Abstract
Purpose
In this study, we examined the correlation between pronucleus size and the potential for human single pronucleus (1PN) zygotes to develop into blastocysts after IVF and ICSI.
Methods
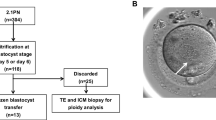
This study included 112 patients who underwent a total of 112 cycles of IVF/ICSI. To evaluate embryo development, 1PN zygotes were compared with 2PN zygotes in the same IVF/ICSI cycle (control cycles) using time-lapse live embryo imaging. To assess the potential for blastocyst formation, cutoff values for pronuclear area and diameter were established through receiver operating characteristic curve analysis, after which 1PN zygotes were classified based on those cutoff values.
Results
Among 1PN zygotes cultured to day 5/6, the rate of embryo development was significantly lower than from 2PN zygotes. However, the rates of blastocyst formation and good quality blastocysts from 1PN zygotes with large pronuclear areas (≥ 710 μm2) or diameters (≥ 31 μm) were significantly higher than from 1PN zygotes with smaller pronuclear areas (≤ 509, 510–609, and 610–709 μm2) or diameters (≤ 24, 25–27,and 28–30 μm) (P < 0.01). Moreover, the results for 1PN zygotes with large pronuclei were similar to those for 2PN zygotes.
Conclusions
The developmental potential of 1PN zygotes with large pronuclear areas (≥ 710 μm2) or diameters (31 μm) appears to be similar to that of 2PN zygotes, and measurement of pronuclear area or diameter in 1PN zygotes is a simple, potentially useful, clinical method.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There have been a number of reports in which normal healthy babies were born after transfer of abnormal single pronucleus (1PN) embryos or blastocysts in human-assisted reproductive technology [1,2,3,4,5,6]. We previously reported the births of nine normal healthy babies without malformation from 1PN zygotes cultured until the blastocyst stage [5]. Moreover, Bradley et al. reported that after transfer of 20 IVF and six ICSI blastocysts derived from 1PN zygotes, six babies with no detectable abnormalities and three ongoing pregnancies were achieved [6]. However, there is no established protocol for safely transferring abnormal 1PN embryos, which makes it crucial to select the normal 1PN embryos.
In a recent investigation of the chromosomal constitution of 1PN embryos using preimplantational genetic screening (PGS), Mateo et al. found that the rate of euploidy among 1PN2PB ICSI zygotes was 17% (15/88) after blastomere biopsy on day 3,while the rate of euploid blastocysts among 1PN2PB ICSI zygotes was only 2.3% (2/88), and the transferred euploid blastocysts developed into healthy babies [7]. Another PGS study performed by Bradley et al. revealed adjusted abnormality rates of about 40% in 1PN blastocysts derived from IVF or ICSI [6]. These results show that 1PN embryos often develop into blastocysts with normal chromosomal compositions. The chromosomal status and genetic composition in 1PN zygotes and embryos have also been analyzed using fluorescence in situ hybridization (FISH) [8,9,10] or immunostaining [11]. The rate of diploid chromosomal constitution among IVF 1PN embryos detected using FISH was significantly higher than in ICSI [8,9,10]. Similarly, the rate of 1PN zygotes with both the maternal and paternal genomes was significantly higher in IVF (86.7%) than ICSI (30.3%) based on histone modification detected by immunostaining [11]. Otsu et al. measured pronuclear diameters and found that whereas the rate of blastocyst formation among large 1PN zygotes (29–34 μm) was 18%, small 1PN zygotes (23–26 μm) did not develop into blastocysts [12]. They suggested that the large size of their pronuclei made it likely that the large 1PN zygotes contained both parental genomes. In that report, however, little embryological data for 1PN embryos was presented.
A non-invasive time-lapse live embryo imaging system was recently developed for prediction of blastocyst formation and selection of high-quality embryos before transfer. This system enables evaluation of pronucleus formation and embryonic development without altering the culture environment [13]. In the present study, we retrospectively examined time-lapse images to determine whether one can more effectively select 1PN embryos with adequate potential to develop into blastocysts based on measurements of pronuclear area or diameter in 1PN zygotes after IVF/ICSI. In addition, to evaluate the developmental potential of 1PN zygotes, we compared the rates of blastocyst formation and good quality blastocysts based on area and diameter of the pronucleus in 1PN zygotes.
