Abstract
Purpose
To assess whether there is an association between extended in vitro culture based on embryo developmental stage at transfer and pre-malignant gestational trophoblastic disease (GTD) risk of molar pregnancy during assisted reproduction.
Methods
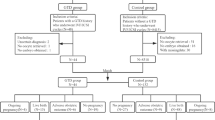
A retrospective study was carried out using Human Fertilization and Embryology Authority (HFEA) anonymized register from 1999 to 2016. A total of 540,376 cycles were eligible to be included in the study after excluding any kind of donor treatment or surrogacy, frozen embryo transfers, and cycles with incomplete data. Subgroup analysis was carried out in subjects with primary infertility aiming to exclude an increased risk in those with a previous GTD. Multivariate logistic regression analysis was used to adjust for possible confounders, and the effect of day of embryo transfer in IVF (in vitro fertilization)/ICSI (intracytoplasmic sperm injection) treatment on a molar pregnancy GTD outcome was analyzed.
Results
The prevalence of a molar pregnancy GTD among the study population was 3.4/10,000 livebirths (53/156,683) with a higher risk in the over 40 age category. No significant difference of pre-malignant GTD incidence was seen between IVF and ICSI (0.01% vs 0.009% respectively). No association was seen with GTD based on type/cause of infertility or number of embryos transferred.
Crude (1.06; 95% CI 0.852–1.31) and adjusted (1.07; 95% CI (0.857–1.32) odds ratios were calculated to see an association between day of embryo transfer and the occurrence of a GTD. There was no association between day of embryo transfer and molar GTD risk after adjusting for age and secondary infertility.
Conclusion
No significant association between pre-malignant molar gestational trophoblastic disease and extended in vitro embryo culture was found after analyzing 540,376 cycles of IVF and ICSI.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gestational trophoblastic disease (GTD) spans a disease spectrum from premalignant conditions such as partial and complete hydatidiform molar pregnancy (commonly referred to as hydatidiform moles) through to malignant conditions such as invasive mole, choriocarcinoma, very rare placental site trophoblastic tumor, and epithelioid trophoblastic tumor. It is a result of abnormal proliferation of the trophoblast with a reported incidence of 1 in 714 live births in the UK [1]. Global prevalence of gestational trophoblastic disease varies vastly with highest-reported incidence of hydatidiform mole being in Southeast Asia with 2 per 1000 pregnancies [2]. The reported incidence is approximately 1–3 in 1000 pregnancies for CHM (complete hydatidiform mole) and 3 in 1000 pregnancies for PHM (partial hydatidiform mole) in North America and Europe [3]. Wide variations in epidemiology of GTD could be due to genetic traits and cultural factors, but accumulating reliable epidemiologic data, such as inconsistencies in case definitions, inability to adequately characterize the population at risk, non-availability of centralized databases, lack of well-chosen control groups against which to compare possible risk factors, and rarity of the diseases, could play a role as well [2].
Complete molar pregnancies are diploid in origin containing androgenic genetic material. Around 80% of the time, complete molar pregnancy (CHM) can arise from impregnation of an inactive oocyte by a haploid sperm with subsequent duplication of paternal genetic material. Less commonly, around 16% of the time, it could occur due to dispermic fertilization of an empty oocyte. In rare occasions, complete moles can also occur by fertilization of a haploid oocyte containing mutated copy of NLRP7 gene or KHDC3L gene by a healthy sperm [4]. It has been reported that 50–80% of recurrent hydatidiform moles in women are due to pathogenic variants in the alleles of NLRP7 or KHDC3L [5]. These molar pregnancies with gene mutations are diploid biparental in origin. Partial moles are triploid in origin, containing two sets of paternal haploid genetic material and one set of maternal haploid genetic material. Partial hydatidiform moles (PHM) usually arise by dispermic fertilization of a haploid oocyte or more rarely by a diploid sperm fertilizing a haploid oocyte. In rare occasions, the PHM have also been reported with other karyotypes (diploid biparental, triploid dyginic, tetraploid triandric) [6].
