Abstract
As an economically important marine seaweed, Pyropia yezoensis can be asexually reproduced via archeospores. However, the molecular mechanism of archeospore formation remains unclear. In this study, we characterized the transcriptomes of two strains of Py. yezoensis, which are strain W′ with abundant archeospores and strain W with scarce archeospores. The 25- and 35-day-old gametophytic blades of both strains were sampled for RNA sequencing. Based on comparative transcriptomic analysis, a large number of differential expressed genes (DEGs) of four pairwise groups between samples were generated. In an effort to identify the genes related to archeospore formation, 415 consensus DEGs reflecting the difference between two strains were investigated. Gene ontology and KEGG analysis revealed that the function for these DEGs was enriched in various processes, such as translation, ribosome assembly, RNA methylation, vesicles assembly, intracellular localization of organelles, and endocytosis. According to functional analysis on 80 selected DEGs, it indicated that genes involved in calcium and MAPK signaling pathway, cytoskeletal movement, and cell wall degradation were related to the formation of archeospores. The accuracy of RNA-seq results was validated by quantitative real-time PCR of 8 selected DEGs. These findings suggest that the molecular mechanism of archeospore formation in Py. yezoensis is sophisticated and our results will serve as an essential foundation for further studies.
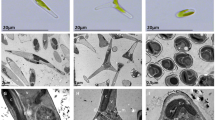
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The red marine alga Pyropia yezoensis is one of the economically important maricultural crops, (Sutherland et al. 2011). This alga has been extensively cultivated in Japan, China, and Korea and has the highest commercial value per unit mass in comparison with other algal cultivars (Kim et al. 2017). Pyropia yezoensis presents several interesting evolutionary, physiological, and ecological characteristics related to its development and reproduction. The life cycle of Py. yezoensis consists of alternation between two heteromorphic stages, the diploid filamentous sporophyte stage and the haploid foliose gametophyte stage. The reproductive ways of Py. yezoensis have been revealed to be versatile. In addition to the classic sexual reproduction, it exhibits asexual reproduction through the production of archeospores from young gametophytic blades (Shimizu et al. 2007; Mikami et al. 2019). Archeospore formation has been observed in several Pyropia species, such as Py. tenera, Py. suborbiculata, and Py. chauhanii (Tseng and Chang 1954; Song and Yan 2015; Yang et al. 2019).
During the process of archeospore formation the color and shape of differentiated vegetative cells of the gametophytic blade change, and then, a single spore is released and germinates into another gametophytic blade (Ma and Shen 1996). Archeospore formation is an important feature of great economic value, which has been explored to develop new techniques and patterns for germling production. In field cultivation archeospores provide a large number of secondary germlings and increase the yield on the cultivating net (Li 1984; Yan et al. 2004). Furthermore, the characteristics of archeospore germlings excel those of conchospore germlings in terms of growth speed and stress tolerance abilities (Li 1984). Nevertheless, there are trade-offs between archeospore formation and blade growth, as too many archeospores will slow the growth of blades. Therefore, a thorough understanding of this biological trait is needed ahead of application.
It has been shown that the formation and release of archeospores can be affected by several culture conditions, including light intensity, photoperiod, water temperature, salinity, and desiccation (Li and Wang 1987, 1984; Li et al. 1988, 1986; Kitade et al. 2008; Xu et al. 2019). Moreover, the production of archeospores is significantly accelerated by the exogenous addition of allantoin and ammonium sulfate (Li and Wang 1987; Mizuta et al. 2003; Sun et al. 2018). Notably, it has been reported that chemical mutagenesis can generate stably inherited mutants with different archeospore-producing abilities (Yan et al. 2004; Zhang and Yan 2014), suggesting that archeospore formation is a genetically controlled trait of Pyropia. Recently, several studies were carried out to explore the physiological and molecular mechanisms for the formation and release of archeospores. For example, Gao et al. (2011) proposed that cell wall and photosynthetic properties were changed during archeospore formation in Py. yezoensis. It has also been found that photosynthesis-dependent Ca2+ influx triggered archeospore reproduction, and they also reported that oxidative stress could promote archeospore formation (Takahashi et al. 2010; Takahashi and Mikami 2017). However, the molecular mechanism of archeospore formation is still obscure as only a few preliminary studies have been carried out (Xu et al. 2003; Kitade et al. 2008; He et al. 2013).
