Abstract
Plant-specific NAM, ATAF, and CUC (NAC) transcription factors (TFs) play clear roles in plant development and abiotic stress responses. Chinese dwarf cherry (Cerasus humilis) is an economically important shrub, that has strong resistance to drought. In this study, we isolated and functionally characterized a novel NAC TF from C. humilis. The ChNAC1 ORF contained 894 nucleotides, encoding 297 amino acid residues. ChNAC1 amino acid sequences had the highest similarity with homologous petunia (Petunia hybrida) and tomato (Solanum lycopersicum) NAM proteins. The ChNAC1 transcripts were most abundant in the leaves of seedlings and significantly up-regulated by drought stress. The nuclear localization and transcriptional activity of the C-terminal domain further confirmed that ChNAC1 functions as a TF. Yeast two-hybrid results showed that ChNAC1 can homodimerize in yeast cells. Next, we transformed ChNAC1 into wild-type Arabidopsis thaliana and found that the ectopic expression of ChNAC1 increased chlorophyll, water, proline, and protein contents as well as higher peroxidase (POD) and superoxide dismutase (SOD) activities while decreasing electrolyte conductivity and reactive oxygen species (ROS) contents compared to wild-type and mutant lines. Further, overexpression of ChNAC1 positively regulated ABA-responsive genes under drought stress and increased ABA sensitivity during root growth. Collectively, these results demonstrate the key role of ChNAC1 in drought stress tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Abiotic stresses including drought, extreme temperature, salinity, and nutrient imbalance can severely perturb plant growth and development, which leads to great losses in agricultural productivity worldwide (Xiong et al. 2002). Plants have evolved a variety of mechanisms that effectively address environmental challenges (He et al. 2018). Among all the key elements of stress-resistance systems in plants, many transcription factors (TFs) have been identified as playing important roles in abiotic stresses (Ma et al. 2019).
The NAM, ATAF, and CUC (NAC) transcription factor (TF) family is one of the largest species-specific TF families that has been identified (Olsen et al. 2005). Members of the NAC family commonly possess a highly conserved NAC domain at their N termini, which serve as sites for both DNA binding and homo- or hetero- dimerization with other NAC proteins (Olsen et al. 2005; Welner et al. 2012). The C-terminal regions, in contrast, are variable in sequence and length and serve as transcriptional activators or repressors (Nuruzzaman et al. 2010). NAC TFs regulate a diverse range of developmental processes in plants, e.g. apical meristem formation (Kim and Park 2007), root growth (Wang et al. 2015), leaf senescence and hormone signaling (Kim et al. 2016), secondary wall formation (Xu et al. 2014) and fruit ripening (Zhu et al. 2014). Moreover, NAC TFs play vital roles in responses to various abiotic stresses such as drought (Wu et al. 2009; He et al. 2016; Thirumalaikumar et al. 2018). The identification of genes encoding NAC proteins with differential expression in response to drought stress has the potential to clarify the current understanding of their roles in plant responses and defense mechanisms.
Cerasus humilis, commonly known as Chinese dwarf cherry, is an economically valuable member of the family Rosaceae and is indigenous to China. The fruits of C. humilis are known as ‘‘calcium fruit’’ because of their relatively high calcium content. Additionally, C. humilis seeds have been used as a traditional Chinese medicine for at least 2000 years (Chau and Wu 2006). In addition, like most perennial dwarf shrubs, C. humilis is highly stress-resistant, especially to drought and cold stresses (Du et al. 1993). NAC family genes have been reported in woody plants, such as Populus and Malus (Zhao et al. 2014; Zhong et al. 2014; Jia et al. 2018). However, there is little information available on NAC genes in C. humilis, especially with respect to their role in stress responses.
Here, we have transformed ChNAC1 into wild-type Arabidopsis plants to examine the protective role of this gene under drought stress conditions. Our data demonstrated that ChNAC1 is a stress-related TF that is potentially useful for engineering abiotic stress tolerance into crops.
Materials and methods
Materials and drought treatment
Cerasus humilis seedlings were imported from Shanxi province, China. We used Arabidopsis ecotype Columbia as the WT plant, ataf1 (T-DNA insertion mutant SALK-314847) as a negative control for studying the ChNAC1 function in this report due to its highest sequence identity (67.42%) with ChNAC1 among Arabidopsis gene bank. Seeds of the Arabidopsis were obtained from TAIR. We confirmed the lack of ATAF1 expression in the homozygous mutants using RT-PCR (Fig. 6c).
Cerasus humilis seedlings were grown under a 12-h photoperiod at approximately 25 °C, with a photosynthetic photon flux density (PPFD) of 600 μmol m−2 s−1 and a relative humidity of 70% in the greenhouse. A. thaliana seedlings were grown in soil-filled pots at 25 °C with 70% humidity under long-day conditions (16 h of light/8 h of dark) in a growth chamber with a light intensity of 600 μmol m−2 s−1 provided by cool-white fluorescent lights.
