Abstract
The genus Stenosoma comprises 13 species in the NE Atlantic, Mediterranean and Black Sea. Recent studies have improved knowledge on the distributional patterns of the genus, reducing species ranges and allowing the identification of cryptic species. Lacking a free-swimming life-stage, Stenosoma typically display small- to medium-sized ranges, with the exception of S. nadejda which occurs in the Atlantic-Mediterranean region. In this study, we build upon previous work and examine phylogeographic patterns of S. nadejda throughout its entire range to assess levels of genetic differentiation and evidence of cryptic species. While Elongation Factor 1α gene sequences revealed an almost absence of genetic and geographic structure across the entire distribution of S. nadejda, the Cytochrome c oxidase subunit I gene sequences revealed a phylogenetic split defining two Mediterranean clades. Our results confirm the wide distribution of S. nadejda in the Atlantic/Mediterranean region and show that the distribution of the two Mediterranean clades does not match the east–west break in genetic continuity typically displayed by many species. This pattern is probably a consequence of sporadic long-distance dispersal by rafting. Conversely, an almost absence of shared haplotypes at the local level, conforms to the recognized poor ability of peracarids for autonomous dispersal.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Superorder Peracarida is one of the richest groups of marine arthropods, comprising roughly one third (∼21,000) of the known crustacean species. One of the most striking traits of peracarids, unique among crustaceans, is the presence of a brood pouch from where adult-like juveniles emerge. With the exception of the Mysida, for which position within Peracarida is debatable (Poore, 2005; Spears et al., 2005; Jenner et al., 2009), the larger peracarida Orders (Cumacea, Tanaidacea, Amphipoda and Isopoda) encompass mostly benthic species which are relatively small-sized. Consequently, for strictly shallow water or coastal species the lack of a planktonic life-stage clearly limits their ability to disperse through all but very short distances. In these species rafting on seaweed or flotsam is thought to be the primary mechanism of long-range dispersal (Thiel & Haye, 2006). One of the strongest evidence of the importance of rafting as a driver for the dispersion of marine organisms comes precisely from the wide, sometimes disjunct, distributions of some species that otherwise lack any means of autonomous long-range dispersal (Thiel & Gutow, 2005a). Examples abound in peracarids, but recently the usage of molecular techniques has challenged the taxonomic status of many such species, showing that they often consist of localized and highly divergent genetic lineages forming complexes of cryptic species (Hurtado et al., 2010; Markow & Pfeiler, 2010; Baratti et al., 2011; Varela & Haye, 2012; Cabezas et al., 2013a, b, 2014; Larsen et al., 2014, Copilaş-Ciocianu & Petrusek, 2015). Notwithstanding, cases of species with large distributional ranges have also been confirmed by means of molecular evidence (Nikula et al., 2010; Haye et al., 2012), although they fall short of the increasing number of widely ranging taxa prompting for a taxonomic revision. Does this difference arise from a true sporadic nature of rafting as a long-range dispersal mechanism in peracarids?
There is clearly a shortage of data to answer unequivocally the previous question. Most of the studies mentioned focused on single or few species within a genus, and are by no means representative of the great diversity of forms and life-history traits found on peracarids. For example, the family Idoteidae is world-wide distributed, comprising roughly 180 shallow-water species (Poore, 2001) from which a single one is putatively cosmopolitan: the obligatory rafter Idotea metallica Fabricius, 1793 (Brusca, 1984). All other species have distributions that range from tens to thousands of kilometres and, despite being reported as common rafters (14 out of 38 isopod species, Thiel & Gutow, 2005b), there is but a single phylogeographic analysis of a widely distributed idoteid—Idotea balthica (Pallas, 1772)—which confirmed its amphi-Atlantic distribution, but did not include a substantial part of its range (Wares & Cunningham, 2001).
In this work, we explore the genetic diversity of Stenosoma nadejda (Rezig, 1989), a common idoteid that can be found in the intertidal and shallow-waters of the Atlantic-Mediterranean region, living among algae or sea-grasses. The phylogeny of the genus was recently assessed, showing that its evolution is strongly linked with the geological and climatic history of the Atlantic-Mediterranean region (Xavier et al., 2012b). This and other works have also contributed to redefine the ranges of most Stenosoma species, particularly by demonstrating that those once thought to occur in the whole region were actually restricted to either the Atlantic or the Mediterranean basins or to smaller areas within them (Santos et al., 2011; Xavier, 2011; Xavier et al., 2012a, b). Hence, S. nadejda remains as the only species of the genus with a large distribution, occurring in the Atlantic, from central Portugal down to Morocco, and throughout the entire Mediterranean (Xavier, 2011).
