Abstract
Anthocyanin has an important antioxidant protective effect on plant resistance to oxidative stress. In this study, an Arabidopsis mutant dpg1 (delayed pale-greening) with a chloroplast development defect was studied. It was found that the anthocyanin accumulation of this mutant had increased during the seedling stage, and the expressions of the anthocyanin biosynthetic and regulatory genes were up-regulated. Further studies showed that exogenous ABA (abscisic acid) treatments significantly promoted the chloroplast development of the dpg1 mutant, and the anthocyanin content was significantly decreased to the level of the wild-type. When using NF (norflurazon) to simulate the oxidative stress treatments of wild-type Arabidopsis, the anthocyanin content had significantly increased compared with the control. However, the exogenous ABA treatments could significantly reduce the anthocyanin accumulation level induced by the oxidative stress. Furthermore, the components ABI1 (abscisic acid insensitive 1) and ABI3 (abscisic acid insensitive 3) of the ABA signaling pathway were found to play important roles during this process. These results indicate that the increases in the anthocyanin accumulation in the dpg1 mutant seedlings could be mediated by oxidative stress. Meanwhile, the ABI1 and ABI3 were involved in the process of the ABA inhibiting anthocyanin accumulation which had been induced by the oxidative stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Anthocyanins are a group of plant pigments responsible for the purple coloration of plant parts at particular developmental stages or under special environmental conditions (Grotewold 2006; Rowan et al. 2009; Hoffmann et al. 2016). The presence of anthocyanin in flowers and fruits is required for attracting pollinators and seed-dispersing animals (Winkel-Shirley 2001). Anthocyanins are also an important class of polyphenols with remarkable antioxidant activity and help to protect plants from oxidative damage caused by abiotic stress (Steyn et al. 2002).
The anthocyanin biosynthetic pathway has been extensively studied in various plant species. Genes encoding enzymes required for the anthocyanin biosynthetic pathway are grouped into two classes (Holton and Cornish 1995). The early biosynthesis genes (EBGs) are involved in the common steps of the different flavonoid subpathways, and mainly include chalcone synthase (CHS), chalcone isomerase (CHI), flavanone 3-hydroxylase (F3H) and flavonoid 3′-hydroxylase (F3′H). The late biosynthesis genes (LBGs) primarily include dihydroflavonol 4-reductase (DFR), leucoanthocyanidin oxygenase (LDOX), anthocyanidin reductase (ANR), and UDP-glucose:flavonoid 3-O-glucosyltransferase (UF3GT) (Pelletier et al. 1997). Expression of these genes is regulated by positive and negative regulatory transcription factors. For example, the WD-repeat independent MYBs (myeloblastosis, MYB11, MYB12, and MYB111) and MYBs/bHLH/WD-repeat complex regulate the expression of EBGs and LBGs, respectively (Mehrtens et al. 2005; Stracke et al. 2007; Gonzalez et al. 2008). In Arabidopsis, transcription factors PIF3 (phytochrome-interacting factor 3) and HY5 (long hypocotyl 5) positively regulate anthocyanin biosynthesis by directly binding to the promoters of anthocyanin biosynthetic genes, including CHS, CHI, F3H, F3′H, DFR and LDOX (Lee et al. 2007; Shin et al. 2007). In contrast to the positive transcription factors mentioned above, the R3-MYB protein MYBL2 (myeloblastosis family transcription factor-like 2) acts as a transcriptional repressor and negatively regulates the biosynthesis of anthocyanin (Dubos et al. 2008; Matsui et al. 2008; Wang et al. 2016; Xie et al. 2016).
The chloroplast is an essential organelle in plant cells and plays important roles in primary metabolism. The coordinated expression of genes distributed between the nuclear and chloroplast genomes is essential for the assembly of functional chloroplasts (Inaba and Schnell 2008). While chloroplast development is largely under nuclear control, developmentally arrested or damaged chloroplasts can regulate nuclear gene expression via retrograde signaling pathways (Leister 2005; Nott et al. 2006; Chen et al. 2018). For example, norflurazon (NF), an inhibitor of carotenoid biosynthesis, causes the photooxidative destruction of chloroplasts and leads to a markedly decreased expression of a large number of nuclear photosynthesis-related genes (Burgess and Taylor 1987; La Rocca et al. 2001). Recently, more and more studies showed that the plastid retrograde signals affected more cellular processes other than photosynthesis-associated nuclear genes repression (Cottage et al. 2010; Cheng et al. 2012; Yu et al. 2012; Feng et al. 2016). Some evidence is available to support that retrograde signaling is involved in anthocyanin biosynthesis (Cottage et al. 2010; Cheng et al. 2012).
