Abstract
Acute pancreatitis (AP) is one of the most common gastrointestinal emergencies, often resulting in self-digestion, edema, hemorrhage, and even necrosis of pancreatic tissue. When AP progresses to severe acute pancreatitis (SAP), it often causes multi-organ damage, leading to a high mortality rate. However, the molecular mechanisms underlying SAP-mediated organ damage remain unclear. This study aims to systematically mine SAP data from public databases and combine experimental validation to identify key molecules involved in multi-organ damage caused by SAP. Retrieve transcriptomic data of mice pancreatic tissue for AP, lung and liver tissue for SAP, and corresponding normal tissue from the Gene Expression Omnibus (GEO) database. Conduct gene differential analysis using Limma and DEseq2 methods. Perform enrichment analysis using the clusterProfiler package in R software. Score immune cells and immune status in various organs using single-sample gene set enrichment analysis (ssGSEA). Evaluate mRNA expression levels of core genes using reverse transcription-polymerase chain reaction (RT-PCR) and immunohistochemistry. Validate serum amylase, TNF-α, IL-1β, and IL-6 levels in peripheral blood using enzyme-linked immunosorbent assay (ELISA), and detect the formation of neutrophil extracellular traps (NETs) in mice pancreatic, liver, and lung tissues using immunofluorescence. Differential analysis reveals that 46 genes exhibit expression dysregulation in mice pancreatic tissue for AP, liver and lung tissue for SAP, as well as peripheral blood in humans. Functional enrichment analysis indicates that these genes are primarily associated with neutrophil-related biological processes. ROC curve analysis indicates that 12 neutrophil-related genes have diagnostic potential for SAP. Immune infiltration analysis reveals high neutrophil infiltration in various organs affected by SAP. Single-cell sequencing analysis shows that these genes are predominantly expressed in neutrophils and macrophages. FPR1, ITGAM, and C5AR1 are identified as key genes involved in the formation of NETs and activation of neutrophils. qPCR and IHC results demonstrate upregulation of FPR1, ITGAM, and C5AR1 expression in pancreatic, liver, and lung tissues of mice with SAP. Immunofluorescence staining shows increased levels of neutrophils and NETs in SAP mice. Inhibition of NETs formation can alleviate the severity of SAP as well as the levels of inflammation in the liver and lung tissues. This study identified key genes involved in the formation of NETs, namely FPR1, ITGAM, and C5AR1, which are upregulated during multi-organ damage in SAP. Inhibition of NETs release effectively reduces the systemic inflammatory response and liver-lung damage in SAP. This research provides new therapeutic targets for the multi-organ damage associated with SAP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
AP is a common inflammatory disease of the exocrine pancreas, characterized by acute inflammation of the pancreas and histological destruction of acinar cells. It is one of the common gastrointestinal emergencies. In recent years, the global incidence of pancreatitis has been on the rise, with a global incidence of 30–40 cases per 100,000 population annually [1]. Approximately 80% of patients present with mild acute pancreatitis (MAP), which is relatively mild and can be rapidly improved with appropriate fluid resuscitation, pain management, and early enteral nutrition. However, about 20% of patients will progress from local involvement to systemic organ and system involvement, becoming SAP [2]. SAP is characterized by critical illness, multiple complications, and a high mortality rate, with hospital mortality rates ranging from 13% to 35% [3]. However, the mechanism of multi-organ damage associated with SAP remains unclear. Therefore, understanding its epidemiology, risk factors, pathophysiology, and potential mediators is crucial for the prognosis and management of AP patients.
Acute lung injury (ALI) is the earliest and most common complication of SAP. It is characterized by diffuse damage to pulmonary microvascular endothelial cells and alveolar epithelial cells, increased permeability of the microvascular basement membrane, and exudation of protein-rich edema fluid into the interstitium or alveoli. Its main clinical manifestations are refractory hypoxemia and respiratory failure, with a mortality rate of approximately 60%–70%, making it one of the leading causes of death in SAP patients [4,5,6,7]. However, the mechanism by which severe pancreatitis leads to ALI remains unclear, and effective interventions to mitigate lung injury are lacking. Additionally, due to the blood leaving the pancreas being processed by the liver before returning to the heart, the liver often suffers damage from extra-pancreatic organs. As early as 1984, Blamey et al. reported liver damage in 80% of AP patients, with the severity of liver damage positively correlated with the progression of AP [8]. Clinically, liver damage is an important indicator of the severity of AP and has significant prognostic value for AP [9, 10]. In SAP models, elevated levels of serum alanine aminotransferase and aspartate aminotransferase have long been considered evidence of liver damage. Although the liver has compensatory function, liver damage remains a serious, and even fatal, complication in the development of severe pancreatitis.
Neutrophils serve as the frontline defense against microbial pathogens, protecting the body from invasion. However, excessive activation of neutrophils can also mediate tissue damage and sterile inflammation. Recent studies have found that neutrophils play a central role in the development of SAP. In the early stages of SAP, the inflammatory response is primarily mediated by neutrophils [11]. During SAP, pancreatic cell damage leads to abnormal activation of pancreatic proteases, which generate sterile inflammatory signals recruiting neutrophils to the pancreas and releasing cytokines and chemokines. Meanwhile, activated neutrophils prolong their lifespan and release reactive oxygen species (ROS) and cytotoxic substances, further exacerbating local pancreatic injury [12]. As inflammation persists, neutrophils undergo a cascade reaction of transendothelial migration, leading to capillary obstruction and microthrombus formation, resulting in local tissue necrosis. This exacerbates local pancreatic inflammation into systemic inflammatory response syndrome (SIRS) and, through overwhelming inflammatory responses, causes distant organ damage and multiple organ dysfunction syndrome (MODS). However, the mechanisms by which neutrophils cause tissue damage require further investigation.
In this study, through analysis of bulk transcriptomic data and single-cell RNA sequencing (scRNAseq) data during acute pancreatitisand SAP, neutrophils are implicated in tissue damage via the formation of NETs. The study also explores the relationship between NETs formation during AP and liver-lung damage, predicts potential traditional Chinese medicine and drug targets, and provides new insights for improving the prognosis of AP and the treatment of associated liver-lung damage.
Method
Data Collection and Processing
Downloaded transcriptomic data from two AP datasets, GSE109227 and GSE65146, from the GEO database; transcriptomic data related to SAP lung injury from dataset GSE151572; transcriptomic data related to SAP liver injury from dataset GSE151927; peripheral blood transcriptomic data during human AP from dataset GSE194331. According to the corresponding data sets, tissue samples were obtained within 24 hours post modeling in mice for GSE109227, GSE65146, GSE151572, and GSE151927. Blood samples were obtained within 24 hours after hospital admission for patients in GSE194331. To increase sample size, merged and batch effects were removed for GSE109227 and GSE65146 using the “Combat” package in R software. Dataset GSE181276 contained single-cell transcriptomic data from pancreatic cells of control mice and mice with AP at 1, 7, and 28 days; dataset GSE198183 contained single-cell transcriptomic data from pancreatic cells of control mice and mice with AP at 2 days and 6 weeks. Control mice from GSE181276 at day 1 of AP and control mice from GSE198183 at day 2 of AP were included as single-cell transcriptomic data for this study. The standard workflow of scRNA-seq data analysis was performed using the “Seurat” package in R. Cells with fewer than 200 or more than 6000 genes, as well as those with mitochondrial gene expression exceeding 15%, were filtered out. The “harmony” package in R was used to reduce batch effects between samples. The “FindVariableFeatures” function was employed to identify the top 3000 variably expressed genes. Cell subtypes were identified by comparing with marker genes from the CellMarker2.0 database.