Several techniques have been described in literature to avoid recurrent molar pregnancy with assisted reproductive technology (ART). These include intracytoplasmic sperm injection (ICSI) to overcome dispermic fertilization, identification of retaining polar bodies during embryo growth, preimplantation genetic testing for aneuploidy (PGT-A), and pre-selection of male embryos to guard against transferring XX embryos created by fertilization of an inactive oocyte by a haploid X-bearing spermatozoon [7]. Despite above technological advances and selection of normally fertilized (2PN) embryos, there are reported cases of pre-malignant GTD following assisted reproduction. It is a known fact that blastocyst transfer compared to cleavage stage transfer is associated with higher pregnancy and live birth rates [8]. However, evidence is lacking whether extended embryo culture to blastocyst stage influences implantation of an abnormal embryo or abnormal trophoblastic proliferation leading to GTD. A previous study done to assess the incidence of molar pregnancy using HFEA data did not address the extended in vitro culture and association with GTD [9].
Although essential determination of the ploidy status is a process that occurs at fertilization, it would be interesting to know how the in vitro culture conditions and morphological selection would affect the risk of molar pregnancy during assisted reproduction. Theoretically, the natural selection of embryos and observation of a more morphologically differentiated stage of embryos might influence the risk of molar GTD during assisted reproduction. Likewise, there is a possibility of alteration in genomic imprinting due to methylation changes post fertilization that can be attributed to the expression of molar pregnancy as well [10].
With this background, the objective of this study was to assess whether extended culture to blastocyst had any effect on the risk of molar pregnancy during IVF and ICSI.
Methods
A retrospective study was carried out using Human Fertilization and Embryology Authority (HFEA) anonymised register data from 1999 to 2016. HFEA holds the longest running register for fertility treatment data in the world and is the national database for fertility treatment data in the UK. The anonymised database is accessible to the researchers and confidentiality of the patients is protected (https://www.hfea.gov.uk/about-us/our-data/).
Women aged 18 to 44 years during treatment were included in the analysis. Cycles with incomplete data where patient’s age at treatment and day of embryo transfer was not available were not included in the analysis. Other exclusion criteria were frozen embryo transfer (FET), donor (egg/sperm/embryo) treatment, and surrogacy. We excluded frozen embryo transfers from our final analysis as we lacked data on extended in vitro culture of these frozen embryos. For frozen embryo transfer cycles, the HFEA-database did not capture whether the embryo thawed was frozen at cleavage stage or blastocyst stage. In addition to the lack of data regarding duration of in vitro culture prior to FET cycle, the age of women at the time of freeze was also not reported in the national database.
Extended culture was defined as embryos cultured in vitro between 5 and 7 days before transfer (blastocyst stage) whilst embryo transfer between 1 and 4 days at cleavage stage was grouped as a standard in vitro culture. Live birth outcomes were recorded to calculate prevalence of GTD. Prevalence of molar pregnancy per 10,000 livebirths and incidence of molar pregnancy per ART cycle were calculated. Incidence of molar pregnancy according to day of embryo transfer was assessed. The terminology used to record pre-malignant GTD outcome on the HFEA database was “molar pregnancy.” A subgroup analysis of those with primary infertility was performed after excluding subjects with secondary infertility in order to exclude a potential bias due to an increased associated risk in those with a previous history of GTD. Multivariate regression analysis was used to adjust for possible confounders and to assess whether there is any association between day of embryo transfer and risk of molar pregnancy. R (4.0.2) statistical software was used for the analysis of data. Crude and adjusted odds ratio (OR) were calculated with 95% confidence intervals (CI).
For unadjusted odds ratios, a logistic regression model with one predictor variable (\({X}_{1}\)) was used with formula given below, where \(p\) is the probability of occurrence of an event (e.g., occurrence of a disease).
The regression coefficient (\({\beta }_{1}\)) above is the estimated increase in the log odds of the outcome per unit increase in the value of the exposure. In other words, the exponential function of the regression coefficient (\({e}^{{\beta }_{1}}\)) is the odds ratio associated with a one-unit increase in the exposure. For adjusted odds ratios, the same logistic regression concept was extended to include multiple variables and was used for the analysis as given below.
where (\({e}^{{\beta }_{1}}\)) becomes the adjusted odds ratio for exposure \({X}_{1}\) whilst keeping other factors \({X}_{2 }, {X}_{3}, \dots\) constant (i.e., adjusted odds ratio after controlling for factors\({X}_{2 }, {X}_{3}, \dots\).).
