Abstract
The flower of Zingiber zerumbet is characterized by a distinctive labellum, a highly modified floral organ believed to be formed by the fusion of several infertile members of the androecial whorl (staminodes). Across the Zingiberaceae, the number of staminodes involved in labellum formation varies from two to four, and these are reflected in the number of lobes that comprise the mature labellum. Research on the flower development in Zingiberaceae has been limited to species with either no labellum lobes or species displaying a bilobed labellum. Zingiber zerumbet is a representative of the genus with a three-lobed labellum, and its flower development remains poorly understood at both morphological and molecular levels. This study aims to give a comprehensive description of its flower development and to identify potential genes related to flower development using morphological and genetic characterization. Our results show that floral organ initiation is sequential with the sepal whorl initiating first, followed by petal and inner androecium together, followed by outer androecium, and finally the initiation of the inferior gynoecium. The three-lobed labellum comprises four androecial members: Two abaxial inner androecial members fuse to form the single central lobe, and two adaxial outer androecial members individually form the two lateral lobes of the labellum. Two developmental stages (floral primordium and organ-differentiated flowers) were selected for transcriptome sequencing. Two-thousand and seventy-five transcription factors were identified. Seven boundary genes and seven organ-specific genes were also discovered. Our study provides fundamental information for further studies on the molecular mechanisms of flower development and evolution across the Zingiberaceae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Zingiberaceae are a family of important and charismatic plants with significant ornamental, culinary, and medicinal value. The flowers of this family, colorful and morphologically diverse, are composed of two distinct and largely symmetric perianth whorls, a single fertile stamen, lateral staminodes, and a central abaxial labellum (Kirchoff 1988a). The calyx and corolla form the symmetrical perianth of Zingiberaceae, however these floral whorls are not conspicuous in the mature flower and form only a minor part of the overall floral display. The labellum, a distinctive organ comprised of 2–4 fused androecial members, is the most distinctive feature of Zingiberaceae flowers. The evolution of the six androecial members of Zingiberales has received extensive attention, especially focusing on petaloidy and fusion (Kirchoff 1983, 1988b, 1991; Specht et al. 2012; Almeida et al. 2013, 2014, 2015a, b; Piñeyro-Nelson et al. 2017; Specht and Almeida 2017). However, the diversity and evolution of organ composition of the labellum across the Zingiberaceae remains unexplored. At present, the most widely accepted interpretation is that the labellum develops from the fusion of two inner androecial members (Lestiboudois 1829; Eichler 1884). However, the labellum of different genera in Zingiberaceae are morphologically varied (Fig. 1), and this interpretation was based solely on research on species of the genus Hedychium, characterized by a two-lobed labellum. Later studies were limited to genera displaying an unlobed labellum, including Alpinia and Amomum, or additional taxa with a bilobed labellum including species of Hedychium and Kaempferia (Schumann 1904; Schachner 1924; Kirchoff 1997, 1998). A systematic understanding of how the three-lobed labellum develops in Zingiberaceae is still lacking.
Zingiber zerumbet is an important medicinal and ornamental plant. Although extensive research has been conducted in Z. zerumbet on its medicinal, chemical, pharmacological uses and its ornamental horticultural potential (Yob et al. 2011), no comprehensive description of its flower development has been reported. Therefore, Z. zerumbet, characterized as a species with a three-lobed labellum, is valuable for investigating how the three-lobed labellum develops in Zingiberaceae at both morphological and molecular levels. Likewise, understanding the molecular genetics underlying the development of the three-lobed labellum will provide insights into the processes involved with labellum formation more generally, including, but not limited to, staminode laminarity and organ fusion.
The molecular basis of floral development has been explored in depth in the model plant Arabidopsis thaliana, supplemented by developmental genetic studies, identifying key regulatory genes involved in the specification of floral organ identity (Bowman et al. 1991; Coen and Meyerowitz 1991; Weigel and Meyerowitz 1994; Theiβen 2001). Most of the key floral identity genes belong to the MADS-box gene family, individually or jointly determining the specification of the identity of floral organs as components of floral whorls. In Zingiberales, petaloid staminodes develop as part of the androecial whorl in four of eight families (Kirchoff et al. 2009), and in Zingiberaceae and Costaceae 2–5 of these staminodes can fuse to form the labellum. Several of the floral identity MADS-box genes have been isolated from Zingiberales to investigate their potential roles in the formation of petaloid staminodes and the resulting labellum (Bartlett and Specht 2010; Song et al. 2010; Yockteng et al. 2013; Almeida et al. 2013, 2015a; Fu et al. 2014). Within Zingiber, investigation of molecular development has focused on Z. officinale (Bartlett and Specht 2010; Yockteng et al. 2013).
As the organ identity of the labellum is thus understood, we can look to genes involved in generating particular organ morphologies (e.g. laminarity; Almeida et al. 2013) that contribute to the overall structure of the labellum. In the Zingiberaceae, it is largely understood that the labellum is formed by the fusion of the primordia of several staminodes belonging to one or both of the androecial whorls. Floral organ fusion provides great potential for flower diversity (Endress 2011). Fusion, more accurately defined as a lack of organ separation, is a process associated with the formation of distinct boundaries during primordia growth and organogenesis (Specht and Howarth 2015). Research in various model species has established that several key genes including CUP-SHAPED COTYLEDON 1–3 (CUC1-3), JAGGED LATERAL ORGANS (JLO), LATERAL ORGAN BOUNDARIES (LOB), KNOTTED1-LIKE HOMEOBOX GENE 6 (KNAT6) and LIGHT-DEPENDENT SHORT HYPOCOTYLS 3 (OBO1/LSH3) contribute to boundary region specification, and loss-of-function mutants in these genes usually demonstrate a phenotype that includes organ fusion (Aida et al. 1997; Belles-boix et al. 2006; Borghi et al. 2007; Takeda et al. 2011; Wang et al. 2016). Additional organ-specific genes may play significant roles in boundary specification during flower development within and between particular whorls. These organ-specific genes are expressed specifically in floral organs or regions between floral organs, and include PETAL LOSS (PTL) (expressed in regions between sepal primordia), RABBIT EARS (RBE) (expressed in petal primordia) and SUPERMAN (SUP) (expressed in stamen primordia) (Sakai et al. 1995; Krizek et al. 2006; Lampugnani et al. 2012). Characterizing the expression of these genes at different stages of floral development in Z. zerumbet can help determine how these genes are involved in the development of the Zingiberaceae labellum.
There are various challenges to study the genetic mechanisms of floral development in non-model systems, however these are the very plants in which interesting morphologies, such as the labellum, often evolve. Although Z. zerumbet is not a traditional model system for plant biology studies, it is a suitable system for the study of stamen polarity and floral organ fusion. RNA-seq has obtained rapid adoption in recent years to reveal relative patterns of gene expression during development, providing a precious opportunity for genomic exploration in non-model plant species that lack inbred and mutant lines for genetic screening. The genome of Musa acuminata (Musaceae; Zingiberales), published in 2012 (D’Hont et al. 2012), provides a reference genome that serves as a tool for the development of genome-based resources for both phylogenetic (Sass et al. 2016) and transcriptome or gene expression studies (Almeida et al. 2018) across the Zingiberales order. The increasing numbers of floral and organ-specific transcriptomes sequenced (Zhang et al. 2013; Huang et al. 2015; Li et al. 2017; Almeida et al. 2018) combined with a reference genome from closely related species M. acuminata (D’Hont et al. 2012) make it possible to study the molecular and genetic mechanisms involved in the development of the Z. zerumbet flower.