Differential Analysis
For microarray data, differential analysis was conducted using the “Limma” package in R software. For raw sequencing data, differential analysis was performed using the “DESeq2” package in R software. Differential genes were defined with a threshold of Fold Change >1.5 and FDR < 0.05.
Homologous ID Conversion
Perform homologous ID conversion for mice-origin genes to match human-origin genes using the g:Profiler online tool (https://biit.cs.ut.ee/gprofiler/orth).
Functional Enrichment Analysis
Perform enrichment analysis using the “clusterProfiler” package in R software. This includes enrichment analysis for Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and Reactome pathways, with a corrected p value <0.05 considered significant enrichment.
Protein-Protein Interaction Network and Identification of Core Genes
The STRING database (https://string-db.org/) integrates all experimentally validated or predicted protein-protein interaction relationships [13]. Protein interaction analysis is conducted using the STRING database with a Confidence Score > 0.4 threshold. The MOCODE plugin in Cytoscape 3.8.0 software is utilized for modular analysis of the protein-protein interaction (PPI) network derived from the STRING database. The Cytohubba plugin in Cytoscape 3.8.0 software is employed to extract core genes from the PPI network using algorithms such as MCC, Degree, MNC, EPC, and Closeness.
Immune Cell Infiltration and Immune Scoring
Calculate the immune cell infiltration scores and immune status scores in each tissue using the ssGSEA algorithm.
Gene-Drug Network
The relationship between traditional Chinese medicine monomers and genes is established using the HERB database [14], and visualization is performed using Cytoscape.
Animal Husbandry and Intervention in Experiments
All experimental procedures were approved by the Ethics Committee of the First Affiliated Hospital of Harbin Medical University. The mice were housed under controlled conditions with a temperature of 18–24 °C and humidity of 50–60%, with a 12-hour light-dark cycle. They were provided with ad libitum access to food and water. Mice were allowed one week to acclimate to the new environment before initiating the dietary intervention, which lasted for 8 weeks. Following random allocation principles, mice were divided into the following groups: control group (NC group), mild acute pancreatitis group (MAP group), severe acute pancreatitis group (SAP group), SAP+Neutrophil depletion group (SAP+Anti-Ly6g group) and SAP+NETs depletion group (SAP+DNase I group). For AP modeling, a physiological saline solution containing 5 μg/ml cerulein was administered via intraperitoneal injection at a dose of 10 ml/kg every hour for 10 consecutive doses, and the mice were euthanized 24 hours later. For SAP modeling, a sterile solution of 8% L-arginine salt (A92600, MiliporeSigma, Burlington, MA) in physiological saline was prepared and adjusted to pH 7.0. Mice received intraperitoneal injections of L-arginine (4 g/kg) every hour, while the control group received intraperitoneal injections of physiological saline [15,16,17]. The neutrophils in mice were depleted using anti-Ly6g antibody (Bio X Cell, clone 1A8, #BP0075–1), specifically by intraperitoneal injection of the antibody at a dosage of 100μg per mouse once every 48 hours for one week prior to euthanasia. DNase I (Merck) was used to inhibit the formation of NETs, administered by intraperitoneal injection at a dosage of 5 mg/kg one day before establishing the AP model in mice.
Real-Time Fluorescence Quantitative PCR
Total RNA was extracted from tissues or cells using the RNA extraction kit from Axygen Scientific Inc. (Silicon Valley, USA). The extracted RNA was then reverse transcribed into cDNA using the Toyobo Reverse Transcription Kit. SYBR GREEN reagent was employed to detect the expression of target genes. RT-qPCR samples were preheated at 95 °C for 10 minutes, followed by 40 cycles of denaturation at 95 °C for 15 seconds and annealing/extension at 60 °C for 1 minute. GAPDH was used as an internal reference gene. Data were analyzed using the 2^-ΔΔCt method. Supplementary Table 1 provides details of the primer information.
ELISA
Peripheral blood was collected using the mice fundus vein puncture method. The collected blood was centrifuged at 3000 rpm for 15 minutes at 4 °C, and the supernatant was collected for enzyme-linked immunosorbent assay (ELISA) analysis. The levels of IL-1β, IL-6, and TNF-α were analyzed according to the manufacturer’s instructions using ELISA kits purchased from Elabscience Biotech Co. (Wuhan, China).
Serum amylase activity was determined using a commercial amylase assay kit. In brief, serum samples were added to wells containing known concentrations of excess substrate (starch). Amylase in the samples hydrolyzed starch, and then iodine was added to react with the unhydrolyzed starch, producing a blue-colored compound. By measuring the absorbance of the blue-colored compound at 660 nm, the amount of starch hydrolyzed by amylase could be inferred, and thus the amylase activity could be calculated. The unit is expressed as U/L.
Hematoxylin and Eosin Staining
The mice pancreas, liver, and lung tissue specimens were fixed in 4% paraformaldehyde solution, dehydrated in a series of alcohol gradients, embedded in paraffin, and then sectioned into continuous slices of 5 μm thickness. The tissue sections were stained with hematoxylin and eosin (H&E), and the tissue morphology was observed using an optical microscope.
Immunohistochemistry
The freshly collected pancreas and liver-lung tissues were fixed overnight in 4% paraformaldehyde, dehydrated, and embedded in paraffin. The paraffin-embedded samples were then sectioned into 4 μm thick slices. Antigen retrieval was performed in a pressure cooker for 2 minutes using Tris-EDTA buffer (pH = 9). Endogenous peroxidase activity was blocked with 3% hydrogen peroxide. The sections were then incubated with 10% goat serum at 37 °C for 30 minutes to block nonspecific binding. Subsequently, the sections were incubated overnight at 4 °C with primary antibodies against ITGAM (SANTA; sc-1186; 1:200), ITGAM FPR1 (SANTA; sc-53,795; 1:200), and FPR1 (abcam;ab113531;1:200). After washing, the sections were incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies at 37 °C for 30 minutes. Finally, the sections were incubated in Diaminobenzidine (DAB) for 5–10 minutes to develop a stable color.
Immunofluorescence
After antigen retrieval in citrate buffer (Hangzhou Hulk Biotechnology Co., Ltd., Hangzhou, China, HK1222) using microwave, pancreas and liver-lung tissue sections were blocked with 5% BSA (Beyotime, Jiangsu, China, ST2254) for 30 minutes. The sections were then incubated overnight with primary antibodies at 4 °C, followed by three washes with PBS and addition of secondary antibodies. Nuclear staining was performed with 4′,6-diamidino-2-phenylindole (DAPI). Images were captured using a microscope (Leica, Wetzlar, Germany, DM2500) and captured by Pannoramic SCAN II (3D HISTECH) and fluorescence microscope (LAS X software, Leica, Wetzlar, Germany).