In higher plants, under the conditions of high intensity illumination or other abiotic stresses, the chloroplast will suffer oxidative stress (Hernandez et al. 1995; Bailey et al. 2002; Sakamoto et al. 2004). Also, the anthocyanin accumulation will be enhanced through the up-regulated expressions of the anthocyanin biosynthetic and regulatory genes, in order to resist the oxidative stress (Maruta et al. 2014). It has been reported that many mutants which were deficient in chloroplast development had increased sensitivity to oxidative stress (Myouga et al. 2008; Huang et al. 2009; Kikuchi et al. 2009; Miura et at. 2010; Gan et al. 2014). For example, when a mutation occurs in the FtsH2 gene encoding metalloproteases in the chloroplasts of Arabidopsis, the var2 (varigated2) mutant showed a green/white variegated phenotype, and the chloroplast development in the white leaf tissue of the var2 mutant was abnormal. However, the expression level of the Cu/Zn superoxide dismutase 2 was found to be increased, which indicated that the tissue which was deficient in chloroplast development in the var2 mutant increased the sensitivity to the oxidative stress (Miura et al. 2010). When plants are growing under adverse environmental conditions, the content of active oxygen in the plants increases significantly, which causes the plants to suffer from oxidative stress (Hammond-Kosack and Jones 1996; Yousuf et al. 2017). However, ABA treatments can be adopted to reduce the degree of oxidative stress injury (Jiang and Zhang 2002; Larkindale and Knight 2002; Hu et al. 2006). On one hand, ABA can induce the expressions of genes which encode antioxidant enzymes in order to enhance plant resistance (Zhu and Scandalios 1994; Bueno et al. 1998; Guan et al. 2000; Azeem et al. 2016). On the other hand, ABA can also enhance antioxidant enzyme activities and the content of non-enzymatic antioxidants in plants, in order to alleviate oxidative stress injuries (Bueno et al. 1998; Gong et al. 1998; Guan and Scandalios 1998; Jiang and Zhang 2002; Sarafraz-Ardakani et al. 2014).
Our previous study showed that AtDPG1 plays an essential role in early chloroplast biogenesis. Its absence triggers plastid retrograde signaling, which ultimately down-regulates the expressions of the nuclear genes which encode chloroplast-localized proteins (Liu et al. 2016). To investigate the relationship between chloroplast development and anthocyanin accumulation, this study characterized the Arabidopsis dpg1 mutant, and found that the disruption of the AtDPG1 will result in the enhancement of the anthocyanin accumulation, accompanied by increased expressions of anthocyanin biosynthetic and regulatory genes. Further studies indicate that the increases in the anthocyanin content in the dpg1 mutant seedlings could be mediated by oxidative stress. ABA plays an important role in regulating the growth, development, and abiotic adaptability of the plants. The results of this study indicated that the ABA could inhibit the anthocyanin accumulation induced by oxidative stress, while the ABI1 and ABI3 played positive regulating roles in this process.
Materials and methods
Plant material and growth conditions
Arabidopsis thaliana L. ecotypes Columbia (Col) and Landsberg (Ler) were used as the wild types in this study. The dpg1 (SALK_004621C) and abscisic acid insensitive mutants (abi1: CS22, abi3: CS24), which are in the Col and Ler background, respectively, were ordered from the Arabidopsis Biological Resource Center. The DPG1/dpg1 transgenic lines are in the Col background as previously described (Liu et al. 2016). The Arabidopsis seeds were surface sterilized in a solution of 25% bleach plus 0.01% Triton X-100 for 10 min and washed with sterilized water for 10–15 times. Following 3 days of stratification in the dark at 4 °C, the seeds were germinated and grown on plates containing 1/2 MS medium [0.8% (w/v) agar, 2% (w/v) sucrose, pH 5.8] at 22 ± 1 °C under a 16-h light/8-h dark photoperiod with a light intensity of 90 µE m−2 s−1 to produce Arabidopsis seedlings.
To examine the effects of ABA and NF on anthocyanin biosynthesis, the sterilized and cold-treated seeds were germinated and grown vertically on 1/2 MS medium without (Mock) or with 0.25 µM ABA, 5 µM NF, or 5 µM NF and 0.25 µM ABA (NF + ABA) for 4 days. The 4-day-old seedlings were harvested for anthocyanin measurement.
RNA extraction, cDNA synthesis, and gene expression analysis
Total RNA was extracted from plant tissues with TRI Reagents, followed by treatment with RNase-free DNase I (TaKaRa, Dalian, China) at 37 °C for 1 h to degrade genomic DNA. First strand cDNA was synthesized from 2.0 µg of total RNA using ImProm-II™ reverse transcriptase (Promega, Madison, WI, USA) in 10.0 µL reaction, following the manufacturer’s instructions with minor modifications. PCR reaction contained 1.0 µL of template, 1.5 U ExTaq (TaKaRa, Dalian, China), dNTPs (0.2 mM each) and the primers in a final volume of 30 µL. Three RT-PCR reactions were repeated independently using ACTIN2 as an internal control. For real-time RT-PCR analysis, first-strand cDNA was synthesized in a same way. The reaction was performed using SYBR Green Perfect mix (TaKaRa, Dalian, China) on a CFX96 (Bio-Rad), following the manufacturer’s instructions. The following standard thermal profile was used for all PCRs: 95 °C for 2 min; 40 cycles of 95 °C for 10 s and 60 °C for 30 s. Gene expression was normalized to that of ACTIN2 by subtracting the CT value of ACTIN2 from the CT value of the gene of interest. Expression ratios were then obtained from the Eq. 2ΔΔCT. Primers for genes of interest are listed in Supplementary Table 1.
