Abstract
Interactions between algae and associated bacterial communities in the phycosphere depend on many factors such as culture age, nutrient availability, and antibiotic production and greatly influence algal survival and growth. The microbial community associated with the marine microalga Tetraselmis suecica F&M-M33 in a laboratory culture and an outdoor mass culture, set up from the laboratory one, run during a whole year was investigated in different seasons through isolation in pure culture of bacteria, amplified ribosomal DNA restriction analysis (ARDRA) of all the isolates, and terminal restriction fragment length polymorphism (T-RFLP) analysis. The total number of bacterial isolates was 152, which clustered in thirty-six 16S rDNA groups by ARDRA. Sequencing of a representative of each ARDRA group permitted identification of bacterial genera never reported before in association with microalgae in outdoor mass cultures, although most of them were previously found to be associated with the marine environment (e.g., seawater, sea sediments). T-RFLP analysis revealed that spring and autumn bacterial community profiles were closely related, while the bacterial laboratory community was considerably different (less than 50 % similarity) from that associated with the outdoor culture in different seasons. T-RFLP results suggest the presence of a core of bacteria that are closely associated with the alga and of a part of the community which varies seasonally.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
During growth, microalgae release a variety of organic compounds, mainly amino acids, peptides, sugars, and polyalcohols, which represent a substantial fraction of total photoassimilated carbon. Percent extracellular release (PER) values are on average about 10 %, especially in nutrient-rich environments and algal cultures (Grossart and Simon 2007; Thornton 2014), whereas PER increases to 30–40 % in oligotrophic waters, mainly when nitrogen is low compared with carbon (Thornton 2014). PER values up to 70–80 % have been reported by some authors (Grossart and Simon 2007; Hulatt and Thomas, 2010). The zone extending out of the algal cell in which bacterial growth is stimulated by algal extracellular products is termed phycosphere, in analogy to plant rhizosphere (Bell and Mitchell 1972). Strong interactions between bacteria and microalgae occur in the phycosphere and are the result of the balance between stimulatory and inhibitory activities (Natrah et al. 2014), which can also be mediated by phytohormones released by bacteria (Amin et al. 2015). These interactions can be species-specific, and strong diurnal variations have been shown to occur in a micro-spatial context (Cole 1982; Azam and Malfatti 2007) or in larger time-scale during different phases of algal blooms (Ramanan et al. 2016).
Non-axenic laboratory cultures of microalgae are often associated with bacterial strains that were present in the sampled natural habitat and that have been co-cultivated with the algal cells during enrichment and isolation. Besides the original bacterial community, microalgal cultures harbor other bacterial strains that can enter the culture as contaminants. In fact, cultures of microalgae, even when carried out in closed systems (photobioreactors), are susceptible to contamination, especially when performed outdoors (Tredici 2004; Chini Zittelli et al. 2013a, b). Our knowledge of the composition and role of the bacterial communities associated with microalgal mass cultures is still in its infancy (Natrah et al. 2014; Ramanan et al. 2016). Fukami et al. (1997) report that biomanipulation of bacterial assemblages (e.g., by the addition of selected bacterial strains) in mass cultures could have a stabilizing effect. In general, the understanding of the specific interactions between bacteria and microalgae could improve culture reliability, for example, in aquaculture facilities (Riquelme and Avendaño-Herrera 2003), where a higher control of pathogens by non-pathogenic bacteria in the presence of algal metabolites was observed (Sharifah and Eguchi 2011). Quorum sensing regulates virulence of different aquaculture pathogens, so microalgae able to interfere with quorum sensing, such as Chlamydomonas, Chlorella, Nannochloropsis, Isochrysis, and Tetraselmis (Teplitski et al. 2004; Natrah et al. 2011) might become interesting biocontrol agents for use in aquaculture (Natrah et al. 2011). Algae-bacteria consortia are already exploited for wastewater treatment, but great improvements can be expected from a deep investigation of algae-bacteria interactions (Ramanan et al. 2016).
The marine microalga Tetraselmis (Chlorodendrophyceae, Chlorophyta) is widely used in aquaculture as food for bivalve molluscs, the larval stages of crustaceans, and the cultivation of rotifers and Artemia (Tredici et al. 2009) and has recently been evaluated as a partial substitute of fish-based feed in sea-bass juveniles (Tulli et al. 2012). Tetraselmis biomass ensures a balanced diet and provides polyunsaturated fatty acids, as it contains eicosapentaenoic acid (EPA). Tetraselmis is used in “green water” and “pseudo-green water” techniques and is known for its antibacterial activity against several aquaculture pathogens (Austin et al. 1992; Regunathan and Wesley 2004; Makridis et al. 2006) and for its potential probiotic action in fish (Irianto and Austin 2002). Tetraselmis, thanks to its high growth rate and productivity (Chini Zittelli et al. 2006), is widely produced in hatcheries and is also commercialized by several companies in the form of concentrated vital paste or lyophilized powder (Tredici et al. 2009). This microalga is also proposed as feedstock for biofuel production (Aquafuels 2011; Bondioli et al. 2012; Yao et al. 2012).
To understand the role of bacteria associated with microalgal growth, the first step is to acquire information on the structure and composition of the associated bacterial community. Therefore, in the present study, we investigated the bacterial community associated with an outdoor mass culture of Tetraselmis suecica strain F&M-M33 in different seasons by terminal restriction fragment length polymorphism (T-RFLP) and through isolation in pure culture and identification through partial sequencing of the 16S ribosomal DNA (rDNA) gene of the associated cultivable bacteria. The bacterial community associated to the laboratory culture from which the outdoor culture was set up was also analyzed to verify whether the bacteria present in the original culture were maintained, associated with a culture derived from them, and kept outdoors for a whole year.