In the present study, comparative transcriptomic analysis between abundant and scarce archeospores release strains of Py. yezoensis was conducted, and the genes related to archeospore formation were analyzed, providing valuable insights into the molecular basis of the archeospore formation in Pyropia.
Materials and methods
Algal materials and culturing conditions
Reliable materials for investigating archeospore characteristics require strains with different archeospore formation abilities and similar genetic backgrounds. To achieve this requirement, Py. yezoensis pure strains of W′ and W were selected from the germplasm strains of our laboratory, with the former continuously producing archeospores and the latter scarcely forming archeospores. These strains were previously established by separating and purifying a spottedly variegated gametophytic blade, produced by progeny of the hybridization between red mutant (fre-1,♀) and wild type (U-511,♂) (Yan et al. 2000; Yan and Aruga 2000). Free-living conchocelis of the two strains were stored in the laboratory as described by Kato and Aruga (1984). The medium used for gametophytes was sterilized seawater containing MES medium (Wang et al. 1986), and the culture conditions were as follows: 19 ± 1 °C, 40 μmol photons m−2 s−1, and 10 L: 14 D photoperiod.
Sampling settings and archeospore quantification
To determine the suitable sampling strategy for investigating the gene expression change during archeospore formation, the archeospore release abilities of the two strains were evaluated. For each strain, 500 individuals were randomly selected from 18-day-old (initialized from the germination of single conchospore) germlings, and the number of individuals releasing archeospores was recorded each day. The proportion of individuals releasing archeospores increased continuously in W′ strain, while all individuals in W strain scarcely released archeospores (Fig. 1a). We sampled 25- and 35-day-old germlings of the two strains for transcriptomic sequencing, named W′25d, W′35d, W25d, and W35d, respectively. For each sample, 0.1–0.3 g of gametophytic blade was washed with distilled water and flash-frozen in liquid nitrogen before being stored in − 80 °C refrigerator.
Meanwhile, 10 individuals from 18-day-old germlings were employed to quantify the released archeospores of two strains. Plastic cups were used to cultivate the gametophytes, and the cups with new culture solution were replaced every day. The released archeospores were attached on the replaced cups, and they could grow into visible germlings after several days culturing. Next, the numbers of the archeospores germlings were counted by scraping them off from the cups, and the quantity of archeospores was equivalent to the overall amount of archeospores germlings. The total number of archeospores per blade was calculated for 7 days, and the experiment was repeated three times.
RNA extraction and sequencing
Total RNA from each algal sample was extracted using Trizol reagent (Thermo Fisher, USA). The absorbance at wavelengths of 260 nm and 280 nm (A260/A280) was measured by NanoDrop 2000, and the integrity was examined by 1% agarose gel electrophoresis and Agilent 2100. The RIN (RNA integrity number) value was confirmed to be greater than 6.0 for all samples, and 2 μg of total RNA was used as input material for sequencing library preparations. RNA sequencing was performed by the Illumina HiSeqXTen sequencing platform (NovelBio Bio-Pharm Technology Co. Ltd., Shanghai) following the standard library preparation procedures.
Transcriptome de novo assembly and annotation
Raw reads were quality filtered using Fast-QC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/), and the clean data were used for the downstream analyses. De novo assembly of a reference transcriptome from the pair-ended reads was carried out using Trinity (Haas et al. 2013), and putative open reading frames were predicted with OrfPredictor (Min et al. 2005). Clean RNA-seq reads were mapped to the reference transcriptome assembly with BWA-MEM (Li and Durbin 2009), and expression levels of the transcripts were quantified as fragments per kilobase of exon per million fragments mapped (FPKM).