Transmission electron microscopy (TEM) analysis
The cotyledons of 7-day-old wild-type and dpg1 seedlings were fixed in 2.5% (w/v) glutaraldehyde in phosphate buffer (pH 7.4) overnight at 4 °C. Thereafter, the samples were rinsed thoroughly with the same buffer 3–6 times and post-fixed with 1% (w/v) osmium tetroxide for 2 h at 4 °C. Then the samples were dehydrated in a graded ethanol series (v/v, 30%, 50%, 70%, 90%) and in 1:1 mixture of 90% ethanol and 90% acetone, at last in acetone 3 times, embedded in Epon812 and polymerized at 35 °C for 16 h, then 48 °C for 24 h, and 65 °C for 48 h. For observation, ultrathin sections of the samples were cut with a diamond knife and collected on 200-mesh copper grids. After contrasting with uranyl acetate and lead citrate, the grids were examined with a JEM-2100 transmission electron microscope.
Anthocyanin measurement
Anthocyanin measurement was performed as described by Chory et al. (1989). The seedlings were grown for 4–7 days after sowing on 1/2 MS medium, and then used for anthocyanin measurement. Seedlings of each genotype were incubated overnight in 0.6 mL of 1% HCl in methanol at 4 °C and extracted using an equal volume of chloroform after the addition of 0.4 mL of water. After centrifugation (12,000 rpm for 10 min at 25 °C), the quantity of anthocyanins was determined by spectrophotometric measurement of the aqueous phase (A530–0.25A657) and normalized to the fresh weight of each sample. 3 independent biological samples were used to measure anthocyanin for each genotype.
Analysis of chlorophyll contents
Chlorophylls were determined as described previously with minor modifications (Lichtenthaler and Wellburn 1983). Fresh plant tissues were homogenized in 80% acetone, and debris was removed by centrifugation at 12,000 rpm for 10 min (4 °C). The absorbance of the supernatant at 663 and 645 nm was measured with a 723B spectrophotometer (Tianjin PURUISI Equipment Co. LTD, China). All measurements were repeated in 3 independent experiments.
ABA determination
The wild type and dpg1 seedlings were grown for 4 days after sowing on 1/2 MS medium supplemented with 2% sucrose, and then used for ABA determination. The seedlings were ground into powder in liquid nitrogen. Quantification of ABA was carried out with ELISA at the Phytohormones Research Institute (China Agricultural University) as previously described by Yang et al. (2001).
Results
Anthocyanin content increased in an Arabidopsis mutant which was defective in chloroplast biogenesis
To investigate the relationship between chloroplast development and anthocyanin accumulation, we focused on the dpg1 phenotype (defective in chloroplast development) in anthocyanin accumulation. In this study, wild-type and dpg1 seeds were germinated on 1/2 MS medium containing 2% sucrose, and allowed to grow for 4, 5, 6, and 7 days. The anthocyanin content was quantified spectrophotometrically following the extraction in acidic methanol. As shown in Fig. 1a, b, the anthocyanin content in the wild type Arabidopsis seedlings had gradually decreased with the development progress. However, the anthocyanin content in the dpg1 mutant seedlings was observed to be significantly higher than that of the wild type at a fixed point in time. Moreover, the anthocyanin accumulation in the dpg1 mutant seedlings was significantly increased in the upper hypocotyls, as well as the edges of the adaxial surfaces of the cotyledons (Fig. 1a).
The anthocyanin content increased in the dpg1 mutant, which was defective in chloroplast biogenesis. a Phenotypes of the representative seedlings of wild-type (Col) and dpg1 seedlings were grown on 1/2 MS medium for 4–7 days. Scale bar = 1 mm. b Time-course of anthocyanin accumulation in wild-type and dpg1 seedlings. Anthocyanins were extracted in acidified methanol from 3 replicate samples of wild-type and dpg1 seedlings grown on 1/2 MS medium. The seedlings were harvested daily 4–7 days after germination. The anthocyanin content was determined from the absorbance of the extract at 530 nm and 657 nm. c Quantitative RT–PCR analysis of mRNA levels was performed for the regulatory genes (PAP1 and MYB11) of the anthocyanin biosynthetic pathway and anthocyanin biosynthetic genes (CHS, CHI, F3′H, DFR, LDOX, UGT78D2, and UF3GT) in 4-day-old seedlings. The data were normalized relative to the ACTIN2 gene, then expressed relative to the wild-type value. The transcript level of each gene was set to 1 in the wild-type. The error bars represent the SDs for the means of 3 independent replicates. The asterisks indicate statistically significant differences compared with the corresponding wild type (Student’s t test: *P < 0.05)
To understand the molecular basis of the changes in the anthocyanin levels, the expression of the regulatory genes (PAP1 and MYB11) in the anthocyanin biosynthetic pathway was first examined by quantitative RT-PCR. As shown in Fig. 1c, the transcriptions of these two genes in the dpg1 mutant were significantly higher than in the wild-type seedlings. The expression of the anthocyanin biosynthetic genes was then monitored, including CHS, CHI, F3′H, DFR, LDOX, UGT78D2, and UF3GT. These genes displayed the same expression pattern as follows: the transcript levels of these genes were higher in the dpg1 mutant than in the wild-type (Fig. 1c). All these results indicated that the increases in the anthocyanin content may have been due to the enhanced expressions of the anthocyanin biosynthetic and regulatory genes in the dpg1 mutant.