Materials and methods
Laboratory and outdoor cultures
Tetraselmis suecica F&M-M33 was maintained in sterile F medium (Guillard and Ryther 1962) under laboratory conditions in 1-L tubes bubbled with a sterile air/CO2 mixture (98:2, v/v) and continuous illumination (200 μmol photons m−2 s−1). The culture was carried out under sterile conditions. One sample was collected from the laboratory batch culture for bacterial community analysis when the culture reached a concentration suitable to set up another culture. Thus, the laboratory culture was used to start an outdoor mass culture kept under a semi-continuous regime. The outdoor culture was carried out from February to November in a 5-m2 panel surface (1 m height × 5 m length), 300-L first-generation Green Wall Panel (GWP®-I) photobioreactor (patented by Tredici and Rodolfi 2004) at the Istituto per lo Studio degli Ecosistemi of the CNR in Sesto Fiorentino (43° 50′ N; 11° 11′ E), near Florence (Italy). Compressed air was bubbled at the bottom of the reactor through a perforated plastic tube for mixing and degassing, while CO2, used as carbon source and for pH regulation, was injected into the culture through a gas diffuser. The dilution rate (fraction of the culture volume that is withdrawn and substituted with fresh culture medium) and the frequency of dilutions were varied according to culture productivity, which changes in different seasons according to the different radiation available and temperature. In winter, a low dilution rate (30 %) and a low dilution frequency (7 days) were adopted, while in the summer, a dilution of 50 % every 2 days was applied; intermediate dilution (40–45 % was applied in spring and autumn every 3 days). In spring and summer, when temperature exceeded the pre-set value of 27 °C, a control unit provided temperature regulation by spraying water on the reactor surface. During the night in spring and summer and during the whole day in the other seasons culture temperature was allowed to equilibrate to ambient. Samples were collected from this culture once per season (early March, mid-May, end of July, early November) and termed Winter, Spring, Summer, and Autumn. All the samples were collected from the volume withdrawn from the culture at the moment of dilution, except the autumn sample that was collected 1 day after dilution.
Tetraselmis seucica quantification in the outdoor culture
Microalgal concentration of the outdoor culture was determined at each sampling by measuring dry weight according to Chini Zittelli et al. (2000) and by counting cells using a Thoma haemocytometer. Algal growth of the laboratory culture was followed daily. Algal cell dimensions were determined on microphotographs using image analysis software (Nikon, Japan), and these were used to calculate the surface area of the longitudinal section, approximating the alga shape to an ellipse.
Determination of number of T. suecica-associated bacteria
Bacterial concentration of the outdoor culture was determined at each sampling by viable cell count. Samples were maintained in agitation at room temperature for 3 h on a magnetic stirrer to allow the flocks to disperse. Serial dilutions of the microalgal culture samples were prepared using a sterile NaCl (9 g L−1) solution. The agar plate count method was used. Aliquots of 100 μL of each culture dilution were spread in triplicate on Marine Agar (Difco Marine Broth 2216) plates, which were incubated at 27 °C. The number of colony forming units (CFU) was evaluated after 6 days of incubation.
Isolation of bacterial strains and ARDRA characterization of the isolates
Bacterial isolates were obtained from the plates used for the viable cell count. Colonies were selected to represent the widest variation of colony characteristics (color, shape, edge, etc.). The number of isolates obtained was 27 for the laboratory culture, 21 for the winter, 25 for the spring, 31 for the summer, and 48 for the autumn sample of the outdoor mass culture.
DNA of bacterial isolates was extracted from colony formed on Marine Agar plates using the InstaGene Matrix kit (Bio-Rad, USA). The extracted DNA was diluted 1:10 in sterile ultra-pure water. Amplified 16S rDNA was obtained using primers 63f (CAGGCCTAACACATGCAAGTC) and 1387r (GGGCGG(AT)GTGTACAAGGC) (Marchesi et al. 1998). Polymerase chain reactions were performed using a GeneAmp PCR System 2700 (Applied Biosystems, USA) in a final volume of 50 μL, each containing DNA template solution, 2.5 U of Taq DNA polymerase (Polymed Srl, Italy), 0.25 μM of each primer, 0.2 mM dNTP, 2 mM MgCl2, and 0.3 mg mL−1 of bovine serum albumin. All reactions included both negative (DNA-free) and positive controls. The PCR consisted of 35 cycles of denaturation at 94 °C for 45 s, annealing at 55 °C for 35 s, extension at 72 °C for 1 min, preceded by 6 min at 94 °C for initial denaturation, and followed by 7 min at 72 °C for final extension. The successful amplification of the expected fragment (about 1300 bp) was checked by electrophoresis in 0.8 % (w/v) agarose gel at 4 V cm−1 for 1 h in TBE buffer (90 mM Tris-borate, 90 mM boric acid, 2 mM EDTA; pH 8.3) using 100 bp DNA ladder (Invitrogen, Italy) as molecular size marker.
Aliquots containing 300 ng of amplified DNA were digested with 5 U of a restriction endonuclease. DNA from each strain was digested using HaeIII or CfoI (New England Biolabs, USA) in separate extractions. Incubations were performed for 3 h at 37 °C to ensure complete digestion and for 20 min at 65 °C to ensure enzyme inactivation. Each digestion was electrophoresed through 2.8 % (w/v) agarose gel stained with ethidium bromide and run in TBE buffer at 5 V cm−1. Gel images were captured with a CCD camera (UVItec Gel Documentation System, Belgium), and analyzed. Bacteria isolates were clustered into operation taxonomic units (ARDRA groups) and then identified through 16S rDNA sequencing. The presence/absence of each ARDRA group in the different samples (laboratory and outdoor cultures) was used to perform a binary matrix. Sørensen similarity coefficient was used to compute a similarity matrix through Primer 6 software (PRIMER-E Ltd, UK). The matrix obtained was analyzed to perform a cluster analysis using average linkages algorithm.