Differential expression analysis
Differential expression analysis was conducted for four comparisons combined by four samples (W′25d, W′35d, W25d, and W35d), which were W35d vs W25d, W′35d vs W′25d, W′25d vs W25d, and W′35d vs W35d. The R package of EBseq was used to identify the differential expression genes (DEGs) with setting the criteria for differential screening at P value < 0.05, FDR ≤ 0.05, and log2|FC| ≥ 1.
Gene ontology and KEGG enrichment analysis of DEGs
GO enrichment analysis for DEGs was performed using the GOseq R package (Young et al. 2010). The GO terms with corrected P values under 0.05 were defined as significantly enriched terms for DEGs. In addition, KEGG annotations were performed based on the KEGG database (http://www.genome.jp/kegg/pathway.html), and pathways with corrected P values under 0.05 were defined as significantly enriched pathways for DEGs.
Expressional validation of DEGs with qRT-PCR
For qRT-PCR validation, eight genes representing different expression patterns were selected from DEGs. The primers were designed in the open reading frame region using NCBI Primer Blast (Table 1). RNA of each sample was reversely transcribed into cDNA using Takara reverse transcription kit (Takara, Japan). qRT-PCR was performed with Takara’s SYBR Green fluorescent dye and Bio-Rad CFX-96 real-time PCR system using cDNA as a template and 18S as an internal reference gene. Three biological replicates were set for each gene, and three technical replicates were set for each biological replicate. The relative expression level of each gene was calculated by the 2-ΔΔCt method.
Availability of data and materials
All RNA sequencing data were submitted to NCBI SRA database (https://www.ncbi.nlm.nih.gov/sra/) under study number SRP191712. The accession numbers of each are as follows: SRR8873477, SRR8873478, SRR8873475, and SRR8873476.
Results
Archeospore formation ability evaluation of two strains
To explore the molecular mechanism of archeospore reproduction, W′ and W strains of Py. yezoensis were selected, and the archeospore formation ability was evaluated. For each strain, the proportions of releasing individuals were documented. When germlings cultured from 18- to 25-days-old, the blades of W′ strain continuously formed archeospores, and nearly one third of the individuals began to release archeospores (Fig. 1a). However, the blades of W′ strain hardly formed archeospores. Moreover, the amount of archeospores during 7 days cultivation was documented for two strains. In total, there were up to 1234 archeospores per blade in W′ strain and only 2 in W strain (Fig. 1b), indicating that the ability of archeospore formation of W′ strain was 617 times than that of W strain.
Sequencing and de novo assembly
Transcriptomic sequencing was performed by the Illumina HiSeqXTen sequencing platform, and approximately 39.5–55.2 million raw reads were produced for four sequencing libraries (Table 2). After filtering the low-quality reads, there were 46,327,654, 33,036,768, 39,605,176, and 34,710,324 clean reads for libraries of W′25d, W′35d, W25d, and W35d, respectively. Based on de novo assembly, a total of 53,689 contigs were obtained, and over 99% reads in four libraries could be mapped to these contigs.
Differential gene expression analysis
Differential expression analysis for pairwise comparison resulted in four groups (Fig. 2). In comparing of 35- to 25-day-old samples, distinct numbers of up- and downregulated differential expression genes (DEGs) were observed in the two strains. In W′ strain, a total of 4705 (W′35d vs. W′25d group) DEGs were identified, the majority of which were upregulated DEGs. In W strain, 2365 DEGs (W35d vs. W25d group) were identified, and the numbers of up- and downregulated DEGs were similar. When comparing samples between different strains of the same age, the DEGs pattern was also distinct. For 25-day-old samples, the number of DEGs was 3031 (W′25d vs. W25d group), with more upregulated than downregulated ones. For 35-day-old samples, 635 DEGs (W′35d vs. W35d group) were identified, the majority of which were upregulated ones.
