Abstract
Four and a half LIM domains 2 (FHL2) is a protein of 279 amino acids in length containing four full LIM-domains and a half LIM-domain at the amino terminus. FHL2 is one transcriptional cofactor that can interact with many different proteins, such as AP-1, BRCA1, IGFBP, and integrin, and involved in organ differentiation, development, cell apoptosis, and carcinogenesis. Recent studies showed that FHL2 could play different roles acting as co-activator or corepressor in different cancer types, depending on the cell types involved. However, no report about FHL2 function in tongue squamous cell carcinoma (TSCC) is available to date. This study aims to determine the FHL2 expression and its biological functions in TSCC via in vitro and in vivo studies. Results show that FHL2 expression was associated with the pathological differentiation of TSCC samples through immunohistochemistry. FHL2 overexpression could stimulate cell proliferation, invasiveness, and metastases investigated by MTT, flow cytometry, Transwell and cell scratch methods. FHL2 could also elevate tumor-related molecule nuclear transcription factor-B (NF-кB) and β-catenin expression levels both at transcriptional and translational levels through real-time PCR and Western blot analyses. The in vivo nude mice experiment showed that the tumorigenicity of FHL2 overexpression group was significantly increased compared with control groups. These results suggest that FHL2 overexpression could contribute to the growth, proliferation, invasiveness, and metastasis of human tongue squamous cell carcinoma; furthermore, its function in TSCC might be related with the upregulation of NF-кB and β-catenin expressions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Head and neck cancer is a malignant tumor with high incidence and known as the 6th frequent carcinoma in the whole body, in which tongue is the easily affected intraoral site (Mignogna et al. 2004; Wu et al. 2013a, b). Tongue squamous cell carcinoma (TSCC) is the most common pathological type of tongue cancer, which is known for its poor prognosis and high mortality rates because of features, such as high invasiveness and metastasis. However, in cases with successful surgery, TSCC frequently leads to severe defects in speech, chewing, and cancer-related death. The common strategies to treat tongue cancer are head and neck surgery combined with adjuvant therapy, but the treatment results are still not ideal until now. Recently, researchers are exploring more cancer-related molecules and pursuing new biological methods to treat TSCC to obtain better results (Duan et al. 2006; Jia et al. 2013; Lim et al. 2014;Yao et al. 2007). Target gene therapy method has become a prospective strategy in cancer field. Many molecules in various signaling pathways, such as microRNAs, nuclear transcription factor-B (NF-кB), and β-catenin (Duan et al. 2006; Jia et al. 2013; Lim et al. 2014; Yao et al. 2007), are involved in the development and prognosis of tongue squamous cell carcinoma. However, the precise mechanism of TSCC is still unknown.
Four and a half LIM domains 2 (FHL2) is a LIM-only protein that belongs to the four and a half LIM-only protein family; it is strongly expressed in cardiac and skeletal muscle cells, but a much lower level was observed in other tissues and cell types (Martin et al. 2002; Lai et al. 2006). FHL2 can function as an adapter or modifier in protein interactions mediated by LIM domains, which allow FHL2 to have diverse functions in the regulation of many cellular processes, such as cell survival, proliferation, differentiation, adhesion, motility, gene expression regulation, and signal transduction (Kleiber et al. 2007; Kurakula et al. 2011; Lai et al. 2006; Tian et al. 2015; Wei et al. 2003; Xia et al. 2013; Zhang et al. 2011). The role of FHL2 in cancer is particularly intriguing because FHL2 binds to different proteins and can function in a cell-type dependent manner because it acts as a transcriptional coactivator of several transcription factors, including androgen receptor, AP-1, CREB, BRCA1, and NF-kB in various transformed cell types, or as transcriptional corepressors of ERK2, PLZF, SRF, and FOXO1 (Cao et al. 2015; Kleiber et al. 2007; Kurakula et al. 2011; Wei et al. 2003; Xia et al. 2013; Zhang et al. 2011; Zienert et al. 2015).