Materials and methods
Patients
This retrospective cohort study included 112 patients who underwent 112 IVF/ICSI treatment cycles at two IVF centers (Kishokai Medical Corporation, Aichi Prefecture, Japan) between January 2016 and August 2017. Cycles in which at least one 1PN embryo was observed were compared to 2PN embryos in the same IVF or ICSI cycles.
Ovarian stimulation, fertilization, embryo culture, and evaluation
Ovarian stimulation, fertilization, embryo culture, and evaluation have been previously described by Itoi et al. [5, 14]. Briefly, the ovarian stimulation protocols were performed using a gonadotrophin-releasing hormone (GnRH) antagonist (Cetrotide; Merck Serono, Tokyo, Japan) or GnRH agonist (buserelin acetate; Sprecure, Mochida Pharmaceutical) in a short protocol with follicle-stimulating hormone (FSH; Merck Serono) and/or human menopausal gonadotrophin (hMG; Ferring Pharmaceuticals, Tokyo, Japan) and clomiphene citrate (Merck Serono). Human chorionic gonadotrophin (Mochida Pharmaceutical, Tokyo, Japan) or GnRH agonist was administered when the leading follicles averaged 18 mm in diameter. Oocyte retrieval was performed 35 h after HCG administration or 34 h after the GnRH agonist dose. Oocytes were fertilized using IVF or ICSI techniques within 3–6 h after retrieval. Fertilization was assessed 18–19 h after IVF or ICSI by two embryologists using a time-lapse live embryo imaging system. They monitored captured images until the pronuclear stage along the time and z-axes. At that time, pronuclear zygotes that were indistinct on the time-lapse imaging were confirmed using an inverted microscope. The 2PN and 1PN zygotes were divided and cultured until day 5/6. Embryo quality was evaluated on day 3 using the grading system of Veeck [15]. Briefly, good quality embryos were defined on day 3 as being seven to nine cells, < 10% fragmentation, and symmetric blastomeres. Blastocysts were evaluated using the grading system of Schoolcraft et al. [16]. Briefly, blastocyst formation was defined as development past the early blastocyst. Good quality blastocysts were defined based on the degree of expansion (full, expanded, hatching, and hatched blastocyst), the morphology of the inner cell mass, and the trophectoderm (A and B).
Time-lapse live embryo imaging
We observed 1PN zygotes after IVF or ICSI using a time-lapse live embryo imaging system (Primo Vision; Vitrolife, Tokyo, Japan) as described previously [14]. Briefly, Metaphase II (MII) oocytes that were denuded of cumulus cells 4–5 h after IVF were transferred to 100 or 150 μl drops of Universal IVF Medium in a microwell dish (Vitrolife). Alternatively, MII oocytes were immediately transferred to a microwell dish following ICSI. The microwell dish was then set on the Primo Vision system, and light images were captured at 10-min intervals until the pronuclear stage.
The measurement of pronucleus size
Pronuclear areas and diameters were measured using Primo Vision software 18–19 h after IVF or ICSI. Thereafter, cutoff values for blastocyst formation and good quality blastocysts based on pronuclear area and diameter in 1PN zygotes were determined based on receiver operating characteristic (ROC) curves calculated using EZR [17].
Statistical analysis
The χ2 test was used to evaluate the significance of differences between proportions (%). Pronuclear sizes were compared using the Student’s t tests. Values of P < 0.05 were considered significant.
Results
As shown in Table 1, the rate of good quality embryos on day 3 among 1PN zygotes (18.5%) was significantly lower than among 2PN zygotes (38.5%; P < 0.01). The rates of blastocyst formation and good quality blastocysts among 1PN zygotes (32.2 and 11.6%, respectively) were also significantly lower than among 2PN zygotes (68.3 and 38.5%, respectively; P < 0.01).
The ROC curve in Fig. 1 shows the cutoff value and AUC for blastocyst formation from 1PN zygotes were 708 μm2 and 0.662 for pronuclear area and 31 μm and 0.661 for pronuclear diameter. For good quality blastocysts from 1PN zygotes, the cutoff value and AUC were 713 μm2 and 0.848 for pronuclear area and 31 μm and 0.827 for pronuclear diameter. In subsequent analyses, the cutoff values were therefore set at 710 μm2 for pronuclear area and 31 μm for pronuclear diameter.