The antibodies used were as follows: Anti-CitH3 (Abcam, ab281584, 1:100), Anti-MPO (Abcam, ab300650, 1:100), Anti-Ly6g (Abcam, ab238132, 1:100), Fluorescein (FITC)-conjugated Affinipure Goat Anti-Rat IgG(H + L) (Proteintech, SA00003–11,1:100), Fluorescein (FITC)-conjugated Affinipure Rabbit Anti-Goat IgG(H + L) (Proteintech, SA00003–4, 1:100).
Statistical Analysis
For the bioinformatics analysis, statistical analysis was conducted using R software (version 4.0.2). For experimental validation, differential analysis was performed using GraphPad Prism 8. Continuous variables between two groups were compared using either the t-test or Mann–Whitney U test, depending on whether the data followed a normal distribution. For comparisons among three groups of samples, the Kruskal-Wallis test was used. Spearman correlation analysis was used for all correlation analyses. The diagnostic ability of molecules was reflected by receiver operating characteristic (ROC) curves. For all statistical analyses, a significance level of P < 0.05 was considered statistically significant (*, P < 0.05; **, P < 0.01; ***, P < 0.001).
Result
Data Processing and Hub Gene Selection
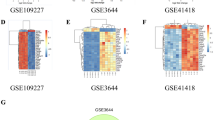
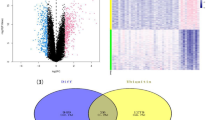
The GSE65146 and GSE109227 datasets were merged and batch effects were successfully removed (Supplementary Fig. 1A-B). Differential analysis of mice pancreatic transcriptome data revealed 3264 upregulated and 1905 downregulated genes in AP pancreatic tissue compared to normal pancreatic tissue (Supplementary Fig. 2A). Compared to normal lung tissue, SAP lung tissue had 651 upregulated and 471 downregulated genes (Supplementary Fig. 2B). Compared to normal liver tissue, SAP liver tissue had 4375 upregulated and 2716 downregulated genes (Supplementary Fig. 2C). There were 106 genes simultaneously upregulated (Supplementary Fig. 2D) and 14 genes simultaneously downregulated (Supplementary Fig. 2E) in pancreatic, lung, and liver tissues. Compared to normal human blood, AP patient peripheral blood had 3149 upregulated and 1768 downregulated genes (Supplementary Fig. 2F). After converting the selected mice-origin genes to human-origin genes in the three groups (pancreas, liver, and lung) and taking the intersection with the differentially expressed genes identified in blood, 45 upregulated and 1 downregulated gene were obtained (Supplementary Fig. 2G-H). Functional enrichment analysis of these differentially expressed genes revealed that they were mainly associated with MAPK, NF-κB, IL-17, cytokine-cytokine receptor interaction pathways based on KEGG enrichment analysis (Supplementary Fig. 3A). Reactome enrichment analysis indicated that these genes were mainly related to IL-4/IL-13, IL-10, IL-1, TOLL-like receptor signaling pathways (Supplementary Fig. 3B). GO-BP enrichment analysis showed that these genes were mainly associated with neutrophil-mediated immunity, neutrophil activation, neutrophil degranulation, and neutrophil migration (Supplementary Fig. 3C). Through the analysis of the results of the three enrichment analyses, it was found that these genes mainly enriched in neutrophil-related biological processes (Supplementary Fig. 3C). Given the important role of neutrophils in inflammation and organ damage associated with SAP [18, 19], we decided to select 12 genes (including C5AR1, CD14, CLEC4D, FPR1, ITGAM, LCN2, LRG1, MMP8, PLAUR, PTPRJ, S100A8, and STOM) out of the 46 genes that could enrich in neutrophil-related processes for further analysis.
Differential Genes Related to Neutrophils Have Diagnostic Potential for Moderate to Severe Pancreatitis
By analyzing the expression of the aforementioned 12 neutrophil-related genes in mild and moderate-severe acute pancreatitis, it was found that compared to normal peripheral blood, the expression of these 12 genes was significantly increased in peripheral blood of AP patients (Supplementary Fig. 4A). Among them, ITGAM, LRG1, and S100A8 showed a significant increase in expression in peripheral blood with the severity of AP (Supplementary Fig. 4A). To establish the potential of these genes in diagnosing mild and moderate-severe AP, ROC curve analysis was performed. The results showed that among the 12 genes, LCN2 (AUC = 0.718), MMP8 (AUC = 0.723), S100A8 (AUC = 0.702), and STOM (AUC = 0.713) performed well in distinguishing between mild and moderate-severe AP and had diagnostic significance (Supplementary Fig. 4B). To further improve diagnostic efficacy, ROC curve analysis was performed for the combination of LCN2, MMP8, S100A8, and STOM. The results showed that the combined AUC value of LCN2, MMP8, S100A8, and STOM in distinguishing between mild and moderate-severe AP was 0.770, further enhancing the diagnostic potential for mild and moderate-severe AP (Supplementary Fig. 4C). Additionally, a protein-protein interaction (PPI) network and a disease-drug-target gene network were constructed, and potential therapeutic Chinese medicines were screened (Supplementary Fig. 5A-B).
AP Can Lead to Multiorgan Immune Dysregulation
The ssGSEA algorithm can assess the levels of immune cells and immune status in organs based on transcriptomic data. We used the ssGSEA algorithm to score the immune cells and immune status in the pancreas and peripheral blood during AP, as well as in the liver and lungs during SAP. We found that compared to the normal pancreas, the infiltration levels of B cells, macrophages, neutrophils, pDCs, Th1, and Th2 cells were significantly increased in the pancreas of the AP group, while the levels of CD8+ T cells, mast cells, and helper T cells were significantly decreased (Fig. 1A). In addition, the inflammatory score, type I IFN response, and type II IFN response scores were higher in the pancreas of the AP group compared to the normal pancreas (Fig. 1B). Compared to normal peripheral blood, the scores of neutrophils and macrophages were significantly increased in the blood of AP patients, while the scores of other cells such as B cells and T cells were significantly decreased (Fig. 1C). Surprisingly, the inflammatory score was decreased in the blood of AP patients compared to normal blood (Fig. 1D). Compared to the normal liver, the levels of macrophages, Tfh cells, Treg cells, inflammatory score, and auxiliary inflammation score were significantly increased in the liver of the SAP group, while pDCs were decreased (Fig. 1E and F). Similarly, compared to the normal lungs, the levels of macrophages, neutrophils, Treg cells, and type II IFN response scores were significantly increased in the lungs of the SAP group, while CD8+ T cell infiltration was decreased (Fig. 1G and H), suggesting immune dysregulation in multiple organs during AP.
Immune cells and immune status scores of each tissue. (A-B) pancreatic immune cell and immune status score. (C-D) peripheral blood immune cells and immune status score. (E-F) liver immune cells and immune status score of severe pancreatitis. (G-H) pulmonary immune cells and immune status scores in severe pancreatitis. *P < 0.05 **P < 0.01 ***P < 0.001
The Correlation between the Expression Levels of the 12 Genes in Various Organs and the Infiltration of Immune Cells and Immune Status Scores
We analyzed the correlation between the expression levels of these 12 genes in various organs and the infiltration of immune cells and immune status scores. It is evident that in the pancreas, peripheral blood, liver, and lung tissues, the expression of hub genes is significantly positively correlated with neutrophil and macrophage infiltration (Fig. 2A-D). Additionally, in pancreatic, liver, and lung tissues, the expression of hub genes is positively correlated with auxiliary inflammatory response scores and type II IFN response scores (Fig. 2A, C-D). Surprisingly, in peripheral blood tissue, hub genes are only positively correlated with the APC co-inhibition response score, macrophages, and neutrophils, while they are negatively correlated with almost all other immune cells and immune status scores (Fig. 2B). Considering the results of previous enrichment analyses, it can be speculated that hub genes may play an intermediary role in highly infiltrated neutrophils and macrophages during SAP-induced damage to other organs.