In addition, consensus DEGs were identified between comparing groups (Fig. 3). According to Venn diagram illustration, the number of common DEGs between groups of W′35d vs. W′25d and W35d vs. W25d was only 64, and these common DEGs could account for the genes involved in normal developmental process. Likewise, there were 415 common DEGs between groups of W′25d vs. W25d and W′35d vs. W35d, which could be regarded as DEGs between the two strains at the same age. Among these 415 common DEGs, 361 were upregulated, and 54 were downregulated in W′ strain, and the expression trends of each DEGs (up- or downregulation) in two comparing groups were identical (Table S2).
GO and pathway enrichment of common DEGs
As the 415 common DEGs were considered as genes reflecting the difference between two strains, the function of them might be related to the archeospore formation of Py. yezoensis. Gene ontology (GO) enrichment analysis showed that the functions were enriched in several basal biological processes, such as translation, RNA activity, actin and vesicle assembly, intracellular localization of Golgi and mitochondria, and peroxisome activity (Table S3). Among them, the GO terms of cytoplasmic translation, RNA methylation, and actin organization were highly enriched (Fig. 4).
Moreover, according to KEGG analysis, approximately 55% DEGs were mapped to 58 KEGG pathways (Table S4). The top significantly enriched pathways included ribosome assembly, endocytosis, carbon and nitrogen metabolism, photooxidation, as well as AGE-RAGE and MAPK signaling pathways (Fig. 5).
Analysis of expression patterns of candidate DEGs
Based on consensus DEGs and functional enrichment screening, a total of 80 DEGs were further selected for hierarchical cluster analysis (Fig. 6). The result showed that the expression profiles of these genes were divergent between W′ and W strain. Most DEGs were specifically upregulated in W′ strain with abundant archeospores, suggesting that these genes may play roles in the process of archeospore formation. In terms of functional annotation, the constitution of these DEGs was as follows: 3 dehydrogenase genes and 17 Rab small GTPase genes, 6 myosin and 11 actin genes, 13 calmodulin (CaM) genes and 5 MAPKK kinase genes, 21 E3 ubiquitin ligase-related genes, 3 chitinase genes, and one gene encoded extracellular matrix protein.
qRT-PCR validation of candidate DEGs
To confirm the RNA-seq results, the expression levels of eight genes were quantified via qRT-PCR, and the expression levels (FPKM) among four samples were compared. The results showed that the expression patterns for all detected genes via qRT-PCR were consistent with RNA-Seq results, suggesting the reliability of transcriptomic data (Fig. 7).
Discussion
DEGs analysis revealed archeospore formation-related genes
The archeospore formation of Py. yezoensis was initialized by specialization of vegetative cells, and these specific cells could release archeospores which eventually germinated into gametophytic blades (Nelson et al. 1999). However, the molecular mechanisms underlying such asexual reproductive process remain unclear. In the present study, as the archeospore formation abilities of W′ strain were 617 times than that of W (Fig. 1b), we deduced that such trait difference could be determined by distinct regulatory mechanism of two strains. By comparative transcriptomic analysis, the dynamic changes of gene expression were investigated. Notably, the 415 common DEGs between two strains could be defined as genes related to archeospore formation, in which 80 DEGs were further selected according to functional screening. Here, we focused on the genes with the function of cellular signaling pathways, cytoskeletal movement, and cell wall degradation, representing three regulatory processes during archeospore formation.