FHL2 is overexpressed in breast cancer, ovarian cancer, osteosarcoma, gastrointestinal cancer, and prostate cancer, but downregulated in malignant rhabdomyosarcoma and liver cancer (Genini et al. 1997). Furthermore, FHL2 increases the expression of E-cadherin but reduces activity of the transcription factor NF-кB in colon HT-29 cells, which make the role of FHL2 in cancer development more elusive (Amann et al. 2010; Brun et al. 2013; Kleiber et al. 2007; Ng et al. 2011; Qiao et al. 2009; Wang et al. 2007). The intriguing aspects of FHL2 acting as oncoprotein or tumor suppressor may be related with its interaction with different partner proteins in different cell types.
In the present study, we aim to investigate the FHL2 expression in TSCC and explore its effects on cell proliferation, invasiveness, metastases, and tumorigenicity; we preliminarily explore its role and possibly related signaling molecules in tongue squamous cell carcinoma development via in vivo and in vitro studies.
Materials and methods
Clinical samples
The study was approved by the ethics committee of School and Hospital of Stomatology, Shandong University. Tumorous tongue tissues and adjacent nontumor tissues (at least 1 cm distance away from the tumor edge) were collected with written consent from patients who underwent surgical resection at the School and Hospital of Stomatology, Shandong University between 2009 and 2012. Tissue samples with definitive demographic and clinicopathological records were used for our analyses. Figure 1 provides the summary of the clinicopathological characteristics of 47 patients. In this study, samples from each patient were stained with hematoxylin and eosin, and classified histologically.
Correlation between four and a half LIM domains 2 (FHL2) expression and clinicopathological parameters in patients with TSCC. a Immunohistochemical staining for FHL2 expression in cancer-proximal tissues and well, moderately, and poorly differentiated tongue squamous cell carcinoma (TSCC). b The associations between FHL2 expression and clinicopathological factors in patients with TSCC. *Significantly different compared with controls (P < 0.05)
Immunohistochemistry
Immunohistochemistry staining was performed according to the instructions of the Power Vision Histostain Streptavidin-Peroxidase kit (Zhongshan, Beijing, China). First, the activity of endogenous peroxidase of tissues was blocked with 3 % H2O2 for 15 min. Antigen retrieval was performed with 0.05 % (w/v) trypsin (Sigma-Aldrich, St. Louis, MO) dissolved in Tris–HCl buffer containing 0.1 % (w/v) CaCl2 at 37 °C for 10 min. After treatment with normal serum for 20 min to block non-specific binding, the sections were incubated with anti-FHL2 monoclonal antibody (1:100 dilution; Santa Cruz, CA) at 4 °C overnight. Negative controls were prepared by replacing the primary antibody with PBS.
For immunohistochemistry, biotinylated goat anti-mouse immunoglobulin G (IgG) (Zhongshan) was applied as secondary antibody for 15 min at 37 °C. Sections were exposed to streptavidin–peroxidase conjugate (Zhongshan) for 10 min at 37 °C and visualized by the application of diaminobenzidine solution (Zhongshan) for 2 min. Finally, sections were lightly counterstained with hematoxylin. To determine the integrated optical density of FHL2 staining quantitatively, Image-Pro Plus 6.0 (IPP 6.0) software was used, and FHL2 expression was determined by randomly selecting five tumor cell areas of each specimen under the same conditions. Subsequently, the function of irregular automated optical inspection was used by IPP 6.0 software to score.
Cell culture and transfection
The human tongue squamous cell carcinoma cell line Tca8113 (China Center for Type Culture Collection, Hubei, China) was cultured in RPMI 1640 medium supplemented with 10 % fetal bovine serum, 2 mM l-glutamine, 50 U/ml of penicillin, and 50 µg of streptomycin. Cells were incubated in an atmosphere of 5 % CO2 and 37 °C temperature.