Identification of bacterial isolates by 16S rDNA sequencing
For each ARDRA group, one representative isolate (see Table 2 for isolate indication) was selected and then identified through 16S rDNA sequencing. The amplified DNA, obtained as described in the isolation paragraph, was purified using the QIAquick PCR Purification kit (Qiagen Inc, USA). The purified DNA was quantified by electrophoresis through 1 % (w/v) agarose gel using the DNA Molecular Weight Marker XIV (Roche, Switzerland). Sequencing was performed using BigDye Terminator Cycle Sequencing Kit on an ABI 3100 DNA sequencer (Applied Biosystems). DNA sequences obtained were viewed, edited, and aligned with BioEdit software (Ibis Biosciences, Abbott Laboratories, USA). Sequence analysis was performed with BlastN program in the GenBank database (National Center for Biotechnology Information, NCBI, http://www.ncbi.nlm.nih.gov).
T-RFLP analysis of bacterial community associated with T. suecica
T-RFLP analysis was performed on the laboratory culture and on the outdoor culture sampled in the different seasons, as well as on an axenic T. suecica F&M-M33 culture to evaluate the possible interference of amplification of microalgal 16S rRNA genes. DNA was extracted from T. suecica cultures using the FastDNA SPIN Kit for Soil (Qbiogene Inc, USA) (Kohli et al. 2015). Extractions were performed in triplicate and nucleic acids concentrations and quality were measured spectrophotometrically and by gel electrophoresis, respectively. Fifty grams of extracted DNA were amplified using 63f primer labeled with 6-carboxyfluorescein (FAM) and 1387r primer (Marchesi et al. 1998). Polymerase chain reactions were performed in a final volume of 50 μL. A touchdown PCR program was performed as described in Tatti et al. (2012). All reactions included both negative and positive controls. PCR products were checked by electrophoresis in 0.8 % (w/v) agarose gel at 5 V cm−1 for 1 h in TBE buffer (90 mM Tris-borate, 90 mM boric acid, 2 mM EDTA; pH 8.3) stained with ethidium bromide and observed under UV light. To minimize stochastic PCR biases (Polz and Cavanaugh 1998), two different amplifications were run for each sample and then the amplified products were pooled and purified with PCR purification kit (Qiagen). About 300 ng of purified amplification products were digested separately with 20U of HaeIII, CfoI, or MspI (Roche) at 37 °C for 3 h. A 60-ng aliquot of digested product was resolved by capillary electrophoresis on an ABI 3730XL Genetic Analyzer (Applied Biosystems) using Gene Scan 500 (Applied Biosystems) as size standard. Terminal restriction fragment sizes between 50 and 500 bp were determined using GeneScan analytical software v1.0 (Applied Biosystems). For each sample, three replicates were analyzed. Peaks which were not present in all replicates were considered PCR artifacts and thus removed, and only peaks whose fluorescence intensity contributed to the sum of all peaks intensity for more than 3 % were considered. Profiles obtained for each sample with three restriction digestions were aligned to perform a binary matrix representing the presence or absence of peaks of a certain length.
Sørensen similarity coefficient was used to compute a similarity matrix through Primer 6 software (PRIMER-E Ltd). The matrix obtained was analyzed to perform a cluster analysis using average linkages algorithm. Diversity indexes, Shannon (H), and equitability (E) were also calculated.
Statistical analysis
Experimental results on microalgal and bacterial concentrations were analyzed with one-way ANOVA followed by Tukey’s multi comparative test.
Results
Total viable count of algae-associated bacteria
The total viable bacteria associated with the outdoor culture of T. suecica F&M-M33 were lower than those observed in the laboratory batch culture, both in terms of number per unit volume and number per unit total biomass (Table 1). Outdoors, the highest bacterial abundance per unit culture volume was found in winter, the season in which the culture dilution was less frequent due to the slower culture growth (Table 1). Similar results were obtained when the bacterial abundance was expressed per unit total biomass (Table 1). Although plate counts allow to evaluate only the cultivable fraction and part of the difference in bacterial counts may result from the presence of a different proportion of non-cultivable bacteria in the different seasons, in previous experiments performed on outdoor and laboratory cultures the difference between total counts by fluorescence microscopy after dyeing with acridine orange and plate counts was no more than 15 % (unpublished data). Thus, we consider the plate counts representative of the algal density of the samples.
