 4-quinolone of the NFLX molecule are located inside the β-CD cavity. The absence of correlation signals between the ethyl and β-CD H3 and H5 protons furthermore indicates that the ethyl group remains outside the cavity. The aromatic H5 and H8 protons yielded the strongest NOE peaks, indicating that the benzene ring of NFLX resides inside the center of the cavity. Driven by either hydrophobic interactions or van der Waals forces, the NFLX piperazine moiety seem to approach the apolar wall near the larger β-CD rim. To allow their inclusion, the piperazine and 4-quinolone groups of NFLX are considered to adopt a folded arrangement inside the β-CD cavity.
Partial contour plots of the ROESY spectrum of the NFLX/β-CD complex [(1) weak correlation peaks between H2 of NFLX and H3 and H6 of β-CD, (2) strongest correlation peaks between H5 of NFLX and H3 and H6 of β-CD, (3) strong correlation peaks between H8 of NFLX with H3 and H6 of β-CD, (4) weak correlation peaks between H7α of NFLX with H5 and H6 of β-CD and (5) strong correlation peaks between H7β of NFLX and H5 and H6 of β-CD]
The NOE results thus indicated that the NFLX molecule enters the cavity of β-CD from the larger rim and that the 4-quinolone group and piperazine groups reside inside the β-CD cavity.
Molecular modeling (geometry optimization and molecular docking)
To obtain further structural information about the inclusion structure of the NFLX/β-CD complex and to rationalize the experimental ROESY NMR spectroscopic results, we performed molecular modeling studies using the AutoDock program. The docking results generated two structures representing two families of similar conformations. The lowest energy conformations are shown in Fig. 6.
In both conformations, the NFLX molecule penetrates the β-CD cavity from the larger rim of the truncated cone. Inside the cavity, the benzene ring of NFLX is located at the center and the piperazine group tilted approaching the larger rim. These molecular modeling results therefore agree well with the results of the ROESY NMR spectroscopic experiments.
Diffusion coefficients of free NFLX, the NFLX/β-CD complex, NFLX in a liposome, and the NFLX/β-CD complex in a liposome
The rate of Brownian motion of NFLX molecules can be expected to be reduced when these molecules form inclusion complexes with β-CD or are incorporated into liposomes. By comparing the diffusion coefficients of free NFLX with the diffusion coefficients resulting after inclusion into β-CD and/or a liposome, information about the state of the drug in the different systems can be obtained.
The large amounts of phospholipids and cholesterol in the investigated liposomes cause the corresponding 1H NMR spectra to be very complex. However, the NFLX molecule has a fluorine atom at the sixth position and none of the other components of the investigated systems contain fluorine atoms. Thus, 19F NMR spectroscopy provides selective information about the state of NFLX without interference of the other compounds present in the mixtures. The D values obtained for free NFLX, for NFLX complexed at the same concentrations by β-CD or incorporated in a liposome, and of the β-CD of NFLX in the liposome were therefore measured using 19F NMR. The results including the standard deviations of the measurements, obtained from three independent measurements, are summarized in Table 1.
The decreasing D values indicate restriction of the molecular motion upon complexation of NFLX or incorporation into liposomes. Taking the free form of NFLX as reference, the molecular motion gradually slows down in the direction NFLX/β-CD complex > NFLX in liposome > NFLX/β-CD complex in liposome.
STD NMR
STD NMR spectroscopy is extremely useful to elucidate the bound conformation of a ligand in a protein/ligand complex [16–31]. If a protein proton is selectively excited by 1H NMR spectroscopy until saturation, spin diffusion occurs quickly dispersing the saturation to other protons, including the ones of the ligand through magnetization transfer at the binding site. Deducting the obtained spectrum from a reference spectrum obtained by setting the irradiating frequency beyond the normal region of the chemical shifts (e.g., −30 ppm) generates the STD NMR spectrum. STD signals reflect proximities of ligand protons to ones of the protein thus help identifying the binding epitopes. In the systems investigated here the same method can be used to obtain information about potential NFLX–β-CD–liposome contacts [16, 32].