Transfection assays were made by using Lipofectamine 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. Briefly, Tca8113 cells were seeded the day before transfection at 1 × 105/well in 24-well plates with 2 ml of RPMI1640 medium without antibiotics. For each plate, about 10 μg of pcDNA3 plasmids encoding the C-terminal flag-tagged FHL2 were mixed with Opti-MEM, and cells transfected with pcDNA3 empty vector were used as control. These above two plasmids were generously endowed by Professor Cheng (Lai et al. 2006). The mixture was then combined with Lipofectamine 2000 (Invitrogen) in Opti-MEM, incubated for 20 min at room temperature, and added into the cells. The cells were incubated with RPMI1640 supplemented with 5 % FBS and 0.4 mg/ml G418 (geneticin) for selection on the growth and survival of successfully transfected cells expressing the neomycin-resistant gene (neo). Cells surviving G418 treatment were pooled and used for subsequent analyses.
In this study, FHL2-overexpressed TSCC cell line (Tca8113/FHL2 cells) was obtained via stable transfection of pcDNA3.0-FHL2-flag plasmid; cell lines transfected by pcDNA3.0 plasmid (Tca8113/pcDNA3 cells) and parental Tca8113 cells (Tca8113/wt cells) were used as controls.
Analysis of cell growth in vitro
The in vitro growth rate of cells was measured using the MTT method. Briefly, cells (1 × 103/well) were seeded in 96-well plates. On the day of harvest, approximately 100 µl of spent medium was replaced with an equal volume of fresh medium containing 10 % MTT 5 mg/ml stock. Plates were incubated at 37 °C for 4 h, and about 100 µl of DMSO was added to each well; afterward, plates were shaken at room temperature for 10 min. The absorbance was measured at 570 nm.
Flow cytometry analysis for cell cycle
The distribution of cells in the cell cycle phases was determined by FACS analysis of the DNA content. Cells were fixed in ice-cold 70 % ethanol and stored at −20 °C. Prior to analysis, cells were washed and resuspended at 1 × 106 cells/ml in PBS buffer, and incubated with 0.1 mg/ml RNase A and 40 µg/ml propidium iodide at 37 °C for 30 min. Samples were analyzed by a FACS scanner (Becton–Dickinson, San Joes, CA, USA) (Duan et al. 2006).
Cell invasion assay
Cell invasion assays were performed using Transwell method, following previous publication (Yu et al. 2014). Briefly, cells in 0.5 ml of serum-free medium were plated into the inserts (Matrigel-coated inserts for invasion assay). Inserts were placed in wells with 0.75 ml of complete medium containing 10 % FBS as a chemoattractant. After culturing for 20–24 h at 37 °C, cells were fixed with methanol for 8 min and stained with Giemsa reagent (Sigma-Aldrich). Cells on the upper sides of the inserts were removed with a cotton swab, and the insert membranes were cut and mounted on glass slides. The numbers of invaded cells on the membranes were determined by counting the cell numbers in the field at 400× magnification under light-field microscope. The relative migration/invasion activity was measured by normalizing the mean of total migrated/invaded cells per insert in overexpressing cells to that in the corresponding controls. The data shown represent the average of at least three repeated experiments.
Cell migration assay
Cell migration ability was monitored using a wound-healing assay according to previous instructions (Zhu et al. 2010). Cells were seeded at a high density on 6-well cell culture plate. After serum-free incubation for 18 h, wounds were made by scraping through the cell monolayer with a sterile micropipette tip. Cells were further incubated in cell culture medium for up to 72 h with or without plasmid transfection. Images were taken under the microscope to measure cell migration using the widths of the wound in cell monolayer.
RNA isolation and quantitative real-time PCR
For gene expression analysis, total RNA was isolated and purified from cells using Trizol reagent according to manufacturer’s instructions (Takara, Otsu, Japan). Reverse transcription was performed using prime script RT reagent kit with gDNA eraser (Takara). Real-time PCR was carried out by using SYBR Green I Master (ROCHE Light Cycle 480, Germany). Table 1 provides a list of the human sequence-specific primers used in this experiment. The ΔΔCt method was used for quantifying gene expression relative to GAPDH.