Results
There were 1,033,588 treatment cycles during the study period resulting in 228,461 livebirths and 78 cases of molar pregnancies reported. This gave an overall GTD prevalence of 4/10,000 live births in all ART cycles performed in the UK. Of the 1,033,588 treatment cycles, only 540,376 cycles met the inclusion criteria of fresh IVF or ICSI resulting in 156,683 live births and 53 cases of molar pregnancies reported. Although FET cycles were excluded from the final analysis, there were 13 reported molar pregnancies in a total of 176,208 FET cycles in women during the same period who had a successful thawed embryo transfer. The prevalence was 3.3 per 10,000 live birth occurrences (13/39,682) following a FET cycle.
The prevalence of molar pregnancy among the study population after meeting inclusion criteria was 3.4/10,000 livebirths (53/156,683) or 1 in every 2956 live births. The incidence was ~ 1 in 10,000 ART cycles when considering all fresh IVF and ICSI cycle outcomes (53/540,376).
Figure 1 shows the age distribution of the study sample along with the number of molar pregnancies reported in each age category. Majority of the patients were in 18–34 category. The prevalence of molar pregnancy was significantly higher in the over 40 age category at 11.6 per 10,000 live births compared to under 35 years (2.6 per 10,000 live births) with odds ratio of 4.0 (95% CI: 1.14–14.19). The prevalence of molar pregnancy was not significant for 35–37 and 36–39 age subgroups at 3.02 and 3.27 per 10,000 live births respectively (95% CI: 0.20–4.95) in comparison to the reference group of under 35 years.
Age distribution of study population and distribution of molar pregnancies of women who had fresh IVF and ICSI cycles in the UK from 1999 to 2016 based on Human Fertilization and Embryology Authority (HFEA) national data. The x-axis denotes the female age range whilst the primary y-axis denotes the total numbers of IVF/ICSI cycles and live births. The secondary y-axis denotes the number of molar pregnancies and the incidence of molar pregnancy per 10,000 live births
Table 1 gives demographic and clinical characteristics of the study sample with odds ratio for each factor given for the incidence of molar pregnancy risk per ART cycle.
Majority of women undergoing IVF/ICSI within the study population had cleavage stage transfer. A total of 24.8% of fresh IVF or ICSI cycles were undertaken following extended culture to day 5–7 (n = 134,299). Table 2 shows the distribution of study population according to day of embryo transfer.
No significant difference of molar pregnancy incidence was seen between IVF (25/247,903) and ICSI (28/289,774) (0.01% vs 0.009% respectively per ART cycle). Out of the 28 molar pregnancies reported among ICSI population, 20 were associated with male factor infertility. No association was identified between GTD and type/cause of infertility or number of embryos transferred.
Crude (1.06; 95% CI 0.852–1.31) and adjusted (1.07; 95% CI 0.857–1.32) odds ratios were calculated to assess an association of molar GTD with extended culture. This did not show any association between day of embryo transfer and GTD risk after adjusting for age.
Subgroup analysis was carried out using subjects with primary infertility in order to exclude a previous history of GTD. There were 16 molar pregnancies among 143,557 fresh ART cycles in women with primary infertility. This also showed no correlation between extended culture of embryos and molar GTD risk (OR: 1.21; 95% CI 0.6–1.92).