For the drought treatment, C. humilis seedlings at the 25–35-leaf stage and 5 week Arabidopsis seedlings were subjected to drought stress by withholding water for the indicated numbers of days (Kim et al. 2014). Seedling leaves were collected at specific times, frozen immediately in liquid nitrogen, and stored at − 80 °C for the later measurement of physiological and biochemical indexes (Cong et al. 2017).
Molecular cloning and sequence analysis of ChNAC1
Total RNA from C. humilis leaves was isolated using the E.Z.N.A.® Plant RNA Kit (Omega Bio-tek, Norcross, GA, USA) in accordance with the manufacturer’s protocol. Approximately 0.5 µg of cDNA was synthesized using the ReverTra Ace® qPCR RT Master Mix with gDNA Remover (Toyobo, Co., Ltd., Osaka, Japan) and was then used as the template for subsequent PCR amplification. Based on the C. humilis transcriptome database constructed by our research group (data not shown), a novel NAC transcription factor ChNAC1 was isolated using a specific primer pair (Table 1).
NCBI Open Reading Frame (ORF) Finder was employed to predict the ChNAC1 ORF. The conserved domain was predicted using the NCBI Conserved Domain Database (CDD). Homology search was performed using the NCBI blastn tool. Nucleotide sequence translation and multiple amino acid sequence alignments were performed using DNAMAN 6.0 software. In addition, a phylogenetic tree was constructed using the neighbor-joining (NJ) method implemented in MEGA 5.1. The putative nuclear localization signal was detected using the protein subcellular localization prediction tool PSORT (Wang et al. 2016).
Subcellular localization analysis for ChNAC1
The full-length coding sequence of ChNAC1 obtained by RT-PCR amplification was extended from the CACC nucleotide sequence at its 5′ end using the proofreading KOD-Plus-Neo enzyme (Toyobo). The fragment was first cloned into the pENTR™/D-TOPO vector using the pENTR™ Directional TOPO® Cloning Kit (Invitrogen, Carlsbad, CA, USA) and then cloned into the pGWB5 vector in frame with the GFP sequence using Gateway® LR ClonaseTM II Enzyme Mix (Invitrogen) according to the manufacturer’s instructions. The 35S::ChNAC1-GFP and the 35S::GFP constructs were transiently transformed into live onion epidermal cells using the particle bombardment method. Then the tissues were incubated in darkness at 25 °C for 48 h and visualized using the Axio Imager Z2 microscope (Zeiss, Oberkochen, Germany) (Wang et al. 2014).
Transcriptional activation assay
The coding regions of the ChNAC1 ORF (1–891 bp), N-terminus (1–480 bp), and C-terminus (481–891 bp) were amplified by PCR with specific primer pairs (Table 1) and then cloned into the pGBKT7 vector to fuse the sequences with a GAL4 DNA binding domain and create the pGBKT7-ChNAC1 (1–297 aa), pGBKT7-ChNAC1-N (1–160 aa), and pGBKT7-ChNAC1-C (161–297 aa) constructs, respectively. Fusion plasmids and the empty pGBKT7 vector were individually transformed into the yeast strain Y2H Gold using the lithium acetate method (Liu et al. 2017). The transformants were diluted or undiluted and then streaked onto the SD medium lacking Trp (SD/-Trp). For self-activation screening, the colonies were transferred to media lacking Trp, His (SD/-Trp/-His), or SD/-Trp/-His plus X-α-Gal media. The plates were incubated at 30 °C for 2–3 days. The yeast growth status was used to evaluate the transactivation activity of ChNAC1 proteins.
Homodimer assay
The coding region of the ChNAC1 ORF (1–891 bp) was inserted into the pGADT7 and pGBKT7 vectors. The pGBKT7 and pGADT7 plasmids were then co-transformed into the yeast strain Y2H Gold using the lithium acetate method. The diluted or undiluted cells were plated onto medium lacking Trp, Leu (SD/-Trp-Leu). For interaction screening, the colonies were transferred to media lacking Trp, Leu, His (SD/-Trp-Leu-His), or SD/-Trp-Leu-His plus X-α-Gal media. The plates were incubated at 30 °C for 2–3 days. Yeast growth status was used to identify homodimers.
Arabidopsis transformation
The 35S::ChNAC1-GFP vector was transformed into wild-type Arabidopsis plants with the Agrobacterium tumefaciens strain EHA105 (TIANDZ) using the floral dipping method (Yin et al. 2012). T0 seeds carrying the ChNAC1 constructs screened on MS medium supplemented with 50 mg L−1 kanamycin were transplanted into soil and the T1 transgenic Arabidopsis plants were verified and screened by PCR and qRT-PCR. The T3 homozygous positive lines were used for all further experiments.