Initially taken as a presumptive Mediterranean invader in the Portuguese coast, S. nadejda was shown to occur naturally in the Atlantic coast of Iberia (Xavier et al., 2009), and contrarily to what was assumed, is likely to have an Atlantic origin (Xavier et al., 2012b). A phylogeographic survey focused in the Alboran Sea resulted in the identification of a new species, Stenosoma stephenseni Santos and Xavier 2011, sympatric with S. nadejda and from which it can be distinguished by a remarkable genetic divergence, but rather subtle morphological differences (Xavier et al., 2011). In this work, we extend the previous mtDNA (based on the cytochrome c oxidase subunit I—COI) phylogeographic analysis eastwards up to the Aegean Sea, and provide additional data on the variation of a nuclear marker (exon1 of the elongation factor—EF1α), to investigate whether S. nadejda “hides” further cryptic species or if it has dispersed farther into the eastern Mediterranean.
Materials and methods
Individuals were captured by submerging and washing collected algae in fresh water and were subsequently preserved in 96% ethanol. Specimens were identified according to a recent key (Santos et al., 2011). A total of 264 individuals were used for the present study. The dataset consisted of 94 newly collected individuals from 14 Mediterranean localities east of the Almeria-Oran Front (AOF), which was complemented with 170 individuals collected at 18 locations from the Atlantic, the Alboran Sea, Algeria and west Tunisia and used in previous works (see Table 1; Fig. 1; Xavier et al., 2009, 2011).
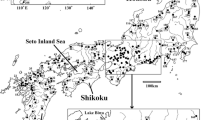
Geographical location of sampling sites and groups defined by BAPS for the COI dataset. The black line represents the Almeria-Oran Front (AOF). Sampling sites are: 1 Baleal; 2 Oliveirinha; 3 Porto de Mós; 4 Olhos d’Água; 5 Chipiona; 6 Rabat; 7 Assilah; 8 Cap Mazari; 9 Calla Iris; 10 Al Hoceima; 11 Marbella; 12 Alboran Island; 13 Cabo da Gata; 14 Villajoyosa; 15 Peñiscola; 16 Tighremt; 17 Tigzirt; 18 Cap Serrat; 19 Bizerte; 20 Cap Bon; 21 Nabeul; 22 Banuyls-sur-Mer; 23 Sestri Levante; 24 Vernazza; 25 Santa Marinella; 26 Capo Colonna; 27 Giovanazzo; 28 Cap Mannu; 29 Villanueva Monteleones; 30 Skala Sykamineas; 31 Molivos; 32 Anaxos. Clusters retrieved with BAPS are coded with combinations of squares or circles and different colours (white, grey and black)
DNA extraction and amplification of the COI were done for all the newly sampled specimens (n = 94) using the methodology previously described (Xavier et al., 2009). As previous works did not include nuclear DNA data, a portion of the nuclear gene Elongation factor 1α (EF1α) was amplified for 95 specimens selected from the 264 available, representative of the entire known distribution of S. nadejda. Primers used to amplify the EF1α were EF1AF- 5′ GAYTTCATYAAGAAACATG 3′ and EF1AR- 5′ GAAWGTCTCYACGCACATGGG 3′; these primer sequences were available at the Crandall lab primer database (http://crandalllab.byu.edu/PrimerDatabase.aspx, however, this database is no longer online. PCR conditions were as follows: initial 4 min denaturation at 94°C, followed by 30 cycles of 45 s at 94°C, 45 s at 55 ∘C and 1 min at 72°C. Final extension was achieved at 72°C for 12 min. PCR reactions were done in a volume of 20 μl with a magnesium concentration of 3 Mm. Platinum Taq (Invitrogen, Carlsbad, CA, USA) was used for all PCR amplifications. PCR products were all sequenced in both directions by a commercial company (High-Throughput Genomics Unit—HTGU, Department of Genome Sciences of the University of Washington).
Estimates of genetic diversity and tests of recombination
Sequences were checked and edited using CodonCode Aligner (CodonCode, Dedham, MA, USA). The software PHASE v.2.1.1 (Stephens et al., 2001; Stephens & Donnelly, 2003; Stephens & Scheet, 2005) was used to resolve EF1α haplotypes. Command line and input files for this software were generated by running SeqPHASE (Flot, 2010). Only haplotypes with probabilities higher than 90% were included in the analysis. Sequences were aligned using ClustalW (Thompson et al., 1994) as implemented in BioEdit (Hall, 1999). COI sequences were uploaded in DNASP (Librado & Rozas, 2009) and translated to aminoacids to obtain the number of synonymous and non-synonymous substitutions, and to search for premature stop codons, which are indicative of the presence of pseudogenes.