Correlation between the expression levels of 12 genes in each organ and immune cell infiltration and immune status score. (A) Pancreatic immune cell infiltration and immune status score; (B) peripheral blood immune cell infiltration and immune status score; (C) liver immune cell infiltration and immune status score; (D) pulmonary immune cell infiltration and immune status score. *P < 0.05 **P < 0.01 ***P < 0.001
The Expression of the 12 Neutrophil-Related Genes at the Single-Cell Level
To better understand the expression of these 12 genes at the single-cell level, we conducted scRNA-seq analysis in a mice AP model (Fig. 3A). We identified a total of 27 cell subtypes in four samples from the control group and the AP group (Fig. 3B). By comparing these 27 cell subtypes with the marker genes of various cell types in mice pancreatic tissue from the CellMarker database, we found that the 27 cell subtypes could be classified into 12 cell types, including acinar cells, fibroblasts, ductal cells, endothelial cells, macrophages, B cells, monocytes, neutrophils, beta cells, dendritic cells, and peptide cells (Fig. 3C). Figure 3D shows the top 5 marker genes for each of the 12 cell types.
Subsequently, we explored and compared the expression of these 12 hub genes across the 12 cell types (Fig. 4). The results showed that C5AR1, CD14, CLEC4D, PLAUR, ITGAM, FPR1, PTPRJ, and S100A8 had the highest expression levels in neutrophils, followed by macrophages (Fig. 4A-H). On the other hand, LRG1 and STOM exhibited the highest expression levels in endothelial cells, while LCN2 was predominantly expressed in ductal cells, and MMP8 was mainly expressed in macrophages (Fig. 4I-L).
Through integrated analysis with the KEGG pathway database, we found that among these 12 genes, FPR1, ITGAM, and C5AR1 were simultaneously identified as key genes involved in the formation of NETs (Supplementary Fig. 5). NETs play a crucial role in mediating the development of AP [9, 20], but their involvement in the multi-organ damage observed in SAP remains unclear [10, 21]. Therefore, we decided to further investigate FPR1, ITGAM, and C5AR1 as targets for experimental validation.
FPR1, ITGAM, and C5AR1 Exhibit Increased Expression in AP Tissues
To validate whether FPR1, ITGAM, and C5AR1 mediate multi-organ damage caused by SAP, we first verified whether they were upregulated in various organs of SAP. We used C57BL/6 mice to establish models of MAP and SAP. HE staining results showed significant inflammation in pancreatic tissues of MAP and SAP mice compared to the control group, with a significant increase in pathological scores and serum amylase levels (Fig. 5A). In contrast, significant inflammatory damage was observed in the lung and liver tissues of the SAP group, while no significant inflammation was observed in the lung and liver tissues of the AP group (Fig. 5A). Compared to the control group, MAP mice showed elevated levels of serum amylase, TNF-α, IL-1β, and IL-6, and these indicators were further increased in the serum of SAP mice (Fig. 5B-E).
Expression levels of FPR1, ITGAM and C5AR1 as key genes in the formation of NETs in SAP organs. (A) Pancreatic tissue inflammation levels, pathological scores and serum amylase levels in MAP and SAP mice. (B-E) serum amylase, TNF-α, IL-1β and IL-6 levels in MAP and SAP mice; (F) mRNA expression levels of FPR1,ITGAM and C5AR1 in pancreas in MAP group; (G) mRNA expression levels of FPR1,ITGAM and C5AR1 in lung tissue in MAP group; (H) mRNA expression levels of FPR1,ITGAM and C5AR1 in liver tissue in MAP group; (I-K) expressions of FPR1,ITGAM and C5AR1 in three tissues by IHC; (L) expressions of Ly6g in three tissues by immunofluorescence, *P < 0.05 **P < 0.01 ***P < 0.001
Subsequently, we validated the expression of FPR1, ITGAM, and C5AR1 in the mice models using qPCR. In pancreatic tissues, mRNA expression of FPR1, ITGAM, and C5AR1 was significantly increased in the MAP group compared to the control group, and this trend was more significant in the SAP group (Fig. 5F). In lung tissues, mRNA expression of FPR1, ITGAM, and C5AR1 was significantly increased in the SAP group compared to the control group, while no significant increase was observed in the MAP group (Fig. 5G). In liver tissues, mRNA expression of FPR1, ITGAM, and C5AR1 was significantly increased in the SAP group compared to the control group, while no significant increase was observed in the MAP group (Fig. 5H). Subsequently, we further validated the differential expression of FPR1, ITGAM, and C5AR1 at the protein level using immunohistochemistry, and the results were consistent with those obtained by qPCR (Fig. 5I-K).
Neutrophiles Increase during AP and Mediate SAP-Related Multi-Organ Damage
To investigate the role of neutrophils in the multi-organ damage associated with SAP, we examined the expression of the neutrophil-specific marker Ly6g through immunofluorescence. The results indicated that, compared to the control group, there was an increase in neutrophils in the pancreas of the MAP group, and this phenomenon was even more significant in SAP (Fig. 5L). In lung tissue, compared to the control and MAP groups, the infiltration of neutrophils was significantly increased in the SAP group, although there was no significant difference between the control and MAP groups (Fig. 5L). Similarly, in liver tissue, the infiltration of neutrophils was significantly increased in the SAP group compared to the control and MAP groups (Fig. 5L).
Subsequently, we depleted neutrophils in mice using an anti-Ly6g antibody, and then established an SAP model to explore the role of neutrophils in SAP-associated multi-organ damage. Immunofluorescence results showed that the infiltration of neutrophils in the pancreas, lungs, and liver of SAP mice was increased, while the anti-Ly6g antibody successfully depleted neutrophils in these organs (Fig. 6A). ELISA results showed that treatment with the anti-Ly6g antibody reduced levels of blood amylase, IL-1β, IL-6, and TNF-α in the peripheral blood of SAP mice, indicating a reduction in systemic inflammation (Fig. 6B-E). H&E staining showed that treatment with the anti-Ly6g antibody reduced the level of tissue damage in the pancreas, lungs, and liver of SAP mice (Fig. 6F).
Neutrophil-mediated multi-organ damage in SAP. (A) Immunofluorescence demonstrated that treatment with the anti-Ly6g antibody reduced neutrophil infiltration in the pancreas, lungs, and liver; (B-E) serum amylase, IL-1β, IL-6, and TNF-α levels in mice treated with the anti-Ly6g antibody were verified by ELISA; (F) H&E staining showed that treatment with the anti-Ly6g antibody alleviated multi-organ damage associated with SAP
NETs Increase during AP and Mediate SAP-Related Multi-Organ Damage
Subsequently, we performed immunofluorescence confocal microscopy to detect the formation of NETs. The results showed that compared to the control group, both MAP and SAP groups exhibited NETs in pancreatic tissues (Fig. 7A). In lung tissues, NETs were not detected in the control and MAP groups, while there was a significant increase in NETs in the lung tissues of the SAP group (Fig. 7B). Similarly, in liver tissues, NETs were not detected in the control and MAP groups, while there was a significant increase in NETs in the liver tissues of the SAP group (Fig. 7C). This indicates that during SAP, there is a significant formation of NETs that contribute to multi-organ damage.