ARDRA analysis and identification of the isolates
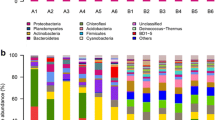
Bacteria isolated from plates used to determine the number of cultivable bacteria associated with the laboratory and outdoor cultures of T. suecica F&M-M33 were subjected to ARDRA using the enzymes CfoI and HaeIII, in separate experiments. The analysis of the ARDRA profiles of the 152 isolates allowed to evidence the high diversity within the 16 typologies of colonies observed and permitted to identify 36 different ARDRA groups (data not shown). Notably, the maximum number of different ARDRA groups was detected in the autumn outdoor culture (17 distinct groups), while the lowest number of groups was obtained from winter and summer outdoor cultures (8) (Fig. 1a). The laboratory culture showed nine ARDRA groups (Fig. 1a). About 19 % of the obtained ARDRA groups were isolated only from the laboratory culture, and only about 8 % of the groups were present both in the laboratory and in at least one season in the outdoor culture (Table 2). Among the 29 ARDRA groups found in the outdoor culture, only one was found in all seasons, about 10 % of the ARDRA groups were found in three different seasons, 28 % were found at least in two seasons, while over 58 % were found only in one season (Table 2). The presence/absence of each ARDRA group in the different samples was used to compare bacterial communities of the laboratory and outdoor cultures (Fig. 1b). The outdoor culture seasonal samples cluster together, even if with low similarity, and apart from the laboratory culture.
a Number of ARDRA groups observed in an outdoor Tetraselmis suecica F&M-M33 culture maintained year round, and its inoculum kept in the laboratory. b Average linkages cluster analysis based on Sørensen similarity coefficient of the bacterial communities associated with the outdoor and the laboratory cultures, obtained from ARDRA data
Bacteria representing the different ARDRA groups were identified through 16S rDNA sequencing (Table 2). The most represented phyla were Proteobacteria (20 representatives of which 7 belonging to the Roseobacter-clade and 4 to Rhizobiales) and Bacteroidetes (14 representatives of which 8 belonging to Flavobacteriales). Only six of the closest relatives of the ARDRA group representatives were previously isolated in association with algae, as it emerges from the information linked to each closest relative retrieved through BlastN by means of the GenBank accession number. ARDRA group 27 (closest relative the Alteromonadales, Marinobacter sp.) was the only one detected in the outdoor culture during the whole cultivation period, but it was not found in the laboratory culture (Table 2).
T-RFLP analysis of total bacterial communities associated with T. suecica cultures
The total bacterial community structure of outdoor and laboratory cultures was determined through T-RFLP analysis. The total number of ribotypes found with three restriction enzymes (HaeIII, CfoI, or MspI) was higher in the laboratory (59 ribotypes) than in the outdoor cultures (Table 3). Summer (53 ribotypes) and spring (51 ribotypes) were the richest seasons in ribotypes (Table 3). The profiles were dissimilar to each other, but the outdoor cultures shared from 35 % (autumn sample digested with CfoI) to 59 % (summer sample digested with MspI) ribotypes with the laboratory culture.
The T-RFLP dendrogram obtained using Sørensen similarity coefficient showed that the communities associated with the outdoor cultures had a higher similarity among themselves than with the community associated with the laboratory culture (Fig. 2a). Among the seasons, the highest similarity was found between spring and summer communities. This result was similar to that obtained by ARDRA analysis performed on the isolated strains (Fig. 1b). The ARDRA dendrogram showed high similarity with the T-RFLP community data. Also according to T-RFLP data, the most similar bacterial communities were those of spring and summer, whereas the main difference with the ARDRA dendrogram was that the autumn instead of the winter sample was the closest to the laboratory culture. These differences are probably related to the colony selection process that led to the bacterial isolates on which the ARDRA analysis was performed and that may have missed some ARDRA groups because very diverse bacteria can form colonies morphologically very similar.
a Average linkages cluster analysis based on Sørensen similarity coefficient of the bacterial communities associated with an outdoor Tetraselmis suecica F&M-M33 culture maintained year round, and its inoculum kept in the laboratory, obtained from T-RFLP data. b Diversity indexes of the same bacterial communities, expressed as Shannon index (H) and equitability (E) calculated from T-RFLP data
Univariate analysis of T-RFLP data in the form of Shannon diversity index (H) and equitability (E) was performed for the bacterial communities. The bacterial community associated with the laboratory culture showed the highest H diversity and the highest E (Fig. 2b). Regarding the outdoor cultures, the lowest diversity was present in autumn, the highest in summer. E showed the same trend as diversity, suggesting the presence of few dominating species during the colder months and a more distributed abundance during the mild season.
The T-RFLP analysis was also conducted on the isolates representing each ARDRA group to highlight those cultivable bacteria with the tightest association with T. suecica F&M-M33. Results obtained showed that only a few ribotypes corresponding to isolates were present in all the analyzed samples (outdoors during the whole cultivation period and in the laboratory): the representatives of ARDRA groups 3 and 33 (Flavobacetriales), 6 (Rhizobiales) and 23 (Roseobacter-clade) (data not shown).
Discussion
Many of the studies concerning the composition of bacterial communities associated with microalgae deal with natural environments, especially in relation to toxic algal blooms, or laboratory cultures (Ramanan et al. 2016). To our knowledge, no literature is available on the seasonal variation of the bacterial community associated with outdoor cultures of the microalga T. suecica. The present study, through the identification of bacterial isolates from T. suecica F&M-M33 outdoor cultures, showed for the first time that seasonal variation in the bacterial flora occurred and that most of the identified bacteria were typically associated with marine algae or found in seawater habitats, suggesting that a strictly associated bacterial flora could remain during the whole cultivation period, even considering that the culture was held a hundred of kilometers away from the sea, thus making contamination by bacteria from seawater aerosol improbable. Of the 26 different bacterial genera to which the isolates here identified belong, 18 had already been reported to be associated with marine microalgae of different classes and genera (Table 4). These identified bacterial genera had previously been found in association with dinoflagellates and green algae (13 bacterial genera), diatoms (8), and Prymnesiophyta (7). Only three of these bacterial genera were previously reported in association with Tetraselmis species (Table 3), of which only two (Roseobacter and Mesorhizobium) were specifically in association with T. suecica (Nicolas et al. 2004). The eight genera that have not been previously reported to grow in association with microalgae, mainly originate from seawater (Suzuki et al. 1999; Nedashkovskaya et al. 2005; Yoon et al. 2007; Hwang and Cho 2008; Kim et al. 2010; Nedashkovskaya et al. 2010; Zhang et al. 2014; Global Catalogue of Microorganisms). The bacterial genera among those found in T. suecica F&M-M33 cultures associated with the largest number of microalgal genera are Roseobacter (21), Marinobacter (20), Pseudomonas (11), and Muricauda (9) (Table 4). Some of the bacterial genera associated with T. suecica F&M-M33 cultures have representatives endowed with algicidal activity: Marivirga (GenBank accession number KC890797.1, closest relative to our ARDRA group 30), Pseudomonas (Dakhama et al., 1993), Halomonas (Su et al. 2011), and some representatives of the Roseobacter clade (Seyedsayamdost et al. 2011; Sule and Belas 2013; Riclea et al. 2012).