Measures of haplotype diversity (Hd) and nucleotide diversity (π) were estimated for COI for 28 localities using Arlequin v3.11 (Excoffier et al., 2005). Localities with less than four individuals or which displayed no polymorphisms were excluded from these analyses. The Maximum Chi-square test of Maynard Smith (Maynard Smith, 1992) and GeneConv (Sawyer, 1989), both available in the software package Recombination Detection Program version 3.44 (Martin et al., 2010), were used to test for recombination in the EF1α dataset.
Estimates of population structure and isolation by distance
Population structure was investigated for the COI and EF1α genes separately using two methods. One is implemented in the software BAPS 5 (Corander et al., 2004). This method uses Bayesian statistics to test mixture and to define clusters of populations, requiring no a priori knowledge about the genetic structure of populations (Corander et al., 2004). The optimal number of clusters in BAPS 5 was selected according to the best output log(ml) value and the posterior probabilities associated with each tested number of cluster (K). Values of K were set to vary between 1 and 20. Similarly, analysis of population structure using the concatenated (nuclear and mitochondrial) dataset was also conducted using BAPS.
The second method is based on the comparison of pairwise Fsts estimated with Arlequin v3.11 (Excoffier et al., 2005) (excluding localities with less than four individuals). For the COI dataset, Fst estimations using were made for the localities east of the Almeria-Oran oceanographic front using Arlequin v3.11 (Excoffier et al., 2005). Significance of pairwise Fst values was tested by performing 300 permutations of haplotypes between locations or clusters, under the null hypothesis of no differentiation. Due to the low level of population structure found for the EF1α gene, isolation by distance (IBD) was only tested for the mtDNA dataset of Mediterranean locations. IBD was tested using a non-parametric Mantel test as incorporated in the IBDWS v. 3.15 (Jensen et al., 2005), using Fst genetic and geographical distances (calculated as the approximate linear distance along shoreline) between population pairs, and 30,000 randomizations.
Uncorrected pairwise p-distances were calculated for all individuals based on COI to acess minimum and maximum genetic divergence distance. Additionally uncorrected p-distances were also conducted between clusters defined by BAPS. Genetic distances were calculated using MEGA5 software (Tamura et al., 2011).
Phylogenetic reconstruction
The phylogenetic tree built for the COI included all unique haplotypes from newly collected individuals, plus an additional set of randomly chosen unique haplotypes representative of the three major clades of S. nadejda found west of Almeria-Oran Front (AOF). The latter included six individuals from Atlantic Morocco (Genbank accession numbers JF915252-3, JF915255, JF915262 and JF915273-4) and 19 individuals from southwest Iberia and the Alboran basin (Genbank accession numbers FJ905064-65, JF915286, JF915288, FJ905079, FJ905077, FJ905074, FJ905070, FJ905067-68, FJ905053-54, FJ905048-49, JF915198-JF915200, JF915207, JF915211) (Xavier et al., 2009, 2011). Additionally, two COI sequences, one from Stenosoma capito (Rathke, 1837) and another from Stenosoma lancifer (Miers, 1881) were used as outgroups (Genbank accession numbers FJ905097 and FJ905098, respectively) (Xavier et al., 2009). The software jModelTest (Posada, 2008) was used to determine the adequate model of sequence evolution using the AIC criteria, which in this case was GTR+I+Γ (−ln 2892).
The methods used for tree reconstruction were Maximum Likelihood (ML) with the software package PhyML (Guindon & Gascuel, 2003), using 1000 bootstraps to estimate branch support, and Bayesian Inference (BI) using MRBAYES 3.1.2 (Huelsenbeck & Ronquist, 2001; Ronquist & Huelsenbeck, 2003). For the BI, two independent runs were performed using 15 × 106 generations. Parameters were sampled every 100 generations with the heating parameter set to 0.1. Majority-rule consensus trees were estimated by combining results from duplicated analyses and a burnin of 10%. Adequate burnin was checked with the software Tracer v1.5 (Rambaut & Drummond, 2007). Haplotype networks reconstructions for the entire EF1α dataset and for the COI dataset including only Mediterranean individuals (collected east of the Almeria-Oran Front) were conducted using TCS (Clement et al., 2000) and output file was run through tcsBU (Santos et al., 2015).
