Abstract
Purpose
This study aimed to evaluate the epigenetic reprogramming of ICR1 (KvDMR1) and ICR2 (H19DMR) and expression of genes controlled by them as well as those involved in methylation, demethylation, and pluripotency.
Methods
We collected germinal vesicle (GV) and metaphase II (MII) oocytes, and preimplantation embryos at five stages [zygote, 4–8 cells, 8–16 cells, morula, and expanded blastocysts (ExB)]. DNA methylation was assessed by BiSeq, and the gene expression was evaluated using qPCR.
Results
H19DMR showed an increased DNA methylation from GV to MII oocytes (68.04% and 98.05%, respectively), decreasing in zygotes (85.83%) until morula (61.65%), and ExB (63.63%). H19 and IGF2 showed increased expression in zygotes, which decreased in further stages. KvDMR1 was hypermethylated in both GV (71.82%) and MII (69.43%) and in zygotes (73.70%) up to morula (77.84%), with a loss of methylation at the ExB (36.64%). The zygote had higher expression of most genes, except for CDKN1C and PHLDA2, which were highly expressed in MII and GV oocytes, respectively. DNMTs showed increased expression in oocytes, followed by a reduction in the earliest stages of embryo development. TET1 was downregulated until 4–8-cell and upregulated in 8–16-cell embryos. TET2 and TET3 showed higher expression in oocytes, and a downregulation in MII oocytes and 4–8-cell embryo.
Conclusion
We highlighted the heterogeneity in the DNA methylation of H19DMR and KvDMR1 and a dynamic expression pattern of genes controlled by them. The expression of DNMTs and TETs genes was also dynamic owing to epigenetic reprogramming.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Assisted reproductive technologies (ARTs) are an often used strategy by infertile couples to achieve pregnancy [1]. In contrast to humans, when used in domestic animals, such as bovines, ARTs are helpful in improving embryo production rate per year, genetic gain, and genetic and phenotypic selection [2, 3]. The in vitro production (IVP) of bovine embryos is the primary biotechnological tool used in both commercial and research laboratories. More than 350 thousand embryos were produced in vitro in 2020 for commercial purpose [4]. Additionally, IVP of bovine embryos is a useful strategy for better understanding of gametogenesis and early embryonic development, including those studies related to animal models of human relevance [5,6,7]. During IVP of embryos, oocytes undergo in vitro maturation until metaphase II (MII) to achieve nuclear and cytoplasmic competence for fertilization and support the initial cleavages during early embryo development prior to embryonic genome activation [8, 9]. Oocyte growth and maturation are accompanied by epigenetic reprogramming, although genome-wide remethylation and imprinting stablishment affect oocyte competence and fertilization, and embryo development [10, 11].
As the embryo develops, the undifferentiated zygote undergoes differentiation to become a blastocyst, with two cell lineage, an inner cell mass (ICM) and a trophectoderm (TE) [12]. In this moment, genes as NANOG, OCT4, and SOX2, are essential to the proper cell differentiation and embryo pluripotency maintenance [13,14,15]. During the embryo in vitro culture, an epigenetic reprogramming occurs, in which, the genome loses its epigenetic marks brought by gametes due to a DNA demethylation wave [16, 17]. The DNA demethylation is mainly active in the maternal and mainly passive in the paternal genome [18, 19]. The ten-eleven translocation (TET) enzymes are primarily responsible for the active wave of DNA demethylation [20, 21], whereas the exclusion of nuclear DNA methyltransferase 1 (DNMT1) is the main mechanism associated with passive DNA demethylation [22, 23]. After genome DNA demethylation, de novo DNA methylation and remethylation are carried out by DNMT enzymes family [18, 19]. At this developmental stage, epigenetic reprogramming occurs throughout the genome except for the genes controlled by genomic imprinting [24].
Genomic imprinting is an important epigenetic mechanism that regulates gene expression, and consequently, embryo development [25]. In general, imprinted genes present an expression pattern based on their parental origin; maternal or paternal imprinted genes are found in clusters and may be regulated by imprinting control regions (ICRs) [17, 24, 26]. Additionally, at least one differentially methylated region (DMR) is found surrounding the ICRs [27]. Modification of epigenetic mechanisms, such as DNA methylation, can imbalance the expression of genes located at these ICRs and lead to the development of syndromes [25, 28].
On the telomeric region of bovine chromosome 29, two ICRs, namely ICR1 and ICR2, seem to be related to developmental disorders such as the large offspring syndrome (LOS) [29,30,31]. H19DMR is found in the ICR1, and it harbors and regulates the expression of H19 and IGF2 genes. The maternal chromosome transcribes H19 but does not express IGF2, and the opposite occurs in the paternal chromosome. This is possible due to a complex imprinting model involving CTCF-binding protein and enhancer competition model [32,33,34]. In ICR2, KvDMR1 is located in intron 10 of KCNQ1 and in the promoter region of its antisense KCNQ1OT1 [35, 36]. Besides KCNQ1 and KCNQ1OT1, several other genes, including CDNQ1C, PHLDA2, and SLC18A22, are influenced by the KvDMR1 [37,38,39].
Previous studies have shown a relationship between ARTs and aberrant epigenetic reprogramming of gene expression during embryo development in cattle [30, 40, 41] as well as humans [42,43,44]. Loss of imprinting due to alterations in DNA methylation and misregulation of imprinted genes have been cited as frequent causes underlying LOS in cattle and Beckwith–Wiedemann syndrome (BWS) in humans [43]. The present study aimed to evaluate the DNA methylation patterns of H19DMR and KvDMR1 as well as the expression of imprinted genes from ICR1 and ICR2 during bovine oocyte maturation and early embryo development; we also evaluated the expression patterns of important genes associated with epigenetic reprogramming (DNMTs and TETs) and pluripotency. In this study, we show the DNA methylation of H19DMR and KvDMR1 and a dynamic expression pattern of the genes controlled by them, besides a global epigenetic reprogramming using DNMTs, TETs, and pluripotency genes in bovine oocytes and embryos.
Material and methods
Ethics statement
The project was approved by the Ribeirao Preto Medical School Animal Ethics Committee (CEUA-FMRP no. 004/2019–1).
In vitro production of bovine embryos
In vitro maturation of oocytes
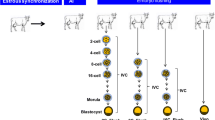
A total of 747 bovine ovaries were collected from a slaughterhouse located near Ribeirao Preto-SP (Brazil). The follicles ranging from 2 to 8 mm were aspirated using an 18-gauge needle connected to a 10-mL syringe [9]. A total of 3960 grade I cumulus-oocyte complexes (COCs) were selected as previously described [45]. The GV oocytes destined for DNA methylation and gene expression analysis were denuded using hyaluronidase solution (4 μg/mL). The denuded oocytes were pooled (n = 40) in 10 μL of phosphate-buffered saline solution (PBS), immersed in liquid nitrogen, and stored at − 80 °C for subsequent DNA methylation and gene expression analysis (Figure S1).