The two most represented groups among the bacteria isolated from T. suecica F&M-M33 were the Flavobacteriales and the Roseobacter clade. The Cytophaga-Flavobacterium-Bacteroides group is among the most widely diffused ones, especially in marine environments where it plays an important role in organic matter decomposition (Kirchman 2002). Its abundance seems related to the high growth rate of its members and their ability to respond quickly to the increase in organic matter availability, particularly when phytoplankton blooms occur (Kirchman 2002).
Roseobacter clade representatives are among the most abundant and well-known marine bacteria, also thanks to their high cultivability (Buchan et al. 2005), and appear to be responsible for 5–10 % of energy generation in the ocean surface layer (Allgaier et al. 2003). This group together with the Rhodobacterales is characterized by the widespread ability to perform aerobic anoxygenic photosynthesis that generates energy without carbon dioxide biofixation. This metabolic ability provides an advantage over competitors, as it allows maintaining a membrane electrochemical potential that can be used either to generate ATP or for active transport, and could thus increase the efficiency of organic substrate use for biosynthesis (Yurkov and Beatty 1998). The association with algal cells, and in this case with the flagellate Tetraselmis, could contribute to keep these bacteria in the photic layer (Yurkov and Beatty 1998).
The T-RFLP analysis of the total bacterial community associated with an outdoor culture of T. suecica F&M-M33 showed a lower bacterial diversity compared to that of the laboratory culture from which the outdoor culture originated, and its composition was strongly influenced by seasons, with the winter sample showing the highest bacterial number and the summer one being that with the highest bacterial diversity. Summer and spring bacterial communities showed a similarity of about 70 %. Interesting, all outdoor samples shared about 50 % of ribotypes with the laboratory culture whatever the restriction enzyme used, suggesting that a core group of bacteria was always present (Table 3). The use of T-RFLP approach to identify the presence of the strains isolated from T. suecica F&M-M33 in all the samples analyzed permitted to bring to light the presence of a cultivable bacterial core constituted by Muricauda, Leeuwenhoekiella, Ponticoccus, and Mesorhizobium. The first three bacteria had never been isolated in association with T. seucica. The availability of cultivable isolates that could be considered symbionts of T. seucica opens new perspectives to develop an artificial bacterial consortium that will permit to reconstruct the complex interactions that take place within the holobiont composed by T. seucica and its symbiotic bacteria.
In conclusion, complex and taxonomically rich communities with several bacterial taxa not previously reported as associated with T. suecica were found. The bacterial community in outdoor cultures of T. suecica F&M-M33 was composed of a core group of strictly associated bacteria (Muricauda, Leeuwenhoekiella, Ponticoccus, never found associated to this alga before, and Mesorhizobium), although seasonal variations in the composition of the bacterial flora occurred. This close association between the alga and its bacterial flora may play a beneficial role in increasing productivity and stability of algal mass cultures. Further studies are necessary to evaluate to which extent these effects occur in T. suecica F&M-M33 cultures, in particular, and in microalgae cultures, in general.
References
Allgaier M, Uphoff H, Felske A, Wagner-Döbler I (2003) Aerobic anoxygenic photosynthesis in Roseobacter clade bacteria from diverse marine habitats. Appl Environ Microb 69:5051–5059
Amin SA, Green DH, Hart MC, Kupper FC, Sundac WG, Carrano CJ (2009) Photolysis of iron–siderophore chelates promotes bacterial–algal mutualism. Proc Natl Acad Sci U S A 106:17071–17076
Amin SA, Hmelo LR, van Tol HM, Durham BP, Carlson LT, Heal KR, Morales RL, Berthiaume CT, Parker MS, Djunaedi B, Ingalls AE, Parsek MR, Moran MA, Armbrust EV (2015) Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature 522:98–101
AquaFUELs (2011) Taxonomy, Biology and Biotechnology. Report of the AquaFUELs Project. www.aquafuels.eu/attachments/079_Merged%20reports%20-Taxonomy_Biology%20&%20Biotechnology.pdf. Accessed 14 Nov 1912.