Western blot analysis
Cellular protein was isolated by cellular protein extraction solution kit (Sigma-Aldrich). Protein concentrations were measured using the BCA Protein Assay Kit (Sigma-Aldrich). Different concentrations of cellular protein were adjusted to the similar concentration for 1 μg/μl, mixed with 4× gel loading buffer, separated on 10 or 12 % sodium dodecyl sulfate–polyacrylamide gels, and transferred onto PVDF membrane (Millipore, Billerica, MA). Nonspecific antibody bindings were blocked by pre-incubation of the membranes in 1× Tris-buffered saline containing 0.1 % Tween-20 and 5 % milk for 2 h at room temperature. Membranes were incubated for 2 h at 37 °C or overnight at 4 °C with antibodies against respective proteins at different dilutions (FHL2, 1:1000; NF-кB and β-catenin, 1:1000; β-actin, 1:2500) (Santa Cruz). After washing, these membranes were incubated with 1:2000 diluted anti-rabbit or anti-mouse IgG antibody (Santa Cruz) for 1 h at 37 °C. Protein bands were visualized by an electrochemoluminescent kit (Santa Cruz).
Tumor formation in nude mice
Fifteen BALB/C nude mice (Shandong Provincial Center for Disease Control and Prevention, Shandong, China) were randomly separated into three groups, and each group had five mice. The mice were 6–8 weeks old, and the average weight was 20 g. One group of mice was injected subcutaneously with 1 × 106/ml Tca8113/FHL2 cell, and the two control groups were injected with Tca8113/pcDNA3 or Tca8113/wt cells. Mice were checked every 3 days to monitor for tumor formation. The volumes were calculated according to the formula: volume = length × (width2)/2.
Statistical analysis
Each experiment was performed at least thrice using independent cell or tissue samples. Data were displayed as the mean ± standard deviation. Data were analyzed by one-way ANOVA or t test by using the SPSS software (SPSS 17.0). Statistical significance was set at P < 0.05.
Results
Expression of FHL2 in TSCC tumor tissues and correlation between FHL2 protein level and clinicopathological parameters in patients with TSCC
The expression status of FHL2 in TSCC tumorous tissues was evaluated and determined by immunohistochemistry. The results showed that the FHL2 protein was positively stained in the TSCC samples, whereas only weakly stained FHL2 expression was found in adjacent nontumor tissues (Fig. 1). Furthermore, its staining in poorly differentiated tumors was higher than that of moderately and well-differentiated tumors.
The clinicopathological analyses showed that FHL2 expression was positively associated with the pathological differentiation of TSCC tissues (Fig. 1, P = 0.00), disease site, and lymph node metastasis; however, no significant associations were determined between FHL2 expression, gender, and age (Fig. 1, P = 0.473, P = 0.345). FHL2 expression levels in cancer tissues were much higher than those in cancer-proximal site. The lower pathological differentiation of TSCC also resulted in the higher FHL2 expression levels. In addition, those patients with high lymph node metastasis also have higher FHL2 expression levels.
FHL2 was stably transfected into Tca8113 cells
Stable cell populations of Tca8113/FHL2 cells were generated after G418 selection. Overexpression of FHL2 mRNA and protein levels was detected by RT-PCR and Western blot analysis; these Tca8113/wt and Tca8113/pcDNA3 cell levels were higher in Tca8113/FHL2 than in the control groups. The flag expression in Tca8113/FHL2 cells also verified the successful transfection outcome, whereas no expression of flag was detected in the Tca8113/wt and Tca8113/pcDNA3 cells (FHL2 overexpression and flag expression by Western blot in Fig. 5). These results showed that FHL2 was stably overexpressed in Tca8113/FHL2 cells.
Effects of FHL2 overexpression on cell proliferation and cell cycle
To further characterize the effects of FHL2 overexpression on TSCC cells, we performed in vitro functional assays, including cell proliferation and cell cycle. MTT assay was used to detect the ability of cell proliferation. Our result showed that the general growth rate of Tca8113/FHL2 cells was increased compared with those of the two control cells (Fig. 2a, P < 0.05). Nevertheless, the effect of FHL2 on cell proliferation was relatively mild. The most significant difference could be observed at day 4 but eventually disappeared at day 7. We also explored whether FHL2 overexpression could affect the cell cycle by FACS to obtain further insights into the mechanism by which FHL2 contributed to TSCC cell proliferation. As shown in Fig. 2b, G0–G1 cells were downregulated in Tca8113/FHL2 cells compared with those in the two control groups. Significant increase in the percentage of cells in the S phase was also found in the Tca8113-FHL2 cells. The results indicated that FHL2 overexpression could result in the stimulation of cell cycle progression from G1 to S phase.