DNase I is a commonly used inhibitor of NETs. To determine whether NETs release is involved in mediating multi-organ damage in SAP, we inhibited the generation of NETs using DNase I and induced a mice SAP model. After applying DNase I, fluorescence confocal microscopy revealed a significant reduction in NETs in pancreatic, lung, and liver tissues of SAP mice, indicating a significant inhibition of NETs formation (Fig. 8A-C). Subsequently, we performed HE staining on liver and lung tissues of SAP mice treated with DNase I to study the severity of SAP and the extent of damage to the pancreas, lung, and liver. The results showed that compared to SAP mice without DNase I treatment, SAP mice treated with DNase I exhibited significantly reduced inflammatory responses and decreased damage to the pancreas, lung, and liver (Fig. 8D). After inhibiting NETs formation and release, the levels of serum amylase, TNF-α, IL-1β, and IL-6 in SAP mice were reduced, indicating a reduction in the severity of systemic inflammation (Fig. 8E-H).
Inflammatory response and NETs formation after NETs inhibitors. (A-C) NETs formation in pancreatic, lung and liver tissues after administration of NETs inhibitors. (D) H&E staining showed that treatment with the NETs inhibitors alleviated multi-organ damage associated with SAP. (E-H) serum amylase, TNF-α, IL-1β and IL-6 levels in mice treated with NETs inhibitors were verified by ELISA, *P < 0.05 **P < 0.01 ***P < 0.001
Discussion
AP leads to significant morbidity and mortality. Globally, the estimated incidence of AP is 33.74 cases per 100,000 person-years (95% CI 23.33–48.81), with a mortality rate due to AP of 1.60 cases per 100,000 person-years (95% CI 0.85–1.58) [22]. The severity of AP can be mild, moderate, or severe, depending on the extent of local pancreatic injury and, more importantly, systemic damage to organs distant from the pancreas [23]. MAP typically lacks obvious local or systemic complications. However, about 20% of AP patients present with a more severe form of the disease, characterized by significant local complications such as necrosis, often due to systemic damage caused by widespread inflammation. In this pathophysiological process, a large release of cytokines and inflammatory mediators activates multiple signaling pathways, causing damage to the body. However, the underlying mechanisms are not fully understood.
Previous studies have long believed that the excessive activation of pancreatic enzymes is the direct pathogenic mechanism of pancreatitis [24, 25]. However, in clinical practice, the condition of SAP patients treated with pancreatic enzyme inhibitors did not improve significantly. Therefore, there are still pathogenic mechanisms of AP independent of excessive pancreatic enzyme activation. In recent years, increasing evidence supports that inflammatory factors are crucial pathogenic mechanisms of pancreatitis independent of pancreatic enzyme activation [26]. The release of inflammatory signals from pancreatic acinar cells can mediate the recruitment and activation of circulating inflammatory cells (monocytes and neutrophils) [27, 28]. The excessive activation of neutrophils can trigger intense local and systemic inflammatory responses, leading to acute respiratory distress syndrome, cardiovascular failure, renal failure, and gastrointestinal bleeding, which are also the reasons for the high mortality rate of SAP [29]. Enrichment analysis revealed that differential genes are mainly associated with neutrophil-mediated immunity, activation, degranulation, migration, and other biological processes. Additionally, the analysis of the expression levels of 12 hub genes in various organs and their correlation with immune cell infiltration and immune status scores revealed a significant positive correlation between hub gene expression and neutrophils and macrophages in pancreatic, peripheral blood, liver, and lung tissues. Furthermore, increased neutrophil infiltration levels in pancreatic and lung tissues in the SAP group suggest that neutrophils participate in the immune dysregulation of multiple organs during SAP. During SAP, the release of TNF-α, IL-1, and IL-6 promotes neutrophil adhesion and migration, increases capillary permeability, exacerbates pancreatic damage, and SIRS [30,31,32]. Additionally, increased complement C3a in peripheral blood can promote neutrophil activation and infiltration, increasing vascular permeability [33]. Moreover, immune complexes, such as IC3b complement deposition on endothelial cell membranes, can serve as signaling mediators for neutrophils, facilitating their long-distance migration in tissues [34]. Although neutrophil recruitment is considered a nonspecific defensive response against invading microorganisms, excessive recruitment and activation of neutrophils may lead to the release of large amounts of pro-inflammatory mediators and ROS. Therefore, SAP patients may experience the progression of pancreatitis and multiple organ failure due to the continuous exacerbation of systemic inflammatory cascades [35].
Under the action of inflammatory mediators and chemotactic factors, neutrophils are the first cells to migrate to the pancreas, releasing inflammatory mediators and triggering local inflammatory reactions [36]. However, neutrophil infiltration into tissues often comes at the expense of damaging host cells, and the level of neutrophil infiltration can reflect the extent of tissue damage to some extent. Under the stimulation of various inflammatory factors, activated neutrophils can release a meshwork of DNA fibers called NETs, which are packed with intracellular substances such as elastase, tissue protease G, and MPO to protect the host from inflammatory damage [37, 38].
Our research suggests that neutrophils and NETs may play significant roles in the occurrence and development of SAP. In mouse models of pancreatic inflammation, significant improvements in tissue damage were observed by inhibiting neutrophil infiltration and NETs generation. Inhibiting NETs can reduce the release of CXCL2 in the pancreas and lungs and the recruitment of neutrophils, a finding worth noting as it suggests that NETs themselves may act as chemoattractants in the form of DAMPs or stimulate the release of chemotactic factors, thereby exacerbating the inflammatory response [39]. This indicates that NETs may act as a double-edged sword in inflammation, with their formation inhibiting the inflammatory response to some extent but amplifying it in certain circumstances, worsening tissue damage. For example, neutrophils accumulated in the pancreas can retrogradely migrate to the circulatory system, causing systemic and local complications of SAP such as multiple organ failure, thrombosis, IPN, and sepsis. Research has also found that NETs can regulate the activity of important molecular mediators such as STAT3 and the activation of pancreatic protease in pancreatic acinar cells during inflammation [39]. Additionally, IL-17A, primarily produced by activated T cells, can promote neutrophil accumulation in pancreatic ducts, while bicarbonate ions and calcium carbonate crystals in pancreatic juice, together with accumulated neutrophils, promote the formation of NETs. This, in turn, exacerbates SAP by blocking pancreatic ducts, leading to acute biliary pancreatitis [40].
At the level of amplifying the inflammatory response, NETs can activate inflammatory cells such as monocytes or macrophages by regulating the assembly of NLRP3 inflammasomes in neutrophils via PAD4 [41, 42], leading to the release of inflammatory factors such as IL-1β [43] and IL-18 [44]. Studies have found that NETs may also induce the secretion of pro-inflammatory chemokines IL-8 and B cell-activating factor (BAFF) through pathways involving Akt, ERK1/2, and p38 phosphorylation [45]. Extracellular AIM2-NET interaction may further promote sustained secretion of IFN-I, thereby amplifying the inflammatory response [46].