Arora M, Chandrashekar Anil A, Delany J, Rajarajan N, Emami K, Mesbahi E (2012) Carbohydrate degrading bacteria closely associated with Tetraselmis indica: influence on algal growth. Aquat Biol 15:61–71
Austin B, Baudet E, Stobie M (1992) Inhibition of bacterial fish pathogens by Tetraselmis suecica. J Fish Dis 15:55–61
Azam F, Malfatti F (2007) Microbial structuring of marine ecosystems. Nat Rev Microbiol 5:782–791
Baker KH, Herson DS (1978) Interactions between the diatom Thalassiosira pseudonanna and an associated pseudomonad in a mariculture system. Appl Environ Microb 35:791–796
Bell W, Mitchell R (1972) Chemotactic and growth responses of marine bacteria to algal extracellular products. Biol Bull 143:265–277
Berg GM, Repeta DJ, Laroche J (2002) Dissolved organic nitrogen hydrolysis rates in axenic cultures of Aureococcus anophagefferens (Pelagophyceae): comparison with heterotrophic bacteria. Appl Environ Microb 68:401–404
Bondioli P, Della Bella L, Rivolta G, Chini Zittelli G, Bassi N, Rodolfi L, Casini D, Prussi M, Chiramonti D, Tredici MR (2012) Oil production by the marine microalgae Nannochloropsis sp. F&M-M24 and Tetraselmis suecica F&M-M33. Bioresource Technol 114:567–572
Bruckner CG, Bahulikar R, Rahalkar M, Schink B, Kroth PG (2008) Bacteria associated with benthic diatoms from Lake Constance: phylogeny and influences on diatom growth and secretion of extracellular polymeric substances. Appl Environ Microb 74:7740–7749
Buchan A, González JM, Moran MA (2005) Overview of the marine Roseobacter lineage. Appl Environ Microb 71:5665–5677
Chini Zittelli G, Pastorelli R, Tredici MR (2000) A Modular Flat Panel Photobioreactor (MFPP) for indoor cultivation of Nannochloropsis sp. under artificial illumination. J Appl Phycol 12:521–526
Chini Zittelli G, Rodolfi L, Biondi N, Tredici MR (2006) Productivity and photosynthetic efficiency of outdoor cultures of Tetraselmis suecica in annular columns. Aquaculture 261:932–943
Chini Zittelli G, Rodolfi L, Bassi N, Biondi N, Tredici MR (2013a) Photobioreactors for microalgae biofuel production. In: Borowitzka MA, Moheimani N (eds) Algae for biofuels and energy. Springer, Dordrecht, pp 115–131
Chini Zittelli G, Biondi N, Rodolfi L, Tredici MR (2013b) Photobioreactors for mass production of microalgae. In: Richmond A, Hu Q (eds) Handbook of microalgal culture: applied phycology and biotechnology, 2nd edn. Wiley, Oxford, pp 225–266
Cole JJ (1982) Interactions between bacteria and algae in aquatic ecosystems. Annu Rev Ecol Syst 13:291–314
Dakhama A, de la Noüe J, Lavoie MC (1993) Isolation and identification of antialgal substances produced by Pseudomonas aeruginosa. J Appl Phycol 5:297–306
Fukami K, Nishijima T, Ishida Y (1997) Stimulative and inhibitory effects of bacteria on the growth of microalgae. Hydrobiologia 358:185–191
Global Catalogue of Microorganisms, http://gcm.wfcc.info/Strain_numberToInfoServlet?strain_number=NBRC%20101843. Accessed 13 July 2016
Green DH, Llewellyn LE, Negri AP, Blackburn SI, Bolch CJS (2004) Phylogenetic and functional diversity of the cultivable bacterial community associated with the paralytic shellfish poisoning dinoflagellate Gymnodinium catenatum. FEMS Microbiol Ecol 47:345–357
Green DH, Shenoy DM, Hart MC, Hatton AD (2011) Coupling of dimethylsulfide oxidation to biomass production by a marine Flavobacterium. Appl Environ Microb 77:3137–3140
Grossart HP, Simon M (2007) Interactions of planktonic algae and bacteria: effects on algal growth and organic matter dynamics. Aquat Microb Ecol 47:163–176
Guillard RRL, Ryther JH (1962) Studies of marine planktonic diatoms. I. Cyclotella nana (Hustedt) and Detonula confervacea (Cleve). Can J Microbiol 8:229–239
Hold GL, Smith EA, Birkbeck TH, Gallacher S (2001) Comparison of Paralytic Shellfish Toxin (PST) production by the dinoflagellates Alexandrium lusitanicum NEPCC 253 and Alexandrium tamarense NEPCC 407 in the presence and absence of bacteria. FEMS Microbiol Ecol 36:223–234
Huang S, Fulbright S, Zeng X, Yates T, Wardle G, Chisholm S, Xu J, Lammers P (2011) Characterization of the bacterial metagenome in an industrial algae bioenergy production system. Poster presented at the 4th Congress of the International Society for Applied Phycology, Halifax, Canada, 19-24/06/2011. http://www.osti.gov/scitech/servlets/purl/1093676. Accessed 26 September 2016
Hulatt CJ, Thomas DN (2010) Dissolved organic matter (DOM) in microalgal photobioreactors: a potential loss in solar energy conversion? Bioresour Technol 101:8690–8697
Hwang CY, Cho BC (2008) Ponticoccus litoralis gen. nov., sp. nov., a marine bacterium in the family Rhodobacteraceae. Int J Syst Evol Micr 58:1332–1338
Hwang CY, Zhang GI, Kang S-H, Kim HJ, Cho BC (2009) Pseudomonas pelagia sp. nov., isolated from a culture of the Antarctic green alga Pyramimonas gelidicola. Int J Syst Evol Micr 59:3019–3024