Overexpression of AtDPG1 significantly decreased the content of anthocyanin in the dpg1 mutant seedling
In order to confirm that the disruption of the AtDPG1 was responsible for the anthocyanin over-accumulation phenotype, a rescue of the phenotype was attempted by introducing a cDNA of the AtDPG1 gene driven by the CaMV 35S promoter back into the heterozygous dpg1 plants. Among the 19 analyzed T1 transgenic lines, 3 lines were found to be homozygous dpg1, and showed the characteristics of a wild-type phenotype (Fig. 2a). Analysis by quantitative RT-PCR revealed that the 3 independent complemented transgenic lines had significantly higher expression levels of AtDPG1 to those of dpg1 seedlings (Fig. 2b). Furthermore, the anthocyanin content in the transgenic seedlings was significantly decreased when compared with the dpg1 mutant, and was found to have almost decreased to the level of the wild-type Arabidopsis seedlings (Fig. 2c).
Overexpression of AtDPG1 significantly decreased the content of anthocyanin in the dpg1 mutant seedling. a Phenotypes of the 4-day-old dpg1 homozygous seedlings were rescued via transformation with a 35S::AtDPG1 transgene. Scale bar = 1 mm. b Quantitative RT-PCR analyses of the AtDPG1 transcript in wild-type seedlings, 3 independent complementation lines (DPG1/dpg1-1, DPG1/dpg1-2 and DPG1/dpg1-3), and mutant (dpg1) seedlings. c Anthocyanins were extracted in acidified methanol from 3 replicate samples of wild-type, dpg1, and rescued seedlings grown on 1/2 MS medium for 4 days. The anthocyanin content was determined from the absorbance of the extract at 530 nm and 657 nm. The error bars represent the SDs for the means of 3 independent replicates. The asterisks indicate statistically significant differences compared with the corresponding wild type (Student’s t test: *P < 0.05)
ABA treatment significantly decreased the anthocyanin content in the dpg1 mutant
ABA has been found to play an important role in regulating plant growth and development, as well as adversity adaptability (Zeevaart and Creelman 1988; Wilkinson and Davies 2002). In this study, exogenous ABA was used to treat wild-type Arabidopsis and dpg1 mutant seedlings, and the anthocyanin content of these samples was then determined. The results are shown in Fig. 3a, b. In the control group, the anthocyanin content was significantly increased in the dpg1 mutant seedlings. However, the anthocyanin content in the dpg1 mutant seedlings was observed to be significantly decreased to the level of wild-type following the ABA treatment. These results indicate that the anthocyanin content had been increased, possibly due to the decrease of the ABA content in the dpg1 mutant seedlings. In order to verify this inference, the ABA content in the wild-type Arabidopsis and dpg1 mutant seedlings was determined. The results showed that there was no significant difference in the ABA content between the wild-type Arabidopsis and dpg1 mutant seedlings (Fig. S1). Previous studies have shown that ABA plays an important role in plant resistance to oxidative stress (Guan and Scandalios 1998; Guan et al. 2000; Jiang and Zhang 2001, 2002). Therefore, it was suspected that the increases in anthocyanin content in the dpg1 mutant were possibly due to the seedlings suffered oxidative stress. The exogenous ABA treatment was able to mitigate the oxidative stress degree of the dpg1 mutant, which resulted in the anthocyanin content becoming significantly decreased. For the purpose of confirming this inference, norflurazon (NF) was used in this study to simulate the oxidative stress conditions. As shown in Fig. 3c, d. The anthocyanin content in the wild-type Arabidopsis seedlings was found to be significantly increased following the NF treatment. However, it was found that the exogenous ABA treatment had the ability to decrease the anthocyanin accumulation induced by oxidative stress in the Arabidopsis seedlings. Similarly, the anthocyanin content in the dpg1 seedlings was observed to be significantly higher than that of the wild-type in the control group, and the NF treatment had further enhanced the accumulation anthocyanin content in the dpg1 mutant seedlings. However, the exogenous ABA treatment also significantly decreased the anthocyanin content induced by the NF in the dpg1 mutant (Fig. 3d).