Effects of FHL2 overexpression on cell proliferation via MTT assay and cell cycle distribution by FACS analyses. a The growth rate of Tca8113-FHL2 cells was increased compared with the control cells. b Cell cycle analyses by FACS showed that G0–G1 cells were downregulated in Tca8113/FHL2 cells compared with those in two control groups. *Significantly different compared with controls (P < 0.05)
Effect of FHL2 overexpression on cell invasion and migration
The effect of FHL2 overexpression on TSCC cell invasion was examined using a Transwell chamber system. More cells have invaded the lower surface of the membrane in FHL2-expressing cells compared with that in control cells (Fig. 3). Similarly, in the migration cell–scratch experiment, we found that Tca8113/FHL2 cells could migrate at a higher rate than the control cells (Fig. 4).
Effects of FHL2 overexpression on cell invasion via Transwell method. a Microscopy image of invading cells in Transwell invasion assay. b More cells have invaded the lower surface of the membrane in FHL2-expressing cells compared with those in control cells. *Significantly different compared with controls (P < 0.05)
Effects of FHL2 overexpression on cell migration via cell scratch experiment. Microscopy image of the cell migration of Tca8113/FHL2, Tca8113/pcDNA3, and Tca8113/wt from one representative experiment. Bar graph represents the percentage of reduced wound width (%). Tca8113/FHL2 cells could migrate at a higher rate than the control cells. *Significantly different compared with controls (P < 0.05)
FHL2 upregulated NF-кB and β-catenin in Tca8113 cells at transcriptional and translational levels
In this study, we performed RT-PCR and Western blot (Fig. 5) techniques to assess the expression levels of NF-кB and β-catenin. FHL2 overexpression could lead to the upregulation of NF-кB and β-catenin expression both at mRNA and protein levels compared with that of control cells.
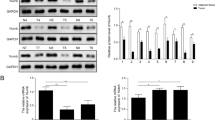
Effects of FHL2 overexpression on NF-kB and β-catenin mRNA and protein levels by real-time PCR and Western blot analysis (including FHL2 overexpression and Flag expression indicating the successful transfection of FHL2). FHL2 overexpression could lead to NF-кB and β-catenin expression upregulation both at mRNA and protein levels compared with that of control cells. *Significantly different compared with controls (P < 0.05)
FHL2 overexpression accelerated tumor formation in nude mice
To observe the effect of FHL2 overexpression on tumor growth, we performed in vivo nude mice experiments. Our results showed that tumor volume gradually increased during the 30 day observation. FHL2 overexpression could significantly induce tumor growth compared with the control groups (Fig. 6).
Effect of FHL2 overexpression on the growth of Tca8113 cells in vivo. Tumor volume gradually increased via observing and measuring every 3 days during the 30 day observation (3, 6, 9, 12, 15, 18, 21, 24, 27, and 30 days). FHL2 overexpression could significantly induce tumor growth compared with the control groups. *Significantly different compared with controls (P < 0.05)
Discussion
To identify the differences in the genetics of cancer and normal cells and explore the cancer-associated genes is important for the development of targeted therapies in cancer treatment. Although active investigations related with oral squamous cell carcinomas have been performed, the pathogenesis and molecular basis for oral tumor cell emergence, survival, and proliferation remain unclear (Duan et al. 2006; Fujii et al. 2011; He et al. 2014; Hu et al. 2015; Huang et al. 2014; Wu et al. 2013a, b; Xia et al. 2011; Yao et al. 2007; Yu et al. 2014; Zhang et al. 2015).