Annotation analysis of cellular signaling pathways
The Ca2+ signaling pathway mainly relies on Ca2+/calmodulin (CaM) and calcineurin to initiate intracellular responses and participate in a variety of biological processes in eukaryotes (Clapham 2007). As the key component of Ca2+ signaling pathway, CaM mediates diverse biological processes, such as cell differentiation, cell movement, cytoskeletal, and enzyme activity regulation (Berridge et al. 2003). It has been reported that extracellular Ca2+ influx plays a critical role in the production of archeospores from Py. yezoensis (Li et al. 2009; Takahashi et al. 2010). Moreover, mitogen-activated protein kinase (MAPK) cascades were the evolutionary conserved signaling modules in all eukaryotes, playing key roles in gene expression and cytoplasmic activity. In yeast, the MAPK cascade regulates many spore formations and cell wall integration processes (Rodríguez-Peña et al. 2010). We found that 13 CaM and 5 MAPKK kinase homologous genes were highly expressed in W′ strain (Fig. 6) and the MAPK signaling pathway was significantly enriched as the top 20 significant signaling KEGG pathways of the common DEGs (Fig. 5). Therefore, the regulatory roles of canalized signaling pathways as Ca2+ and MAPK are worthy of investigation.
Annotation analysis of cytoskeletal movement
The narrow cytoskeleton mainly referred as microfilaments, microtubules, and intermediate fibers, and the combination of actin and myosin can achieve cell movement in animals or cytoplasmic flow in plants. Pyropia species capable of dispersing archeospores can achieve cytoplasmic flow and cell morphological changes during the formation of archeospores. During the transformation of the vegetative cells into archeospores, the shape of cell gradually became rounded by regular polygons and leads to the separation of protoplasts from the cell wall in the form of “water drop,” leaving only a transparent gelatinous layer. Thus, it is possible that cytoskeletal movement is involved in such a process. We found that the expression of some actin and myosin homologous genes was upregulated in W′ strain (Fig. 6), suggesting their roles in regulating the cytoskeletal changes of vegetative cells and causing the separation of protoplasts.
Annotation analysis of cell wall degradation
Enzymes isolated from marine animals and microorganisms, such as sea snail and Pseudomonas, can effectively hydrolyze the cell wall of algal vegetative cells (Han et al. 1997). The escape of archeospores from the constraint of the cell wall is required during the process of archeospore formation, although it was still unclear whether Pyropia can produce enzymes to digest the cell wall. In the present study, we found four chitinase homologous genes, and 17 Rab small GTPase genes were highly expressed in W′35d sample (Fig. 6). Chitinase belongs to the family of glycosyl hydrolases, which can effectively degrade the conidial wall of fungi (Adams 2004). As chitin is not the component of Pyropia cell wall, the role of these chitinases homologous in the degradation of Py. yezoensis cell wall is still debatable. On the other hand, the roles of Rab small GTPases in cell wall metabolism have been reported in land plants (Yang 2002; Lycett 2008), indicating that the potential function of these genes in archeospore formation should be investigated.
Conclusion
The comparative transcriptomic characteristics of Py. yezoensis strains were carried out for the first time. By exploring the DEGs between samples from two strains with different archeospore formation abilities, we suggest that the formation of archeospores was a complex regulatory process and multiple genes and pathways might be involved. The present study provides an important gene resource for deciphering the molecular mechanisms of archeospore formation, and further molecular, physiological, and phenotypic studies are required to better understand the asexual reproduction in Pyropia.