To this end, we conducted de novo transcriptome sequencing of two floral developmental stages for Z. zerumbet and used comparative transcriptomics to investigate gene expression patterns associated with floral development and specifically the formation of the labellum. To our knowledge, this is the first comprehensive transcriptomic study of flower development for Z. zerumbet, providing important bioinformatic resources for further investigation of genes involved in flower development in this species, and building a foundation for investigating the role of these genes and gene networks in the evolution of floral diversity across the Zingiberales.
Materials and methods
Plant materials
Young inflorescences of Z. zerumbet were collected from April to September in 2017 (for transcriptome sequencing and qRT-PCR) and 2018 (for scanning electron microscopy, SEM) from South China Botanical Garden, Chinese Academy of Sciences (SCBG, CAS), Guangzhou, China.
Scanning electron microscopy
Bracts and larger floral organs were removed under a dissecting microscope, and the floral buds were preserved in 2.5% glutaraldehyde and 2% paraformaldehyde overnight. All floral buds were washed in phosphate buffer three times in 2 h and dehydrated in an alcohol series (30%, 50%, 70%, 80%, 90%, 100%, 100%, 100%). The materials were freeze-dried with a Leica EM 300, mounted on stubs, gold-coated in a Leica EM ACE600, and observed under a JSM-6360LV SEM (JEOL) operated at 10 kV.
RNA extraction and quality verification
Samples of flower primordia (Zp) and flowers showing organ differentiation (Zd) were removed from inflorescences, flash frozen in liquid nitrogen and maintained at − 80 °C for subsequent RNA extraction. Each sample was combined from three inflorescences to reduce the effect of variation between individuals. Three biological replicates were performed for each Zp and Zd sample respectively.
Total RNA used for transcriptome sequencing was extracted with a mirVana™ miRNA Isolation Kit (Thermo Fisher Scientific, USA) according to the manufacturer’s protocol and treated with DNase I (TianGen, China). The integrity of total RNA was verified by gel electrophoresis and with an Agilent Bioanalyzer 2100 (Agilent Technologies, USA) following the criteria of RIN values (RNA Integrity Number) greater than 9.0. The amount of RNA was quantified with a Nanodrop 2000 (Thermo Fisher Scientific, USA).
Library construction and sequencing
After total RNA extraction from floral primordia or organ-differentiated flower tissue, eukaryotic mRNA was enriched by Oligo(dT) beads and fragmented using divalent cations under elevated temperature in Illumina proprietary fragmentation buffer, followed by reverse transcription into cDNA with random primers. Second-strand cDNA were synthesized by DNA polymerase I, RNase H, dNTP and buffer. The resulting cDNA fragments were purified with a QiaQuick PCR extraction kit. End repair, poly(A) addition, and ligation to sequencing adapters were performed (He and Jiao 2014). The ligation products were size selected by agarose gel electrophoresis, PCR amplified (He and Jiao 2014). The libraries were sequenced using Illumina HiSeq™ 4000 by Gene Denovo Biotechnology Co (Guangzhou, China).
Transcriptome de novo assembly and functional annotation
Raw sequence reads were processed by removing adapters, unknown nucleotides, and low-quality sequences, and the remaining cleaned reads were assembled using Trinity v2.1.0 as previously described for de novo transcriptome assembly in the absence of a reference genome (Grabherr et al. 2011).
Unigenes were annotated by using BLASTx program with an E-value threshold of 1e-5 to NCBI non-redundant protein (Nr) database (https://www.ncbi.nlm.nih.gov), Swiss-prot protein (Swiss-prot) database (https://www.expasy.ch/sprot), Clusters of orthologous groups for eukaryotic complete genomes (KOG) database (https://www.ncbi.nlm.nih.gov/COG) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (https://www.genome.jp/kegg). Gene Ontology (GO) annotation was analyzed by Blast2GO 2.3.5 program based on Nr annotation (Conesa et al. 2005).
Analysis of differential gene expression
Gene abundances were calculated as RPKM (Reads per Kilobase per Million mapped reads). The differentially expression analysis between two groups was operated using the edgeR 3.12.1 (https://www.r-project.org/) (Robinson et al. 2009). Genes with a fold change ≥ 2 and the false discovery rate (FDR) < 0.05 was considered as the threshold to judge the significantly differential expression. GO enrichment analysis and KEGG pathway analysis of DEGs were implemented using an online website OmicShare tools (https://www.omicshare.com/tools/).
Real-time quantitative PCR validation of RNA-seq data
Twelve unigenes were selected for validation using Real-time quantitative PCR (qRT-PCR). Total RNA was extracted from flowers at the same two stages as for RNA-seq using a mirVana™ miRNA Isolation Kit (Thermo Fisher Scientific, USA). First-stand cDNA was synthesized using a cDNA Synthesis SuperMix with gDNA Remover (TransGen Biotech, China). PCR primers were designed with Integrated DNA Technologies PrimerQuest tool (https://sg.idtdna.com/primerquest/home/index) and are listed in Supplementary Table S1. qRT-PCR reactions were performed using Top Green qPCR SuperMix (TransGen Biotech, China) in a total volume of 10 μl reaction mixture containing 5 μl of Top Green qPCR SuperMix, 0.2 μl (10 μmol/l) of each primer, 0.2 μl Passive Reference Dye II (50×), 4 μl of water, and 0.4 μl of cDNA template. The qRT-PCR amplification was performed with ABI Prism 7500 Fast Real-time PCR Detection system under the program of 95 °C for 30 s, followed by 40 cycles of 95 °C for 15 s and 60 °C for 35 s. Dissociation stage condition was set at 95 °C for 15 s, 60 °C for 60 s, 95 °C for 15 s, and 60 °C for 15 s to test the specificity of primers. Each reaction was performed in three technical replicates. The β-Actin gene was used as an internal control for normalization. The comparative Ct method (2−∆∆Ct) was used to calculate the relative quantities of transcripts (Livak and Schmittgen 2001).
Results
Flower morphology
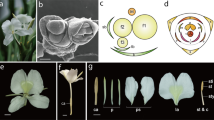
The conical inflorescence of Z. zerumbet is green when young and crimson when aged, formed by closely imbricate bracts (Fig. 2a, b). Three to four pale yellow flowers emerge synchronously and can reach anthesis on the same day (Fig. 2a). Flowers are borne in the axil of the spirally arranged bracts with a single flower produced per bract (Fig. 2c; B). Each single flower is surrounded by a membranous bracteole (Fig. 2c; b). Zingiber zerumbet has a zygomorphic floral structure resulting from the monosymmetric androecial whorls (Fig. 2d). The floral structures include a fused (synsepalous) calyx tube (Fig. 2c, e; ca), a corolla tube with three petal lobes (Fig. 2d, e; p1, p2 and p3), a labellum (Fig. 2d, e; la) with a subobovate central lobe (Fig. 2f; cl) and two obovate lateral lobes (Fig. 2f; ll), a fertile stamen (Fig. 2e, g; st) comprising two thecae and a connective with one beaklike stamen appendage (Fig. 2g; sa), and a pistil (Fig. 2e; pi, Fig. 2g; sty, sti).