Measurement of physiological and biochemical indexes
Total chlorophyll content was measured using a CCM-200 chlorophyll meter (Opti-Sciences Inc., Hudson, NH, USA); the electrolyte conductivity in Arabidopsis leaves was assayed using a conductivity meter; other indexes were assessed as described by Li (2000), with slight modification.
qRT-PCR analysis
The extraction of total RNA and synthesis of cDNA were conducted as noted above. The qRT-PCR reactions were conducted using UltraSYBR Mixture (CWBIO, Beijing, China) with the Roche Light Cycler 480 II real-time PCR System (Roche, Basel, Switzerland) according to the manufacturer’s instructions. Expression levels for all candidate genes were determined using the 2−ΔΔCT method; the relative transcript levels were calculated and normalized to Actin transcript levels. All the specific primer pairs are shown in Table 1.
Measurement of root length
Arabidopsis seeds were simultaneously sown onto MS medium containing 0, 100, 200, or 300 mM mannitol or 0, 1, 2, or 3 μM ABA, respectively. Then, the plates were incubated for 3 days before being vertically placed in a growth room for germination and culture for 15 days (Liu et al. 2016). Approximately 20 plants were used in each experiment, and the root length of each plant was measured individually.
Statistical analysis
All experiments were repeated independently at least three times and statistical significance of differences in measured parameters were tested using Student’s t-tests or ANOVA using GraphPad Prism 6.0 (GraphPad Software, San Diego, CA, USA). Data visualized in the figures are the mean ± SE values of three independent experiments, with significant differences noted at the *P ≤ 0.05, **P ≤ 0.01, and ***P ≤ 0.001 levels.
Results
Isolation and sequence analysis of ChNAC1
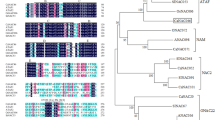
Gene sequence analysis indicated that ChNAC1 (GenBank accession number AKI23600.1) contains a 997-bp ORF corresponding to a predicted protein that consists of 297 amino acids. There is a typical NAM conserved domain between amino acids 9–132 (Fig. 1a). The ChNAC1 protein contains a NAC conserved domain at its N-terminal region, which is further divided into five subdomains (A–E); whereas the C-terminal regions are generally less conserved among the proteins (Fig. 1b). ChNAC1 shares high similarity with homologues, including 81% similarity with NAM from petunia and tomato (Fig. 1c). Subsequently, we discovered that the probabilities of nuclear, cytoplasmic, peroxisomal, and mitochondrial ChNAC1 localization are 82.6%, 8.7%, 4.3%, and 4.3%, respectively, as inferred using PSORTII software (data not shown).
Sequence analysis of ChNAC1. a Nucleotide sequence and the deduced amino acid sequence of ChNAC1. NAM conserved domain was indicated with underline. b Multiple sequences alignment of ChNAC1 and other NAC proteins known. NAC domains were indicated by black box and the five highly conserved subdomains (a–e) are indicated on bottom of the sequences. c The phylogenetic tree was generated using MEGA 5.1 software by the Neighbor-Joining method. The tree was constructed with a 1000-boorstrap replication support and the numbers at the nodes indicate the bootstrap values
Expression pattern analysis of ChNAC1
ChNAC1 was expressed in root, stem, leaf, and petal tissues of seedlings, with the highest expression in leaf tissue (Fig. 2a). ChNAC1 expression displayed a significant up-regulation after 8 days of drought, peaking on day 12 of drought, with an approximately 1.5-fold increase compared to the control (Fig. 2b). These findings indicated that ChNAC1 is strongly expressed in leaves and that ChNAC1 may be involved in physiological responses to drought stress.
Tissue-specific expression of ChNAC1 in Cerasus humilis and the effect of drought stress on the relative expression level of ChNAC1 in leaves. a qRT-PCR analysis of ChNAC1 in the root, stem, leaf and petal in Cerasus humilis. b qRT-PCR analysis of ChNAC1 expression in Cerasus humilis leaves under drought treatment. The relative expression levels were normalized to 1 in stem and 0 h as control, respectively
Nuclear localization of the ChNAC1 protein
Micrographs showed that protoplast cells transformed with the ChNAC1-GFP vector exhibited GFP fluorescence signals in the nucleus specifically (Fig. 3d–e). Whereas, the control GFP protein was located throughout the cells (Fig. 3a–c). Therefore, we summarized that ChNAC1 was localized to the cell nucleus.