ABA treatment decreased the anthocyanin content in the dpg1 mutant. a Phenotypes of the representative seedlings of 4-day-old wild-type (Col) and dpg1 grown on 1/2 MS medium supplemented without (Mock) or with 0.25 µM ABA. Scale bar = 1 mm. b Anthocyanin contents of the 4-day-old wild-type (Col) and dpg1 seedlings grown on 1/2 MS medium supplemented without (Mock) or with 0.25 µM ABA. c Phenotypes of the representative seedlings of 4-day-old wild-type (Col) and dpg1 grown on 1/2 MS medium supplemented without (Mock) or with combinations of 5 µM NF and 0.25 µM ABA (NF + ABA). Scale bar = 1 mm. d Anthocyanin contents of the 4-day-old wild-type (Col) and dpg1 seedlings grown on 1/2 MS medium supplemented without (Mock) or with combinations of 5 µM NF and 0.25 µM ABA (NF + ABA). The error bars represent the SDs for the means of 3 independent replicates. The asterisks indicate statistically significant differences compared with the corresponding wild-type (Student’s t test: *P < 0.05)
ABA treatment promoted chloroplast development in the dpg1 mutant
In order to explore the effects of the ABA treatment on the chlorophyll content and chloroplast development in the dpg1 mutant seedlings, the exogenous ABA was used to treat the wild-type and dpg1 mutant seedlings. The results showed that the cotyledons of dpg1 mutant seedlings were greener than that of the control group following the ABA treatment (Fig. 4a). Furthermore, the content of chlorophyll a, chlorophyll b, and total chlorophyll were found to be significantly higher than those of the control group (Fig. 4b). Similarly, the chloroplast size of the cotyledon of the dpg1 mutant seedling was small, and the thylakoid components had a low abundance and a lower stacking degree of thylakoids in the control group. However, the ABA treatment was able to increase the chloroplast size of the cotyledons of the dpg1 mutant seedling, and significantly increased the abundance and stacking degree of the thylakoids (Fig. 4c). These findings indicated that the exogeneous ABA treatment could significantly promote the chloroplast development in the dpg1 mutant seedlings.
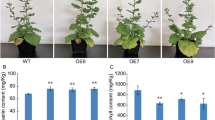
ABA treatment promoted chloroplast development in the dpg1 mutant. a Phenotypes of the representative seedlings of wild-type (Col) and dpg1 grown on 1/2 MS medium supplemented without (Mock) or with 0.25 µM ABA for 5–7 days. Scale bar = 2 mm. b The chlorophyll contents in the 7-day-old wild-type (Col) and dpg1 seedlings grown on 1/2 MS medium supplemented without (Mock) or with 0.25 µM ABA. The asterisks indicate statistically significant differences compared with the corresponding wild type (Student’s t test: *P < 0.05). c Chloroplast structures from 7-day-old wild-type (Col) and dpg1 seedlings grown on 1/2 MS medium supplemented without (Mock) or with 0.25 µM ABA. The arrowheads indicate chloroplasts in the wild-type and dpg1 seedlings. Scale bar = 1 µm
ABI1 and ABI3 play important roles in the ABA inhibition of the anthocyanin accumulation induced by the oxidative stress
The results of our study show that ABA treatment significantly decreased the anthocyanin accumulation induced by the oxidative stress in the Arabidopsis seedlings. To further explore which components in the ABA signaling pathway participated in this process, the oxidative stress condition was simulated by a norflurazon (NF) treatment, and the anthocyanin content was determined in the wild-type and ABA insensitive mutants (abi1 and abi3) seedlings of both the control and treatment group. The results are detailed in Fig. 5. In the control group, the anthocyanin content in the abi1 and abi3 mutants was found to be significantly lower than that of the wild type. These findings indicated that the ABI1 and ABI3 played positive roles in the anthocyanin accumulation of the Arabidopsis seedlings. When the NF treatment was used to simulate the oxidative stress in the wild type and ABA insensitive (abi) mutant seedlings, the anthocyanin contents in both the wild type and abi mutant seedlings were significantly higher than that of the control group. However, there were no significant differences observed in the anthocyanin content between the abi mutants and the wild-type seedlings. These results indicated that, although the ABI1 and ABI3 mutations could potentially influence the anthocyanin content in the Arabidopsis seedlings under normal growth conditions, they did not influence the anthocyanin accumulation induced by the oxidative stress in the Arabidopsis seedlings. When both the NF and ABA were used to treat the abi mutant and wild-type seedlings, the anthocyanin content in the wild type seedlings was significantly decreased when compared with the NF treatment group. However, the anthocyanin contents in the abi1 and abi3 seedlings were not significantly reduced (Fig. 5), which indicated that ABI and ABI3 play important roles in the ABA inhibition of the anthocyanin accumulation induced by the oxidative stress in the Arabidopsis seedlings.