The STD NMR spectra of NFLX incorporated into the liposome, the free liposome, and the 1H NMR spectrum of NFLX are shown in Fig. 7(1). No differences were observed between the spectra of the liposome containing NFLX and that of the free liposome indicating that the NFLX molecules do not have direct contact with the liposome membrane. Thus, NFLX is not entrapped within the phospholipid bilayer and has no close contact to the bilayer inside the liposome.
(1) STD NMR spectra of NFLX in the liposome, of the free liposome, and the 1H NMR spectrum of NFLX, (2) STD NMR spectra of β-CD in the liposome, of the free liposome, and the 1H NMR spectrum of β-CD, (3) STD NMR spectra of the NFLX/β-CD complex in the liposome, of the free liposome, and the 1H NMR spectrum of the NFLX/β-CD complex
The STD NMR spectra of β-CD in the liposome, the free liposome, and the 1H NMR spectrum of β-CD are shown in Fig. 7(2). Except for H1, which is shielded by the carbohydrate rings, all other protons of β-CD exhibit STD signals. The broad line shapes and shifted resonances indicate that the β-CD molecules are in close contact with the liposome membrane.
The STD NMR spectra of the NFLX/β-CD complex incorporated into the liposome, the free liposome, and the 1H NMR spectrum of the NFLX/β-CD complex are shown in Fig. 7(3). The STD signals of the β-CD moiety of the NFLX complex are similar to those of free β-CD in liposome. Unlike in the spectrum of NFLX inside the liposome, some protons of NFLX generated clear and strong STD signals, including the H5, H2, and the methyl protons. The H7β atoms of piperazine exhibit weak STD signals, while other protons do not generate any signal. The relative intensities of the observed signals are Hmethyl:H5:H2:Hpiperazine7β = 100:66:52:35.
STD NMR signals originate from the NOE transfer, which depended on spatial proximity. The results therefore suggest that after incorporation into the liposome, the inclusion complex between β-CD and NFLX is maintained and the NFLX/β-CD complex is completely entrapped. When irradiating the signals of the phospholipid molecules, protons H2 and H5 of the NFLX molecule that are in close contact with the internal wall of the β-CD molecule accepted the NOE transfer. Thus, magnetization occurs from the phospholipid molecules to β-CD, and then to NFLX. The methyl group of NFLX outside the β-CD cavity has residual degrees of freedom through rotation around the C–C bond leading to a weak STD NMR signal. It is possible that the methyl group approaches the phospholipid hydrophobic chain and accepts the NOE transfer directly from the phospholipid molecules. These STD NMR results thus provide strong evidence for the presence of the ternary structure of the NFLX/β-CD complex incorporated into the liposome.
Transmission electron micrograph
In the transmission electron micrographs shown in Fig. 8, the black spots indicate the presence of hollow vesicles of spherical liposomes. Since both free liposomes and NFLX liposomes were filtered by using a 0.22 µm filter membrane, particle sizes of the liposomes were below 220 nm. In contrast the particle sizes of the liposomes containing only β-CD or the NFLX/β-CD complex are larger, which could be a consequence of the larger size of β-CD molecules with respect to NFLX.
Conclusions
In this work, the inclusion complex between NFLX and β-CD was characterized by various techniques. The measurements indicated that the complex has a 1:1 stoichiometry and they provided information about complex structure. In addition, the NFLX/β-CD complex was incorporated into liposomes using MLV. Diffusion coefficients measured by 19F NMR spectroscopy provided strong evidence about incorporation of this complex into the liposomes. In addition, STD NMR was used to obtain structural information about the ternary system formed. Notably, these measurements showed that the NFLX/β-CD is retained in the liposome and located close to the bilayer membrane.