Nuclear localization of the ChNAC1–GFP protein. Onion epidermal cells were bombarded with plasmids harboring the GFP coding region 35S::GFP (a–c), or an 35S::ChNAC1–GFP fusion construct (d–f). The 35S::GFP was used as a negative control. The GFP signals, bright-filed, an overlay of the GFP and bright-filed images are shown from up to down
The C-terminal domain of ChNAC1 has transcriptional activity in yeast cells
All transformants grew well on the SD/-Trp medium, indicating that all constructs had been successfully transformed. Yeast cells transformed with the full-length or C-terminal region of ChNAC1 showed strong growth on SD/-Trp/-His medium even when dilutions were used. However, the yeast cells transformed with the pGBKT7 vector or the N-terminal region of ChNAC1 did not exhibit the same result. Furthermore, the yeast cells that grew well on the SD/-Trp/-His medium turned blue in the presence of X-α-Gal (Fig. 4). These results indicate that ChNAC1 is a transcription activator and that its transactivation domain is located at the C-terminus.
Transcription activity of the ChNAC1 protein in Yeast. a Schematic diagrams of the full-length and truncated ChNAC1, which were fused to the pGBKT7. The numbers on the right indicate the last residues of polypeptides. b Growth of yeast cells, diluted or undiluted, transformed with different constructs on selective medium, using pGBKT7 as a negative control
ChNAC1 self-association in yeast cells
All the diluted or undiluted yeast transformants grew normally on SD/-Trp-Leu medium, indicating that the recombinant plasmids were successfully transformed into the yeast strain for further verification. The positive control pGBD-53 × pGAD-T grew normally on SD/-Trp-Leu-His medium and turned blue in the presence of X-α-Gal, but the negative controls pGBD-Lam × pGAD-T, pGBD × pGAD-NAC, and pGBD-N-NAC × pGAD failed to grow (Fig. 5). Surprisingly, pGBD-F-NAC × pGAD grew well as a negative control (Fig. 5), but we can explain this phenomenon by the observed transcriptional activity of ChNAC1 (Fig. 4). The growth of pGBD-F-NAC × pGAD-NAC and pBD-C-NAC × pAD-NAC further demonstrated self-activation. Finally, the normal growth of pBD-N-NAC × pAD-NAC (Fig. 5) indicated that ChNAC1 can homodimerize in yeast cells.
Morphological characteristics of ChNAC1 OXs
A total of 3 lines (Line1, Line2 and Line4) of ChNAC1 overexpression lines (Oxs) containing kanamycin resistance were obtained. Line1 and Line4 plants, which had single copy insertions (Fig. 6a) and higher transcripts abundance (Fig. 6b), were selected for further study.
The identification and selection of overexpression and mutant Arabidopsis. a Genomic DNA PCR analysis of ChNAC1 gene from WT, ataf1, putative OXs with Actin as an internal control. bChNAC1 transcript levels in OXs as revealed by qRT-PCR. c Genomic DNA PCR analysis of ATAF1 gene from ataf1 homogeneous mutant Arabidopsis plants with Actin as an internal control
In the presence of mannitol for the indicated number of days, the root lengths of the ChNAC1 OXs were significantly longer than those of WT and ataf1 plants (Fig. 7a, b). By the end of the drought treatment (12 days), the majority of WT and ataf1 lines presented wilting and discoloration symptoms even near death. However, ChNAC1 OXs were only slightly withered. After 3 days of resumed watering, ChNAC1 OXs recovered to normal levels, but WT and ataf1 plants hardly recovered (Fig. 7c). ChNAC1 expression was significantly induced by drought at different time points (Fig. 7d, e). These data implied that ChNAC1 enhanced plant tolerance to drought stress at maturity.
The effect of ChNAC1 overexpression on drought tolerance in Arabidopsis. a Growth of Arabidopsis seedlings on MS plates with different concentration mannitol. b Quantification of primary root length. c Performance of different genotypes Arabidopsis plants after 0, 5, 9, 12-day drought treatment and recovery 3-day. d Determination of ChNAC1 expression in Line1 under drought treatments by qRT-PCR. ChNAC1 transcription level at 0d was used as the calibrator. e Determination of ChNAC1 expression in Line4 under drought treatments by qRT-PCR. ChNAC1 transcription level at 0d was used as the calibrator. f–h Comparison of chlorophyll content (f), relative water content (h) and electrolyte conductivity (g) in different lines under drought condition
The chlorophyll contents of ChNAC1 OXs were higher than those of WT and ataf1 lines throughout the whole drought treatment, corresponding with the phenotypic difference (Fig. 7f). Both ChNAC1 OX lines showed a gradual decreasing trend. The water contents of ChNAC1 OXs were higher than those of WT and ataf1 plants (Fig. 7g). During the drought treatment, the electrolyte conductivities of the four lines differed to various degrees. For example, the conductivity of ChNAC1 OXs by day 12 had changed little, while the conductivities of ataf1 plants significantly increased, reaching a peak on day 12. However, Line1 and Line4 plants were relatively stable (Fig. 7h), which indicated that ChNAC1 OXs suffered from less severe membrane damage and exhibited higher drought tolerance.