ABI1 and ABI3 play important roles in the ABA inhibition of the anthocyanin accumulation induced by the oxidative stress. a Phenotypes of the representative seedlings of the 4-day-old wild-type (Ler), abi1 (CS22), and abi3 (CS24) grown on 1/2 MS medium supplemented without (Mock) or with combinations of 5 µM NF and 0.25 µM ABA (NF + ABA). Scale bar = 1 mm. b The anthocyanin contents of the 4-day-old wild-type (Ler), abi1 (CS22), and abi3 (CS24) seedlings grown on 1/2 MS medium supplemented without (Mock) or with combinations of 5 µM NF and 0.25 µM ABA (NF + ABA). The error bars represent the SDs for the means of 3 independent replicates. The asterisks indicate statistically significant differences compared with the corresponding wild-type (Student’s t test: *P < 0.05)
Discussion
Our previous data showed that disruption of AtDPG1 led to the Arabidopsis seedlings showing the characteristics of the albino phenotype, and the chloroplast development had been blocked (Liu et al. 2016). This study found that the anthocyanin content in the dpg1 mutant seedlings was significantly increased when compared with the wild-type (Fig. 1b). Also, the anthocyanin had mainly accumulated in the upper hypocotyls, as well as the edges of the adaxial surfaces of the cotyledons (Fig. 1a). The quantitative RT-PCR analyses revealed that the expressions of the anthocyanin biosynthetic genes (such as CHS, CHI, F3′H, DFR, LDOX, UGT78D2, and UF3GT) were significantly increased (Fig. 1c), in which the CHS, CHI, and F3′H were determined to be the early biosynthetic genes for the anthocyanin in Arabidopsis, while the DFR, LDOX, UGT78D2, and UF3GT were the late biosynthetic genes. The results of the previous studies showed that the MYB11 and PAP1, regulated the expression of the early and late biosynthetic genes, respectively, for the anthocyanin in the Arabidopsis (Borevitz et al. 2000; Teng et al. 2005; Stracke et al. 2007). It was showed that the expressions of both genes in the dpg1 mutant were significantly higher than that of the wild-type (Fig. 1c), which indicated that the increased expressions of the anthocyanin biosynthetic genes in the dpg1 mutant were possibly due to the up-regulated expressions of the PAP1 and MYB11 genes. The above-mentioned results showed that disruption of the AtDPG1 had led to the increases in anthocyanin content of the Arabidopsis seedlings, possibly due to the enhanced expressions of the anthocyanin biosynthetic and regulatory genes.
To confirm that the disruption of the AtDPG1 was responsible for the anthocyanin over-accumulation phenotype, we attempted to rescue the phenotype by introducing a cDNA of the AtDPG1 gene driven by the CaMV 35S promoter back into the heterozygous dpg1 plants. As expected, the transgenic lines showed the characteristics of a wild-type phenotype (Fig. 2a). Furthermore, the content of anthocyanin in the transgenic seedlings was found to be significantly decreased when compared with that of the dpg1 mutant (Fig. 2c). It is interesting to note that overexpression of the AtDPG1 gene only resulted in the anthocyanin content in the dpg1 mutant seedlings decreasing to the level of the wild-type, without decreasing to a level significantly lower than the wild-type (Fig. 2c). These findings indicated that the AtDPG1 may not have directly regulated the anthocyanin biosynthesis in the Arabidopsis seedlings. It was possible that the AtDPG1 participated in the regulation of the anthocyanin accumulation by influencing the chloroplast development, or through plastid retrograde signaling. The previous studies showed that the plastid retrograde signaling regulated the expression of the photosynthetic genes when the chloroplast development was blocked, or the function was abnormal (Leister 2005; Nott et al. 2006). With the deepening of the related research in recent years, it has been determined that the plastid retrograde signaling also regulated the anthocyanin biosynthesis, flowering, and stress resistance in higher plants (Cottage et al. 2010; Cheng et al. 2012; Yu et al. 2012; Feng et al. 2016). Our previous data showed that the loss-of-function mutation in the AtDPG1 led to defects in the chloroplast development, and also stimulated the plastid retrograde signaling (Liu et al. 2016). The anthocyanin content in the dpg1 mutant seedlings was significantly increased (Fig. 1b), which indicated that the increases in the anthocyanin content of the dpg1 mutant seedlings were possibly mediated by the plastid retrograde signaling.