Inflorescence and flower structure of Z. zerumbeta Green conical inflorescence of Z. zerumbet (lateral view). Three to four flowers emerge and open almost simultaneously. b Crimson inflorescence when aged (apical view). c Lateral and frontal view of pre-anthesis flower showing bract, bracteole and calyx. d A mature flower at anthesis. e Dissected floral organs of a mature flower. f Three-lobed labellum of Z. zerumbet. g Stamen and pistil structure. B bract, b bracteole, ca calyx, cl central lobe, f flower, la labellum, ll lateral lobe, p petal, pi pistil, sa stamen connective appendage, st stamen, sti stigma, sty style, t theca. Scale bars = 0.5 cm in f and 1 cm in c, e, g
Inflorescence and flower development
Reproductive growth began with the differentiation of bracts along the vegetative shoot. The bract primordia initiated acropetally in a spiral manner around the inflorescence axis with the inflorescence meristem (IM) on the top (Fig. 3a). Flower development began following the initiation of the first bract. A floral buttress (fb) appeared in the axil of the bract, subtended by each bract primordium (Fig. 3a). The floral primordium (fp) then separated from the main apex, enlarged, flattened apically and assumed a rounded, obdeltoid appearance in the polar view (Fig. 3b). After the floral primordium was initiated, a bulge gradually formed at the base of the adaxial side of the floral primordium, which later developed into a crescent shaped bracteole primordium (bp) (Fig. 3b). Development of the floral organs began with the initiation of sepal primordia (se). Three sepal primordia appeared sequentially, either clockwise or counterclockwise, from the three angle corners of the obdeltoid region of floral primordium (Fig. 3c). Thereafter, the sepal primordia extended through intercalary growth until the margins of the adjacent sepal primordia became confluent, gradually separating from the central part of floral primordium to produce a synsepalous calyx (Fig. 3d; ca). During sepal initiation, a bulge initiated at the adaxial side of the floral primordium, defined as a common primordium (Fig. 3d; cp1). This adaxial common primordium enlarged rapidly. Subsequently, two other common primordia (cp2, cp3) formed almost synchronously at the abaxial side of the floral primordium (Fig. 3e) alternate to the position of the sepal primordia. These three common primordia were triangularly arranged, and the unequal development resulted in the adaxial common primordium (cp1) to become obviously larger than the two abaxial common primordia (cp2, cp3) which appeared to be of equal size (Fig. 3e). The triangular center surrounded by three common primordia formed a depression which became deeper and deeper with the enlargement and separation of the common primordia, later developing into a “flower cup” (Fig. 3e, f; fc) from which the gynoecial primordium later emerge. Separation of the three common primordia, in sequential order, commenced from the differentiation of the adaxial common primordium (cp1). The dorsal part of the adaxial common primordium (cp1) developed into the adaxial petal (p1) while the ventral part developed into a single fertile functional stamen (Fig. 3f; s). The fertile stamen bears two thecae (Fig. 3g; t). Each lateral abaxial common primordia subsequently separated to produce a lateral petal (p2, p3) to the exterior and a lateral inner androecial member (ia1, ia2) to the interior (Fig. 3g). Later, the two lateral inner androecial primordia (ia1, ia2) merged through intercalary growth and fused partially with each other from the base to constitute the central lobe (cl) of the labellum (Fig. 3h; cl).
Flower development of Z. zerumbet. a Apical view of a conical shaped inflorescence of Z. zerumbet. Bract primordia initiated upwards in a spiral manner around the inflorescence axis with the inflorescence meristem (IM) on the top. Floral buttress (fb) appeared. b Initiation of floral primordium (fp) and formation of bracteole primordium (bp). c Sepal primordia (se) initiation. d Common primordium(cp1) initiation and formation of ring calyx primordium (ca). e Common primordia (cp2 and cp3) initiation and formation of flower cup (fc). f Separation of adaxial common primordium (cp1) to form petal (p1) and fertile stamen (s) of the inner androecial whorl. g Separation of two abaxial/lateral common primordium (cp2, cp3) to form petal (p2, p3) and inner androecial members (ia1, ia2). The fertile stamen comprised two thecae (t). h Formation of outer androecial members (oa1, oa2). Fusion of two inner androecial members (ia1, ia2) to form central lobe of labellum (cl). i Calyx gradually enclosed other floral organs. j *Indicates the initiation position of gynoecial primordium. k Frontal view of flower with calyx and petals removed; gynoecial primordium forming the stigma (sti). l Oblique view of stamen with stigma removed. m Lateral view of flower showing extending style (sty) and outer androecial member (oa). Central lobe of labellum (cl) further developed with emarginate apex. n Frontal view of flower showing extending style (sty). Stamen appendage (sa) emerged. o Part of n showing stamen appendage (sa). p Lateral view showing further elongated style (sty) and stamen appendage (sa). q The frontal view of p showing beaklike stamen appendage (sa) with style (sty) removed. r Lateral view of p showing theca (t) with beaklike stamen appendage (sa). s Development of outer androecial member (oa) to form lateral lobe (ll) of labellum. t Part of s showing the base of lateral lobe (ll) was connected with central lobe of labellum (cl). ab abaxial side, ad adaxial side, bp bracteole primordium, ca calyx, cl central lobe of labellum, cp common primordium, fb floral buttress, fc flower cup, fp floral primordium, ia inner androecial primordium, IM inflorescence meristem, ll lateral lobe of labellum, oa outer androecial primordium, p petal, s stamen, sa stamen appendage, se sepal primordium, sti stigma, sty style, t theca. Scale bars = 200 μm in (a, n, p, r), 100 μm in (i, j, k, l, m, o, q, s, t) and 50 μm in (b, c, d, e, f, g, h)
After the differentiation of the inner androecium from the common primordia with the petals, two adaxial/lateral outer androecial primordia (oa1, oa2) initiated on either side of the fertile stamen (Fig. 3h). As the floral organs developed, the synsepalous calyx gradually enclosed the internal floral organs (Fig. 3i). At this stage, fertile stamen and petals appeared to become independent and distinct (Fig. 3i).
Following the initiation of almost all other floral organ primordia, the gynoecial primordium initiated from the depression in the central part of the floral primordium (Fig. 3j). The growth rate of the gynoecium was significantly faster than that of fertile stamen and the central lobe of labellum after initiation, extending longitudinally and soon forming the stigma (sti) and style (sty) (Fig. 3k–n, p) (See supplementary materials Fig. S1 for a more complete development of flower cup and carpel). With the stamen development to some extent (Fig. 3k–m), the stamen connective appendage (sa) began to differentiate upwards and ultimately formed a beaklike appendage which enclosed part of the style (Fig. 3n–r).