Discussion
Assisted reproductive technology (ART) is a helpful tool to study early human embryo development prior to implantation. The advantage of morphological assessment of early embryos at fertilization check may aid in embryo selection by exclusion of 3PN embryos for transfer thereby minimizing the risk of GTD. Our study based on HFEA’s large national ART registry in the UK confirms that the incidence of molar pregnancy is substantially lower with fresh IVF and ICSI compared to general population [1]. Similar observations were made by a previously published study by Nickkho-Amiry et al. [9] with an incidence of molar pregnancy with fresh ICSI at 1/3709–1/4302 and an incidence of molar pregnancy with fresh IVF cycles at 1/4333. However, they did not specifically address the impact of extended in vitro culture conditions to blastocyst stage on GTD risk. In comparison to their reporting of the incidence of molar pregnancy per number of pregnancies regardless of cleavage or blastocyst stage transfer, our calculations were based on fresh ART cycle and live birth outcomes based on extended in vitro culture of the embryos. In their study, they have included fresh and frozen embryo transfers and also donor treatments as well. We excluded frozen embryo transfers from our analysis as the HFEA database was restricted in reporting of stage at which embryos were frozen and the age of women at the time of freezing that may influence the risk of molar pregnancy. In addition, we wished to mitigate the unknown influence of freezing and thawing on genomic imprinting and abnormal trophoblastic proliferation. Donor treatments were excluded to minimize variables as this does not reflect the GTD risk factors specific to the couples. With the use of intracytoplasmic sperm injection, albeit one would expect a lower prevalence of molar pregnancy than IVF, such difference was not seen between the two fertilization methods. ICSI technique invariably protects against dispermy; however, it might not be able to prevent injection of diploid sperm or other rare mechanisms of molar pregnancy formation [11]. In healthy men, diploid sperm account for about 0.2–0.3%, but it may be as high as 1.9% in males with oligozoospermia [12]. As ICSI is usually performed for male factor problem with sub optimal sperm quality, it is plausible that the higher possibility of diploid sperm errors in these men could be one possible explanation of GTD with its use. In this retrospective cohort, 20 out of 28 molar pregnancies seen with ICSI were associated with male factor infertility. Another possible explanation for molar pregnancy following ICSI is when an oocyte with mutations in the maternal-effect genes NLRP7 and KHDC3L is injected with a healthy sperm. These are termed biparental complete hydatidiform mole. Although there is a maternal copy of genome, this is not expressed due to widespread loss of methylation [10]. Although, diploid biparental allelic pathogenic variants in NLRP7 and KHDC3L are linked to familial or recurrent reproductive losses due to molar pregnancies, a study of Danish women did not identify a pathogenic association [5, 13].
There is a possibility that the embryos considered to be diploid had a third pronucleus (PN) which was not visible at the specific time of morphological assessment of embryos. The possibility of human errors identifying 3PN embryos should also be borne in mind in this situation. Triploid zygotes have also been observed when rescue ICSI is used following failed fertilization with IVF [9]; however, this technique is not practiced in the UK. Theoretically, there is a possibility that timelapse imaging could be useful in reducing the human errors and inter observer variations in this situation; however, we did not come across any studies assessing the use of time lapse imaging to prevent molar pregnancies. A study done by Grossmann et al. suggested that most of the tripronucleate zygotes after ICSI were due to non-extrusion of the second polar body [14]. Maternal triploidy is commonly not shown to be associated with GTD [15]. The selection of euploid embryos by preimplantation genetic testing (PGT) may be considered beneficial in ruling out abnormally fertilized embryos based on ploidy status. However, molar pregnancy has been reported following transfer of a euploid blastocyst after PGT [16].
Even though the manifestation of a molar pregnancy is determined at the fertilization based on uniparental ploidy status, uncertainty exists regarding extended in vitro culture to blastocyst stage and correlation with implantation of an abnormal embryo or an abnormal trophoblastic proliferation leading to GTD. There has been concerns about the occurrence of epimutations with assisted reproductive techniques [10]. Studies have reported alteration in early placental methylation [17, 18] as well as differential genomic and transcriptomic expression [19] in placental tissue in ART conceptions compared to natural conceptions thereby raising the question of predisposition to GTD-risk. Ploidy status to diagnose molar pregnancies using traditional diagnostic tools such as ultrasound, histopathology, and cytogenetics may underdiagnose true prevalence that can be picked up by genomic sequencing. A large retrospective cohort study of 26,101 products of conception analysis using newer genomic technology, single nucleotide polymorphism (SNP) array identified a significantly higher detection of GTD in 3.1% of all reproductive losses [20]. According to this study, approximately 71% of partial molar pregnancies and 30% of complete moles were missed by traditional standard techniques.
As far as we know, no studies have been done to assess an effect of culture condition on trophoblastic proliferation and risk of subsequent gestational trophoblastic disease. Our findings concluded that the selection of blastocyst stage embryo after extended culture did not alter the likelihood of having a pre-malignant GTD compared to cleavage stage embryo.
Demographic data showed a higher prevalence of molar pregnancy per 10,000 livebirths in above 40 age category, in par with known higher risk of molar pregnancy in women above 40 in the general population. Of interesting note, although the incidence of molar pregnancy was significantly lower in women undergoing assisted reproduction, this may suggest increased risk with advancing age is not totally eliminated with treatment.