Oocytes were in vitro matured in microdrops (20 COCs/microdrop) of in vitro maturation (IVM) medium [TCM199 with Earle’s salts, glutamine, NaHCO3 pyruvate (22 μg/mL), 10% FBS, FSH (0.5 mg/mL), LH (50 μg/mL), amikacin (83 μg/mL), and estradiol (1.0 μg/mL)], for 22–24 h in maximum humidity, at 38.8 °C and 5% CO2. After IVM, MII oocytes were used for in vitro fertilization or molecular analysis. Those destined for DNA methylation and gene expression analysis were denuded using hyaluronidase solution (4 μg/mL). The denuded oocytes were pooled (n = 40) in 10 μL of PBS, immersed in liquid nitrogen, and stored at − 80 °C for subsequent DNA methylation and gene expression analysis (Figure S1).
In vitro fertilization
Semen samples from a single bull and from the same batch were used for each experiment. Semen was centrifuged (342 × g) using a Percoll (Sigma-Aldrich, USA) gradient (45:90) for 30 min. After centrifugation, the pellet was analyzed for sperm concentration and motility. Matured COCs were placed with spermatozoids (20 COCs/microdrop) in in vitro fertilization (IVF) medium [TALP supplemented with heparin (10 mg/mL), pyruvate (22 μg/mL), BSA FAF (Fatty acid free) (6 μg/mL), PHE solution (2 μM of penicillamine, 1 μM of hypotaurine, and 0.25 μM of epinephrine), and amikacin (83 μg/mL)]. Each drop contained approximately two million sperm per milliliter (2 × 106 sperms/mL). Oocytes and sperms were left together in the IVF medium for 18 h in maximum humidity at 38.8 °C and 5% CO2.
In vitro culture of embryos
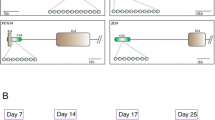
At the end of the fertilization period, presumptive zygotes (single-cell embryos; D0) were transferred to CR2 culture medium [46] (10 embryos/microdrop) with modifications [47]. The embryos were maintained in maximum humidity at 38.8 °C and 5% CO2 for 7 days for preimplantation embryonic development. The cleavage rate was measured 48 h after IVF. On day 4 (D4), the embryos were evaluated and those considered degenerated were discarded; a part of the in vitro culture medium was replaced with fresh medium. On day 7 of development (D7), embryos were classified according to the International Embryo Transfer Society (IETS) as initial blastocysts, blastocysts, expanded blastocysts (ExB), and hatched blastocysts according to blastocoel size, inner cell mass (ICM) position, and trophectoderm (TE) formation. Embryo production was recorded based on viable embryos on D7 in relation to the zygotes cultured on D0. The collection period was based on the method described by Hafez and Hafez [48], and pools were collected as follows: zygotes (n = 40), 4–8 cells (n = 30), 8–16 cells (n = 30), morula (n = 20), and ExB (n = 10) (Figure S1). Samples were pooled in 10 μL of PBS, immersed in liquid nitrogen and stored at − 80 °C for subsequent DNA methylation and gene expression analysis.
Analysis of gene expression using real-time quantitative PCR (qPCR)
RNA was extracted using the PureLink™ RNA Mini Kit (ThermoFisher Scientific, Waltham, MA, USA) following the manufacturer’s protocol. Total RNA was quantified using a NanoDrop 2000 spectrophotometer (ThermoFisher Scientific, Waltham, MA, USA). The complementary DNA (cDNA) synthesis was performed using the SuperScript® IV First-strand Synthesis System Kit (ThermoFisher Scientific, Waltham, MA, USA) using oligo dT and random hexamer primers (70/30 proportion), following the manufacturer’s instructions.
Gene expression analysis was performed using a StepOne Real-Time PCR System (Applied Biosystems, USA) containing SYBR® Green PCR Master Mix, 2 pmol of each primer, and 2 μL of 1:2 diluted cDNA in a final volume of 10 μL. The amplification conditions were as follows: 95 °C for 10 min, followed by 45 cycles of 95 °C for 15 s and 65 °C for 1 min. The reactions were performed and analyzed in triplicates, and only those with a standard deviation > 0.3 were included. Negative controls were used to detect any contamination. The primer sequences for IGF2, KCNQ1OT1, and the reference gene GAPDH were as described by Verruma et al. [49]. The primer sequences for H19, PHLDA2, CDKN1C, KCNQ1, DNMT1, DNMT3a, DNMT3b, TET1, TET2, TET3, OCT4, and NANOG genes were designed using the Primer 3Plus software based on bovine genome reference (assembly ARS-UCD1.3) obtained using NCBI database (ncbi.nlm.nih.gov). ACTB reference gene primer sequence was previously described by Rios et al. [50]. The primer sequences are listed in supplementary table S1. Primer efficiencies were obtained using linear regression (5 points, 1:2 dilution range), and those with efficiencies between 90 and 105% were considered suitable for gene expression analysis, following the manufacturer’s recommendations. Relative gene expression was evaluated using the method described by Pfaffl [51].
Genomic DNA isolation and bisulfite modification
Genomic DNA was extracted using lysis protocol with 2 μL Proteinase K (20 ng/µL) (Invitrogen, USA) and 18 µL Tris HCl (10 mM) for 1 h at 55 °C followed by 10 min at 95 °C for enzyme inactivation. DNA concentration and integrity were evaluated using a Nanodrop 2000 spectrophotometer (ThermoFisher Scientific, Waltham, MA, USA). Bisulfite conversion of DNA was carried out using the EpiTect Bissulfite Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. The DNA was stored at − 20 °C until further use.
The CpG islands at H19DMR and KvDMR1 as well as the primer sequences were predicted using the MethPrimer software [52]; sequences are listed in Supplementary Table S1. The amplification was carried out using a Bio-Rad T100 Thermal Cycler (Bio-Rad Laboratories, USA); each reaction tube contained 10 × PCR buffer II, MgCl2 (25 mM), 5 mM of dNTPs (10 mM/μL), 5 pmol of each primer (10 pmol/μL), 0.04 IU of AmpliTaq Gold® DNA Polymerase (5 U/μL; ThermoFisher Scientific, Waltham, MA, USA), 3 μL of bissulfite converted DNA, and ultrapure water in a final volume of 25 μL. Amplification conditions were as follows: 5 min at 95 °C, followed by 50 cycles of 45 s at 94 °C, 45 s at 59 °C, and 45 s at 72 °C. The final extension step was performed for 10 min at 72 °C. The PCR products were stored at − 20 °C until next generation sequencing.