FHL2 is a transcriptional cofactor that performs tumor-promoting or -suppressing activities depending on the types of tumor cells (Chen et al. 2012; Kleiber et al. 2007; Ng et al. 2011). Expression of FHL2 is altered in many cancer cases, and the expression pattern is different among different cancer types. However, whether or how FHL2 is involved in tongue tumorigenesis remains unclear. In this study, we used in vitro and in vivo experiments to investigate the biological function of FHL2 in human tongue squamous cell carcinoma. From the immunohistochemical protein expression analyses, we found that FHL2 was upregulated in most of the clinical tumor samples. The expression status of FHL2 in clinical tissue samples and clinicopathological analyses indicated that FHL2 might participate in the development of TSCC, and FHL2 expression may be used in the diagnosis process of TSCC.
Biological behaviors, including cell proliferation, cell invasion, metastases, and tumor growth, are important factors that affect tumor development, progression, and prognosis. We subsequently evaluated the function of FHL2 by constructing the stable transfectants expressing FHL2 proteins. FHL2 overexpression could stimulate the Tca8113 cell proliferation, cell invasion, metastases, and tumor growth significantly. Our results and information indicated that FHL2 could play important roles in the development of TSCC and act as a cancer-promoting factor. We postulate that target therapies against FHL2 gene or protein might be an optional strategy that can contribute to the treatment of TSCC and improve the patients’ life qualities.
β-Catenin is a central control factor in Wnt signaling, and aberrant activation of the Wnt/β-catenin signaling is associated with basal cell carcinoma of head and neck (Duan et al. 2006; Lee et al. 2014; Yao et al. 2007; Yu et al. 2014). Furthermore, the NF-кB expression was increased in cancers and during tumor angiogenesis. Once activated, NF-кB will translocate into nuclear and upregulate a number of genes necessary for the angiogenesis of tumors, which have кB binding sites in their promoter regions (Duan et al. 2006; Yao et al. 2007). Our real-time PCR and Western blot results showed that FHL2 overexpression could upregulate β-catenin and NF-кB expression levels significantly in Tca8113/FHL2 cells. These results indicated that β-catenin and NF-кB might be the potentially important players in FHL2-mediated growth control in tongue squamous cells and possibly in FHL2-mediated oncogenesis. However, this study is only a preliminary research about FHL2 expression and function in the development of TSCC; the precise role and mechanism of FHL2 and its signaling pathway in TSCC need further investigation.
In summary, our study has shown that FHL2 expression is associated with the pathological differentiation of TSCC and could be considered a potential predictor of clinical outcome in patients with TSCC. The results suggest that FHL2 may be a candidate tumor activator and act as a potential and novel biomarker of TSCC.
References
Amann T, Egle Y, Bosserhoff AK, Hellerbrand C (2010) FHL2 suppresses growth and differentiation of the colon cancer cell line HT-29. Oncol Rep 23:1669–1674
Brun J, Dieudonne FX, Marty C, Muller J, Schule R, Patino-Garcia A, Lecanda F, Fromigue O, Marie PJ (2013) FHL2 silencing reduces Wnt signaling and osteosarcoma tumorigenesis in vitro and in vivo. PLoS One 8:e55034