Almost simultaneously, the two adaxial/lateral outer androecial primordia (oa1, oa2) continued to enlarge and developed into two lateral lobes (ll) which became adnate to the central lobe of the labellum (Fig. 3s, t), ultimately forming a three-lobed labellum.
Description of developmental events
To describe flower development of Z. zerumbet more accurately, we divided the continuous process of flower development into 11 stages (as shown in Fig. 4) using a series of landmark events based on our observations. We fully considered the uniqueness of development process in Z. zerumbet flower while referring to the stages division of early flower development in A. thaliana (Smyth et al. 1990). Stage 1 (corresponding to Fig. 3a) begins with the initiation of a floral buttress on the flank of inflorescence meristem (IM). Stage 2 (corresponding to Fig. 3b) commences when the floral primordium (fp) separates from the meristem with the bracteole primordium (bp) forms. Sepal primordia (se1, se2, se3) then arise (Fig. 3c; Fig. 4 Stage 3). Adaxial common primordium (cp1) arise and the synsepalous calyx (ca) forms (Fig. 3d; Fig. 4 Stage 4). Two abaxial common primordia (cp2, cp3) arise and the “flower cup” (fc) appears (Fig. 3e; Fig. 4 Stage 5). During Stage 6, the adaxial common primordium (cp1) develops into an adaxial petal (p1) and the fertile stamen (Fig. 3f). In Stage 7, the abaxial common primordia (cp2, cp3) separate into the abaxial petal (p2, p3) and the inner androecial members (ia1, ia2) (Fig. 3g). The two inner androecial members (ia1, ia2) later form the central lobe (cl) of the labellum, and the outer androecial primordia initiate (Figs. 3h, 4 Stage 8). When the calyx gradually encloses all internal floral organs, the carpel primordia (c) initiate from the flower cup region to form the style (Figs. 3i–l, 4 Stage 9). During Stage 10, the style grows rapidly and becomes longer than the stamen and the central lobe (cl), and the stamen connective appendage (sa) begins to differentiate (Fig. 3m–r). At this point, the outer androecial primordia (oa1, oa2) form the two lateral lobes (ll) of the labellum, generating the three-lobed labellum (Fig. 3s, t; Fig. 4 Stage 11). During this last stage, all floral organs become mature.
Eleven developmental stages of Z. zerumbet flower. Stage 1: flower buttress arises. Stage 2: floral primordium and bracteole primordium form. Stage 3: sepal primordia arise. Stage 4: synsepalous calyx and adaxial common primordium arise. Stage 5: abaxial common primordia and flower cup arise. Stage 6: adaxial petal and fertile stamen arise. Stage 7: abaxial petals and inner androecial primordia arise. Stage 8: central lobe forms and outer androecial primordia arise. Stage 9: calyx enclose inner floral organs and carpel primordium arises. Stage 10: style grows rapidly and stamen connective appendage differentiates. Stage 11: three-lobed labellum forms and all floral organs mature. ab abaxial side, ad adaxial side, B bract (primordium), bp bracteole primordium, c carpel (primordia), ca calyx, cl central lobe of labellum, cp common primordium, fb flower buttress, fc flower cup, fp floral primordium, IM inflorescence meristem, ia inner androecial primordium, ll lateral lobe of labellum, oa outer androecial primordium, p petal, s stamen, sa stamen connective appendage, se sepal primordium, *indicates aborted stamen. The color of carpel represents its longitudinal depth. The darker the color, the longer the style
Transcriptome sequencing and assembly
To acquire a comprehensive knowledge of the flower development process, two distinct developmental stages including a floral primordium stage (Zp, Fig. 3b) and a differentiated flower stage (Zd, Fig. 3c–m) were selected as materials to perform transcriptome sequencing. A total of 451,572,534 raw reads with a length of 2 × 150 bp each were generated from six RNA-seq libraries (3 × floral primordium and 3 × organ-differentiated flower for three biological replicates each). After removing low quality reads and adapters, a total of 431,451,702 high quality clean reads were processed for assembly using Trinity (Grabherr et al. 2011). A total of 96,980 unigenes with a mean size of 877 bp and N50 length of 1521 bp were obtained from de novo assembly, the length of which ranges from 201 to 15,876 bp. Sixteen-thousand six-hundred and seventeen (16,617) unigenes were ≥ 1.5 Kb in length and 4181 unigenes were ≥ 3 Kb in length. The length distribution of unigenes is shown in Supplementary Table S2.
Gene annotation and functional classification
To predict and analyze the function of the assembled unigenes, BLASTx was used to perform a homology search against public databases including Nr, Swiss-prot, KOG and KEGG database. A total of 41,278 unigenes were annotated by aligning them with known genes in these databases and the distribution in databases are shown in Fig. S2a. Among 96,980 unigenes, 41,027(42.3%) unigenes were annotated to Nr database, of which 10,192 unigenes showed an E-value less than 1E-150. Species similarity based on BLASTx analysis of unigenes in Nr database demonstrated that the highest ratio of matched sequences was derived from matches to Musa acuminata (47.5%, 19,493 unigenes, Fig. S2b), of the same order (Zingiberales) as Zingiber. The next-closest matches were monocot taxa Elaeis guineensis (4.5%, 1854 unigenes), Phoenix dactylifera (4.2%,1724 unigenes) and Anthurium amnicola (3.8%, 1558 unigenes) (Fig. S2b).
Annotations of total unigenes against the Swiss-prot database revealed that 27,642 (28.5%) unigenes showed significant hits against known sequences. 22,547 (23.2%) unigenes were mapped to KOG database and classified into 25 functional categories (Fig. S2c). The general function prediction category accounted for the highest percentage of genes (38.2%) among all categories, followed by signal transduction mechanisms (18.1%).
A total of 14,885 (15.3%) unigenes were assigned to 127 different pathways according to the KEGG database, which emphasizes on biochemical pathways. The details can be found in Supplementary Table S3. These pathways provide valuable basic information for exploring specific processes and functions for future research. A total of 7455 assembled unigenes were identified by the Gene Ontology and divided into three major functional categories (biological process, cellular component and molecular function) and 46 subcategories based on sequence homology, as shown in Supplementary Table S4.
Analysis of differential gene expression during flower development
Comparison of gene expression between Zp and Zd (Zp vs Zd) revealed that 868 unigenes displayed significant changes in expression (FDR < 0.05 and |log2Fold Change|> 1) in Zp vs Zd. Compared with Zp, 519 unigenes were up-regulated and 349 were down-regulated. GO term enrichment analysis was conducted for all DEGs (Fig. S3a). In biological process category, DEGs were largely enriched in metabolic process, cellular process and single-organism process. In molecular function category, catalytic activity and binding were the two dominant enriched terms. In cellular component category, the most represented items were cell, cell part and membrane. A total of 868 DEGs were assigned to 59 KEGG pathways. The most represented ten pathways are shown in Fig. S3b. The pathway with the largest proportion of represented terms was the pathway for plant hormone signal transduction. Of the total 20 DEGs enriched in this pathway, six were identified as auxin-response genes, indicating that auxin may have a large effect during the growth and differentiation of floral primordium.