Physiological and biochemical identification of drought resistance in transgenic Arabidopsis
In general, ChNAC1 OXs had lower H2O2 and O2− levels compared with the wild-type and ataf1 plants throughout the drought stress (Fig. 8a, b). ChNAC1 OXs also showed higher peroxidase (POD) activity levels while the ataf1 showed lower levels compared to control plants with under prolonged drought times (Fig. 8c). Superoxide dismutase (SOD) activity followed a pattern similar to that of POD. SOD activities of ChNAC1 OXs were always the highest, followed by wild-type and ataf1 lines (Fig. 8d). We observed that the proline contents of ChNAC1 OXs increased, although not very significantly throughout the drought period overall (Fig. 8e). The contents of soluble protein varied slightly among the different genotypes. For example, in the early stage of drought (i.e., day 5), both ChNAC1 OXs increased by about 20% compared with day 0. In contrast, wild-type and ataf1 plants exhibited a slight decrease. The protein levels of ChNAC1 OXs remained higher relative to wild-type and ataf1 plants during the whole treatment period (Fig. 8f).
Expression analysis of drought-related genes under drought treatment
We analyzed the expression patterns of several drought- and ABA-responsive markers, including COR47 (Xiang et al. 2018), RD22 (Abe et al. 1997), KIN1, KIN2 (Kurkela and Borg-Franck 1992), and RD20 (Shinozaki and Yamaguchi-Shinozaki 2007), during drought stress. Our analysis revealed that all candidate genes in this study showed significant differences in expression levels between wild-type and other plants under drought conditions except RD22. The expression levels of the core stress-related genes such as COR47 and genes in the ABA signaling pathway, including KIN1, KIN2, and RD20, were apparently down-regulated in ataf1 plants, but up-regulated in OXs compared with wild-type plants (Fig. 9), indicating that ChNAC1 may mediate drought stress signaling via an ABA-dependent pathway.
Overexpression of ChNAC1 increased ABA sensitivity
Phenotype analysis showed that root elongation of all plants was substantially inhibited by exogenous ABA. Specifically, the root growth inhibition in ChNAC1 OXs was more severe than that in other plants (Fig. 10a). Statistically, the root lengths of OXs were significantly inhibited compared with wild-type plants, while ataf1 plants were significantly enhanced on medium with 0.2 μM and 0.4 μM ABA (Fig. 10b). Thus, ChNAC1 increased the sensitivity of Arabidopsis to ABA.
Discussion
As a consequence of the dramatic climate change that has occurred in recent years, many regions of the world have been frequently affected by drought. In order to develop drought-resistant plant varieties by using genetic engineering methods, it is essential to clarify the molecular mechanisms that control and transduce drought stress signals (Sanchez et al. 2011). TFs are central regulators of the many genes that are affected by drought, including members of the NAC family (Wu et al. 2016). Here, we reported a new NAC TF, ChNAC1, which plays a positive role in protecting plants against drought stress.
Comparison of amino acid sequences between ChNAC1 and NAC proteins from other plant species suggests that ChNAC1 may be clustered into the typical NAC group (Fig. 1b). Notably, the C-terminus of the NAC protein is less conserved, which may account for the distinct functions of different proteins across various clades (Jensen et al. 2010). The ChNAC1 protein sequence is relatively similar to those of petunia and tomato NAM proteins (Fig. 1c). The typical NAM conserved domain prediction (Fig. 1a) and phylogenetic tree analysis revealed that ChNAC1 belongs to the NAM subfamily. NAM is essential in petunia embryo development and flower pattern formation (Jensen et al. 2010). Wang et al. (2018) also reported that NAM proteins can regulate plant resistance. These results indicated that ChNAC1 is a putative NAM member of the NAC gene family. The expression of ChNAC1 was tissue-specific (Fig. 2a), which is in line with the results of Seok et al. (2017), who found that AtNAP transcripts were more abundant in leaves than in other organs. The expression levels of ChNAC1 in both Arabidopsis (Fig. 7d, e) and C. humilis (Fig. 2b) provide a clue that ChNAC1 might be involved in drought stress responses (Liu et al. 2017).