The 1PN zygotes were then divided into the following four groups each based pronuclear areas and diameters: ≤ 509, 510–609, 610–709, and ≥ 710 μm2; or ≤ 24, 25–27, 28–30, and ≥ 31 μm. As shown in Table 2, there was a significant correlation between the pronuclear area in 1PN zygotes and embryological outcomes. The proportion of good quality embryos on day 3 did not significantly differ among the four groups. On the other hand, when 1PN zygotes were cultured into blastocysts, the rates of blastocyst formation and good quality blastocysts among embryos with pronuclear areas ≥ 710 (56.1 and 36.6%) were significantly higher than in the groups with smaller pronuclear areas (17.9–24.3 and 0.0–5.4%, respectively; P < 0.05). Moreover, the results obtained with 1PN embryos with pronuclear areas ≥ 710 were similar to the results with 2PN embryos. Notably, the pronuclear area of ≥ 710 μm2 was significantly larger than average for 1PN embryos and also larger than the pronuclei in 2PN embryos (P < 0.05) (Fig. 2).
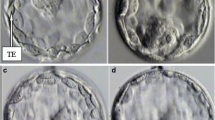
a 1PN and 2PN zygotes at the pronuclear stage. (a) 1PN zygote with a large pronucleus, (b) 1PN zygote with a small pronucleus. (c) 2PN zygote. The scale bar represents 50 μm. b Average pronuclear areas in 1PN and 2PN zygotes. In 2PN zygotes, gray and light gray indicate the average of the small PN and large PN, respectively
We next assessed the correlation between pronuclear diameter in 1PN embryos and embryological outcomes (Table 3). The blastocyst formation rate among 1PN embryos with pronuclear diameters ≥ 31 (60.0%) was significantly higher than among embryos with smaller pronuclear diameters (16.0–26.5%; P < 0.05). Moreover, the rate of good quality blastocysts increased with increases in the diameter of the pronucleus (≤ 24, 0.0%; 25–27, 2.0%; 28–30,10.8 and ≥ 31, 34.3%).
We also examined the rates of blastocyst formation and good quality blastocysts among large 1PN embryos (≥ 710 μm2 and ≥ 31 μm). With respect to the pronuclear area, the rates of blastocyst formation and good quality blastocysts were respectively 59.4 (19/32) and 40.6% (13/32) with IVF and 44.4 (4/9) and 22.2% (2/9) with ICSI. With respect to the pronuclear diameter, the rates of blastocyst formation and good quality blastocysts were respectively 60.7 (17/28) and 35.7% (10/28) with IVF and 57.1 (4/7) and 28.6% (2/7) with ICSI. There were no significant differences in pronuclear area or diameter between IVF and ICSI.
Discussion
Using ROC curve analysis, we have established pronuclear area and diameter cutoff values for the prediction of blastocyst formation and development of good quality blastocysts from 1PN zygotes. We found that 1PN zygotes with large pronuclear areas or diameters (≥ 710 μm2 or ≥ 31 μm, respectively) were more likely to develop into good quality blastocysts than those with smaller pronuclear areas or diameters. We suggest it is possible to select the 1PN zygotes with high potential for development into blastocysts based on pronuclear area or diameter.
Although reports have described live births from 1PN embryos, the safety of transferring 1PN embryos has not yet been established [1,2,3,4,5,6]. It is therefore necessary to establish a method for selecting 1PN embryos with a complete parental genome. Genetic screening is now widely used to investigate the chromosomal constitution of embryos and blastocysts. Capalbo et al. (2017) analyzed 1PN blastocysts in which ploidy assessment was possible and reported that 69.2% of the 1PN blastocysts were diploid [18]. On the other hand, Otsu et al. reported that the rate of blastocyst formation from large 1PN zygotes (29–34 μm) was higher than from small 1PN zygotes (23–26 μm) and that FISH showed blastocysts derived from large 1PN zygotes were more than likely diploid [12]. However, that report provided little embryological data on the 1PN embryos, did not use time-lapse live embryo imaging, and did not compare the rates of blastocyst formation from 1PN and 2PN zygotes; they only measured pronuclear diameter.