Identification of transcription factors
In this study, 2075 transcription factors were identified, accounting for 2.14% of all unigenes and falling into 57 TF families classified by transcription factor database. Among the transcription factors, 22 were specifically expressed in Zp stage and 29 were specifically expressed in Zd stage. The stage-specific expressions can be found in Supplementary Table S5. The top ten transcription factors are shown in Fig. 5a. Among the detected transcription factor gene families, the ERF gene family accounted for the largest proportion (200, 9.6%), followed by bHLH (181, 8.7%), MYB_related (120, 5.8%) and C2H2 (116, 5.6%). Most notably, a total of 56 MADS-box genes, which encode 28 MIKC and 28 M-type transcription factors, were identified in transcriptome sequences and their expression patterns are presented in Fig. 5b. Among these MADS-box genes, several genes, including APETALA1 (AP1), CAULIFLOWER (CAL), APETALA2 (AP2), APETALA3 (AP3), AGAMOUS (AG), SEPALLATA3 (SEP3) and SEPALLATA4 (SEP4) are widely implicated in floral organ identity.
Predicted transcription factors. a Top ten percentages of predicted transcription factors. b Expression pattern of MADS-box transcription factors. Gene IDs corresponding to the expression pattern (right). M-type and MIKC-type categories represented in orange and blue bars, respectively (left). Genes were clustered by RPKM value with color scale representing the log-transformed RPKM value. *Indicate A-class (orange), B-class (blue), C-class (green) and E-class (purple) genes. c Expression pattern of differentially expressed transcription factors during flower development. Gene IDs listed on the right side, clustered by RPKM values. The color scale at right side represents the log-transformed RPKM value. Color bars in left side represent different transcription factors family and the corresponding colors refer to the TFs family identification (legend right)
A total of 67 DEGs encoding transcription factors associated with flower development were identified (Fig. 5c). These differentially expressed TFs were assigned to different transcription factor families including MYB (7), MYB-related (4), MIKC-type (13), M-type (6), YABBY (12), LBD (7), TALE (7), AP2 (5), NAC (3), WOX (2) and TCP (1). These include candidate genes such as AP1 (unigene0000116) and AP2 (unigene0080007 and unigene0057364) which were lower in expression at the Zd stage compared to early development, while SOC1 (unigene0034376), AG (unigene0049214), SEP3 (unigene0068145), YAB1 (unigene0036118 and unigene0042917), YAB2 (unigene0033130, unigene0041500 and unigene0061291) showed higher expression at the Zd stage compared to early development.
Identification of genes related to floral organ boundary specification
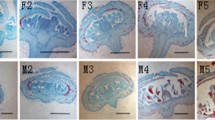
Genes related to floral organ boundary specification were also identified based on our de novo assembly and annotation. In all, seven boundary-specific genes including CUC2, NAM, JLO, KNAT6 and LSH3 were discovered (Fig. 6a). The expression of CUC2 (unigene0051169), NAM (unigene0051170), JLO-1(unigene0041785), JLO-2 (unigene0036065), JLO-3 (unigene0036066) and KNAT6 (unigene0048229) showed higher expression in the floral meristem primordia (Zp) than during floral organ differentiation (Zd) stage, while the expression of LSH3 (unigene0064476) increased during floral development. In particular, LOB4, 6, 18, 40, 41 which were previously identified in at least some members of the Zingiberales (Almeida et al. 2018) were also retrieved in the transcriptome data. Additionally, seven organ-specific genes including RBE, SUP and PTL were identified (Fig. 6b). The expression of RBE-1 (unigene0001424), RBE-2 (unigene0035273), SUP-1 (Unigene0019275) and SUP-3 (Unigene0035251) demonstrated a significant increase at the later developmental stage, while SUP-2 (unigene0091259) showed only a slight increase later in development. The expression of PTL-1 (Unigene0037058) and PTL-2 (Unigene0040470) showed a slight decrease during floral organ differentiation.
Expression of genes related to floral organ boundary specification. a Expression of boundary-specific genes and b expression of organ-specific genes. Clustering based on expression. Circles of different colors and sizes represent log-transformed RPKM expression values. The name of putative gene is assigned by sequence similarity and true orthologs should be further cloned and analyzed
qRT-PCR
To verify the accuracy and reproducibility of the transcriptome analysis, twelve unigenes were selected for validation in two developmental stages (Zp and Zd) using qRT-PCR (Fig. S4; three homologues of CUC and nine randomly selected DEGs). The expression patterns of all twelve genes analyzed by qRT-PCR were consistent in the overall trend with data obtained by RNA-Seq. The expression patterns of three CUC-like genes including unigene0051169, unigene0051170 and unigene0032755 showed reduced expression in Zd compared to Zp, suggesting that CUC-like genes may function early in flower development.
Discussion
Inflorescence and flower development
The development of six androecial members in Zingiberaceae, and especially the composition of the androecial labellum, is of fundamental interest from both evolutionary and developmental perspectives. The interpretation first proposed by Lestiboudois and slightly modified by Eichler remains the most widely accepted (Lestiboudois 1829; Eichler 1884); that the labellum of Zingiberaceae is formed by the fusion of two inner androecial members. The ventral, or abaxial, outer androecial member that initiates between the two ventral inner androecial members soon ceases to grow, contributing little to the formation of the labellum formed by the fusion of the two ventral inner androecial members. The adaxial, or dorsal, inner androecial member develops into a fertile stamen, while the two remaining outer whorl androecial primordia form two lateral staminodes. Developmental work conducted by Schachner and Kirchoff supported this interpretation of the labellum (Schachner 1924; Kirchoff 1997, 1998). This interpretation is mainly based on research in Hedychium which is characterized by a two-lobed labellum; however, the labellum is morphologically variable across the Zingiberaceae and can have zero, two or three lobes. Schumann (1904) proposed an interpretation building upon Eicher’s, in which the bilobed labellum of genera including Hedychium and Kaempferia comprises two fused inner whorl androecial members, while the unlobed labellum of genera including Alpinia and Amomum comprises a single outer whorl androecial member. In a thorough investigation of the vascularization of the labellum in seven Zingiberaceae genera (Burbidgea, Curcuma, Amomum, Hornstedtia, Hedychium, Kaempferia, and Alpinia), Costerus concluded that the labellum consistently consisted of two inner androecial members and one outer androecial member, represented by the medial vein of the labellum (Costerus 1915), regardless of whether the labellum was unlobed or bilobed.
Zingiber is characterized by a three-lobed labellum, which had not been considered in previous characterizations. The results presented in this paper partially support the general interpretation of labellum proposed by Eichler, with some differences. The two abaxial inner androecial members are joined through intercalary growth, fusing basally, and leaving a cleft at apex, to produce the central lobe of the labellum. The two adaxial outer androecial members form two lateral petaloid staminodes with their base adnate to the central lobe, thus developing into two lateral lobes on each side of the central lobe (Fig. 7a). A third abaxial outer androecial primordium, which should appear opposite the abaxial sepal, was not observed and is likely to either never initiate or to abort early during flower development. From this point of view, the three-lobed labellum of Z. zerumbet comprises four androecial members: two abaxial inner androecial and two adaxial outer androecial members. This result provides new insights into the origin and evolution of the six androecial members and the interpretation of the labellum in Zingiberaceae.