DNA methylation analysis using BisSeq
Targeted bisulfite sequencing was performed using the MiSeq (Illumina, USA) platform, covering 10 CpG sites in H19DMR (chr29: 51,168,375–51,168,468) and 18 CpG sites in KvDMR1 (chr29:50,580,053–50.580,210), as shown on Fig. 1. PCR products containing the adapters were barcoded using the Illumina Nextera XT library preparation kit, and sequencing was performed using the 600 bp V3 reagents kit according to the manufacturer’s instructions. The FASTQ files for individual samples were generated using the Illumina pipeline (bcl2fastq2-v2-20). Trimmomatic v0.38.124 [53] was used to remove the adapters and indices from the sequences. The paired read sequences were merged using the default settings of FLASH v1.2.11.425 [54] and aligned to the bisulfite-converted genome using the Bismark v0.18.2 [55] with setting–ambig_bam26, which was also used to count the reads with different DNA methylation patterns. Methylated CpGs were visualized using the UCSC Genome Browser.
Schematic representation of telomeric region of BTA29, highlighting the H19DMR (in the ICR1) and KvDMR1 (in the ICR2) sequenced regions and the genes controlled by theses ICRs. Genes with paternal expression (maternal imprinting) are represented in blue (IGF2, in the ICR1, and KCNQ1OT1, in the ICR2) and maternal expression (paternal imprinting) are represented in pink (H19, in the ICR1, and CDKN1C, KCNQ1, and PHLDA2, in the ICR2)
Statistical analysis
The statistical analysis was performed using the RStudio software (v. 1.4.1717). The data distribution was estimated using the Shapiro–Wilk test. The expression profiles of each gene during the seven stages were compared using ANOVA followed by Tukey’s post hoc test, with a level of significance set at 5% (p < 0.05).
Results
Thirteen replicates were performed to collect all samples during oocyte maturation and embryo preimplantation development. A total of 2478 zygotes were initially cultured in vitro. We observed a cleavage rate of 86.50% ± 5.0% and embryo production at D7 of 36.26% ± 10.7%. Preimplantation developmental stages were selected according to the method described by Hafez and Hafez [42] during oocyte maturation (GV and MII) and embryo preimplantation development before (zygotes and 4-8 cells embryos), during (8-16 cells embryos), and after (morula and ExB) bovine genome activation, as well as during the embryo differentiation into two cell lineages, the ICM and TE.
H19DMR and KvDMR1 methylation pattern and gene expression
In ICR1, the H19DMR controls the H19 and IGF2 genes. Ten CpGs were analyzed in H19DMR. H19DMR methylation increased during oocyte maturation (from 68.04% in GV to 98.05% in MII oocytes) and remained higher in zygote (85.83%) and 4–8 cells embryo (83.23%). Subsequently, DNA methylation at H19DMR decreased in 8–16 cells (69.15%) and morula (61.65%), while maintaining its DNA methylation level in ExB (63.63%) as observed in Fig. 2A.
ICR1 analysis during oocyte maturation and embryo preimplantation development. A H19DMR methylation pattern, measure in percentage that alters between hypomethylation (blue) to hypermethylation (red). B and C H19 and IGF2 gene expression, respectively, where pink marks represent gene that present maternal monoallelic expression (paternal imprinting) and blue marks represent gene that present paternal monoallelic expression (maternal imprinting). Different letters in different developmental stages indicate statistical difference (p < 0.05)
There are no changes in H19 and IGF2 expression during oocyte maturation. (Fig. 2B and C). After oocyte maturation, both H19 (F(6) = 41.89; p < 0.01) and IGF2 (F(6) = 44.2; p < 0.01) showed an upregulation in the zygote stage, the highest expression. After upregulation in the zygotic stage, both genes were downregulated at 4–8 cells stages, thereafter, maintaining low levels with no significant difference in the expression until the blastocyst stage. Notably, the expression of H19 was detected at all evaluated stages; however, the IGF2 expression was not detected in the ExB stage (Fig. 2B and C).
In ICR2, the KvDMR1 controls a higher number of genes when compared to the H19DMR. A total of 18 CpGs were analyzed in the KvDMR1 (Fig. 3A). The KvDMR1 methylation remained stable between GV (71.82%) and MII oocytes (69.43%). During embryo development, the hypermethylation status was maintained between zygote and morula stage (~ 74.44%). Between morula and ExB, it was observed a reduction in the KvDMR1 methylation and ExB was hypomethylated (36.64%) (Fig. 3A).
ICR2 analysis during oocyte maturation and embryo preimplantation development. A KvDMR1 methylation pattern, measure in percentage that alters between hypomethylation (blue) to hypermethylation (red). B, C, D, and E CDKN1C, KCNQ1, KCNQ1OT1, and PHLDA2 gene expression, respectively, where pink marks represent genes that present maternal monoallelic expression (paternal imprinting) and blue marks represent gene that present paternal monoallelic expression (maternal imprinting). Different letters in different developmental stages indicate statistical difference (p < 0.05)
Several genes are influenced by the KvDMR1 methylation pattern, including CDKN1C, KCNQ1, KCNQ1OT1, and PHLDA2 (Fig. 3B, C, D, and E, respectively). Similar to IGF2 and H19, zygote was the stage with higher expression in the KCNQ1 (F(6) = 5.96; p < 0.01) and KCNQ1OT1 (F(6) = 20.04; p < 0.01) (Fig. 3C and D). CDKN1C showed a major expression in MII oocytes (F(6) = 14.95; p < 0.01), whereas GV was the higher expression in PHLDA2 (F(6) = 44.2; p < 0.01), as shown in the Fig. 3B and E. All genes, except CDKN1C, were downregulated in the 4–8 cells. CDKN1C, KCNQ1OT1, and PHLDA2 were detected at all the evaluated stages. KCNQ1 showed the lowest expression among all genes of ICR2; its expression was not detected during GV oocyte, morula, and ExB stages.
Expression of DNA methylation, demethylation, and pluripotency-related genes
The expression of DNMTs (DNMT1, DNMT3a, and DNMT3b) and TETs (TET1, TET2, and TET3) was detected at all evaluated stages. In the DNMT family, the MII oocyte showed the highest expression, followed by the GV stage (Fig. 4A, B, and C). Downregulation was observed between oocyte maturation and zygote for DNMT1 (F(6) = 84.43; p < 0.01) and DNMT3b (F(6) = 153.2; p < 0.01), and between MII and 4–8 cells for DNMT3a (F(6) = 63.22; p < 0.01).Although in different proportions, the expression of DNMT3a and DNMT3b presented a similar pattern during oocyte maturation and preimplantation embryo development (Fig. 4B and C).