References
Loftsson, T., Brewester, M.: Pharmaceutical applications of cyclodextrins. 1. Drug solubilization and stabilization. J. Pharm. Sci. 85, 1017–1025 (1996)
Song, L.X., Bai, L., Xu, X.M., He, J., Pan, S.Z.: Inclusion complexation, encapsulation interaction and inclusion number in cyclodextrin chemistry. Coord. Chem. Rev. 253, 1276–1284 (2009)
Martin Del Valle, E.M.: Cyclodextrins and their uses: a review. Process Biochem. 39, 1033–1046 (2004)
McCormack, B., Gregoriadis, G.: Drugs in cyclodextrins in liposomes: a novel concept in drug delivery. Int. J. Pharm. 112, 249–258 (1994)
McCormack, B., Gregoriadis, G.: Entrapment of cyclodextrin drug complexes into liposomes: potential advantages in drug delivery. J. Drug Target. 2, 449–454 (1994)
Loukas, Y.L., Vraka, V., Gregoriadis, G.: Drugs in cyclodextrins in liposomes: a novel approach to the chemical stability of drugs sensitive to hydrolysis. Int. J. Pharm. 162, 137–142 (1998)
Cabec, L.F., Figueiredo, I.M., Paula, E., Marsaioli, A.J.: Prilocaine–cyclodextrin–liposome: effect of pH variations on the encapsulation and topology of a ternary complex using 1HNMR. Magn. Reson. Chem. 49, 295–300 (2011)
Loukas, Y.L., Jayasekera, P., Gregoriadis, G.: Novel liposome-based multicomponent systems for the protection of photolabile agents. Int. J. Pharm. 117, 85–94 (1995)
Schmidt, A.K., Cottaz, S., Driguez, H., Schulz, G.E.: Structure of cyclodextrin glycosyltransferase complexed with a derivative of its main product β-cyclodextrin. Biochemistry 37, 5909–5915 (1998)
Frisch, M.J., Trucks, G.W., Schlegel, H.B., Scuseria, G.E., Robb, M.A., Cheeseman, J.R., et al.: Gaussian 03, Revision-B.03. Gaussian, Inc., Pittsburgh (2003)
Morris, G.M., Goodsell, D.S., Halliday, R.S., Huey, R., Hart, W.E., Belew, R.K., Olson, A.J.: Automated docking using a Lamarckian genetic algorithm and empirical binding free energy function. J. Comput. Chem. 19, 1639–1662 (1998)
Job, P.: Formation and stability of inorganic complexes in solution. Anal. Chim. 9, 113–203 (1928)
Johnson, C.S.: Diffusion ordered nuclear magnetic resonance spectroscopy: principles and applications. Prog. Nucl. Reson. Spectrosc. 34, 203–256 (1999)
Cameron, K.S., Fielding, L.: NMR diffusion coefficient study of steroid–cyclodextrin inclusion complexes. Magn. Reson. Chem. 40, S106–S109 (2002)
Jullian, C., Miranda, S., Zapata-Torres, G., Mendizabal, F., Olea-Azar, C.: Studies of inclusion complexes of natural and modified cyclodextrin with (+) catechin by NMR and molecular modeling. Bioorg. Med. Chem. 15, 3217–3224 (2007)
Meinecke, B., Meyer, B.: Determination of the binding specificity of an integral membrane protein by saturation transfer difference NMR: RGD peptide ligands binding to integrin αIIbβ3. J. Med. Chem. 44, 3059–3065 (2001)
Mayer, M., Meyer, B.: Group epitope mapping by saturation transfer difference NMR to identify segments of a ligand in direct contact with a protein receptor. J. Am. Chem. Soc. 123, 6108–6117 (2001)
Mayer, M., James, T.L.: Detecting ligand binding to a small RNA target via saturation transfer difference NMR experiments in D2O and H2O. J. Am. Chem. Soc. 124, 13376–13377 (2002)
Sandstro, C., Berteau, O., Gemma, E., Oscarson, S., Kenne, L., Gronenborn, A.M.: Atomic mapping of the interactions between the antiviral agent cyanovirin-N and oligomannosides by saturation-transfer difference NMR. Biochemistry 43, 13926–13931 (2004)