Aldabbous has demonstrated that MPO derived from NETs stimulates the activation, proliferation, and migration of endothelial cells in a model of pulmonary arterial hypertension [47]. Activated endothelial cells, in turn, promote NETosis via the IL-8 signal. This could potentially form a positive feedback loop of tissue damage and inflammation. Additionally, the negatively charged DNA within NETs, along with serine proteases and neutrophil elastase, can promote platelet aggregation. Through the NETs-platelet-coagulation axis, this promotes thrombus formation, thereby contributing to tissue damage and amplifying the inflammatory process [48]. Although NETs play a crucial role in inflammation, we must still recognize that the exacerbation of inflammation is the result of the combined action of multiple factors, including Netosis. Our findings also indicate that inhibiting the formation of NETs can only partially alleviate the severity of inflammation.
Through combined analysis, we identified genes associated with NETs, including C5AR1, FPR1, and ITGAM, as NETs are their common downstream. ITGAM encodes the α chain of integrin αM, an integrin that has been shown to be crucial for promoting the adhesion and transmembrane migration of neutrophils and monocytes to activated endothelial cells, primarily through ICAM-1 on the activated endothelial cells, thus directing these immune cells to migrate to sites of infection or inflammation [49]. Various integrins play a pathogenic role in MAP/SAP [50]. During acute and severe pancreatitis, the levels of ITGAM in the peripheral blood of patients are significantly increased [51, 52]. ITGAM can independently promote the generation of reactive oxygen species (ROS) through activating signaling pathways such as Src kinase, Syk, and PI3K δ, thereby facilitating the release of NETs [53]. C5aR1 was initially found in neutrophils, monocyte macrophages, and mast cells and is one of the most important components of the complement cascade with multiple pro-inflammatory effects. C5a/C5aR1 can directly trigger neutrophil activation but does not participate in myeloid cell infiltration into the lungs [50]. We studied its possible role in myeloid cell activation, focusing mainly on neutrophils. In Alzheimer’s disease, neutrophils can be attracted to amyloid plaques by several pro-inflammatory factors such as C5a, and release NETs [54]. Although these data suggest an association between C5a/C5aR1 and NETs formation, we cannot rule out the indirect effects of C5a on NETs formation. Indeed, evidence suggests that NETs can amplify the inflammatory process by promoting tissue damage and the production of additional cytokines/chemokines. NETs play a crucial role in the interaction between neutrophils and macrophages during the early acute phase of ALI. They promote ALI by facilitating macrophage polarization towards the M1 phenotype. The gene FPR1 encodes the G protein-coupled receptor expressed on macrophages, mediating their response to microbial invasion of the host. The macrophage-SCIMP-FPRs-neutrophil axis plays a critical role in the immune process of ALI [55]. Our research also confirms the upregulation of FPR1 expression in multiple organs during SAP.
Although this study found that NETs act as a common downstream effector promoting SAP-related inflammation for ITGAM, FPR1, and C5AR1, they do not solely exert their pro-inflammatory effects through NETs. During viral infections, extracellular dsRNA can activate many immune cells, including macrophages [56]. As a pattern recognition receptor (PRR) on the cell surface, ITGAM can detect extracellular dsRNA. Extracellular dsRNA enhances TLR3-dependent inflammatory oxidative signaling by activating ITGAM, and triggers inflammation signaling that is dependent on ITGAM but not on TLR3 [57]. FPR1 can recognize bacterial and host-derived N-formyl peptides, such as N-formylmethionyl-leucyl-phenylalanine (fMLF) released during bacterial infections. When FPR1 binds these ligands, it activates downstream inflammatory signaling pathways, promoting the activation of the NLRP3 inflammasome and the production of IL-1β, IL-6, or TNF-α [58, 59]. C5aR1 is often associated with sterile inflammation. When complement activation is enhanced, the binding of C5a to C5AR1 can cause contraction and increased permeability of vascular endothelial cells, thereby facilitating the ingress of inflammatory mediators and immune cells through the vascular wall into the interstitial tissue [60]. The activation of C5aR1 on endothelial cells induces an inflammatory state in the endothelium, while its activation on innate immune cells promotes antigen uptake, tissue infiltration, and the induction of a pro-inflammatory effector phenotype [61], together constituting chronic pathological inflammation.
Based on previous research findings and the discoveries of our study, we boldly speculate that during SAP, various damage-related molecular patterns activate neutrophils within the pancreas. Activated neutrophils release pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6, causing neutrophil migration to peripheral blood, lungs, and liver, as well as increased NETs. Subsequently, NETs further exacerbates the cascade of inflammation. Meanwhile, C5AR1, FPR1, and ITGAM participate in this process, exacerbating the inflammatory response through the elevation of NETs levels and other pathways. Therefore, targeting key therapeutic targets such as FPR1, ITGAM, and C5AR1 is likely to alleviate SAP and SAP-related ALI and AHI by inhibiting the activation of neutrophils and the formation of NETs. However, further experiments are needed to validate our speculation.
Conclusion
We found that the formation of NETs can exacerbate liver and lung damage associated with SAP, while inhibiting NETs release can effectively reduce the severity of systemic inflammatory responses and liver and lung injury during AP. Through screening and studying the core genes involved in NETs formation, we identified FPR1, ITGAM, and C5AR1 as closely related to NETs formation during SAP, potentially serving as common biomarkers among MAP/SAP, SAP-ALI, and SAP-AHI, laying the theoretical groundwork for future research. Additionally, by constructing a disease-drug-gene network, we predicted potential traditional Chinese medicines and drug targets, offering new avenues for the treatment of liver and lung damage associated with AP.
Data Availability
Data is provided within the manuscript or supplementary information files.
References
Petrov, M.S., and D. Yadav. 2019. Global epidemiology and holistic prevention of pancreatitis. Nature Reviews Gastroenterology & Hepatology 16: 175–184. https://doi.org/10.1038/s41575-018-0087-5.
Gardner, T.B. 2021. Acute Pancreatitis. Annals of Internal Medicine 174: Itc17–itc32. https://doi.org/10.7326/aitc202102160.
Trikudanathan, G., et al. 2019. Current concepts in severe acute and necrotizing pancreatitis: An evidence-based approach. Gastroenterology 156: 1994–2007.e1993. https://doi.org/10.1053/j.gastro.2019.01.269.
Zhu, C.J., et al. 2021. Calycosin attenuates severe acute pancreatitis-associated acute lung injury by curtailing high mobility group box 1 - induced inflammation. World Journal of Gastroenterology 27: 7669–7686. https://doi.org/10.3748/wjg.v27.i44.7669.
Liang, X.Y., T.X. Jia, and M. Zhang. 2021. Intestinal bacterial overgrowth in the early stage of severe acute pancreatitis is associated with acute respiratory distress syndrome. World Journal of Gastroenterology 27: 1643–1654. https://doi.org/10.3748/wjg.v27.i15.1643.
Xin, Y., et al. 2022. Advances in research on the effects of platelet activation in acute lung injury (review). Biomedical Reports 16: 17. https://doi.org/10.3892/br.2022.1500.
Wu, J., et al. 2021. Treatment of severe acute pancreatitis and related lung injury by targeting Gasdermin D-mediated Pyroptosis. Frontiers in Cell and Developmental Biology 9: 780142. https://doi.org/10.3389/fcell.2021.780142.