In the TET family, TET1 (F(6) = 128.8; p < 0.01) showed the highest expression in the morula stage (Fig. 4D), whereas TET2 (F(6) = 22.53; p < 0.01) and TET3 (F(6) = 73.99; p < 0.01) showed the highest expression in the GV stage among all the studied stages (Fig. 4E and F). TET1 gene maintained a low expression until the 4–8 cells stage, and then, it was upregulated until the morula stage, followed by a downregulation until the ExB stage. TET2 and TET3 presented similar expression patterns during oocyte maturation and early embryonic development, with a downregulation between GV oocyte and 4–8 cells, and then remaining relatively low until the ExB stage (Fig. 4E and F).
The expression of NANOG gene was not detected in oocytes (GV and MII) and zygote. Its expression was detected only from the 4–8 cells onwards. From 4-8 cells until morula stage, its expression was upregulated (F(6) = 11.8; p < 0.01), followed by a significant downregulation in the ExB stage (Fig. 4G). Different of NANOG, the expression of OCT4 was detected in oocytes and embryos. The OCT4 showed two moments of upregulation (F(6) = 115.9; p < 0.01), first during oocyte maturation and the second during the morula stage. After upregulation in the morula stage, a downregulation was observed in the ExB stage (Fig. 4H).
Discussion
The early embryo development is crucial for pregnancy success. The IVP of bovine embryos is a useful tool for obtaining biological material for research related to gametogenesis and early embryo development. Imprinted genes are essential during this period, and in our study, their expression pattern changed during oocyte maturation and early embryo development. In addition, genes related to DNA methylation and demethylation also showed a dynamic expression. Epigenetic processes, including genomic imprinting, are fundamental to fetal and placental development, and some imprinted genes may be embryo, tissue, and/or specie specific, making their analysis difficult [5, 56, 57].
In ICR1 (H19DMR), our study showed hypermethylation during oocyte maturation. The hypermethylation in MII oocytes corroborates with previous data obtained by our research group (unpublished data), as well as in recent data published by Vargas et al. [58]. However, the literature describes an hypomethylation status in mammal MII oocytes [59,60,61]. Contradictory data are frequently found when DNA methylation and gene expression patterns are compared between in vitro and in vivo oocytes and embryos [62, 63]. The hypermethylation at H19DMR found in our results may be related to some factors, such as the region selected for sequencing. Al-Khtib et al. [64] observed that human MII oocytes were generally unmethylated in the H19DMR after IVM; however, some CpG islands in this region showed a higher methylation status. In addition, the in vitro maturation environment should also be considered. It is known that the in vitro manipulation could influence the DNA methylation pattern [65, 66].
A reduction in the H19DMR methylation status was observed between 4–8 and 8–16 cells, maintained stable until ExB (~ 65%). After fertilization, the embryo undergoes to an epigenetic reprogramming that erases the epigenetic marks brought by the parental genomes [24]. However, ICRs and imprinted genes maintained its marks and, due to the parental of origin, the ICRs are usually 50% (± 10%) methylated [67, 68]. In bovine, around 8–16 cells occur the genome activation and new epigenetic marks are stablished in the embryo [19, 69].
IGF2 and H19, both imprinted genes, are found in the neighboring region of H19DMR. IGF2 exhibits maternal imprinting and low monoallelic expression during early bovine development [70,71,72,73]. In the current study, the IGF2 expression was detected at all stages, with the exception of ExB; the zygote has shown the highest expression [71,72,73]. In addition to its importance during early embryo development, a recent study has demonstrated its relevance in the later stages of pregnancy, wherein fetal IGF2 seems to be involved in placental vascularization [74]. The H19, which is found near IGF2, produces a conserved long noncoding RNA (lncRNA), the first functional lncRNA described in the literature [75, 76]. H19 plays an oppositive role of IGF2, repressing embryo weight, growth, and modifications in the H19 expression during early embryo development may lead to phenotypic alterations [73, 77,78,79]. Our results showed that at all evaluated stages, the expression of H19 was higher than IGF2; however, both genes presented similar patterns, with higher expression in the zygote, followed by a downregulation (p < 0.05) and stable expression during the remainder of the preimplantation development.
In ICR2, the KvDMR1 influences a higher number of genes that modulate imprinting, including CDKN1C, KCNQ1, KCNQ1OT1, PHLDA2, and SLC22A18 [31], and its DNA methylation pattern regulates the expression of these genes [29, 35, 80, 81]. In our study, the KvDMR1 was found to be hypermethylated during oocyte maturation. In accordance to our results in MII oocytes, the KvDMR1 is hypermethylated in oocytes [82, 83]. Contrary to the hypermethylation found in the oocytes, spermatozoa is hypomethylated in the KvDMR1 [84]. The hypermethylation status was maintained until the morula stage, with ExB being the only hypomethylated stage (36.64%). A hypomethylation status in KvDMR1 was also observed by Khoueiry et al. [85] in human embryos considered suitable for transfer.
The KCNQ1 belongs to a large family involved in potassium channel formation, which is fundamental to biological processes, including ion exchange for cell volume maintenance [86]. In our study, this gene presented a higher expression level in the zygote. After that, a downregulation was observed between the 4–8 and 8–16-cells stages. Its expression was undetectable in morula and ExB stages. Antisense of KCNQ1, the KCNQ1OT1 is paternally expressed and influences the activity of several surrounding genes. In our study, its expression was stable during oocyte maturation and embryo development, presenting an increase only in zygotes, followed by a downregulation in the next stage. KCNQ1OT1 transcribe an lncRNA that regulates chromatin and the nucleus, influencing the expression of several genes [38, 87].
The highest variation in gene expression was observed for CDKN1C. Between oocyte maturation and embryo development, we observed two upregulations in this gene, first in GV oocytes and second in 8–16-cells embryos, followed by a downregulation. CDKN1C is a cyclin-dependent kinase complex (CDKs) controlled by lncRNAs. It regulates the cell cycle and encodes a protein (p57Kip2) whose function is largely associated with correct embryo development and pregnancy evolution [38, 88]. In bovines, Driver et al. [37] have demonstrated that silencing CDKN1C leads to a reduction in bovine embryo production rate. The p57Kip2 may be related to the nutrients provided to the fetus by the placenta and, when associated with other genes, such as PHLDA2, it plays crucial roles during early embryo development [37, 89, 90]. Contrary to the other KvDMR1 genes, the PHLDA2 was the only gene that showed a downregulation during oocyte maturation. Similar to the results described by Jiang et al. [56], in the present study, PHLDA2 was found to be upregulated after embryo genome activation until the morula stage, followed by a reduction of its expression in the ExB, with a relative expression close to zero.