In the present study, the developmental potential of 1PN zygotes was significantly lower than 2PN zygotes (Table 1), which is consistent with earlier reports [5, 19]. This study also revealed a correlation between pronuclear size in 1PN zygotes and the rate of embryological developmental to blastocysts, which is consistent with an earlier study [12]. Our results further show that the rates of both development to blastocysts and formation of good quality blastocysts from 1PN zygotes with large pronuclear areas (≥ 710 μm2) and diameters (≥ 31 μm) were significantly higher than from 1PN zygotes with smaller pronuclear areas and diameters. Moreover, the developmental potential of 1PN embryos with large pronuclei was similar to that of 2PN embryos. Specifically, these results show that it is highly likely that 1PN zygotes with large pronuclear areas (≥ 710 μm2) or diameters (≥ 31 μm) include both the female and the male genomes, regardless of insemination method. In earlier studies, it was shown that formation of a 1PN zygote can involve enclosure of juxtaposed male and female nuclei by a common pronuclear envelope [20] and that the male chromatin is fused in the vicinity of the female chromatin before nuclear envelope formation [21]. The pronuclear areas and diameters in such 1PN zygotes were not known, however. It was thought that the rate of embryo development from 1PN zygotes with small pronuclear areas and diameters was due to parthenogenesis and failure of decondensation of the sperm head [22]. We therefore suggest that pronuclear areas or diameters of all 1PN zygotes should be measured and the zygotes cultured to the blastocyst stage.
In this study, we did not perform genetic analyses. Although the developmental potential of large 1PN zygotes appears similar to that of 2PN zygotes, one cannot be certain whether these embryos are euploid without performing PGS. We therefore also suggest that, if possible, PGS should be performed with 1PN blastocysts and that further studies are needed to determine whether the size of the PN is associated with aneuploidy.
Because there was no significant difference in the ROC-AUC cutoff for pronuclear area and diameter in 1PN zygotes, we suggested that measurement of either is equally effective, though measurement of pronuclear diameters may be simpler for clinical use.
In summary, we have shown that the developmental potential of 1PN zygotes with large pronuclear areas (≥ 710 μm2) or diameters (≥ 31 μm) is similar to that of 2PN zygotes. However, these findings must be confirmed through examination of additional 1PN zygotes to further assess the relation between pronuclear size and clinical outcomes.
References
Staessen C, Janssenswillen C, Devroey P, Van Steirteghem AC. Cytogenetic and morphological observations of single pronucleated human oocytes after in-vitro fertilization. Hum Reprod. 1993;8:221–3.
Reichman DE, Jackson KV, Racowsky C. Incidence and development of zygotes exhibiting abnormal pronuclear disposition after identification of two pronuclei at the fertilization check. Fertil Steril. 2010;94:965–70.
Gras L, Trounson AO. Pregnancy and birth resulting from transfer of a blastocyst observed to have one pronucleus at the time of examination for fertilization. Hum Reprod. 1999;14:1869–71.
Dasig D, Lyon J, Behr B, Milki AA. Monozygotic twin birth after the transfer of a cleavage stage embryo resulting from a single pronucleated oocyte. J Assist Reprod Genet. 2004;21:427–9.
Itoi F, Asano Y, Shimizu M, Honnma H, Murata Y. Birth of nine normal healthy babies following transfer of blastocysts derived from human single-pronucleate zygotes. J Assist Reprod Genet. 2015;32:1401–7.
Bradley CK, Traversa MV, Hobson N, Gee AJ, McArthur SJ. Clinical use of monopronucleated zygotes following blastocyst culture and preimplantation genetic screening, including verification of biparental chromosome inheritance. Reprod BioMed Online. 2017;34:567–74.
Mateo S, Vidal F, Parriego M, Rodríguez I, Montalvo V, Veiga A, et al. Could monopronucleated ICSI zygotes be considered for transfer? Analysis through time-lapse monitoring and PGS. J Assist Reprod Genet. 2017;34:905–11.
Staessen C, Van Steirteghem AC. The chromosomal constitution of embryos developing from abnormally fertilized oocytes after intracytoplasmic sperm injection and conventional in-vitro fertilization. Hum Reprod. 1997;12:321–7.