Cao CY, Mok SW, Cheng VW, Tsui SK (2015) The FHL2 regulation in the transcriptional circuitry of human cancers. Gene 572:1–7
Chen YH, Wu ZQ, Zhao YL, Si YL, Guo MZ, Han WD (2012) FHL2 inhibits the Id3-promoted proliferation and invasive growth of human MCF-7 breast cancer cells. Chin Med J (Engl) 125:2329–2333
Duan L, Yao J, Wu X, Fan M (2006) Growth suppression induced by Notch1 activation involves Wnt–β-catenin down-regulation in human tongue carcinoma cells. Biol Cell 98:479–490
Fujii M, Katase N, Lefeuvre M, Gunduz M, Buery RR, Tamamura R, Tsujigiwa H, Nagatsuka H (2011) Dickkopf (Dkk)-3 and β-catenin expressions increased in the transition from normal oral mucosal to oral squamous cell carcinoma. J Mol Histol 42:499–504
Genini MSP, Scholl FA, Remppis A, Mattei MG, Schäfer BW (1997) Subtractive cloning and characterization of DRAL, a novel LIM-domain protein down-regulated in rhabdomyosarcoma. DNA Cell Biol 16:433–442
He KF, Zhang L, Huang CF, Ma SR, Wang YF, Wang WM, Zhao ZL, Liu B, Zhao YF, Zhang WF, Sun ZJ (2014) CD163+ tumor-associated macrophages correlated with poor prognosis and cancer stem cells in oral squamous cell carcinoma. Biomed Res Int 2014:838632
Hu FW, Lee SS, Yang LC, Tsai CH, Wang TH, Chou MY, Yu CC (2015) Knockdown of S100A4 impairs arecoline-induced invasiveness of oral squamous cell carcinomas. Oral Oncol 51:690–697
Huang CF, Yu GT, Wang WM, Liu B, Sun ZJ (2014) Prognostic and predictive values of SPP1, PAI and caveolin-1 in patients with oral squamous cell carcinoma. Int J Clin Exp Pathol 7:6032–6039
Jia LF, Wei SB, Gan YH, Guo Y, Gong K, Mitchelson K, Cheng J, Yu GY (2013) Expression, regulation and roles of MiR-26a and MEG3 in tongue squamous cell carcinoma. Int J Cancer 135:2282–2293
Kleiber K, Strebhardt K, Martin BT (2007) The biological relevance of FHL2 in tumour cells and its role as a putative cancer target. Anticancer Res 27:55–61
Kurakula K, van der Wal E, Geerts D, van Tiel CM, de Vries CJ (2011) FHL2 protein is a novel co-repressor of nuclear receptor Nur77. J Biol Chem 286:44336–44343
Lai CF, Bai S, Uthgenannt BA, Halstead LR, McLoughlin P, Schafer BW, Chu PH, Chen J, Otey CA, Cao X, Cheng SL (2006) Four and half lim protein 2 (FHL2) stimulates osteoblast differentiation. J Bone Miner Res 21:17–28
Lee SH, Koo BS, Kim JM, Huang S, Rho YS, Bae WJ, Kang HJ, Kim YS, Moon JH, Lim YC (2014) Wnt/β-catenin signalling maintains self-renewal and tumourigenicity of head and neck squamous cell carcinoma stem-like cells by activating Oct4. J Pathol 234:99–107
Lim W, Choi H, Kim J, Kim S, Jeon S, Ni K, Song SY, Oh HK, Im Y, Lee G, Lee JY, Moon YL, You JW, Kim O (2014) Expression of cancer stem cell marker during 4-nitroquinoline 1-oxide-induced rat tongue carcinogenesis. J Mol Histol 45:653–663
Martin B, Schneider R, Janetzky S, Waibler Z, Pandur P, Kuhl M, Behrens J, von der Mark K, Starzinski-Powitz A, Wixler V (2002) The LIM-only protein FHL2 interacts with β-catenin and promotes differentiation of mouse myoblasts. J Cell Biol 159:113–122
Mignogna MD, Fedele S, Lo Russo L (2004) The World Cancer Report and the burden of oral cancer. Eur J Cancer Prev 13:139–142
Ng CF, Ng PK, Lui VW, Li J, Chan JY, Fung KP, Ng YK, Lai PB, Tsui SK (2011) FHL2 exhibits anti-proliferative and anti-apoptotic activities in liver cancer cells. Cancer Lett 304:97–106
Qiao L, Wang Y, Pang R, Wang J, Dai Y, Ma J, Gu Q, Li Z, Zhang Y, Zou B, Lan HY, Wong BC (2009) Oncogene functions of FHL2 are independent from NF-κBIα in gastrointestinal cancer. Pathol Oncol Res 15:31–36