Blamey, S.L., C.W. Imrie, J. O'Neill, W.H. Gilmour, and D.C. Carter. 1984. Prognostic factors in acute pancreatitis. Gut 25: 1340–1346. https://doi.org/10.1136/gut.25.12.1340.
Liu, H.B., N.Q. Cui, D.H. Li, and C. Chen. 2006. Role of Kupffer cells in acute hemorrhagic necrotizing pancreatitis-associated lung injury of rats. World Journal of Gastroenterology 12: 403–407. https://doi.org/10.3748/wjg.v12.i3.403.
Gloor, B., et al. 2000. Kupffer cell blockade reduces hepatic and systemic cytokine levels and lung injury in hemorrhagic pancreatitis in rats. Pancreas 21: 414–420. https://doi.org/10.1097/00006676-200011000-00013.
Ke, L., et al. 2014. Predictors of critical acute pancreatitis: A prospective cohort study. Medicine 93: e108. https://doi.org/10.1097/md.0000000000000108.
Kolaczkowska, E., and P. Kubes. 2013. Neutrophil recruitment and function in health and inflammation. Nature Reviews. Immunology 13: 159–175. https://doi.org/10.1038/nri3399.
Szklarczyk, D., et al. 2021. Correction to 'The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets'. Nucleic Acids Research 49: 10800. https://doi.org/10.1093/nar/gkab835.
Fang, S., et al. 2021. HERB: A high-throughput experiment- and reference-guided database of traditional Chinese medicine. Nucleic Acids Research 49: D1197–D1206. https://doi.org/10.1093/nar/gkaa1063.
Long, L., et al. 2020. P-selectin-based dual-model Nanoprobe used for the specific and rapid visualization of early detection toward severe acute pancreatitis in vivo. ACS Biomaterials Science & Engineering 6: 5857–5865. https://doi.org/10.1021/acsbiomaterials.0c00596.
Liu, X., et al. 2018. Isoliquiritigenin ameliorates acute pancreatitis in mice via inhibition of oxidative stress and modulation of the Nrf2/HO-1 pathway. Oxidative Medicine and Cellular Longevity 2018: 7161592. https://doi.org/10.1155/2018/7161592.
Yang, J., X. Tang, Q. Wu, P. Ren, and Y. Yan. 2022. A severe acute pancreatitis mouse model transited from mild symptoms induced by a "two-hit" strategy with L-arginine. Life (Basel, Switzerland) 12. https://doi.org/10.3390/life12010126.
Tokoro, T., et al. 2020. Interactions between neutrophils and platelets in the progression of acute pancreatitis. Pancreas 49: 830–836. https://doi.org/10.1097/mpa.0000000000001585.
Zhang, D., et al. 2022. Novel insight on marker genes and pathogenic peripheral neutrophil subtypes in acute pancreatitis. Frontiers in Immunology 13: 964622. https://doi.org/10.3389/fimmu.2022.964622.
Gea-Sorlí, S., R. Guillamat, A. Serrano-Mollar, and D. Closa. 2011. Activation of lung macrophage subpopulations in experimental acute pancreatitis. The Journal of Pathology 223: 417–424. https://doi.org/10.1002/path.2814.
Zhu, L., et al. 2017. Pharmacokinetics and pharmacodynamics of Shengjiang decoction in rats with acute pancreatitis for protecting against multiple organ injury. World Journal of Gastroenterology 23: 8169–8181. https://doi.org/10.3748/wjg.v23.i46.8169.
Xiao, A.Y., et al. 2016. Global incidence and mortality of pancreatic diseases: A systematic review, meta-analysis, and meta-regression of population-based cohort studies. The Lancet Gastroenterology & Hepatology 1: 45–55. https://doi.org/10.1016/s2468-1253(16)30004-8.
Banks, P.A., et al. 2013. Classification of acute pancreatitis--2012: Revision of the Atlanta classification and definitions by international consensus. Gut 62: 102–111. https://doi.org/10.1136/gutjnl-2012-302779.
Halangk, W., et al. 2000. Role of cathepsin B in intracellular trypsinogen activation and the onset of acute pancreatitis. The Journal of Clinical Investigation 106: 773–781. https://doi.org/10.1172/jci9411.
Singh, V.P., and S.T. Chari. 2005. Protease inhibitors in acute pancreatitis: Lessons from the bench and failed clinical trials. Gastroenterology 128: 2172–2174. https://doi.org/10.1053/j.gastro.2005.03.087.
Sah, R.P., R.K. Dawra, and A.K. Saluja. 2013. New insights into the pathogenesis of pancreatitis. Current Opinion in Gastroenterology 29: 523–530. https://doi.org/10.1097/MOG.0b013e328363e399.
Noel, P., et al. 2016. Peripancreatic fat necrosis worsens acute pancreatitis independent of pancreatic necrosis via unsaturated fatty acids increased in human pancreatic necrosis collections. Gut 65: 100–111. https://doi.org/10.1136/gutjnl-2014-308043.
Montecucco, F., et al. 2014. Treatment with Evasin-3 abrogates neutrophil-mediated inflammation in mouse acute pancreatitis. European Journal of Clinical Investigation 44: 940–950. https://doi.org/10.1111/eci.12327.
Guo, Z.Z., P. Wang, Z.H. Yi, Z.Y. Huang, and C.W. Tang. 2014. The crosstalk between gut inflammation and gastrointestinal disorders during acute pancreatitis. Current Pharmaceutical Design 20: 1051–1062. https://doi.org/10.2174/13816128113199990414.
Nieminen, A., et al. 2014. Circulating cytokines in predicting development of severe acute pancreatitis. Critical Care (London, England) 18: R104. https://doi.org/10.1186/cc13885.
Perides, G., et al. 2011. TNF-alpha-dependent regulation of acute pancreatitis severity by Ly-6C(hi) monocytes in mice. The Journal of Biological Chemistry 286: 13327–13335. https://doi.org/10.1074/jbc.M111.218388.
Hoque, R., et al. 2011. TLR9 and the NLRP3 inflammasome link acinar cell death with inflammation in acute pancreatitis. Gastroenterology 141: 358–369. https://doi.org/10.1053/j.gastro.2011.03.041.
Conway Morris, A., et al. 2009. C5a mediates peripheral blood neutrophil dysfunction in critically ill patients. American Journal of Respiratory and Critical Care Medicine 180: 19–28. https://doi.org/10.1164/rccm.200812-1928OC.
Lämmermann, T., et al. 2013. Neutrophil swarms require LTB4 and integrins at sites of cell death in vivo. Nature 498: 371–375. https://doi.org/10.1038/nature12175.
Lankisch, P.G., M. Apte, and P.A. Banks. 2015. Acute pancreatitis. Lancet 386: 85–96. https://doi.org/10.1016/s0140-6736(14)60649-8.
Kang, H., et al. 2022. Role of neutrophil extracellular traps in inflammatory evolution in severe acute pancreatitis. Chinese Medical Journal 135: 2773–2784. https://doi.org/10.1097/cm9.0000000000002359.
Lee, K.H., et al. 2017. Neutrophil extracellular traps (NETs) in autoimmune diseases: A comprehensive review. Autoimmunity Reviews 16: 1160–1173. https://doi.org/10.1016/j.autrev.2017.09.012.