Results
All sequences were translated to aminoacids and no stop codons were detected. The COI dataset alignment was 568 bp long, and included 156 haplotypes. A total of 42 haplotypes were found in the 94 individuals collected specifically for this work. The latter comprised 60 variable sites, of which 42 were parsimony informative. The amplified segment of EF1α comprised an intronic region with considerable length polymorphism that hindered the haplotype phase estimates. Therefore, this region was excluded from the final EF1α alignment that remained 687 bp long, with 23 haplotypes in a total of 95 individuals. EF1α haplotypes comprised 18 variable sites, from which 16 were parsimony informative. Haplotype diversity was generally high for COI (>0.8) with only a few exceptions: Baleal, Porto de Mós, Banuyls-sur-Mer, Villanueva Monteleones, Cap Serrat, Nabeul and Molivos. The highest values of nucleotide diversity were found at Santa Marinella. Diversity was also considerably high within the population clusters defined by BAPS, with lowest values (Hd = 0.71 and π = 0.0049) encountered in Cluster III (Table 1). Additionally haplotype sharing between localities was rare. On the contrary, for the EF1 α haplotype sharing was high (Table 2).
Finally, for the the EF1 α, no recombination events were detected at the significance level of p = 0.05.
Population structure and isolation by distance
For the COI dataset, uncorrected p-distance varied between 0 and 3.2% if only Mediterranean locations were analysed.
BAPS defined six population clusters with log(ml) of −3920 and with 100% posterior probability. The Atlantic-Alboran region was divided in three population clusters: Cluster I-, including populations from Atlantic North Africa; Cluster II,-including populations from Southwest Iberia (including Northern Alboran) and Cluster III, including Moroccan population from South Alboran. Within the Mediterranean, east of the Almeria-Oran front (AOF), another three clusters were also defined: cluster IV, including most of the southwestern sites but extending into the Adriatic Sea; cluster V, including most sites from the northwest and a few from the southeast; and cluster VI, including eastern locations (Greece) but also present in the northwest, at Santa Marinella, Italy (see Fig. 1).
For the EF1α dataset, BAPS defined three clusters with log(ml) of −730 and 99% posterior probability. Cluster I contains most of the individuals from Atlantic locations and some from the northern and southern Mediterranean. Cluster II contains most of the individuals from the northern Mediterranean as well as a few from the Atlantic-Alboran region and the southern Mediterranean. Cluster III only comprises individuals from the southern Mediterranean (Fig. 2).
Results from the best partition with BAPS analysis for the EF1α gene. Localities were coded with numbers as indicated in Fig. 1 and different clusters are coded by colours: Cluster I—light grey; Cluster II—dark grey; Cluster III–white
Results obtained with BAPS for the concatenated datasets revealed three population clusters, with a log(ml) value was -5480 and 100% posterior probability: (1) North African Atlantic cluster, (2) Iberian Atlantic plus Alboran Sea cluster and (3) a Mediterranean cluster (Fig. 3). These population clusters agree with the major clades defined by the phylogenetic analysis of the COI dataset (see below).
Results from the best partition with BAPS analysis for the concatenated dataset (COI + EF1α). Localities were coded with numbers as indicated in Fig. 1
A pattern of isolation by distance was rejected for the COI dataset (r 2 = 1.47E−02, P > 0.05). For COI, high levels of differentiation were observed between all Mediterranean localities (Table 3). Although for EF1α, levels of differentiation were lower than those of COI, most pairwise Fst values were significant at α = 0.05 (Table 4). Uncorrected p-distances between the clusters defined by BAPS for the COI dataset are summarized in Table 5. The highest genetic distance values were found between Atlantic North African cluster and all the others (3.5–4.9%). P-distance between Atlantic and Mediterranean localites was also high (3–4.1%). .
Phylogenetic analysis
Phylogenetic reconstruction of the COI dataset (Fig. 4) showed three groups: (1) comprising all individuals from North African Atlantic (Group A), (2) a monophyletic group comprising individuals from Atlantic Iberia plus Alboran Sea and finally (Group B) and (3) a group containing all individuals from the Mediterranean (Group C). The three main groups recovered by the phylogenetic analyses correspond to the population clusters defined by BAPS using the concatenated dataset (Fig. 3). The Mediterranean group is divided in two clades, both being present in the western and the eastern basins. The three Mediterranean population clusters obtained by BAPS based on COI are well evident in the haplotype network (Fig. S1 of supplementary material), as are the high numbers of private haplotypes. On the contrary, the network of EF1α haplotype revealed three central haplotypes that are frequent and widespread, and from which several tip haplotypes diverge mostly by a single mutation (Fig S2 of supplementary material).