The retrospective analysis of anonymised HFEA database limited adjustments for confounders not recorded in the national database such as smoking, previous history of GTD, and ethnicity that predispose to GTD risk. This is a key limitation of our study. It is known that a previous history of GTD could be a risk factor for future occurrence. A recent retrospective study carried out in patients undergoing IVF/ICSI with a prior history of GTD has shown a reduced number of good quality embryos compared to secondary infertile women without a prior history of GTD [21]. This study consequentially reported a significant association of low live birth rate of 34% in those with previous history of GTD compared to control cohort (66%) post IVF/ICSI cycles. We tried to overcome this possible confounder by performing a subgroup analysis for women with primary infertility and no previous reported pregnancies thereby excluding possibility of previous GTD. Another drawback of our study was that the type of culture media used and morphological grading of the embryos was not known before the transfer. Lastly, caution needs to be exercised for under-reporting of GTD to HFEA, and the lack of information on the type of pre-malignant GTD identified as HFEA dataset does not differentiate between partial and complete molar pregnancies. Our study focussed on association of extended in vitro culture and GTD risk by analyzing the anonymized national UK database. Unfortunately, the retrospective data restricts access to information not recorded in the database.
Wider implications of the findings
Though GTD cannot be completely prevented by IVF/ICSI, the incidence is significantly low and extended culture is not associated with higher risk of abnormal trophoblastic proliferation or molar GTD occurrence with IVF/ ICSI treatment. These findings would aid informed implication counselling and reassurance of patients during assisted reproduction treatments. Future research is recommended to determine whether women at high risk for molar pregnancies or its recurrence would benefit from ART to minimize risk of GTD. In the UK, there are three regional GTD referral centers where all GTD cases are registered. It would be advisable to record in there and similar databases globally about ART conceptions and GTD occurrence for future studies. Lastly, with the genomic era, it is vital that the future of GTD risk prediction, prevention, and education utilizes new tools of reproductive genomics and artificial intelligence.
Data availability
The data underlying this article were accessed from HFEA anonymised register [https://www.hfea.gov.uk/about-us/our-data/]. The derived data generated in this research will be shared on reasonable request to the corresponding author.
References
The management of gestational trophoblastic disease. Green top guideline number 38. Royal College of Obstetricians. 2020. https://www.rcog.org.uk/en/guidelines-research-services/guidelines/gtg38. Accessed 13 June 2021.
Lurain JR. Gestational trophoblastic disease I: epidemiology, pathology, clinical presentation and diagnosis of gestational trophoblastic disease, and management of hydatidiform mole. Am J Obstet Gynecol. 2010. https://doi.org/10.1016/j.ajog.2010.06.073.
Stevens FT, Katzorke N, Tempfer C, Kreimer U, Bizjak GI, Fleisch MC, Fehm TN. Gestational trophoblastic disorders: an update in 2015. Geburtshilfe Frauenheilkd. 2015. https://doi.org/10.1055/s-0035-1558054.
Candelier JJ. The hydatidiform mole. Cell Adhes Migr. 2016; https://doi.org/10.1080/19336918.2015.1093275
Nguyen NMP, Khawajkie Y, Mechtouf N, et al. The genetics of recurrent hydatidiform moles: new insights and lessons from a comprehensive analysis of 113 patients. Mod Pathol. 2018. https://doi.org/10.1038/s41379-018-0031-9.
Slim R, Wallace EP. NLRP7 and the genetics of hydatidiform moles: recent advances and new challenges. Front Immunol. 2013; https://doi.org/10.3389/fimmu.2013.00242
Reubinoff BE, Lewin A, Verner M, Safran A, Schenker JG, Abeliovich D. Intracytoplasmic sperm injection combined with preimplantation genetic diagnosis for the prevention of recurrent gestational trophoblastic disease. Hum Reprod. 1997. https://doi.org/10.1093/humrep/12.4.805.
Glujovsky D, Farquhar C, Quinteiro Retamar A, Alvarez Sedo C, Blake D. Cleavage stage versus blastocyst stage embryo transfer in assisted reproductive technology. Cochrane Database Syst Rev. 2016. https://doi.org/10.1002/14651858.CD002118.pub5.
Nickkho-Amiry M, Horne G, Akhtar M, Mathur R, Brison DR. Hydatidiform molar pregnancy following assisted reproduction. J Assist Reprod Genet. 2019. https://doi.org/10.1007/s10815-018-1389-9.
Anvar Z, Chakchouk I, Demond H, Sharif M, Kelsey G, Van den Veyver IB. DNA methylation dynamics in the female germline and maternal-effect mutations that disrupt genomic imprinting. Genes. 2021. https://doi.org/10.3390/genes12081214.