Irianto A, Austin B (2002) Probiotics in aquaculture. J Fish Dis 25:633–642
Jasti S, Sieracki ME, Poulton NJ, Giewat MW, Rooney-Varga JN (2005) Phylogenetic diversity and specificity of bacteria closely associated with Alexandrium spp. and other phytoplankton. Appl Environ Microb 71:3483–3494
Kaczmarska I, Ehrman JM, Bates SS, Green DH, Léger C, Harris J (2005) Diversity and distribution of epibiotic bacteria on Pseudo-nitzschia multiseries (Bacillariophyceae) in culture, and comparison with those on diatoms in native seawater. Harmful Algae 4:725–741
Kim B-S, Kim O-S, Moon EY, Chun J (2010) Vitellibacter aestuarii sp. nov., isolated from tidal-flat sediment, and an emended description of the genus Vitellibacter. Int J Syst Evol Micr 60:1989–1992
Kirchman DL (2002) The ecology of Cytophaga-Flavobacteria in aquatic environments. FEMS Microbiol Ecol 39:91–100
Kohli GS, John U, Figueroa RI, Rhodes LL, Harwood DT, Groth M, Bolch CJS, Murray SA (2015) Polyketide synthesis genes associated with toxin production in two species of Gambierdiscus (Dinophyceae). BMC Genomics 16:410
Kuo RC, Lin S (2013) Ectobiotic and endobiotic bacteria associated with Eutreptiella sp. isolated from Long Island Sound. Protist 164:60–74
Lakaniemi A-M, Intihar VM, Tuovinen OH, Puhakka JA (2012a) Growth of Chlorella vulgaris and associated bacteria in photobioreactors. Microb Biotechnol 5:69–78
Lakaniemi A-M, Intihar VM, Tuovinen OH, Puhakka JA (2012b) Growth of Dunaliella tertiolecta and associated bacteria in photobioreactors. J Ind Microbiol Biot 39:1357–1365
Lakaniemi A-M, Hulatt CJ, Wakeman KD, Thomas DN, Puhakka JA (2012c) Eukaryotic and prokaryotic microbial communities during microalgal biomass production. Bioresource Technol 124:387–393
Le Chevanton M, Garnier M, Bougaran G, Schreiber N, Lukomska E, Bérard J-B, Fouilland E, Bernard O, Cadoret J-P (2013) Screening and selection of growth-promoting bacteria for Dunaliella cultures. Algal Res 2:212–222
Makridis P, Alves Costa R, Dinis MT (2006) Microbial conditions and antimicrobial activity in cultures of two microalgae species, Tetraselmis chuii and Chlorella minutissima, and effect on bacterial load of enriched Artemia metanauplii. Aquaculture 255:76–81
Makridis P, Ferreira T, Kokou F, Tsigenopoulos CS, Divanach P (2012) Quantitative and qualitative aspects of bacterial communities associated with cultures of Chlorella minutissima. J World Aquacult Soc 43:571–578
Marchesi JR, Sato T, Weightman AJ, Martin TA, Fray JC, Hiom SJ, Wade WG (1998) Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microb 64:795–799.
Natrah FMI, Kenmegne MM, Wiyoto W, Sorgeloos P, Bossier P, Defoirdt T (2011) Effects of micro-algae commonly used in aquaculture on acyl-homoserine lactone quorum sensing. Aquaculture 317:53–57
Natrah FMI, Bossier P, Sorgeloos P, Yusoff FMD, Defoirdt T (2014) Significance of microalgal–bacterial interactions for aquaculture. Rev Aquacult 6:48–61
Nedashkovskaya OI, Vancanneyt M, Dawyndt P, Engelbeen K, Vandemeulebroecke K, Cleenwerck I, Hoste B, Mergaert J, Tan T-L, Frolova GM, Mikhailov VV, Swings J (2005) Reclassification of [Cytophaga] marinoflava Reichenbach 1989 as Leeuwenhoekiella marinoflava gen. nov., comb. nov. and description of Leeuwenhoekiella aequorea sp. nov. Int J Syst Evol Micr 55:1033–1038
Nedashkovskaya OI, Vancanneyt M, Kim SB, Bae KS (2010) Reclassification of Flexibacter tractuosus (Lewin 1969) Leadbetter 1974 and ‘Microscilla sericea’ Lewin 1969 in the genus Marivirga gen. nov. as Marivirga tractuosa comb. nov. and Marivirga sericea nom. rev., comb. nov. Int J Syst Evol Micr 60:1858–1863
Nicolas J-L, Corre S, Cochard J-C (2004) Bacterial population association with phytoplankton cultured in a bivalve hatchery. Microbial Ecol 48:400–413
Oh J-I, Kim M-J, Lee J-Y, Ko I-J, Kim W, Kim SW (2011) Isolation and characterization of algicidal bacteria from Cochlodinium polykrikoides culture. Biotechnol Bioproc E 16:1124–1133
Otsuka S, Abe Y, Fukui R, Nishiyama M, Senoo K (2008) Presence of previously undescribed bacterial taxa in non-axenic Chlorella cultures. J Gen Appl Microbiol 54:187–193
Polz MF, Cavanaugh CM (1998) Bias in template-to-product ratios in multitemplate PCR. Appl Environ Microb 64:3724–3730
Ramanan R, Kim B-H, Cho D-H, Oh H-M, Kim H-S (2016) Algae–bacteria interactions: evolution, ecology and emerging applications. Biotechnol Adv 34:14–29
Regunathan C, Wesley SG (2004) Control of Vibrio spp. in shrimp hatcheries using the green algae Tetraselmis suecica. Asian Fish Sci 17:147–158
Riclea R, Gleitzmann J, Bruns H, Junker C, Schulz B, Dickschat JS (2012) Algicidal lactones from the marine Roseobacter clade bacterium Ruegeria pomeroyi. Beilstein J Org Chem 8:941–950