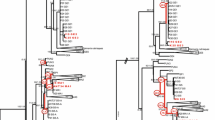
Bayesian consensus tree for the mtDNA COI gene. Node values correspond to Bayesian posterior probabilities and bootstrap support, respectively. Clusters defined by BAPS are also identified (see Fig. 1 for details)
Discussion
The present analyses of DNA sequence variation reveals no evidence for the existence of cryptic speciation between geographically segregated lineages, thus confirming the large geographical distribution of Stenosoma nadejda. Whereas nuclear DNA data showed no evidence for genetic structure, mtDNA data revealed two levels of intraspecific phylogeographic structure in the Mediterranean: local and regional. At the local level, an almost absence of shared haplotypes is in line with the recognized poor ability of peracarids for autonomous dispersal (Thiel & Gutow, 2005a) and with previously published results on this and other species of the same genus (Xavier et al., 2009, 2011; Xavier, 2011). At the regional level, the existence of two distinct but overlapping clades within the Mediterranean region suggests a time of allopatry in the past history of S. nadejda but brings to light an unusual pattern of geographic distribution of genetic variability inside the Mediterranean Sea: instead of the east–west break in genetic continuity typically displayed by many Mediterranean species (Rolland et al., 2007; Arnaud-Haond et al., 2007; Calvo et al., 2009; Sá- Pinto et al., 2012), both mtDNA clades of S. nadejda are present in the western and the eastern Mediterranean basins.
Phylogeography of Stenosoma nadejda throughout its range
Contrary to what could be expected taking into account the poor autonomous dispersal ability of peracarids and the accumulated data on S. nadejda’s mtDNA phylogeography (Xavier et al., 2009, 2011), the analyses of the nuclear marker EF1α revealed an almost absence of phylogeographic structure from the Atlantic to the eastern Mediterranean. While the clustering analysis revealed three groups of individuals, as depicted in Fig. 2, these are admixed across regions. Sharing of EF1α variants across large areas of the distribution is also evident from the haplotype network (Fig. S2 of supplementary material), which also reveals a shallow phylogenetic structure, as haplotypes are closely related differing mostly by a single mutation.
Similar patterns of genetic homogeneity of the nuclear marker EF1α had already been observed for two other species of genus Stenosoma, in the northeast Atlantic. Yet, for S. acuminatum Leach, 1814 and S. lancifer (Miers, 1881) the pattern of genetic variation of EF1α was matched by the one of the mtDNA marker COI. Absence of genetic structure, significant deviations from neutrality and star-like haplotype networks have been observed for both COI and EF1α for S. lancifer as well as for the COI of S. acuminatum, these results being attributed to a recent colonization and demographic expansion (Xavier et al., 2012a). In the present study, however, there are no strong signs of selection or demographic expansion and the phylogeographic pattern of COI is highly structured (Figs. 1, 4, S1).
The COI phylogeographic pattern observed in the present work for locations to the east of the AOF are in line with those previously obtained from the Atlantic and Alboran regions (Xavier et al., 2009, 2011) with almost no haplotype sharing between sampled localities and a marked genetic structure, with several distinct haplogroups. The estimates of genetic differentiation between locations are also similar to what was previously found, suggesting low population connectivity at the local scale. However, the divergence found between Mediterranean specimens using uncorrected p-distances is inferior to the divergence found between Stenosoma species for the same gene fragment (less than 5–18%). Hence, despite the high structure of haplotype distribution, these results combined with the evidence from the EF1α data conform with the hypothesis of a single species that ranges from the Atlantic to the eastern Mediterranean.
Phylogeographic patterns suggest allopatry followed by long-distance dispersal
One of the most interesting results from the present work is the unusual geographic distribution of the two Mediterranean mtDNA clades (Figs. 4). While at the local scale connectivity seems to be very restricted, the two main Mediterranean clades were found to be present in both the western and the eastern basins. The pattern could be explained by a fairly long time of historical allopatry in two regions. The dating of the Stenosoma phylogenetic tree (Xavier et al., 2012b), which included Mediterranean samples of S. nadejda, suggests that diversification within this species was likely caused by isolation during the Pleistocene glaciations. If this is the case, and taking into account present mtDNA data, it is possible that there were two southern Mediterranean glacial refugia, as a west–east genetic break is present in the African coast (Figs. 1, 4). After glaciations, the spread of both clades across these two basins did not follow a coastal stepwise path, and must have been attained by sporadic long-distance dispersal through rafting.
The putative capacity of S. nadejda for long-distance dispersal through rafting seems to be unique within the genus, which comprises other 12 species inhabiting the same area. The two “true” Atlantic Stenosoma species (S. acuminatum and S. lancifer) range from southern UK to Morocco (∼4000 km along the coast line) but do not enter the Mediterranean, while most of the remaining species are restricted to smaller areas, such as the Gulf of Cadiz, the Alboran Sea, Algeria or Tunisia (Xavier, 2011). The only exception is, for now, S. capito which has a similar distributional range (from the Black Sea into the Atlantic). However, molecular and morphological evidence suggests that S. capito is actually a complex of two species whose ranges do not even overlap: one inhabits the Black Sea and the eastern Mediterranean basin, whereas the other extends from the western Mediterranean basin into the Atlantic, reaching northern Morocco and southern Portugal (Xavier et al., 2012b).