Von Meyenn and Reik [91] have shown that epigenetic reprogramming is fundamental to mammalian development and appears to be conserved among several species. In our study, we observed alterations in the expression of DNMT1, DNMT3a, and DNMT3b before (GV) and after (MII) oocyte maturation and in the five evaluated periods during embryo development. These findings corroborate with previously published data [92, 93]. Oocyte growth and maturation are accompanied by increased expression of DNMT3a and DNMT3b [11]. During oocyte maturation and early embryo development, DNMT genes exhibit a dynamic pattern, with upregulation and downregulation depending on the developmental stage [93]. Although in different proportions, in our study, the DNMTs demonstrated similar behavior between GV oocytes and 4–8-cells embryos. An upregulation was observed in the DNMTs expression during oocyte maturation, followed by downregulation in the zygote (higher expression rate) and 4–8 cells. In bovines, the epigenetic reprogramming that occurs during early embryo development has been shown to be due to the action of TET genes in addition to the lower expression of DNMT1 in a passive demethylation process [16, 94]. Reduced DNMT1 expression was also observed in our study, with lower expression levels in 4–8 and 8–16 cells embryos, prior to genome activation. Using murine model, Uysal et al. [95] demonstrated the importance of Dnmt1 and Dnmt3a during preimplantation development. Their knockdown upregulated pluripotency genes, such as NANOG, and increased apoptosis and reactive oxygen species levels.
Analysis of the results showed that TET1 had higher expression rates in the later stages of embryo preimplantation development (morula and ExB), whereas for TET2 and TET3, higher expression was observed during oocyte maturation (GV and MII). Bovine TET3 is believed to be involved in maternal DNA demethylation and, in addition to the oxidation process, is capable of controlling DNA methylation levels, preventing the addition of new methyl groups during de novo DNA methylation [18, 96]. In accordance with our data, Wossidlo et al. [97] showed in bovine that TET3 transcript is elevated in the oocytes and rapidly decreases in the early embryo development.
In the murine model, the Tet1 seems to be related to ICM specification, besides the pluripotency maintenance, influencing NANOG gene regulation, for example [98]. Our data showed that the expression of NANOG was detected only after 4–8 cells. In bovine embryos, this gene is not expressed during early stages and is only detected in 8-cells embryos.[99, 100]. In addition, the expression patterns observed for both OCT4 and NANOG during oocyte maturation and embryo development corroborate with the literature [99, 101]. NANOG seems to be active by the activity of other pluripotency genes, such as the OCT4 gene [13, 101].
Graphical analysis showed that OCT4 was upregulated twice, first during the oocyte maturation and then during the morula (higher expression) and ExB stages and, similar pattern was described by Khan et al. [99]. Simmet et al. [13] demonstrated that embryos produced by somatic cell nuclear transfer, whose donor cell was OCT4 knockout, presented maternal OCT4 mRNA in the morula, demonstrating the importance of its high expression level in the oocytes. The second stage of increased expression could be related to its main function, as this gene is responsible for cell differentiation. Between the stages of morula and blastocyst occurs the blastocoel cavitation and cell differentiation into the first two lineages, ICM and TE [102, 103].
A limitation of this study was the lack of H19DMR and KvDMR1 methylation analysis in semen. However, the use of only one bull was a fixed variable to evaluate possible variation in oocytes and embryos. The main strength of our study is that we evaluated H19DMR and KvDMR1 DNA methylation, expression of genes controlled by them, and global epigenetic reprogramming using DNMTs, TETs, and pluripotency genes at multiple stages during preimplantation development of bovine embryos. Evaluating five different stages allows us to track gene expression as the embryo develops. These data could be helpful and assist to clarify some gaps in embryonic development and the influence of ARTs on these crucial stages. Furthermore, it would be interesting to compare our results with the in vivo production of bovine embryos in future experiments.
ICR1 and ICR2 are highly conserved in different species, including humans, dogs, and cattle [67], and the DNMTs and TETs are crucial for DNA methylation maintenance and epigenetic reprogramming during embryo preimplantation development. In the current study, H19DMR and KvDMR1 were hypermethylated during almost all evaluated stages, and their neighboring imprinted genes presented a dynamic expression pattern during the main embryo preimplantation development. Zygotes presented the highest expression rate compared to other embryonic stages; this could be due to maternal storage during the final oocyte maturation, which is consumed during the first cleavages [104, 105]. In addition, the expression pattern of pluripotency-related genes, owing to their increase at the beginning of cell differentiation, indicated that the embryo development was appropriate. Our findings should assist future studies on epigenetic reprogramming and the influence of ARTs on bovine embryos.
Data Availability
All data is available from the corresponding author and can be made available on request.
References
René C, Landry I, de Montigny F. Couples’ experiences of pregnancy resulting from assisted reproductive technologies: a qualitative meta-synthesis. Int J Nurs Stud Adv. 2022;4.
Grupen CG. The evolution of porcine embryo invitro production. Theriogenology. 2014.
Sirard MA. The influence of in vitro fertilization and embryo culture on the embryo epigenetic constituents and the possible consequences in the bovine model. J Dev Orig Health Dis. 2017;8:411–7.
Viana JH. Statistics of embryo production and transfer in domestic farm animals [Internet]. Embryo Technol. Newsl. 2021. Available from: https://www.iets.org/Portals/0/Documents/Public/Committees/DRC/IETS_Data_Retrieval_Report_2020.pdf
Ménézo YJR, Hérubel F. Mouse and bovine models for human IVF. Reprod Biomed Online [Internet]. 2002;4:170–5. Available from: https://doi.org/10.1016/S1472-6483(10)61936-0
Miller W, Rosenbloom K, Hardison RC, Hou M, Taylor J, Raney B, et al. 28-Way vertebrate alignment and conservation track in the UCSC Genome Browser. Genome Res. 2007;17:1797–808.
Sylvestre EL, Robert C, Pennetier S, Labrecque R, Gilbert I, Dufort I, et al. Evolutionary conservation of the oocyte transcriptome among vertebrates and its implications for understanding human reproductive function. Mol Hum Reprod. 2013;19:369–79.
Wrenzycki C. In vitro culture systems: how far are we from optimal conditions? Anim Reprod. 2016;13:279–82.
Ferré LB, Kjelland ME, Strøbech LB, Hyttel P, Mermillod P, Ross PJ. Review: Recent advances in bovine in vitro embryo production: reproductive biotechnology history and methods. Animal [Internet]. Elsevier; 2020;14:991–1004. Available from: https://doi.org/10.1017/S1751731119002775
Tomizawa SI, Nowacka-Woszuk J, Kelsey G. DNA methylation establishment during oocyte growth: Mechanisms and significance. Int J Dev Biol. 2012;56:867–75.
Sendžikaitė G, Kelsey G. The role and mechanisms of DNA methylation in the oocyte. Essays Biochem. 2019;63:691–705.
Sawai K. Roles of cell differentiation factors in preimplantation development of domestic animals. J Reprod Dev. 2021;1–27.
Simmet K, Zakhartchenko V, Philippou-Massier J, Blum H, Klymiuk N, Wolf E. OCT4/POU5F1 is required for NANOG expression in bovine blastocysts. PNAS. 2018;115:2770–5.