Sultan KM, Munne S, Palermo GD, Alikani M, Cohen J. Chromosomal status of uni-pronuclear human zygotes following in-vitro fertilization and intracytoplasmic sperm injection. Hum Reprod. 1995;10:132–6.
Yan J, Li Y, Shi Y, Feng HL, Gao S, Chen ZJ. Assessment of sex chromosomes of human embryos arising from monopronucleus zygotes in in vitro fertilization and intracytoplasmic sperm injection cycles of Chinese women. Gynecol Obstet Investig. 2010;69:20–3.
van der Heijden GW, van den Berg IM, Baart EB, Derijck AA, Martini E, de Boer P. Parental origin of chromatin in human monopronuclear zygotes revealed by asymmetric histone methylation patterns, differs between IVF and ICSI. Mol Reprod Dev. 2009;76:101–8.
Otsu E, Sato A, Nagaki M, Araki Y, Utsunomiya T. Developmental potential and chromosomal constitution of embryos derived from larger single pronuclei of human zygotes used in in vitro fertilization. Fertil Steril. 2004;81:723–4.
Kaser DJ, Racowsky C. Clinical outcomes following selection of human preimplantation embryos with time-lapse monitoring: a systematic review. Hum Reprod Update. 2014;5:617–31.
Itoi F, Asano Y, Shimizu M, Nagai R, Saitou K, Honnma H, et al. Clinical outcomes after IVF or ICSI using human blastocysts derived from oocytes containing aggregates of smooth endoplasmic reticulum. Reprod BioMed Online. 2017;34:337–44.
Veeck LL. An atlas of human gametes and conceptuses: an illustrated reference for assisted reproductive technology. New York: Parthenon Publishing; 1999.
Schoolcraft WB, Gardner DK, Lane M, Schlenker T, Hamilton F, Meldrum DR. Blastocyst culture and transfer: analysis of results and parameters affecting outcome in two in vitro fertilization programs. Fertil Steril. 1999;72:604–9.
Kanda Y. Investigation of the freely available easy-to-use software ‘EZR’ for medical statistics. Bone Marrow Transplant. 2013;48:452–8.
Capalbo A, Treff N, Cimadomo D, Tao X, Ferrero S, Vaiarelli A, et al. Abnormally fertilized oocytes can result in healthy live births: improved genetic technologies for preimplantation genetic testing can be used to rescue viable embryos in in vitro fertilization cycles. Fertil Steril. 2017;108:1007–15.
Mateo S, Parriego M, Boada M, Vidal F, Coroleu B, Veiga A. In vitro development and chromosome constitution of embryos derived from monopronucleated zygotes after intracytoplasmic sperm injection. Fertil Steril. 2013;99:897–902.
Levron J, Munne S, Willadsen S, Rosenwaks Z, Cohen J. Male and female genomes associated in a single pronucleus in human zygotes. Biol Reprod. 1995;52:653–7.
Krukowska A, Tarkowski AK. Mouse zygotes with one diploid pronucleus formed as a result of ICSI can develop normally beyond birth. Mol Reprod Dev. 2005;72:346–53.
Payne D, Flaherty SP, Barry MF, Matthews CD. Preliminary observations on polar body extrusion and pronuclear formation in human oocytes using time-lapse video cinematography. Hum Reprod. 1997;12:532–41.
Acknowledgements
We thank the embryology staff of the Kishokai Medical Corporation for their assistance in preparing this manuscript and for their expert technical help.
Funding
This work was no external funding.
Author information
Authors and Affiliations
Contributions
EA acquired the data, performed the statistical analysis, and helped to draft the manuscript. FI conceived and designed the study, acquired the data, performed the statistical analysis, and drafted the manuscript. YA acquired the data and helped to draft the manuscript. HH performed the statistical analysis and drafted the manuscript. HO drafted the manuscript. KN participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
This study was approved by the Institutional Review Board of Kishokai Medical Corporation and was conducted in accordance with the principals expressed in the Declaration of Helsinki. All patients provided consent to all treatment procedures and agreed to anonymous use of their data for studies.
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Araki, E., Itoi, F., Honnma, H. et al. Correlation between the pronucleus size and the potential for human single pronucleus zygotes to develop into blastocysts. J Assist Reprod Genet 35, 817–823 (2018). https://doi.org/10.1007/s10815-018-1137-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-018-1137-1