Yipp, B.G., and P. Kubes. 2013. NETosis: How vital is it? Blood 122: 2784–2794. https://doi.org/10.1182/blood-2013-04-457671.
Merza, M., et al. 2015. Neutrophil extracellular traps induce trypsin activation, inflammation, and tissue damage in mice with severe acute pancreatitis. Gastroenterology 149: 1920–1931.e1928. https://doi.org/10.1053/j.gastro.2015.08.026.
Ren, J., I. Dimitrov, A.D. Sherry, and C.R. Malloy. 2008. Composition of adipose tissue and marrow fat in humans by 1H NMR at 7 tesla. Journal of Lipid Research 49: 2055–2062. https://doi.org/10.1194/jlr.D800010-JLR200.
Münzer, P., et al. 2021. NLRP3 Inflammasome assembly in neutrophils is supported by PAD4 and promotes NETosis under sterile conditions. Frontiers in Immunology 12: 683803. https://doi.org/10.3389/fimmu.2021.683803.
Paget, C., E. Doz-Deblauwe, N. Winter, and B. Briard. 2022. Specific NLRP3 Inflammasome assembling and regulation in neutrophils: Relevance in inflammatory and infectious diseases. Cells 11. https://doi.org/10.3390/cells11071188.
Lachowicz-Scroggins, M.E., et al. 2019. Extracellular DNA, neutrophil extracellular traps, and Inflammasome activation in severe asthma. American Journal of Respiratory and Critical Care Medicine 199: 1076–1085. https://doi.org/10.1164/rccm.201810-1869OC.
Kahlenberg, J.M., C. Carmona-Rivera, C.K. Smith, and M.J. Kaplan. 2013. Neutrophil extracellular trap-associated protein activation of the NLRP3 inflammasome is enhanced in lupus macrophages. Journal of Immunology 190: 1217–1226. https://doi.org/10.4049/jimmunol.1202388.
Dömer, D., T. Walther, S. Möller, M. Behnen, and T. Laskay. 2021. Neutrophil extracellular traps activate Proinflammatory functions of human neutrophils. Frontiers in Immunology 12: 636954. https://doi.org/10.3389/fimmu.2021.636954.
Antiochos, B., et al. 2022. The DNA sensors AIM2 and IFI16 are SLE autoantigens that bind neutrophil extracellular traps. eLife 11. https://doi.org/10.7554/eLife.72103.
Aldabbous, L., et al. 2016. Neutrophil extracellular traps promote angiogenesis: Evidence from vascular pathology in pulmonary hypertension. Arteriosclerosis, Thrombosis, and Vascular Biology 36: 2078–2087. https://doi.org/10.1161/atvbaha.116.307634.
McDonald, B., et al. 2017. Platelets and neutrophil extracellular traps collaborate to promote intravascular coagulation during sepsis in mice. Blood 129: 1357–1367. https://doi.org/10.1182/blood-2016-09-741298.
Torres-Gomez, A., C. Cabañas, and E.M. Lafuente. 2020. Phagocytic Integrins: Activation and signaling. Frontiers in Immunology 11: 738. https://doi.org/10.3389/fimmu.2020.00738.
Ren, Y., et al. 2021. Milk fat globule EGF factor 8 restores mitochondrial function via integrin-medicated activation of the FAK-STAT3 signaling pathway in acute pancreatitis. Clinical and Translational Medicine 11: e295. https://doi.org/10.1002/ctm2.295.
Wereszczynska-Siemiatkowska, U., A. Siemiatkowski, A. Swidnicka-Siergiejko, B. Mroczko, and A. Dabrowski. 2015. The imbalance between matrix metalloproteinase 9 and tissue inhibitor of metalloproteinase 1 in acute pancreatitis. Zeitschrift fur Gastroenterologie 53: 199–204. https://doi.org/10.1055/s-0034-1385705.
Vitale, D.S., et al. 2022. Matrix metalloproteinases and their inhibitors in pediatric severe acute pancreatitis. PLoS One 17: e0261708. https://doi.org/10.1371/journal.pone.0261708.
Silva, J.C., et al. 2020. Mac-1 triggers neutrophil DNA extracellular trap formation to aspergillus fumigatus independently of PAD4 histone citrullination. Journal of Leukocyte Biology 107: 69–83. https://doi.org/10.1002/jlb.4a0119-009rr.
Kretzschmar, G.C., et al. 2021. Neutrophil extracellular traps: A perspective of Neuroinflammation and complement activation in Alzheimer's disease. Frontiers in Molecular Biosciences 8: 630869. https://doi.org/10.3389/fmolb.2021.630869.
Pei, X., et al. 2024. Exosomal secreted SCIMP regulates communication between macrophages and neutrophils in pneumonia. Nature Communications 15: 691. https://doi.org/10.1038/s41467-024-44714-4.
Majde, J.A. 2000. Viral double-stranded RNA, cytokines, and the flu. Journal of Interferon & Cytokine Research 20: 259–272. https://doi.org/10.1089/107999000312397.
Zhou, H., et al. 2013. CD11b/CD18 (mac-1) is a novel surface receptor for extracellular double-stranded RNA to mediate cellular inflammatory responses. Journal of Immunology 190: 115–125. https://doi.org/10.4049/jimmunol.1202136.
Chen, G., et al. 2022. Structural basis for recognition of N-formyl peptides as pathogen-associated molecular patterns. Nature Communications 13: 5232. https://doi.org/10.1038/s41467-022-32822-y.
Wang, H.L., et al. 2023. HCH6-1, an antagonist of formyl peptide receptor-1, exerts anti-neuroinflammatory and neuroprotective effects in cellular and animal models of Parkinson's disease. Biochemical Pharmacology 212: 115524. https://doi.org/10.1016/j.bcp.2023.115524.
Guo, R.F., and P.A. Ward. 2005. Role of C5a in inflammatory responses. Annual Review of Immunology 23: 821–852. https://doi.org/10.1146/annurev.immunol.23.021704.115835.
Höpken, U.E., B. Lu, N.P. Gerard, and C. Gerard. 1996. The C5a chemoattractant receptor mediates mucosal defence to infection. Nature 383: 86–89. https://doi.org/10.1038/383086a0.
Acknowledgments
We thank the reviewers and editors for their assistance. We thank the following organizations for their support: this work was supported by the National Natural Science Foundation of China (No. 82170654, 82100675), Key Research and Development Program of Heilongjiang Province (No. 2022ZX06C06).
Funding
This research was funded by National Natural Science Foundation of China (No. 82170654, No. 82370650), Key Research and Development Program of Heilongjiang Province (No. 2022ZX06C06).
Author information
Authors and Affiliations
Contributions
X.L. designed the study and analyzed the data. Y.Z. and H.W. wrote the manuscript. X.L. and Z.M. prepared the images and tables. D.X reviewed and revised the manuscript. Y.Z. supervised the research. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of First Affiliated Hospital of Harbin Medical University (No: 2021091).
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, X., Zheng, Y., Meng, Z. et al. Gene Regulation of Neutrophils Mediated Liver and Lung Injury through NETosis in Acute Pancreatitis. Inflammation (2024). https://doi.org/10.1007/s10753-024-02071-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10753-024-02071-w