Riquelme CE, Avendaño-Herrera RE (2003) Microalgae and bacteria interaction in the aquatic environment and their potential use in aquaculture. Rev Chil Hist Nat 76:725–736
Sapp M, Schwaderer AS, Wiltshire KH, Hoppe H-G, Gerdts G, Wichels A (2007) Species-specific bacterial communities in the phycosphere of microalgae? Microbial Ecol 53:683–699
Schäfer H, Abbas B, Witte H, Muyzer G (2002) Genetic diversity of ‘satellite’ bacteria present in cultures of marine diatoms. FEMS Microbiol Ecol 42:25–35
Seyedsayamdost MR, Case RJ, Kolter R, Clardy J (2011) The Jekyll-and-Hyde chemistry of Phaeobacter gallaeciensis. Nat Chem 3:331–335
Sharifah EN, Eguchi M (2011) The phytoplankton Nannochloropsis oculata enhances the ability of Roseobacter clade bacteria to inhibit the growth of fish pathogen Vibrio anguillarum. PLoS One 6:e26756
Sison-Magnus MP, Jiang S, Tran KN, Kudela RM (2014) Host-specific adaptation governs the interaction of the marine diatom, Pseudo-nitzschia and their microbiota. ISME J 8:63–76
Su J, Yang X, Zhou Y, Zheng T (2011) Marine bacteria antagonistic to the harmful algal bloom species Alexandrium tamarense (Dinophyceae). Biol Control 56:132–138
Sule P, Belas R (2013) A novel inducer of Roseobacter motility is also a disruptor of algal symbiosis. J Bacteriol 195:637–646
Suzuki T, Muroga Y, Takahama M, Nishimura Y (1999) Roseivivax halodurans gen. nov., sp. nov and Roseivivax halotolerans sp. nov, aerobic bacteriochlorophyll-containing bacteria isolated from a saline lake. Int J Syst Bacteriol 49:629–634
Tatti E, Decorosi F, Viti C, Giovannetti L (2012) Despite long-term compost amendment seasonal changes are main drivers of soil fungal and bacterial population dynamics in a Tuscan vineyard. Geomicrobiol J 29:506–519
Teplitski M, Chen H, Rajamani S, Gao M, Merighi M, Sayre RT, Robinson JB, Rolfe BG, Bauer WD (2004) Chlamydomonas reinhardtii secretes compounds that mimic bacterial signals and interfere with quorum sensing regulation in bacteria. Plant Physiol 134:137–146
Thornton DCO (2014) Dissolved organic matter (DOM) release by phytoplankton in the contemporary and future ocean. Eur J Phycol 49:20–46
Tosteson TR, Ballantine DL, Tosteson CG, Hansley V, Bardales AT (1989) Associated bacterial flora, growth, and toxicity of cultured benthic dinoflagellates Ostreopsis lenticularis and Gambierdiscus toxicus. Appl Environ Microb 55:137–141
Tredici MR (2004) Mass production of microalgae: photobioreactors. In: Richmond A (ed) Handbook of microalgal culture: biotechnology and applied phycology. Blackwell Science, Oxford, pp 178–214
Tredici MR, Rodolfi L (2004) Reactor for industrial culture of photosynthetic micro-organisms. PCT Patent WO2004/074423 [2 Sept 2004].
Tredici MR, Biondi N, Chini Zittelli G, Ponis E, Rodolfi L (2009) Advances in microalgal culture for aquaculture feed and other uses. In: Burnell G, Allan G (eds) New technologies in aquaculture: improving production efficiency, quality and environmental management. Woodhead Publishing, Cambridge, pp 610–676
Tulli F, Chini Zittelli G, Giorgi G, Poli BM, Tibaldi E, Tredici MR (2012) Effect of the inclusion of dried Tetraselmis suecica on growth, feed utilization, and fillet composition of European sea bass juveniles fed organic diets. J Aquat Food Prod Tech 21:188–197
Yao C, Ai J, Cao X, Xue S, Zhang W (2012) Enhancing starch production of a marine green microalga Tetraselmis subcordiformis through nutrient limitation. Bioresour Technol 118:438–444
Yoon J, Yasumoto-Hirose M, Matsuo Y, Nozawa M, Matsuda S, Kasai H, Yokota A (2007) Pelagicoccus mobilis gen. nov., sp. nov., Pelagicoccus albus sp. nov. and Pelagicoccus litoralis sp. nov., three novel members of subdivision 4 within the phylum ‘Verrucomicrobia’, isolated from seawater by in situ cultivation. Int J Syst Evol Micr 57:1377–1385
Yurkov VV, Beatty JT (1998) Aerobic anoxygenic phototrophic bacteria. Microbiol Mol Biol Rev 62:695–724
Zhang Z, Chen Y, Fu Y, Jiao N (2014) Draft genome sequence of the novel exopolysaccharide-producing bacterium Altibacter lentus strain JLT2010 T, isolated from deep seawater of the South China Sea. Genome Announc 2:e00954–14
Acknowledgments
The authors are indebted to Dr Graziella Chini Zittelli of the Intitute of Ecosystem Study, CNR, Sesto Fiorentino, Florence, Italy, for providing the algal culture samples from which the isolation of the bacteria was carried out.
Author information
Authors and Affiliations
Corresponding author
Additional information
Natascia Biondi and Giulia Cheloni contributed equally to this work.
Rights and permissions
About this article
Cite this article
Biondi, N., Cheloni, G., Tatti, E. et al. The bacterial community associated with Tetraselmis suecica outdoor mass cultures. J Appl Phycol 29, 67–78 (2017). https://doi.org/10.1007/s10811-016-0966-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-016-0966-5