Under abiotic stress conditions, anthocyanin accumulation is increased to assist in alleviating oxidative damages in plants (Hernandez et al. 1995; Chalker-Scott 1999; Bailey et al. 2002; Sakamoto et al. 2004; Page et al. 2012; Maruta et al. 2014). A large number of previous studies have shown that the ABA can enhance the capacity of plants to resist stress-induced oxidative damages through the generation of reaction oxygen species (ROS), increased expressions of genes which encode antioxidant protective enzymes, and the enhancement of the activities of antioxidant protective enzymes (Bueno et al. 1998; Kaminaka et al. 1999; Guan et al. 2000; Jiang and Zhang 2001; Sarafraz-Ardakani et al. 2014). In this study, it was found that the anthocyanin content was significantly increased when the NF was used to simulate the oxidative stress conditions, and treat the wild-type Arabidopsis seedlings. However, ABA treatments were found to significantly reduce the level of anthocyanin accumulation induced by the NF (Fig. 3c, d), which indicated that the ABA could reduce the anthocyanin accumulation induced by oxidative stress may through enhancing the capacity of the plants to resist the oxidative stress. Many mutants hypersensitive to high light are liable to accumulate more anthocyanin than wild type under various stresses (Kleine et al. 2007; Frenkel et al. 2009; Youssef et al. 2010). It has also been reported that several mutants (var2, clpr4, and sig6), which are defective in chloroplast development and are hypersensitive to high light, produced higher content of anthocyanin than wild type (Gan et al. 2014). The anthocyanin content in the dpg1 mutant was observed to be significantly increased under the normal growth conditions, and it could be significantly decreased to the level of the wild-type following the exogenous ABA treatment (Fig. 3b). It is possible that the anthocyanin over-accumulation is a photoprotective mechanism in dpg1 mutant, while the exogenous ABA was able to significantly decrease the anthocyanin content by enhancing the capacity of the dpg1 seedlings to resist photooxidative stress. Related research literature revealed that the expression of the Cu/Zn superoxide dismutase 2 gene had been up-regulated in many mutants with abnormal chloroplast biogenesis due to experiencing oxidative stress, which indicated that the occurrence of oxidative stress was a common phenomenon in mutants with abnormal chloroplast biogenesis (Myouga et al. 2008; Huang et al. 2009; Kikuchi et al. 2009; Miura et al. 2010; Gan et al. 2014). It was determined that the ABA enhanced the capacity of the plants to resist oxidative stress through inducing the expressions of the genes which encoded antioxidant protective enzymes in higher plants, such as the Mn-superoxide dismutase, Fe-superoxide dismutase, and Cu/Zn-superoxide dismutase genes (Zhu and Scandalios 1994; Bueno et al. 1998; Guan and Scandalios 1998; Kaminaka et al. 1999; Guan et al. 2000). The results of our previous data showed that the AtDPG1 mutation led to the Arabidopsis seedling displaying the characteristics of the albino phenotype, and the chloroplast development had been blocked (Liu et al. 2016). In this study, we found that the anthocyanin content in the dpg1 seedlings was significantly increased when compared with the wild-type (Fig. 1). When the vector harboring the wild-type AtDPG1 gene was constructed and transformed into heterozygous dpg1 plants, it was able to obtain progeny homozygous for dpg1 yet with the wild type phenotype (Fig. 2a). The further phenotype analysis of the rescued plants indicated that the chloroplast development in these transgenic seedlings was normal (Liu et al. 2016), and the anthocyanin content was significantly decreased (Fig. 2c). These findings implied that the dpg1 mutant had possibly suffered oxidative stress due to the blocking of the chloroplast development, which eventually led to increases in the anthocyanin accumulation. It was also found that the exogeneous ABA treatment was able to promote the chloroplast development in the dpg1 mutant (Fig. 4c), and the anthocyanin content was obviously lower than that of the control group (Fig. 3). These results further implied that the exogenous ABA could decrease the anthocyanin accumulation by enhancing the capacity of the dpg1 seedlings to resist the oxidative stress.
ABI1 and ABI3 are important components in the ABA signaling pathway. During the various stages of growth and development of the plants, the abi1 and abi3 mutants had displayed insensitive phenotype to the ABA treatment (Gazzarrini and McCourt 2001; Finkelstein et al. 2008). In this study, the anthocyanin content in the abi1 and abi3 seedlings was observed to be significantly decreased under the normal growth conditions (Fig. 5), which indicated that ABI1 and ABI3 played a positive regulatory role in the anthocyanin accumulation in the Arabidopsis seedlings. When NF was used for the simulation of the oxidative stress to treat the Arabidopsis seedlings, the anthocyanin content in the wild-type, as well as in the abi1 and abi3 seedlings, was significantly higher than that of the control group. In contrast, there were no significant differences observed in the anthocyanin content between the abi mutants and wild-type seedlings, indicating that the ABI1 and ABI3 could not have been involved in the anthocyanin accumulation induced by the oxidative stress in the Arabidopsis seedlings. To further explore the roles of the ABI1 and ABI3 involved in the ABA inhibition of the anthocyanin accumulation induced by oxidative stress, the abi mutants and wild-type seedlings were treated with NF together with ABA. The results showed that the anthocyanin content in wild-type seedlings was obviously lower than that in the NF treatment group. However, the anthocyanin content in the abi1 and abi3 seedlings had not been significantly decreased (Fig. 5). Since the abi1 and abi3 were mutants which were insensitive to the ABA signal, it indicated that the ABI1 and ABI3 played positive regulatory roles in the ABA inhibition of the anthocyanin accumulation induced by the oxidative stress.