Jerabek S, Merino F, Schöler HR, Cojocaru V. OCT4: Dynamic DNA binding pioneers stem cell pluripotency. Biochim Biophys Acta [Internet]. Elsevier B.V.; 2014;1839:138–54. Available from: https://doi.org/10.1016/j.bbagrm.2013.10.001
Cao Y. Regulation of germ layer formation by pluripotency factors during embryogenesis. Cell Biosci. 2013;3:1–10.
Milazzotto MP, de Lima CB, da Fonseca AM, dos Santos EC, Ispada J. Erasing gametes to write blastocysts: metabolism as the new player in epigenetic reprogramming. Anim Reprod. 2020;17:1–23.
Messerschmidt DM. Should I stay or should I go: protection and maintenance of DNA methylation at imprinted genes. Epigenetics. 2012;7:969–75.
Zhang J, Zhang S, Wang Y, Cheng H, Hao L, Zhai Y, et al. Effect of TET inhibitor on bovine parthenogenetic embryo development. PLoS One. 2017;12:1–13.
Ross PJ, Sampaio RV. Epigenetic remodeling in preimplantation embryos: cows are not big mice. Anim Reprod. 2018;15:204–14.
Stower H. Epigenetics: reprogramming with TET. Nat Rev Genet. Nature Publishing Group; 2014;15:66.
Li D, Guo B, Wu H, Tan L, Lu Q. TET family of dioxygenases: crucial roles and underlying mechanisms. Cytogenet Genome Res. 2015;146:171–80.
Howell CY, Bestor TH, Ding F, Latham KE, Mertineit C, Trasler JM, et al. Genomic imprinting disrupted by a maternal effect mutation in the Dnmt1 gene. Cell. 2001;104:829–38.
Branco MR, Oda M, Reik W. Safeguarding parental identity: Dnmt1 maintains imprints during epigenetic reprogramming in early embryogenesis. Genes Dev. 2008;22:1567–71.
Reik W, Dean W, Walter J. Epigenetic reprogramming in mammalian development. Science (80- ). 2001;293:1089–93.
Magee DA, Spillane C, Berkowicz EW, Sikora KM, Machugh DE. Imprinted loci in domestic livestock species as epigenomic targets for artificial selection of complex traits. Anim Genet. 2014;45:25–39.
Lewis A, Reik W. How imprinting centres work. Cytogenet Genome Res. 2006;113:81–9.
Pervjakova N, Kasela S, Morris AP, Kals M, Metspalu A, Lindgren CM, et al. Imprinted genes and imprinting control regions show predominant intermediate methylation in adult somatic tissues. Epigenomics. 2016;8:789–99.
Tian XC. Genomic imprinting in farm animals. Annu Rev Anim Biosci. 2014;2:23–40.
Hori N, Nagai M, Hirayama M, Hirai T, Matsuda K, Hayashi M, et al. Aberrant CpG methylation of the imprinting control region KvDMR1 detected in assisted reproductive technology-produced calves and pathogenesis of large offspring syndrome. Anim Reprod Sci. Elsevier B.V.; 2010;122:303–12.
Chen Z, Hagen DE, Elsik CG, Ji T, Morris CJ, Moon LE, et al. Characterization of global loss of imprinting in fetal overgrowth syndrome induced by assisted reproduction. PNAS. 2015;112:4618–23.
Robbins KM, Chen Z, Wells KD, Rivera RM. Expression of KCNQ1OT1, CDKN1C, H19, and PLAGL1 and the methylation patterns at the KvDMR1 and H19/IGF2 imprinting control regions is conserved between human and bovine. J Biomed Sci. 2012;19:1–10.
Murrell A, Heeson S, Cooper WN, Douglas E, Apostolidou S, Moore GE, et al. An association between variants in the IGF2 gene and Beckwith-Wiedemann syndrome: interaction between genotype and epigenotype. Hum Mol Genet. 2004;13:247–55.
Kim TH, Abdullaev ZK, Smith AD, Ching KA, Loukinov DI, Green RDD, et al. Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell. 2007;128:1231–45.
Nordin M, Bergman D, Halje M, Engström W, Ward A. Epigenetic regulation of the Igf2/H19 gene cluster. Cell Prolif. 2014;47:189–99.
Beatty L, Weksberg R, Sadowski PD. Detailed analysis of the methylation patterns of the KvDMR1 imprinting control region of human chromosome 11. Genomics. 2006;87:46–56.
Wang M, Li D, Zhang M, Yang W, Cui Y, Li S. Methylation of KvDMR1 involved in regulating the imprinting of CDKN1C gene in cattle. Anim Genet. 2015;46:354–60.
Driver AM, Huang W, Kropp J, Peñagaricano F, Khatib H. Knockdown of CDKN1C (p57kip2) and PHLDA2 results in developmental changes in bovine pre-implantation embryos. PLoS One. 2013;8.
Stampone E, Caldarelli I, Zullo A, Bencivenga D, Mancini FP, Della Ragione F, et al. Genetic and epigenetic control of CDKN1C expression: Importance in cell commitment and differentiation, tissue homeostasis and human diseases. Int J Mol Sci. 2018;19:1–24.
O’Neill MJ. The influence of non-coding RNAs on allele-specific gene expression in mammals. Hum Mol Genet. 2005;14:113–20.
Smith LC, Therrien J, Filion F, Bressan F, Meirelles FV. Epigenetic consequences of artificial reproductive technologies to the bovine imprinted genes SNRPN, H19/IGF2 and IGF2R. Front Genet. 2015;5:1–6.
Li Y, Donnelly CG, Rivera RM. Overgrowth syndrome. Vet Clini North Am Food Anim Pr. 2019;35:265–76.
Gomes MV, Huber J, Ferriani RA, Amaral Neto AM, Ramos ES. Abnormal methylation at the KvDMR1 imprinting control region in clinically normal children conceived by assisted reproductive technologies. Mol Hum Reprod. 2009;15:471–7.
Mussa A, Molinatto C, Cerrato F, Palumbo O, Carella M, Baldassarre G, et al. Assisted reproductive techniques and risk of Beckwith-Wiedemann syndrome. Pediatrics. 2017;140.
Hattori H, Hiura H, Kitamura A, Miyauchi N, Kobayashi N, Takahashi S, et al. Association of four imprinting disorders and ART. Clin Epigenetics Clin Epigenetics. 2019;11:1–12.
Leibfried L, First NL. Characterization of bovine follicular oocytes and their ability to mature in vitro. J Anim Sci. 1979;48:76–86.
Rosenkrans CF, Zeng GQ, McNamara GT, Schoff PK, First NL. Development of bovine embryos in vitro as affected by energy substrates 1. Biol Reprod. 1993;49:459–62.