Savage P, Sebire N, Dalton T, et al. Partial molar pregnancy after intracytoplasmic sperm injection occurring as a result of diploid sperm usage. J Assist Reprod Genet. 2013. https://doi.org/10.1007/s10815-013-0002-5.
Rosenbusch BE. Mechanisms giving rise to triploid zygotes during assisted reproduction. Fertil Steril. 2008. https://doi.org/10.1016/j.fertnstert.2007.06.031.
Andreasen L, Christiansen OB, Niemann I, Bolund L, Sunde L. NLRP7 or KHDC3L genes and the etiology of molar pregnancies and recurrent miscarriage. Mol Hum Reprod. 2013. https://doi.org/10.1093/molehr/gat056.
Grossmann M, Calafell JM, Brandy N, Vanrell JA, Rubio C, Pellicer A, Egozcue J, Vidal F, Santalo J. Origin of tripronucleate zygotes after intracytoplasmic sperm injection. Hum Reprod. 1997. https://doi.org/10.1093/humrep/12.12.2762.
Zaragoza MV, Surti U, Redline RW, Millie E, Chakravarti A, Hassold TJ. Parental origin and phenotype of triploidy in spontaneous abortions: predominance of diandry and association with the partial hydatidiform mole. Am J Hum Genet. 2000. https://doi.org/10.1086/302951.
Zhou B, Anglin HP, Quaas AM. Molar pregnancy after in vitro fertilization with euploid single embryo transfer. F&S reports. 2021; https://doi.org/10.1016/j.xfre.2021.01.003
Chi F, Zhao M, Li K, Lin AQ, Li Y, Teng X. DNA methylation status of imprinted H19 and KvDMR1 genes in human placentas after conception using assisted reproductive technology. Ann Transl Med. 2020; https://doi.org/10.21037/atm-20-3364.
Savage P, Monk D, Hernandez Mora JR, van der Westhuizen N, Rauw J, Tinker A, Robinson W, Song Q, Seckl MJ, Fisher RA. A case of intraplacental gestational choriocarcinoma; characterised by the methylation pattern of the early placenta and an absence of driver mutations. BMC Cancer. 2019. https://doi.org/10.1186/s12885-019-5906-8.
Yang S, Zheng W, Yang C, Zu R, Ran S, Wu H, Mu M, Sun S, Zhang N, Thorne RF, Guan Y. Integrated analysis of hub genes and MicroRNAs in human placental tissues from in vitro fertilization-embryo transfer. Front Endocrinol (Lausanne). 2021. https://doi.org/10.3389/fendo.2021.774997.
Maisenbacher M, Merrion K, Kutteh W. Single nucleotide polymorphism (SNP) microarray detects molar pregnancies in 3% of miscarriages. Fertil Steril. 2019. https://doi.org/10.1016/j.fertnstert.2019.06.015.
Cai X, Zhang M, Huang C, Jiang Y, Zhou J, Xu M, Yan G, Sun KN. Association between gestational trophoblastic disease (GTD) history and clinical outcomes in in vitro fertilization/intracytoplasmic sperm injection (IVF/ICSI) cycles. Reprod Biol Endocrinol. 2022. https://doi.org/10.1186/s12958-022-00898-2.
Author information
Authors and Affiliations
Contributions
MC contributed to the study conception, design, data interpretation, supervision, and critical revision of the manuscript. Material preparation, data collection, and analysis were performed by IB and RB contributed to data interpretation and analysis. The first draft of the manuscript was written by IB and all authors commented on previous versions of the manuscript. PM contributed to analysis of frozen embryo transfer data for revised version. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The data output in this article utilized publically available data from the Human Fertilization and Embryology Authority (HFEA) national data registry. Since an anonymized version of the HFEA database is freely available, no ethical approval was required for this study. The Health Research Authority tool http://www.hra-decisiontools.org.uk/research/ confirmed this was not considered research by the National Health Service in the UK and hence was exempt from requiring ethics approval.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bambaranda, B.G.I.K., Bomiriya, R., Mehlawat, P. et al. Association of extended culture to blastocyst and pre-malignant gestational trophoblastic disease risk following IVF/ICSI-assisted reproduction cycles: an analysis of large UK national database. J Assist Reprod Genet 39, 2317–2323 (2022). https://doi.org/10.1007/s10815-022-02583-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-022-02583-0