Anecdotal evidence suggests that the striking difference between the distributional range of S. nadejda and those of the remaining species of the genus is likely related with its more efficient use of rafting for long-distance dispersal. In the southern Portuguese coast, where it coexists with S. lancifer and S. accuminatum, S. nadejda is by far the most common species found on low-shore intertidal algae. In this region, it is frequent to observe many individuals attached to drifting algae, especially on the invasive species Asparagopsis armata Harvey 1855 which often forms floating masses of considerable size. Experimental work has demonstrated that some idoteids, such as I. balthica and I. granulosa, can actively swim to colonise floating material such as detached seaweeds (Clarkin et al., 2012). Interestingly, I. balthica is the only idoteid for which molecular evidence confirms a large (amphi-Atlantic) distribution (Wares & Cunningham, 2001), although some degree of intraspecific differentiation exists between both sides of the Atlantic (Wares, 2001).
It remains to be known the degree to which S. nadejda is adapted to rafting conditions, since specific traits, such as feeding strategies, metabolic rates, or even the capacity to reproduce “en route”, ultimately determine the success of long-distance dispersal (Gutow et al., 2006). However, the low levels of genetic connectivity found at the local scale suggest that rafting is not frequent in this species.
References
Arnaud-Haond, S., E. Migliaccio, D.-A. Mand, S. Teixeira, M. S. van de Vliet, F. Alberto, G. Procaccini, C. M. Duarte & E. A. Serrão, 2007. Vicariance patterns in the Mediterranean Sea: east-west cleavage and low dispersal in the endemic seagrass Posidonia oceanica. J Biogeogr 34: 963–976. doi:.
Baratti, M., M. Filippelli & G. Messana, 2011. Complex genetic patterns in the mangrove wood-borer Sphaeroma terebrans Bate, 1866 (Isopoda, Crustacea, Sphaeromatidae) generated by shoreline topography and rafting dispersal. Journal of Experimental Marine Biology and Ecology 398: 73–82.
Brusca, R. C., 1984. Phylogeny, evolution and biogeography of the marine isopod subfamily Idoteinae (Crustacea: Isopoda: Idoteidae). Transactions of San Diego Society of Natural History 20: 99–134.
Cabezas, M. P., P. Cabezas, A. Machordom & J. M. Guerra-García, 2013a. Hidden diversity and cryptic speciation refute cosmopolitan distribution in Caprella penantis (Crustacea: Amphipoda: Caprellidae). Journal of Zoological Systematics and Evolutionary Research 51: 85–99.
Cabezas, M. P., C. Navarro-Barranco, M. Ros & J. M. Guerra-García, 2013b. Long-distance dispersal, low connectivity and molecular evidence of a new cryptic species in the obligate rafter Caprella andreae Mayer, 1890 (Crustacea: Amphipoda: Caprellidae). Helgoland Marine Research 67: 483–497.
Cabezas, M. P., R. Xavier, M. Branco, A. M. Santos & J. M. Guerra-García, 2014. Invasion history of Caprella scaura Templeton, 1836 (Amphipoda: Caprellidae) in the Iberian Peninsula: multiple introductions revealed by mitochondrial sequence data. Biological Invasions 16: 2221–2245.
Calvo, M., J. Templado, M. Oliverio & A. Machordom, 2009. Hidden Mediterranean biodiversity: molecular evidence for a cryptic species complex within the reef building vermetid gastropod Dendropoma petraeum (Mollusca: Caenogastropoda). Biological Journal of the Linnean Society 96: 898–912.
Clarkin, E., C. A. Maggs, G. Arnott, S. Briggs & J. D. R. Houghton, 2012. The colonization of macroalgal rafts by the genus Idotea (sub-phylum Crustacea; order Isopoda): an active or passive process? Journal of the Marine Biological Association UK 92: 1273–1282.
Clement, M., D. Posada & K. Crandall, 2000. TCS: a computer program to estimate gene genealogies. Molecular Ecology 9: 1657–1660.
Copilaş-Ciocianu, D. & A. Petrusek, 2015. The southwestern Carpathians as an ancient centre of diversity of freshwater gammarid amphipods: insights from the Gammarus fossarum species complex. Molecular Ecology 24: 3980–3992.
Corander, J., P. Waldmann, P. Marttinen & M. J. Sillanpää, 2004. Baps 2: enhanced possibilities for the analysis of genetic population structure. Bioinformatics 20: 2363–2369.
Excoffier, L., G. Laval & S. Schneider, 2005. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online 1: 47–50.
Flot, J. F., 2010. Seqphase: a web tool for interconverting phase input/output files and fasta sequence alignments. Molecular Ecology Resources 10: 162–166.