Verruma CG, Mioranza A, Petta T, Vila RA, Furtado CLM, de Moraes LAB, et al. Partial replacement of fetal bovine serum during in vitro culture decreases phospholipid content in in vitro produced bovine embryos. Rev Bras Reprodução Anim. 2020;44:108–15.
Hafez ES., Hafez B. Reprodução Animal. 7a edição. São Paulo: Manole; 2004.
Verruma CG, Eiras MC, Fernandes A, Vila RA, Libardi C, Furtado M, et al. Folic acid supplementation during oocytes maturation influences in vitro production and gene expression of bovine embryos. Zygote. 2021;29:342–249.
Rios ÁFL, Lemos DC, Fernandes MB, Andrea MV, Gomes MVM, Lôbo RB, et al. Expression of the CTCF gene in bovine oocytes and preimplantation embryos. Genet Mol Biol. 2007;30:1202–5.
Pfaffl MW. A new mathematical for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29:16–21.
Li L, Dahiya R. MethPrimer: designing primers for methylation PCRs. Bioinformatics. 2022;18:1427–31.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Magoč T, Salzberg SL. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics. 2011;27:2957–63.
Krueger F, Andrews SR. Bismark: A flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011;27:1571–2.
Jiang Z, Dong H, Zheng X, Marjani SL, Donovan DM, Chen J, et al. mRNA levels of imprinted genes in bovine in vivo oocytes, embryos and cross species comparisons with humans, mice and pigs. Sci Rep Nat Publ Group. 2015;5:1–10.
Chen Z, Robbins KM, Wells KD, Rivera RM. Large offspring syndrome: a bovine model for the human loss-of-imprinting overgrowth syndrome Beckwith-Wiedemann. Epigenetics. 2013;8:591–601.
Suzuki J, Therrien J, Filion F, Lefebvre R, Goff AK, Perecin F, et al. Loss of methylation at H19 DMD is associated with biallelic expression and reduced development in cattle derived by somatic cell nuclear transfer. Biol Reprod. 2011;84:947–56.
Zhang S, Chen X, Wang F, An X, Tang B, Zhang X, et al. Aberrant DNA methylation reprogramming in bovine SCNT preimplantation embryos. Sci Rep [Internet]. Nature Publishing Group; 2016;6:1–11. Available from: https://doi.org/10.1038/srep30345
O’Doherty AM, Magee DA, O’Shea LC, Forde N, Beltman ME, Mamo S, et al. DNA methylation dynamics at imprinted genes during bovine pre-implantation embryo development. BMC Dev Biol. 2015;15:1–12.
Vargas LN, Caixeta FMC, Dode MAN, Caetano AR, Franco MM. DNA methylation profile of single in vitro matured bovine oocytes. Mol Reprod Dev. 2023;90:227–35.
Urrego R, Rodriguez-Osorio N, Niemann H. Epigenetic disorders and altered gene expression after use of assisted reproductive technologies in domestic cattle. Epigenetics. 2014;9:803–15.
Adona PR, Leal CLV, Biase FH, De Bem TH, Mesquita LG, Meirelles FV, et al. In vitro maturation alters gene expression in bovine oocytes. Zygote. 2016;24:624–33.
Al-Khtib M, Perret A, Khoueiry R, Ibala-Romdhane S, Blachre T, Greze C, et al. Vitrification at the germinal vesicle stage does not affect the methylation profile of H19 and KCNQ1OT1 imprinting centers in human oocytes subsequently matured in vitro. Fertil Steril. 2011;95:1955–60.
Osman E, Franasiak J, Scott R. Oocyte and embryo manipulation and epigenetics. Semin Reprod Med. 2018;36.
Park CH, Kim HS, Lee SG, Lee CK. Methylation status of differentially methylated regions at Igf2/H19 locus in porcine gametes and preimplantation embryos. Genomics [Internet]. Elsevier Inc.; 2009;93:179–86. Available from: https://doi.org/10.1016/j.ygeno.2008.10.002
Fang F, Hodges E, Molaro A, Dean M, Hannon GJ, Smith AD. Genomic landscape of human allele-specific DNA methylation. Proc Natl Acad Sci U S A. 2012;109:7332–7.
Costa NS, Silveira MM, Vargas LN, Caetano AR, Rumpf R, Franco MM. Epigenetic characterization of the H19/IGF2 locus in calf clones placenta. Pesqui Vet Bras. 2020;40:1063–72.
Meirelles FV, Caetano AR, Watanabe YF, Ripamonte P, Carambula SF, Merighe GK, et al. Genome activation and developmental block in bovine embryos. Anim Reprod Sci. 2004;82–83:13–20.
Kawase Y, Moriki A, Minato Y, Hamano S, Matsukawa K, Kono T. Expression by RT-PCR in Bovine Fetuses. J Mamm Ova Res. 2000;17:124–7.
Murrell A, Heeson S, Reik W. Interaction between differentially methylated regions partitions the imprinted genes Igf2 and H19 into parent-specific chromatin loops. Nat Genet. 2004;36:889–93.
Smith FM, Garfield AS, Ward A. Regulation of growth and metabolism by imprinted genes. Cytogenet Genome Res. 2006;113:279–91.
Renfree MB, Suzuki S, Kaneko-Ishino T. The origin and evolution of genomic imprinting and viviparity in mammals. Philos Trans R Soc B Biol Sci. 2013;368:1–11.
Sandovici I, Georgopoulou A, Pérez-García V, Hufnagel A, López-Tello J, Lam BYH, et al. The imprinted Igf2-Igf2r axis is critical for matching placental microvasculature expansion to fetal growth. Dev Cell. 2022;57:63-79.e8.
Brannan CI, Dees EC, Ingram RS, Tilghman SM. The product of H19 gene may function as an RNA. Mol Cel Biol. 1990;10:28–36.
Suzuki J, Therrien J, Filion F, Lefebvre R, Goff AK, Perecin F, et al. Loss of methylation at H19 DMD is associated with biallelic expression and reduced development in cattle derived by somatic cell nuclear transfer 1. Biol Reprod. 2011;84:947–56.
Smits G, Mungall AJ, Griffiths-Jones S, Smith P, Beury D, Matthews L, et al. Conservation of the H19 noncoding RNA and H19-IGF2 imprinting mechanism in therians. Nat Genet. 2008;40:971–6.
Kawahara M, Morita S, Takahashi N, Kono T. Defining contributions of paternally methylated imprinted genes at the Igf2-H19 and Dlk1-Gtl2 domains to mouse placentation by transcriptomic analysis. J Biol Chem. 2009;284:17751–65.
Bouckenheimer J, Assou S, Riquier S, Hou C, Philippe N, Sansac C, et al. Long non-coding RNAs in human early embryonic development and their potential in ART. Hum Reprod Update. 2016;23:19–40.