References
Adams DJ (2004) Fungal cell wall chitinases and glucanases. Microbiology 150:2029–2035
Berridge MJ, Bootman MD, Roderick HL (2003) Calcium signalling: dynamics, homeostasis and remodelling. Nat Rev Mol Cell Biol 4:517–529
Clapham DE (2007) Calcium Signaling. Cell 131:1047–1058
Gao S, Wang GC, Yang RL, Xie XJ, Pan GH, Xu P, Zhu JY (2011) Variations in the cell walls and photosynthetic properties of Porphyra yezoensis (Bangiales, Rhodophyta) during archeospore formation. J Phycol 47:839–845
Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, Couger MB, Eccles D, Li B, Lieber M, MacManes MD, Ott M, Orvis J, Pochet N, Strozzi F, Weeks N, Westerman R, William T, Dewey CN, Henschel R, LeDuc RD, Friedman N, Regev A (2013) De novo transcript sequence reconstruction from RNA-seq using the trinity platform for reference generation and analysis. Nat Protoc 8:1494–1512
Han BQ, Liu WS, Wang H, Dai JX (1997) Studies on the enzymes of algal cell wall hydrolysis. Mar Sci 21:47–49
He LW, Zhu JY, Lu QQ, Niu JF, Zhang BY, Lin AP, Wang GC (2013) Genetic similarity analysis within Pyropia yezoensis blades developed from both conchospores and blade archeospores using AFLP. J Phycol 49:517–522
Kato M, Aruga Y (1984) Comparative studies on the growth and photosynthesis of the pigmentation mutants of Porphyra yezoensis in laboratory culture. Jpn J Phycol 32:333–347
Kim JK, Yarish C, Hwang EK, Park M, Kim Y (2017) Seaweed aquaculture: cultivation technologies, challenges and its ecosystem services. Algae 32:1–13
Kitade Y, Asamizu E, Fukuda S, Nakajima M, Ootsuka S, Endo H, Tabata S, Saga N (2008) Identification of genes preferentially expressed during asexual sporulation in Porphyra yezoensis gametophytes (Bangiales, Rhodophyta). J Phycol 44:113–123
Li SY (1984) The ecological characteristics of monospores of Porphyra yezoensis Ueda and their use in cultivation. Hydrobiologia 116:255–258
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 25:1754–1760
Li SY, Wang JC (1984) The influence of the light intensity on the formation, discharge and adherence of monospores. Mar Sci 8:41–43
Li SY, Wang JC (1987) The influence of the various concentrations of (NH4)2SO4 on the growth of thalli of Porphyra yezoensis and the formation, discharge and adherence of monopores. Mar Sci 11:35–39
Li SY, Xu P, Wang M (1986) The influence of seawater on the discharge and adherence of monospores of Porphyra yezoensis Ueda. Mar Sci 10:38–40
Li SY, Xu P, Wang M (1988) Effect of light period on the growth of thalli and the formation, discharge and adherence of monospores of Porphyra yezoensis Ueda. Mar Sci 12:58–61
Li L, Saga N, Mikami K (2009) Ca2+ influx and phosphoinositide signalling are essential for the establishment and maintenance of cell polarity in monospores from the red alga Porphyra yezoensis. J Exp Bot 60:3477–3489
Lycett G (2008) The role of Rab GTPases in cell wall metabolism. J Exp Bot 59:4061–4074
Ma JH, Shin JA (1996) A study on monospores and leafy thalli of gametophytes of Porphyra tenuipedalis. J Fish China 20:132–138
Mikami K, Li CZ, Irie R, Hama Y (2019) A unique life cycle transition in the red seaweed Pyropia yezoensis depends on apospory. Commun Biol 2:299–308
Min XJ, Butler G, Storms R, Tsang A (2005) OrfPredictor: predicting protein-coding regions in EST-derived sequences. Nucleic Acids Res 33:W677–W680
Mizuta H, Yasui H, Saga N (2003) A simple method to mass produce monospores in the thallus of Porphyra yezoensis Ueda. J Appl Phycol 15:351–353
Nelson WA, Brodie J, Guiry MD (1999) Terminology used to describe reproduction and life history stages in the genus Porphyra (Bangiales, Rhodophyta). J Appl Phycol 11:407–410
Rodríguez Peña JM, García R, Nombela C, Arroyo J (2010) The high-osmolarity glycerol (HOG) and cell wall integrity (CWI) signalling pathways interplay: a yeast dialogue between MAPK routes. Yeast 27:495–502