In the cone-shaped spiral inflorescence of Z. zerumbet, flowers differentiate synchronously (Fig. 7b). The flowers differentiate acropetally, from the base towards the apex, along the inflorescence. This flowering characteristic forms a unique ornamental character: flowers in the lower part of the inflorescence have withered and the bracts appear crimson, while flowers in the upper part of the inflorescence are emerging and the bracts remain green. This unique flowering feature prolongs the florescence and improves the ornamental value to some extent, which makes Z. zerumbet a popular tropical garden ornamental and a high-grade cut flower.
Genes related to floral organ identity specification
Based on our transcriptome data, A-class genes AP1 and AP2 show higher expression in the floral meristem primordia (Zp) stage, while the expression of C-class gene AG and E-class gene SEP3 are significantly up-regulated in the organ-differentiated flower (Zd) stage. A possible explanation for this might be that A-class genes are involved in floral meristem maintenance and drive the formation of sepals, which is the ground state of the flower that initiates earlier than other floral organs (Causier et al. 2010). As such, A-class genes begin to function prior to the initiation of sepal primordia defining the floral meristem, evidenced by higher expression in Zp. In Arabidopsis, C-class genes function in the formation of stamen and carpel whorls while E-class genes play a role in the formation of all four whorls (Pelaz et al. 2000; Theiβen 2001; Ditta et al. 2004). Our finding that the expression of C- and E-class genes in organ-differentiated flower (Zd) is higher than in floral meristem primordia (Zp) is consistent with studies in model organisms in which these genes are involved in the development of organ identity and the definition of organ placement along the floral meristem. Our results showing higher expression of YAB1,2 in differentiated flowers (Zd) may be indicative of their active role in the determination of floral organ abaxial–adaxial polarity during floral organ differentiation (Morioka et al. 2015).
The mechanism of MADS-box genes for floral organ identity specification has been well established in model plants including A. thaliana and Antirrhinum majus (Coen and Meyerowitz 1991; Weigel and Meyerowitz 1994). It has been shown that three classes of homeotic genes (A-class, B-class and C-class) determine the identity of the four concentric whorls (sepals, petals, stamens and carpels) in flowers. A-class genes specify sepal identity in whorl 1; A-class genes and B-class genes play a joint role in specifying petal identity in whorl 2; B-class genes and C-class gene codetermine stamen identity in whorl 3; Carpel identity in whorl 4 is conferred by C-class gene alone. In Zingiberales, there are two concentric stamen whorls, and petaloid staminodes replace many of the fertile stamens in four out of eight families. This type of homeotic conversion was hypothesized to be possibly caused by differential expression of B- and C-class genes in the petaloid staminodes of ginger families in Zingiberales causing a shift in organ identity (Wake et al. 2011). Research in Alpinia hainanensis (Zingiberaceae) and Canna indica (Cannaceae) provide evidence that C-class gene is expressed in both petaloid staminode and fertile stamen (Song et al. 2010; Almeida et al. 2013; Fu et al. 2014; Tian et al. 2016) and that shifts in expression may be less critical than differential selection or alterations in gene function (Almeida et al. 2013). Additionally, comparative evolutionary studies of B-class genes across Zingiberales demonstrates shifts in patterns of expression not necessarily correlated with stamen fertility or stamen v. petal identity (Bartlett and Specht 2010). Instead, patterns of gene duplication and gene family evolution show differential recruitment of gene throughout the evolution of the order, with development proceeding in a relatively canalized manner despite selection on gene function and the diversification of particular expression patterns (Yockteng et al. 2013). Recent data indicates that genes involved in developmental processes other than organ identity, such as organ polarity (Almeida et al. 2014; Fu et al. 2014; Morioka et al. 2015) and fusion (Specht and Howarth 2015) are more likely to result in morphologically differences observed throughout the evolution and diversification of Zingiberales floral form (Yockteng et al. 2014). While expression patterns and selection on genes can provide insights into the evolution of development across the order, we still lack the ability to transform taxa across the Zingiberales and test for phenotypes associated with loss of function in genes critical for floral development. Efforts in generating a virus-induced gene silencing system for Z. zerumbet (Mahadevan et al. 2015) and within the Zingiberales (Renner et al. 2009) combined with efforts on tissue culture and transformation for CRISPR, provide a preliminary basis for future studies on gene function during development.
Boundary and organ-specific genes may be involved in floral organ fusion during the formation of the synsepalous calyx and the three-lobed labellum
The whole flower transcriptomes of six exemplar species across the Zingiberales, including species from Musaceae, Lowiaceae, Zingiberaceae, Costaceae, Marantaceae and Cannaceae, have been analyzed in previous studies (Almeida et al. 2018). LOB40, 41 and 6 homologs retrieved in those Zingiberales floral transcriptomes were also identified in the transcriptomes of Z. zerumbet flower. LOB4, recovered previously in Z. officinale, was also retrieved in Z. zerumbet. What difference from previous findings is that LOB18, previously recovered only in the Cannaceae-Marantaceae lineage, was here retrieved in Z. zerumbet. LOB genes have been reported to play roles in defining organ boundaries in Arabidopsis floral organs through negative regulation of the accumulation of brassinosteriod (Shuai et al. 2002; Bell et al. 2012). Additionally, CUC2, also recovered only in the Cannaceae-Marantaceae lineage, was here retrieved in Z. zerumbet while PTL, recovered previously in Z. officinale, was also identified in the floral transcriptome of Z. zerumbet.
Fusion of floral organs exists in various forms in angiosperm, and even in closely related plants, the fusion of different organs or organs from different whorls may produce significant patterns contributing to morphological diversity of floral forms (Endress 2011; Specht and Howarth 2015). Several boundary genes, including CUC1-3, JLO, LOB, KNAT6, OBO1/LSH3 and OBO4/ LSH4, with redundant yet specialized functions, appear to be active during the development of different whorls of floral organs, contributing to fusion within (connation) or between (adnation) organ whorls. Additionally, some organ-specific genes, including PTL and SUP, may be involved in organ-specific boundary formation. The expression and interactions of these organ-specific and boundary genes in floral primordia and organs during floral development were summarized in a proposed boundary formation gene regulatory network (GRN) (Specht and Howarth 2015). If this proposed GRN functions as a conserved boundary-specification mechanism across angiosperms, the genes involved in this network may also be responsible for floral fusion phenotypes occurring in Z. zerumbet. Under this GRN, PTL, which encodes a trihelix transcription factor, is a boundary gene expressed in boundaries between sepal primordia in the outer whorl (Brewer et al. 2004). PTL plays a role in constraining the size of inter-sepal zone by inhibiting growth between developing sepals in A. thaliana (Lampugnani et al. 2012). The ptl mutants demonstrate a phenotype of fusion of adjacent sepals (Lampugnani et al. 2012). Likewise, cuc1cuc2 double mutants also show a high level of fusion between adjacent sepals (Aida 1997). These three genes share their functions in limiting inter-sepal growth by different mechanisms, and may play joint roles in the formation of synsepalous calyx of Z. zerumbet. Results presented here demonstrate the expression of CUC2 and PTL is down-regulated in differentiated flowers in comparison with the floral primordium, indicating these two genes may function in the early stage of floral development. In addition, SUP, a zinc finger protein expressed specifically in stamen in Arabidopsis, acts to maintain a boundary between adjacent groups of cells after differentiation (Sakai et al. 1995). Two SUP candidate genes are significantly down-regulated in differentiated Z. zerumbet flowers (Zd), suggesting that the low expression of SUP may be responsible for the loss of boundary between the four stamen primordia during the formation of three-lobed labellum.