Guindon, S. & O. Gascuel, 2003. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Systematic Biology 52: 696–704.
Gutow, L., J. Strahl, C. Wiencke, H. D. Franke & R. Saborowski, 2006. Behavioural and metabolic adaptations of marine isopods to the rafting life style. Marine Biology 149: 821–828.
Hall, T. A., 1999. Bioedit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/nt. In Nucleic Acids Symposium Series 41: 95–98.
Haye, P. A., A. I. Varela & M. Thiel, 2012. Genetic signatures of rafting dispersal in algal-dwelling brooders Limnoria spp. (Isopoda) along the SE Pacific (Chile). Marine Ecology Progress Series 455: 111–122.
Huelsenbeck, J. P. & F. Ronquist, 2001. Mrbayes: Bayesian inference of phylogeny. Bioinformatics 17: 754–755.
Hurtado, L. A., M. Mateos & C. A. Santamaria, 2010. Phylogeography of supralittoral rocky Intertidal Ligia Isopods in the Pacific region from Central California to Central Mexico. PLoS ONE 5: e11633.
Jensen, J. L., A. J. Bohonak & S. T. Kelley, 2005. Isolation by distance, web service. BMC Genetics 6: 13.
Jenner, R. A., C. N. Dhubhghaill, M. P. Ferla & M. A. Wills, 2009. Eumalacostracan phylogeny and total evidence: limitations of the usual suspects. BMC Evolutionary Biology 9: 21.
Larsen, K., F. Tuya & E. Froufe, 2014. Genetic divergence of tanaidaceans (Crustacea: Peracarida) with low dispersal ability. Scientia Marina 78: 81–90.
Librado, P. & J. Rozas, 2009. Dnasp v5: a software for comprehensive analysis of dna polymorphism data. Bioinformatics 25: 1451–1452.
Markow, T. A. & E. Pfeiler, 2010. Mitochondrial DNA evidence for deep genetic divergences in allopatric populations of the rocky intertidal isopod Ligia occidentalis from the eastern Pacific. Molecular Phylogenetics and Evolution 56: 468–473.
Martin, D. P., P. Lemey, M. Lott, V. Moulton, D. Posada & P. Lefeuvre, 2010. Rdp3: a flexible and fast computer program for analyzing recombination. Bioinformatics 26: 2462–2463.
Maynard Smith, J. M., 1992. Analyzing the mosaic structure of genes. Journal of Molecular Evolution 34: 126–129.
Nikula, R., C. Fraser, H. Spencer & J. Waters, 2010. Circumpolar dispersal by rafting in two subantarctic kelp-dwelling crustaceans. Marine Ecology Progress Series 405: 221–230.
Poore, G., 2001. Isopoda Valvifera: diagnoses and relationships of the families. Journal of Crustacean Biology 21: 205–230.
Poore, G., 2005. Peracarida: monophyly, relationships and evolutionary success. Nauplius 13: 1–27.
Posada, D., 2008. jmodeltest: phylogenetic model averaging. Mol Biol Evol 25: 1253–1256.
Rambaut A. & A. J. Drummond, 2007. Tracer v1.4. http://tree.bio.ed.ac.uk/software/tracer.
Rolland, J. L., F. Bonhomme, F. Lagardere, M. Hassan & B. Guinand, 2007. Population structure of the common sole (Solea solea) in the Northeastern Atlantic and the Mediterranean Sea: revisiting the divide with EPIC markers. Marine Biology 151: 327–341.
Ronquist, F. & J. P. Huelsenbeck, 2003. Mrbayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19: 1572–1574.
Sá- Pinto, A., M. Branco, P. B. Alexandrino, M. C. Fontaine & S. J. E. Baird, 2012. Barriers to gene flow in the marine environment: insights from two common intertidal limpet species of the Atlantic and Mediterranean. PLoS One 7(12): e50330.
Santos, A., R. Xavier, S. Zenboudji, T. Branco & M. Branco, 2011. Stenosoma stephenseni sp. n. (Isopoda, Idoteidae), from the southwestern Mediterranean, with a note on the nomenclatural status of Synisoma Collinge, 1917. ZooKeys 141: 29–44.
Santos, A. M., P. Cabezas-Rodríguez, A. I. Tavares, R. Xavier & M. Branco, 2015. tcsBU: a tool to extend TCS network layout and visualization. Bioinformatics. doi:10.1093/bioinformatics/btv636
Sawyer, S. A., 1989. Statistical tests for detecting gene conversion. Molecular Biology and Evolution 6: 526–538.
Spears, T., R. W. Debry, L. G. Abele & K. Chodyla, 2005. Peracarida monophyly and interordinal phylogeny inferred from nuclear small-subunit ribosomal DNA sequences (Crustacea: Malacostraca: Peracarida). Proceeding of the Biological Society of Washington 118: 117–157.