Shimizu A, Morishima K, Kobayashi M, Kunimoto M, Nakayama I (2007) Identification of Porphyra yezoensis (Rhodophyta) meiosis by DNA quantification using confocal laser scanning microscopy. J Appl Phycol 20:83–88
Song JT, Yan XH (2015) Cytological studies on Pyropia chauhanii from Indian. J Fish China 39:1479–1486
Sun D, Ding HC, Yan XH (2018) The influence of allantoin on monospores release and somatic cell differentiation in gametophytic blades of Pyropia chauhanii. J Fish China 42:534–543
Sutherland JE, Lindstrom SC, Nelson WA, Brodie J, Lynch MDJ, Hwang MS, Choi HG, Miyata M, Kikuchi N, Oliveira MC, Farr T, Neefus C, Mols-Mortensen A, Milstein D, Mueller KM (2011) A new look at an ancient order: generic revision of the Bangiales (Rhodophyta). J Phycol 47:1131–1151
Takahashi M, Mikami K (2017) Oxidative stress promotes asexual reproduction and apogamy in the red seaweed Pyropia yezoensis. Front Plant Sci 8:62–67
Takahashi M, Saga N, Mikami K (2010) Photosynthesis–dependent extracellular Ca2+ influx triggers an asexual reproductive cycle in the marine red macroalga Porphyra yezoensis. Am J Plant Sci 1:1–11
Tseng CK, Chang TJ (1954) Studies on Porphyra I. Life history of Pyropia tenera Kjellm. Acta Bot Sin 3:287–302
Wang SJ, Zhang XP, Xu ZD, Sun YL (1986) A study on the cultivation of the vegetative cells and protoplasts of P. haitanensis I. Oceanol Limnol Sinica 17:217–221
Xu L, Zhou YH, Wang SJ, Wang YX, Ma LB, Zhou ZG (2003) Ultrastructural and molecular evidence for monospore formation in the thallus of Porphyra yezoensis. J Fish China 27:519–527
Xu D, Song XW, Li FT, Pan XH, Zhu WF, Zhang XC (2019) Effects of desiccation, diurnal temperature changes and irradiance on archeospore production of Pyropia yezoensis. Aquaculture 509:167–170
Yan XH, Aruga Y (2000) Genetic analysis of artificial pigmentation mutants in Porphyra yezoensis Ueda (Bangiales, Rhodophyta). Phycol Res 48:177–187
Yan XH, Fujita Y, Aruga Y (2000) Induction and characterization of pigmentation mutants in Porphyra yezoensis (Bangiales, Rhodophyta). J Appl Phycol 12:69–81
Yan XH, Fujita Y, Aruga Y (2004) High monospore-producing mutants obtained by treatment with MNNG in Porphyra yezoensis Ueda (Bangiales, Rhodophyta). Hydrobiologia 512:133–140
Yang Z (2002) Small GTPases: versatile signaling switches in plants. Plant Cell 14:S375–S388
Yang XW, Ding HC, Yan XH (2019) Genetic analysis of the period of meiosis and morphogenesis of the gametophytic blades in Pyropia suborbiculata (Bangiales, Rhodophyta). J Appl Phycol 31:4061–4068
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14–R25
Zhang C, Yan XH (2014) Isolation and characterization of new strains of Pyropia chauhanii (Bangiales, Rhodophyta). J Fish China 38:1457–1465
Acknowledgments
The authors would like to thank Mr. Qingjie Zhang for technical assistance with experiments.
Funding
This study is supported in part by the National Key Research and Development Program of China (2018YFD0900606), National Natural Science Foundation of China (31072208), and Science and Technology Planning Project of Jiangsu Province, China (BE2018335).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Table S1
FPKM values generated in four sequencing samples. (xlsx 3514 kb) (XLSX 3513 kb)
Table S2
Common DEGs generated by comparisons between W′ and W strains. (xlsx 47 kb) (XLSX 46 kb)
Table S3
Detailed GO enrichment information of common DEGs. (xlsx 55 kb) (XLSX 54 kb)
Table S4
Detailed KEGG enrichment information of common DEGs. (xlsx 14 kb) (XLSX 13 kb)
Rights and permissions
About this article
Cite this article
Song, S., Gao, D. & Yan, X. Transcriptomic exploration of genes related to the formation of archeospores in Pyropia yezoensis (Rhodophyta). J Appl Phycol 32, 3295–3304 (2020). https://doi.org/10.1007/s10811-020-02174-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-020-02174-5