Chatterjee, C., Majumder, B., Mukhopadhyay, C.: Pulsed-field gradient and saturation transfer difference NMR study of enkephalins in the ganglioside GM1 micelle. J. Phys. Chem. B 108, 7430–7436 (2004)
Soubias, O., Gawrisch, K.: Probing specific lipid–protein interaction by saturation transfer difference NMR spectroscopy. J. Am. Chem. Soc. 127, 13110–13111 (2005)
Herfurth, L., Ernst, B., Wagner, B., Ricklin, D., Strasser, D.S., Magnani, J.L., et al.: Comparative epitope mapping with saturation transfer difference NMR of sialyl Lewis a compounds and derivatives bound to a monoclonal antibody. J. Med. Chem. 48, 6879–6886 (2005)
Schauff, S., Friebolin, V., Grynbaum, M.D., Meyer, C., Albert, K.: Monitoring the interactions of tocopherol homologues with reversed-phase stationary HPLC phases by 1H suspended-state saturation transfer difference high-resolution magic angle spinning NMR spectroscopy. Anal. Chem. 79, 8323–8326 (2007)
Milojevic, J., Esposito, V., Das, R., Melacini, G.: Understanding the molecular basis for the inhibition of the Alzheimer’s Aβ-peptide oligomerization by human serum albumin using saturation transfer difference and off-resonance relaxation NMR spectroscopy. J. Chem. Soc. 129, 4282–4290 (2007)
Shirzadi, A., Simpson, M.J., Xu, Y., Simpson, A.J.: Application of saturation transfer double difference NMR to elucidate the mechanistic interactions of pesticides with humic acid. Environ. Sci. Technol. 42, 1084–1090 (2008)
Feher, K., Groves, P., Batta, G., Jimenez-Barbero, J., Golland, C.M., Kover, K.E.: Competition saturation transfer difference experiments improved with isotope editing and filtering schemes in NMR-based screening. J. Am. Chem. Soc. 130, 17148–17153 (2008)
Huang, H., Milojevic, J., Melacini, G.: Analysis and optimization of saturation transfer difference NMR experiments designed to map early self-association events in amyloidogenic peptides. J. Phys. Chem. B 112, 5795–5802 (2008)
Pereira, A., Pfeifer, T.A., Grigliatti, T.A., Andersen, R.J.: Functional cell-based screening and saturation transfer double-difference NMR have identified haplosamate A as a cannabinoid receptor agonist. ACS Chem. Biol. 4, 139–144 (2009)
Szczygiel, A., Timmermans, L., Fritzinger, B., Martins, J.C.: Widening the view on dispersant–pigment interactions in colloidal dispersions with saturation transfer difference NMR spectroscopy. J. Am. Chem. Soc. 131, 17756–17758 (2009)
Longstaffe, J., Simpson, M., Maas, W., Simpson, A.: Identifying components in dissolved humic acid that bind organofluorine contaminants using 1H{19F} reverse heteronuclear saturation transfer difference NMR spectroscopy. Environ. Sci. Technol. 44, 5476–5482 (2010)
Viegas, A., Manso, J.O., Corvo, M.C., Manuel, M., Marques, B., Cabrita, E.J.: Binding of ibuprofen, ketorolac, and diclofenac to COX-1 and COX-2 studied by saturation transfer difference NMR. J. Med. Chem. 54, 8555–8562 (2011)
Cabec, L.F., Fernandes, S.A., Paula, E., Marsaioli, A.J.: Topology of a ternary complex (proparacaine–β-cyclodextrin–liposome) by STD NMR. Magn. Reson. Chem. 46, 832–837 (2008)
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Liu, Y., Yuan, X. Preparation and characterization of a ternary inclusion complex comprising the norfloxacin/β-cyclodextrin complex incorporated in a liposome. J Incl Phenom Macrocycl Chem 82, 311–321 (2015). https://doi.org/10.1007/s10847-015-0483-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10847-015-0483-1