In comparison with early floral developmental stages of A. thaliana flower, observed differences in floral organ development of Z. zerumbet begin at Stage 4, with the emergence of the three common primordia. While the existence of common primordia, such as those that emerge during the early floral development of many Zingiberales plants (e.g. Zingiberaceae, Heliconiaceae and Costaceae; Kirchoff 1988b, 1997; Kirchoff et al. 2009), are characteristic of many floral lineages, the gene expression patterns of these primordia and the gene expression patterns underlying differentiation of floral organs from these common primordia has not been studied. It is possible that boundary genes are required to make the demarcation between floral organ identity specification that ultimately results in the emergence of differentiated floral organs from the common primordia. Genes such as CUC, PTL and SUP may play a role in the differentiation of petal and stamen organs, for example, from the common primordium of Z. zerumbet. These may precede genes involved in other morphogenetic processes, such as abaxial/adaxial polarization leading to laminar structures such as the petals or petaloid staminodes. Further analyses are necessary to establish a gene regulatory network involved in the formation of ultimate organ differentiation resulting from common primordia structures. The findings presented in this paper provide a direction for future studies into the molecular mechanisms of floral development and diversification within Zingiber and even across the Zingiberales.
In conclusion, this study presents a comprehensive flower development description of Z. zerumbet and adds new knowledge to the understanding of the development of the labellum in Zingiberaceae using comparative transcriptomic methods. Although further research is needed to investigate what specific roles the identified developmental genes play during flower development and organ differentiation, the transcriptome data in this study provides critical information for future studies on the mechanism of flower development in Z. zerumbet by providing a global list of genes that may be involved in floral organ identity specification and floral organ boundary formation. Data for Z. zerumbet also provides information for evolutionary studies aimed at identifying the processes involved in the development and evolution of organ identity, organ morphology, and floral organ boundary formation across the Zingiberales and in other lineages of flowering plants.
Data availability
The sequencing data were deposited in the National Center for Biotechnology Information Short Read Archive (NCBI-SRA) database (https://www.ncbi.nlm.nih.gov/Traces/sra) under accession number PRJNA540419.
References
Aida M, Ishida T, Fukaki H et al (1997) Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9:841–857
Almeida AMR, Brown A, Specht CD (2013) Tracking the development of the petaloid fertile stamen in Canna indica: insights into the origin of androecial petaloidy in the Zingiberales. AoB Plants 5:1–7
Almeida AMR, Yockteng R, Schnable J et al (2014) Co-option of the polarity gene network shapes filament morphology in angiosperms. Sci Rep 4:6194
Almeida AMR, Yockteng R, Otoni WC, Specht CD (2015a) Positive selection on the K domain of the AGAMOUS protein in the Zingiberales suggests a mechanism for the evolution of androecial morphology. Evodevo 6:1–15
Almeida AMR, Yockteng R, Specht CD (2015b) Evolution of petaloidy in the Zingiberales: an assessment of the relationship between ultrastructure and gene expression patterns. Dev Dyn 244:1121–1132
Almeida AMR, Pineyro-Nelson A, Yockteng R et al (2018) Comparative analysis of whole flower transcriptomes in the Zingiberales. PeerJ 6:e5490
Bartlett ME, Specht CD (2010) Evidence for the involvement of GLOBOSA-like gene duplications and expression divergence in the evolution of floral morphology in the Zingiberales. New Phytol 187:521–541
Bell EM, Lin WC, Husbands AY et al (2012) Arabidopsis LATERAL ORGAN BOUNDARIES negatively regulates brassinosteroid accumulation to limit growth in organ boundaries. Proc Natl Acad Sci USA 109:21146–21151
Belles-Boix E, Hamant O, Witiak SM et al (2006) KNAT6: an Arabidopsis homeobox gene involved in meristem activity and organ separation. Plant Cell 18:1900–1907
Borghi L, Bureau M, Simon R (2007) Arabidopsis JAGGED LATERAL ORGANS is expressed in boundaries and coordinates KNOX and PIN activity. Plant Cell 19:1795–1808
Bowman JL, Smyth DR, Meyerowitz EM, Meyerowitzt EM (1991) Genetic interactions among floral homeotic genes of Arabidopsis. Development 112:1–20
Brewer PB, Howles PA, Dorian K et al (2004) PETAL LOSS, a trihelix transcription factor gene, regulates perianth architecture in the Arabidopsis flower. Development 131:4035–4045
Causier B, Schwarz-Sommer Z, Davies B (2010) Floral organ identity: 20 years of ABCs. Semin Cell Dev Biol 21:73–79
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
Conesa A, Götz S, García-Gómez JM et al (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Costerus JC (1915) Das Labellum und das Diagram der Zingiberaceen. Ann Jardin Botanique Buitenzorg II 14:95–108
D’Hont A, Denoeud F, Aury JM et al (2012) The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488:213–217
Ditta G, Pinyopich A, Robles P et al (2004) The SEP4 gene of Arabidopsis thaliana functions in floral organ and meristem identity. Curr Biol 14:1935–1940
Eichler AW (1884) Uber den bluthenbau der Zingiberaceen. Sitzungsber der Königlich Preuss. Akad Wiss 26:585–600
Endress PK (2011) Evolutionary diversification of the flowers in angiosperms. Am J Bot 98(3):370–396
Fu Q, Liu H, Almeida AMR et al (2014) Molecular basis of floral petaloidy: insights from androecia of Canna indica. AoB Plants 6:490–552
Grabherr MG, Haas BJ, Yassour M et al (2011) Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat Biotechnol 29(7):644–652
He J, Jiao Y (2014) Next-generation sequencing applied to flower development: RNA-seq. In: Riechmann JL, Wellmer F (eds) Flower development: methods and protocols, vol 1110. Springer, New York, pp 401–411
Huang JZ, Lin CP, Cheng TC et al (2015) A de novo floral transcriptome reveals clues into Phalaenopsis orchid flower development. PLoS ONE 10:1–20
Kirchoff BK (1983) Floral organogenesis in five genera of the Marantaceae and in Canna (Cannaceae). Am J Bot 70:508–523
Kirchoff BK (1988a) Floral ontogeny and evolution in the ginger group of the Zingiberales. In: Leins P, Tucker SC, Endress PK (eds) Aspects of floral development. Cramer, Berlin, pp 45–56
Kirchoff BK (1988b) Inflorescence and flower development in Costus scaber (Costaceae). Can J Bot 62:339–345