Tian X, Wang Q, Wang X (2015) Four and a half LIM domain protein 2 enhances differentiation and mineralization of human dental pulp cells. J Endod 41:513–519
Wang J, Yang Y, Xia HH, Gu Q, Lin MC, Jiang B, Peng Y, Li G, An X, Zhang Y, Zhuang Z, Zhang Z, Kung HF, Wong BC (2007) Suppression of FHL2 expression induces cell differentiation and inhibits gastric and colon carcinogenesis. Gastroenterology 132:1066–1076
Wei Y, Renard CA, Labalette C, Wu Y, Levy L, Neuveut C, Prieur X, Flajolet M, Prigent S, Buendia MA (2003) Identification of the LIM protein FHL2 as a coactivator of β-catenin. J Biol Chem 278:5188–5194
Wu T, Jia J, Xiong X, He H, Bu L, Zhao Z, Huang CF, Zhang WF (2013a) Increased expression of Lin28B associates with poor prognosis in patients with oral squamous cell carcinoma. PLoS One 8:e83869
Wu X, Cao W, Wang X, Zhang J, Lv Z, Qin X, Wu Y, Chen W (2013b) TGM3, a candidate tumor suppressor gene, contributes to human head and neck cancer. Mol Cancer 12:151
Xia J, Wang J, Chen N, Dai Y, Hong Y, Chen X, Cheng B (2011) Expressions of CXCR7/ligands may be involved in oral carcinogenesis. J Mol Histol 42:175–180
Xia T, Levy L, Levillayer F, Jia B, Li G, Neuveut C, Buendia MA, Lan K, Wei Y (2013) The four and a half LIM-only protein 2 (FHL2) activates transforming growth factor β (TGF-β) signaling by regulating ubiquitination of the E3 ligase Arkadia. J Biol Chem 288:1785–1794
Yao J, Duan L, Fan M, Yuan J, Wu X (2007) Overexpression of BLCAP induces S phase arrest and apoptosis independent of p53 and NF-κB in human tongue carcinoma : BLCAP overexpression induces S phase arrest and apoptosis. Mol Cell Biochem 297:81–92
Yu T, Liu K, Wu Y, Fan J, Chen J, Li C, Yang Q, Wang Z (2014) MicroRNA-9 inhibits the proliferation of oral squamous cell carcinoma cells by suppressing expression of CXCR4 via the Wnt/β-catenin signaling pathway. Oncogene 33:5017–5027
Zhang W, Wang J, Zou B, Sardet C, Li J, Lam CS, Ng L, Pang R, Hung IF, Tan VP, Jiang B, Wong BC (2011) Four and a half LIM protein 2 (FHL2) negatively regulates the transcription of E-cadherin through interaction with Snail1. Eur J Cancer 47:121–130
Zhang L, Zhang W, Wang YF, Liu B, Zhang WF, Zhao YF, Kulkarni AB, Sun ZJ (2015) Dual induction of apoptotic and autophagic cell death by targeting survivin in head neck squamous cell carcinoma. Cell Death Dis 6:e1771
Zhu Z, Xu X, Yu Y, Graham M, Prince ME, Carey TE, Sun D (2010) Silencing heat shock protein 27 decreases metastatic behavior of human head and neck squamous cell cancer cells in vitro. Mol Pharm 7:1283–1290
Zienert E, Eke I, Aust D, Cordes N (2015) LIM-only protein FHL2 critically determines survival and radioresistance of pancreatic cancer cells. Cancer Lett 364:17–24
Acknowledgments
Supported by Shandong Natural Science Foundation of China (Grant No. ZR2014HQ075), Medical Health and Development Research Foundation of Shandong Province (Grant No. 2013WS0214) and Science and Technology Development Project of Jinan City (Grant No. 201121042).
Author information
Authors and Affiliations
Corresponding author
Additional information
Qiang Wang and Xiaoying Wang are the co-first authors and have contributed equally to this study.
Rights and permissions
About this article
Cite this article
Wang, Q., Wang, X., Tian, X. et al. Four and a half LIM domains 2 contributes to the development of human tongue squamous cell carcinoma. J Mol Hist 47, 105–116 (2016). https://doi.org/10.1007/s10735-016-9654-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10735-016-9654-7