The validation of nuclear localization and transcriptional activation at the C-terminus is required to identify a novel TF (Nakashima et al. 2012; Wu et al. 2016). Although we could not find the nuclear location signal in the ChNAC1 protein sequence, we demonstrated that the nuclear localization of ChNAC1 (Fig. 3) was consistent with the subcellular localization prediction. However, not all TFs are located in nuclei. For example, Duan et al. (2017) identified a lipid-anchored NACsa TF in sickle alfalfa. We also verified the transcriptional activity of ChNAC1 using a yeast one-hybrid system (Fig. 4). Yeast two-hybrid results showed that ChNAC1 can homodimerize (Fig. 5), which will help clarify the intrinsic transcriptional regulation mechanism of ChNAC1 in future research. It has been established that some NAC proteins can form homodimers or heterodimers, e.g., AtNAC2 (Ernst et al. 2004). Additionally, we discovered that the removal of the C-terminus does not affect dimer conformation (Fig. 5). Thus, we proposed that the C-terminus of ChNAC1 is not required for dimerization, which is consistent with the structural features of NAC. A subdomain in the N-terminal is associated with this dimerization, while the C-terminus is responsible for transcriptional regulation (Ernst et al. 2004).
Drought stress affects plant growth and development. In our study, the characterization of the ChNAC1 OXs indicated that the transgenic Arabidopsis demonstrated stronger and healthier growth at both the seedling and mature stages in response to drought stress (Fig. 7a–c), as was observed for DgNAC1 and MusaNAC042 (Tak et al. 2017; Wang et al. 2017).
Reactive oxygen species (ROS) at low concentrations can act as secondary signals in plant cell signal transduction pathways while ROS overproduction is a critical factor that causes membrane damage (Wang et al. 2017). The dual roles of ROS are entirely dependent on the delicate balance between ROS production and elimination by antioxidant enzymes (Olsen et al. 2013). In the present study, the significantly lower ROS contents (Fig. 7a, b) and higher ROS scavenging enzymes activities (Fig. 7c, d) of the ChNAC1 OXs implied that less oxidative damage occurred in them relative to wild-type and mutant plants. Plants also usually accumulate solutes, such as proline and proteins, that alleviate the damage of abiotic stress (Ashraf and Foolad 2007). The proline and protein contents of transgenic plants were slightly higher than those of controls (Fig. 8e, f), indicating that ChNAC1 improved plant drought tolerance, mainly by maintaining ROS balance. The finding was consistent with a recent publication by Wang et al. (2018) who suggested that CsATAF1 positively regulates drought stress tolerance by ROS scavenging in cucumber.
ABA has pivotal roles in plant growth and adaptive responses to various environmental stresses (Takagi et al. 2017). NACs regulate the expression of both ABA-dependent and ABA-independent genes even during abiotic stress responses (Puranik et al. 2011, 2012). We found that ChNAC1 conferred tolerance to drought in transgenic plants through the ABA-dependent signaling pathway (Figs. 9, 10). These findings are consistent with those for several other NAC transcription factors (Shen et al. 2017; Wang et al. 2018).
Conclusions
In summary, we cloned a novel NAC transcription factor from C. humilis and performed functional analyses. This research offers the novel insight that overexpression of ChNAC1 may strengthen drought tolerance, as demonstrated in Arabidopsis. This has the potential to provide tools for improving tolerance in crops under abiotic stresses. Studies on the transcriptional regulatory network mediated by ChNAC1 will further increase the current understanding of the transcriptional regulation of drought tolerance.
Abbreviations
- WT:
-
Wild type
- OX:
-
Overexpression
- PCR:
-
Polymerase chain reaction
- RT-PCR:
-
Reverse transcription PCR
- qRT-PCR:
-
Quantitative real-time PCR
- RW3:
-
Rewater 3 days
- GFP:
-
Green fluorescent protein
- His:
-
Histidine
- Trp:
-
Tryptophan
- Leu:
-
Leucine
- ABA:
-
Abscisic acid
References
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9(10):1859–1868
Ashraf M, Foolad MR (2007) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59(2):206–216
Chau CF, Wu SH (2006) The development of regulations of Chinese herbal medicines for both medicinal and food uses. Trends Food Sci Technol 17(6):313–323