Mancini-DiNardo D, Steele SJS, Levorse JM, Ingram RS, Tilghman SM. Elongation of the Kcnq1ot1 transcript is required for genomic imprinting of neighboring genes. Genes Dev. 2006;20:1268–82.
Donato M, Hussain T, Rodulfo H, Peters SO, Imumorin IG, Thomas BN. Conservation of repeats at the mammalian KCNQ1OT1-CDKN1C region suggests a role in genomic imprinting. Evol Bioinforma. 2017;13:1–14.
Tycko B, Morison IM. Physiological functions of imprinted genes. J Cell Physiol. 2002;192:245–58.
Wyss P, Song C, Bina M. Along the Bos taurus genome, uncover candidate imprinting control regions. BMC Genomics. 2022;23:1–21.
Geuns E, Hilven P, Van Steirteghem A, Liebaers I, De Rycke M. Methylation analysis of KvDMR1 in human oocytes. J Med Genet. 2007;44:144–7.
Khoueiry R, Ibala-Romdhane S, Al-Khtib M, Blachère T, Lornage J, Guérin JF, et al. Abnormal methylation of KCNQ1OT1 and differential methylation of H19 imprinting control regions in human ICSI embryos. Zygote. 2012;21:129–38.
Abbott GW. Biology of the KCNQ1 Potassium Channel. New J Sci. 2014;2014:1–26.
Mitsuya K, Meguro M, Lee MP, Katoh M, Al E. LIT1, an imprinted antisense RNA in the human KvLQT1 locus identified by screening for differentially expressed transcripts using monochromosomal hybrids. Hum Mol Genet. 1999;8:1209–17.
Suntharalingham JP, Ishida M, Buonocore F, del Valle I, Solanky N, Demetriou C, et al. Analysis of CDKN1C in fetal growth restriction and pregnancy loss. F1000Res. 2019;8:90.
Tunster SJ, Van De Pette M, John RM. Fetal overgrowth in the Cdkn1c mouse model of Beckwith-Wiedemann syndrome. DMM Dis Model Mech. 2011;4:814–21.
Tunster SJ, Van De Pette M, John RM. Impact of genetic background on placental glycogen storage in mice. Placenta. 2012;33:124–7.
Von Meyenn F, Reik W. Forget the parents: epigenetic reprogramming in human germ cells. Cell [Internet]. Elsevier Inc.; 2015;161:1248–51. Available from: https://doi.org/10.1016/j.cell.2015.05.039
Lodde V, Modina SC, Franciosi F, Zuccari E, Tessaro I, Luciano AM. Localization of DNA methyltransferase-1 during oocyte differentiation, in vitro maturation and early embryonic development in cow. Eur J Histochem. 2009;53:199–208.
Uysal F, Akkoyunlu G, Ozturk S. Dynamic expression of DNA methyltransferases (DNMTs) in oocytes and early embryos. Biochimie [Internet]. Elsevier Ltd; 2015;116:103–13. Available from: https://doi.org/10.1016/j.biochi.2015.06.019
Messerschmidt DM, Knowles BB, Solter D. DNA methylation dynamics during epigenetic reprogramming in the germline and preimplantation embryos. Genes Dev. 2014;28:812–28.
Uysal F, Sukur G, Bozdemir N, Cinar O. DNA methyltransferase (Dnmt) silencing causes increased Cdx2 and Nanog levels in surviving embryos. Int J Dev Biol. 2023;67:1–8.
Peat JR, Dean W, Clark SJ, Krueger F, Smallwood SA, Ficz G, et al. Genome-wide bisulfite sequencing in zygotes identifies demethylation targets and maps the contribution of TET3 oxidation. Cell Rep. 2014;9:1990–2000.
Wossidlo M, Nakamura T, Lepikhov K, Marques CJ, Zakhartchenko V, Boiani M, et al. 5-Hydroxymethylcytosine in the mammalian zygote is linked with epigenetic reprogramming. Nat Commun. 2011;2.
Ito S, Dalessio AC, Taranova O V., Hong K, Sowers LC, Zhang Y. Role of tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature [Internet]. Nature Publishing Group; 2010;466:1129–33. Available from: https://doi.org/10.1038/nature09303
Khan DR, Dubé D, Gall L, Peynot N, Ruffini S, Laffont L, et al. Expression of pluripotency master regulators during two key developmental transitions: EGA and early lineage specification in the Bovine Embryo. PLoS One. 2012;7:1–12.
Graf A, Krebs S, Heininen-Brown M, Zakhartchenko V, Blum H, Wolf E. Genome activation in bovine embryos: review of the literature and new insights from RNA sequencing experiments. Anim Reprod Sci [Internet]. Elsevier B.V.; 2014;149:46–58. Available from: https://doi.org/10.1016/j.anireprosci.2014.05.016
Aguila L, Osycka-Salut C, Treulen F, Felmer R. Pluripotent core in bovine embryos: a review. Animals. 2022;12:1–20.
Berg DK, Smith CS, Pearton DJ, Wells DN, Broadhurst R, Donnison M, et al. Trophectoderm lineage determination in cattle. Dev Cell [Internet]. Elsevier Inc.; 2011;20:244–55. Available from: https://doi.org/10.1016/j.devcel.2011.01.003
Carreiro LE, Dos SGS, Luedke FE, Goissis MD. Cell differentiation events in pre-implantation mouse and bovine embryos. Anim Reprod. 2021;18:1–15.
Schulz KN, Harrison MM. Mechanisms regulating zygotic genome activation. Nat Rev Genet. Springer US; 2018;20:221–34.
Sha QQ, Zheng W, Wu YW, Li S, Guo L, Zhang S, et al. Dynamics and clinical relevance of maternal mRNA clearance during the oocyte-to-embryo transition in humans. Nat Commun [Internet]. Springer US; 2020;11. Available from: https://doi.org/10.1038/s41467-020-18680-6
Funding
This study was supported by the Coordenação de Aperfeiçoamento de Pessoal do Nível Superior (CAPES), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, Grant no. 310310/2019–9 and C.L.M.F. no. 437037/2018–5), Fundação de Apoio ao Ensino e Pesquisa e Assistência do Hospital das Clínicas da Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo (FAEPA, Grant no. 455/2019 and no. 42/2021), and Associação Nacional de Criadores e Pesquisadores (ANCP).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10815_2023_3011_MOESM1_ESM.tiff
Supplementary file1 Figure S1. Flowchart representing the experimental design, sample collection and (epi)genetic analysis (TIFF 65543 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Verruma, C.G., Santos, R.S., Marchesi, J.A.P. et al. Dynamic methylation pattern of H19DMR and KvDMR1 in bovine oocytes and preimplantation embryos. J Assist Reprod Genet 41, 333–345 (2024). https://doi.org/10.1007/s10815-023-03011-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-023-03011-7