Stephens, M. & P. Donnelly, 2003. A comparison of bayesian methods for haplotype reconstruction from population genotype data. American Journal of Human Genetics 73: 1162–1169.
Stephens, M. & P. Scheet, 2005. Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. American Journal of Human Genetics 76: 449–462.
Stephens, M., N. J. Smith & P. Donnelly, 2001. A new statistical method for haplotype reconstruction from population data. American Journal of Human Genetics 68: 978–989.
Tamura, K., D. Peterson, N. Peterson, G. Stecher, M. Nei & S. Kumar, 2011. Mega5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution 28: 2731–2739.
Thiel, M. & L. Gutow, 2005a. The ecology of rafting in the marine environment. I. The floating substrata. Oceanography and Marine Biology - An Annual Review 42: 181–264.
Thiel, M. & L. Gutow, 2005b. The ecology of rafting in the marine environment. II. The rafting organisms and community. Oceanography and Marine Biology - An Annual Review 43: 279–418.
Thiel, M. & P. A. Haye, 2006. The ecology of rafting in the marine environment. III. Biogeographical and evolutionary consequences. Oceanography and Marine Biology – An Annual Review 44: 323–429.
Thompson, J. D., D. G. Higgins & T. J. Gibson, 1994. Clustal w: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research 22: 4673–4680.
Varela, A. I. & P. Haye, 2012. The marine brooder Excirolana braziliensis (Crustacea: Isopoda) is also a complex of cryptic species on the coast of Chile. Revista Chilena de Historia Natural 85: 495–502.
Wares, J. P., 2001. Intraspecific variation and geographic isolation in Idotea balthica (Isopoda: Valvifera). Journal of Crustacean Biology 21: 1007–1013.
Wares, J. P. & C. W. Cunningham, 2001. Phylogeography and historical ecology of the North Atlantic intertidal. Evolution 55: 2455–2469.
Xavier, R., 2011. Phylogeny and phylogeography of the marine isopod genus Stenosoma: Taxonomical revision and analysis of changes in species’ distributions. Dissertation, University of Porto.
Xavier, R., A. M. Santos, F. P. Lima & M. Branco, 2009. Invasion or invisibility: using genetic and distributional data to investigate the alien or indigenous status of the Atlantic populations of the peracarid isopod, Stenosoma nadejda (Rezig 1989). Molecular Ecology 18: 3283–3290.
Xavier, R., S. Zenboudji, F. P. Lima, D. J. Harris, A. M. Santos & M. Branco, 2011. Phylogeography of the marine isopod Stenosoma nadejda (Rezig, 1989) in North African Atlantic and western Mediterranean coasts reveals complex differentiation patterns and a new species. Biological Journal of the Linnean Society 104: 419–431.
Xavier, R., A. M. Santos & M. Branco, 2012a. MtDNA and nuclear data reveal patterns of low genetic differentiation for the isopods Stenosoma lancifer and Stenosoma acuminatum, with low dispersal ability along the northeast Atlantic coast. Scientia Marina 76: 133–140.
Xavier, R., A. M. Santos, D. J. Harris, M. Sezgin, M. Machado & M. Branco, 2012b. Phylogenetic analysis of the north-east Atlantic and Mediterranean species of the genus Stenosoma (Isopoda, Valvifera, Idoteidae). Zoologica Scripta 41: 386–399.
Acknowledgments
The authors would like to thank D. J. Harris, S. Zenboudji, M. Carretero, D. Salvi and C. Rato for their collaboration during field work. C. Rato and D. J. Harris also provided helpful comments on methodology. This study was funded by FEDER (through COMPETE program), Project “Biodiversity, Ecology and Global Change” cofinanced by North Portugal Regional Operational Programme 2007/2013 (ON.2—O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF), and FCT—Fundação para a Ciência e a Tecnologia funds through the projects FCOMP-01-0124-FEDER-020733 and PTDC/MAR/118205/2010; RX and MB were financed by post-doctoral grants awarded by FCT (SFRH/BPD/75851/2011 and SFRH/BPD/40073/2007, respectively).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This work did not involve endangered or protected species, and no specific permissions were required to collect them. Most specimens were obtained through regular exchange with other institutions/co-workers.
Additional information
Handling editor: Diego Fontaneto
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xavier, R., Branco, M. & dos Santos, A.M. Using a phylogeographic approach to investigate the diversity and determine the distributional range of an isopod (Crustacea: Peracarida), Stenosoma nadejda (Rezig, 1989) in the Atlantic-Mediterranean region. Hydrobiologia 768, 315–328 (2016). https://doi.org/10.1007/s10750-015-2559-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-015-2559-8