Cong J, Li KQ, Xu XY, Zhang HP, Chen HX, Chen YH, Hao J, Wang Y, Huang XS, Zhang SL (2017) A novel NAC transcription factor, PbeNAC1, of Pyrus betulifolia confers cold and drought tolerance via interacting with PbeDREBs and activating the expression of stress-responsive genes. Front Plant Sci 8:1049
Du JJ, Yang H, Chi J (1993) A preliminary study of selected strains of the Chinese dwarf cherry tree. China Fruits 3:23–24
Duan M, Zhang R, Zhu F, Zhang Z, Guo L, Wen J, Dong J, Wang T (2017) A lipid-anchored NAC transcription factor is translocated into the nucleus and activates Glyoxalase I expression during drought stress. Plant Cell 29(7):1748–1772
Ernst HA, Olsen AN, Larsen S, Lo Leggio L (2004) Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors. EMBO Rep 5(3):297–303
He X, Zhu L, Xu L, Guo W, Zhang X (2016) GhATAF1, a NAC transcription factor, confers abiotic and biotic stress responses by regulating phytohormonal signaling networks. Plant Cell Rep 35(10):2167–2179
He L, Wu Y, Zhao Q, Wang B, Liu Q, Zhang L (2018) Chrysanthemum DgWRKY2 gene enhances tolerance to salt stress in transgenic Chrysanthemum. Int J Mol Sci 19(7):2062
Jensen MK, Kjaersgaard T, Nielsen MM, Galberg P, Petersen K, O’Shea C, Skriver K (2010) The Arabidopsis thaliana NAC transcription factor family: structure-function relationships and determinants of ANAC019 stress signalling. Biochem J 426(2):183–196
Jia DF, Gong XQ, Li MJ, LiC Chao, Sun TT, Ma FW (2018) Overexpression of a novel apple NAC transcription factor gene, MdNAC1, confers the dwarf phenotype in transgenic apple (Malus domestica). Genes 9(5):229
Kim SG, Park CM (2007) Membrane-mediated salt stress signaling in flowering time control. Plant Signal Behav 2(6):517–518
Kim Y, Wang M, Bai Y, Zeng ZH, Guo F, Han N, Bian HW, Wang JH, Pan JW, Zhu MY (2014) Bcl-2 suppresses activation of VPEs by inhibiting cytosolic Ca2+ level with elevated K+ efflux in NaCl-induced PCD in rice. Plant Physiol Biochem 80:168–175
Kim HJ, Nam HG, Lim PO (2016) Regulatory network of NAC transcription factors in leaf senescence. Curr Opin Plant Biol 33:48–56
Kurkela S, Borg-Franck M (1992) Structure and expression of kin2, one of two cold- and ABA-induced genes of Arabidopsis thaliana. Plant Mol Biol 19(4):689–692
Li HS (2000) The experiment principle and technique for plant physiology and biochemistry. Beijing, China
Liu Y, Sun J, Wu Y (2016) Arabidopsis ATAF1 enhances the tolerance to salt stress and ABA in transgenic rice. J Plant Res 129(5):955–962
Liu YM, Yu XW, Liu SS, Peng H, Mijiti C, Wang Z, Zhang H, Ma H (2017) A chickpea NAC-type transcription factor, CarNAC6, confers enhanced dehydration tolerance in Arabidopsis Plant. Mol Biol Rep 35:83–96
Ma QB, Xia ZL, Cai ZD et al (2019) GmWRKY16 enhances drought and salt tolerance through an ABA-mediated pathway in Arabidopsis thaliana. Front Sci 9:1–18
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozakibv K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819:97–103
Nuruzzaman M, Manimekalai R, Sharoni AM et al (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465(1–2):30–44
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10(2):79–87
Olsen LF, Issinger O, Guerra B (2013) The Yin and Yang of redox regulation. Redox Rep 18:245–252
Puranik S, Bahadur RP, Srivastava PS, Prasad M (2011) Molecular cloning and characterization of a membrane associated NAC family gene, SiNAC from foxtail millet [Setaria italica (L.) P. Beauv]. Mol Biotechnol 49(2):138–150
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17(6):369–381
Sanchez DH, Pieckenstain FL, Szymanski J et al (2011) Comparative functional genomics of salt stress in related model and cultivated plants identifies and overcomes limitations to translational genomics. PLoS ONE 6(2):e17094
Seok HY, Woo DH, Nguyen LV, Tran HT, Tarte VN, Lee SY, Moon YH (2017) Arabidopsis AtNAP functions as a negative regulator via repression of AREB1 in salt stress response. Planta 245(2):329–341
Shen J, Lv B, Luo L, He J, Mao C, Xi D, Ming F (2017) The NAC-type transcription factor OsNAC2 regulates ABA-dependent genes and abiotic stress tolerance in rice. Sci Rep 7:40641
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227
Tak H, Negi S, Ganapathi TR (2017) Banana NAC transcription factor MusaNAC042 is positively associated with drought and salinity tolerance. Protoplasma 254(2):803–816
Takagi H, Ishiga Y, Watanabe S, Konishi T, Egusa M, Akiyoshi N, Matsuura T, Mori IC, Hirayama T, Kaminaka H, Shimada H, Sakamoto A (2017) Allantoin, a stress-related purine metabolite, can activate jasmonate signaling in a MYC2-regulated and abscisic acid-dependent manner. J Exp Bot 68(17):2519–2532