Kirchoff BK (1991) Homeosis in the flowers of the Zingiberales. Am J Bot 78:833–837
Kirchoff BK (1997) Inflorescence and flower development in the Hedychieae (Zingiberaceae): Hedychium. Can J Bot 75:581–594
Kirchoff BK (1998) Inflorescence and flower development in the Hedychieae (Zingiberaceae): Scaphochlamys kunstleri (Baker) Holttum. Int J Plant Sci 159(2):261–274
Kirchoff BK, Lagomarsino LP, Newman WH et al (2009) Early floral development of Heliconia latispatha (Heliconiaceae), a key taxon for understanding the evolution of flower development in the Zingiberales. Am J Bot 96:580–593
Krizek BA, Lewis MW, Fletcher JC (2006) RABBIT EARS is a second-whorl repressor of AGAMOUS that maintains spatial boundaries in Arabidopsis flowers. Plant J 45:369–383
Lampugnani ER, Kilinc A, Smyth DR (2012) PETAL LOSS is a boundary gene that inhibits growth between developing sepals in Arabidopsis thaliana. Plant J 71:724–735
Lestiboudois T (1829) Notice sur le genre Hedychium de la famille des Musacees (Balisiers et Bananiers). Annu Sci Natl Premium Service 17:113–139
Li W, Zhang L, Ding Z et al (2017) De novo sequencing and comparative transcriptome analysis of the male and hermaphroditic flowers provide insights into the regulation of flower formation in andromonoecious Taihangia rupestris. BMC Plant Biol 17:1–19
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆Ct method. Methods 25:402–408
Mahadevan C, Jaleel A, Deb L et al (2015) Development of an efficient virus induced gene silencing strategy in the non-model wild ginger-Zingiber zerumbet and investigation of associated proteome changes. PLoS ONE 10:1–17
Morioka K, Yockteng R, Almeida AMR et al (2015) Loss of YABBY2-Like gene expression may underlie the evolution of the laminar style in Canna and contribute to floral morphological diversity in the Zingiberales. Front Plant Sci 6:1106
Pelaz S, Ditta GS, Baumann E et al (2000) B and C floral organ identity functions require SEPALLATA MADS-box genes. Nature 405:200–203
Piñeyro-Nelson A, Almeida AMR, Sass C et al (2017) Change of fate and staminodial laminarity as potential agents of floral diversification in the Zingiberales. J Exp Zool B 328:41–54
Renner T, Bragg J, Driscoll HE et al (2009) Viral induced gene silencing as a tool for investigating floral developmental genetics in the Zingiberales. Mol Plant 3:1–11
Robinson MD, McCarthy DJ, Smyth GK (2009) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Sakai H, Medrano LJ, Meyerowitz EM (1995) Role of SUPERMAN in maintaining Arabidopsis floral whorl boundaries. Nature 378:199–203
Sass C, Iles WJD, Barrett CF et al (2016) Revisiting the Zingiberales: using multiplexed exon capture to resolve ancient and recent phylogenetic splits in a charismatic plant lineage. PeerJ 4:e1584
Schachner J (1924) Beitrage zur Kenntnis der Bluten und Samenentwicklung der Scitamineen. Flora 117:16–40
Schumann K (1904) Zingiberaceae. Pflanzenreich 20(4):46
Shuai B, Reynaga-Pena CG, Springer PS (2002) The LATERAL ORGAN BOUNDARIES gene defines a novel, plant-specific gene family. Plant Physiol 129:747–761
Smyth DR, Bowman JL, Meyerowitz EM (1990) Early flower development in Arabidopsis. Plant Cell 2:755–767
Song JJ, Ma W, Tang YJ et al (2010) Isolation and characterization of three MADS-box Genes from Alpinia hainanensis (Zingiberaceae). Plant Mol Biol Report 28:264–276
Specht CD, Almeida AMR (2017) A process-based approach to the study of flower morphological variation. In: Nuño de la Rosa L, Müller G (eds) Evolutionary developmental biology. Springer, Cham, pp 1–15
Specht CD, Howarth DG (2015) Adaptation in flower form: a comparative evodevo approach. New Phytol 206:74–90
Specht CD, Yockteng R, Almeida AMR et al (2012) Homoplasy, pollination, and emerging complexity during the evolution of floral development in the tropical gingers (Zingiberales). Bot Rev 78:440–462
Takeda S, Hanano K, Kariya A et al (2011) CUP-SHAPED COTYLEDON1 transcription factor activates the expression of LSH4 and LSH3, two members of the ALOG gene family, in shoot organ boundary cells. Plant J 66:1066–1077
Theiβen G (2001) Development of floral organ identity: stories from the MADS house. Curr Opin Plant Biol 4:75–85
Tian X, Yu Q, Liu H et al (2016) Temporal-spatial transcriptome analyses provide insights into the development of petaloid androecium in Canna indica. Front Plant Sci 7:1–11
Wake DB, Wake MH, Specht CD (2011) Homoplasy: from detecting pattern to determining process and mechanism of evolution. Science 331:1032–1035
Wang Q, Hasson A, Rossmann S et al (2016) Divide et impera: boundaries shape the plant body and initiate new meristems. New Phytol 209:485–498
Weigel D, Meyerowitz EM (1994) The ABCs of floral homeotic genes. Cell 78:203–209
Yob NJ, Jofrry SM, Affandi MM et al (2011) Zingiber zerumbet (L.) Smith: a review of its ethnomedicinal, chemical, and pharmacological uses. Evidence-based Complement Altern Med 2011:1–12
Yockteng R, Almeida AMR, Morioka K et al (2013) Molecular evolution and patterns of duplication in the SEP/AGL6-like lineage of the Zingiberales: a proposed mechanism for floral diversification. Mol Biol Evol 30:2401–2422
Yockteng R, Almeida AMR, Piñeyro-Nelson A, Specht CD (2014) Adaptation of floral form: an evo-devo approach to study adaptive evolution in floral morphology. In: Laitinen R (ed) Molecular mechanisms in plant adaptation. Wiley, New York, pp 171–192
Zhang J, Wu K, Zeng S et al (2013) Transcriptome analysis of Cymbidium sinense and its application to the identification of genes associated with floral development. BMC Genomics 14:279
Acknowledgements
This work is supported by the National Natural Science Foundation of China (Grant Nos. 31670336, 31200246, 31271318) and Key Laboratory of Plant Resources Conservation and Sustainable Utilization, South China Botanical Garden, Chinese Academy of Sciences (Grant No. Y821171001). We thank Haoran Ding for his assistance with part of the experiments.
Author information
Authors and Affiliations
Contributions
TZ performed the experiments, analyzed the data and wrote the manuscript. HFL designed the research and performed the experiments. CDS and ZCD performed parts of the data analysis and CDS helped write parts of the discussion. YSY and JPL provided assistance with samples collection and offered some photos of Z. zerumbet. All authors contributed to the manuscript revision.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, T., Specht, C.D., Dong, Z. et al. Transcriptome analyses provide insights into development of the Zingiber zerumbet flower, revealing potential genes related to floral organ formation and patterning. Plant Growth Regul 90, 331–345 (2020). https://doi.org/10.1007/s10725-020-00575-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-020-00575-7