Thirumalaikumar VP, Devkar V, Mehterov N, Ali S, Ozgur R, Turkan I, Mueller-Roeber B, Balazadeh S (2018) NAC transcription factor JUNGBRUNNEN1 enhances drought tolerance in tomato. Plant Biotechnol J 16(2):354–366
Wang L, Qin L, Liu W, Zhang D, Wang Y (2014) A novel ethylene responsive factor from Tamarix hispida, ThERF1, is a GCC-box- and DRE motif binding protein that negatively modulates abiotic stress tolerance in Arabidopsis. Physiol Plant 152(1):84–97
Wang F, Lin R, Feng J, Chen W, Qiu D, Xu S (2015) TaNAC1 acts as a negative regulator of stripe rust resistance in wheat, enhances susceptibility to Pseudomonas syringae, and promotes lateral root development in transgenic Arabidopsis thaliana. Front Plant Sci 6:108
Wang G, Zhang S, Ma X, Wang Y, Kong F, Meng Q (2016) A stress associated NAC transcription factor (SlNAC35) from tomato plays a positive role in biotic and abiotic stresses. Plant Physiol 158(1):45–64
Wang K, Zhong M, Wu YH, Bai ZY, Liang QY, Liu QL, Pan YZ, Zhang L, Jiang BB, Jia Y, Liu GL (2017) Overexpression of a chrysanthemum transcription factor gene DgNAC1 improves the salinity tolerance in chrysanthemum. Plant Cell Rep 36(4):571–581
Wang JF, Zhang L, Cao YY, Qi CD, Li ST, Liu L, Wang GL, Mao AJ, Ren SX, Guo YD (2018) CsATAF1 positively regulates drought stress tolerance by an ABA-dependent pathway and by promoting ROS scavenging in cucumber. Plant Cell Physiol 59(5):930–945
Welner DH, Lindemose S, Grossmann JG, Mollegaard NE, Olsen AN, Helgstrand C, Skriver K, Lo LL (2012) DNA binding by the plant-specific NAC transcription factors in crystal and solution: a firm link to WRKY and GCM transcription factors. Biochem J 444(3):395–404
Wu YR, Deng ZY, Lai JB, Zhang YY, Yin BJ, Zhao QZ, Zhang L, Li Y, Yang CW, Xie Q (2009) Dual function of Arabidopsis ATAF1 in abiotic and biotic stress responses. Cell Res 19(11):1279–1290
Wu H, Fu B, Sun P, Xiao C, Liu JH (2016) A NAC transcription factor represses putrescine biosynthesis and affects drought tolerance. Plant Physiol 172(3):1532–1547
Xiang JH, Zhou XY, Zhang XW, Zhang XW, Liu AL, Xiang YC, Yan ML, Peng Y, Chen XB (2018) The Arabidopsis AtUNC-93 acts as a positive regulator of abiotic stress tolerance and plant growth via modulation of ABA signaling and K+ homeostasis. Front Plant Sci 9:718
Xiong LM, Schumaker KS, Zhu JK (2002) Cell signaling during cold, drought, and salt stress. Plant Cell 14:165–183
Xu B, Ohtani M, Yamaguchi M, Toyooka K, Wakazaki M, Sato M, Kubo M, Nakano Y, Sano R, Hiwatashi Y, Murata T, Kurata T, Yoneda A, Kato K, Hasebe M, Demura T (2014) Contribution of NAC transcription factors to plant adaptation to land. Science 343(6178):1505–1508
Yin ZP, Shang ZW, Ren J, Song XS (2012) Foliar sprays of photosynthetic bacteria improve the growth and anti-oxidative capability on Chinese dwarf cherry seedlings. J Plant Nutr 35(6):840–853
Zhao Y, Sun J, Xu P, Rui Zhang, Li LG (2014) Intron-mediated alternative splicing of WOOD-ASSOCIATED NAC TRANSCRIPTION FACTOR1B regulates cell wall thickening during fiber development in populus species. Plant Physiol 164(2):765–776
Zhong R (2014) Functional characterization of poplar wood-associated NAC domain transcription factors. Plant Physiol 152(4):1044–1055
Zhu MK, Chen GP, Zhang JL, Zhang YJ, Xie QL, Zhao ZP, Zong YP, Hu L (2014) The abiotic stress-responsive NAC-type transcription factor SlNAC4 regulates salt and drought tolerance and stress-related genes in tomato (Solanum lycopersicum). Plant Cell Rep 33(11):1851–1863
Acknowledgements
This work was supported by Fundamental Research Funds for the Central Universities (Grant No. 2572018CG02), the National Natural Science Foundation of China (Grant No. 31170569) and the Innovation Project of State Key Laboratory of Tree Genetics and Breeding (Northeast Forestry University).
Author information
Authors and Affiliations
Contributions
XSS conceived and designed the study. FW carried out the main experiments and wrote the manuscript. JWW cloned the ChNAC1 gene. LJS performed the gene expression analysis. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, F., Wang, J.W., Sun, L.J. et al. The molecular cloning and functional characterization of ChNAC1, a NAC transcription factor in Cerasus humilis. Plant Growth Regul 89, 331–343 (2019). https://doi.org/10.1007/s10725-019-00536-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-019-00536-9