References
Azeem S, Li Z, Zheng H, Lin W, Arafat Y, Zhang Z, Lin X, Lin W (2016) Quantitative proteomics study on Lsi1 in regulation of rice (Oryza sativa L.) cold resistance. Plant Growth Regul 78:307–323
Bailey S, Thompson E, Nixon PJ, Horton P, Mullineaux CW, Robinson C, Mann NM (2002) A critical role for the Var2 FtsH homologue of Arabidopsis thaliana in the photosystem II repair cycle in vivo. J Biol Chem 277:2006–2011
Borevitz JO, Xia Y, Blount J, Dixon RA, Lamb C (2000) Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 12:2383–2393
Bueno P, Piqueras A, Kurepa J, Savoure A, Verbruggen A, Montagu MV, Inze D (1998) Expression of antioxidant enzymes in response to abscisic acid and high osmoticum in tobacco BY-2 cell cultures. Plant Sci 138:27–34
Burgess D, Taylor W (1987) Chloroplast photooxidation affects accumulation of cytosolic mRNAs encoding chloroplast proteins in maize. Planta 170:520–527
Chalker-Scott L (1999) Environmental significance of anthocyanins in plant stress responses. Photochem Photobiol 70:1–9
Chen P, Hu H, Zhang Y, Wang Z, Dong G, Cui Y, Qian Q, Ren D, Guo L (2018) Genetic analysis and fine-mapping of a new rice mutant, white and lesion mimic leaf1. Plant Growth Regul 85:425–435
Cheng J, Yuan S, Zhang ZW, Zhu F, Tang H, Xu F, Feng H, Xie HF, Xu WL, Lin HH (2012) Plastid-signalling-mediated anthocyanin accumulation in mature Arabidopsis rosettes. Plant Growth Regul 68:223–230
Chory J, Peto C, Feinbaum R, Pratt L, Ausubel F (1989) Arabidopsis thaliana mutant that develops as a light-grown plant in the absence of light. Cell 58:991–999
Cottage A, Mott EK, Kempster JA, Gray JC (2010) The Arabidopsis plastid-signalling mutant gun (genomes uncoupled1) shows altered sensitivity to sucrose and abscisic acid and alterations in early seedling development. J Exp Bot 61:3773–3786
Dubos C, Le JG, Baudry A, Huep G, Lanet E, Debeaujon I, Routaboul JM, Alboresi A, Weisshaar B, Lepiniec L (2008) MYBL2 is a new regulator of flavonoid biosynthesis in Arabidopsis thaliana. Plant J 55:940–953
Feng P, Guo H, Chi W, Chai X, Sun X, Xu X, Ma J, Rochaix JD, Leister D, Wang H, Lu C, Zhang L (2016) Chloroplast retrograde signal regulates flowering. Proc Natl Acad Sci 113:10708–10713
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415
Frenkel M, Kulheim C, Jankanpaa HJ, Skogstrom O, Dall’Osto L, Agren J, Bassi R, Moritz T, Moen J, Jansson S (2009) Improper excess light energy dissipation in Arabidopsis results in a metabolic reprogramming. BMC Plant Biol 9:12–29
Gan Y, Li H, Xie Y, Wu W, Li M, Wang X, Huang J (2014) THF1 mutations lead to increased basal and wound-induced levels of oxylipins that stimulate anthocyanin biosynthesis via COI1 signaling in Arabidopsis. J Integr Plant Biol 56:916–927
Gazzarrini S, McCourt P (2001) Genetic interactions between ABA, ethylene and sugar signaling pathways. Curr Opin Plant Biol 4:387–391
Gong M, Li YJ, Chen SZ (1998) Abscisic acid-induced thermotoleranee in maize seedlings is mediated by calcium and associated with antioxidant systems. J Plant Physiol 153:488–496
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53:814–827
Grotewold E (2006) The genetics and biochemistry of floral pigments. Annu Rev Plant Biol 57:761–780
Guan L, Scandalios JG (1998) Two structurally similar maize cytosolic superoxide dismutase genes, Sod4 and Soc4A, respond differentially to abscisic acid and high osmoticum. Plant Physiol 117:217–224
Guan L, Zhao J, Scandalios JG (2000) Cis-elements and transfactors that regulate expression of the maize Catl antioxidant gene in response to ABA and osmotic stress: H2O2 is the likely intermediary signaling molecule for the response. Plant J 22:87–95
Hammond-Kosack KE, Jones JD (1996) Resistance gene-dependent plant defense response. Plant Cell 8:1773–1791
Hernandez JA, Olmos E, Corpas FJ, Sevilla F, del Rio LA (1995) Salt-induced oxidative stress in chloroplasts of pea plants. Plant Sci 105:151–167
Hoffmann AM, Noga G, Hunsche M (2016) Alternating high and low intensity of blue light affects PSII photochemistry and raises the contents of carotenoids and anthocyanins in pepper leaves. Plant Growth Regul 79:275–285
Holton T, Cornish E (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7:1071–1083
Hu X, Jiang M, Zhang A (2006) Calcium-calmodulin is required for abscisic acid-induced antioxidant defense and functions both upstream and downstream of H2O2 production in leaves of maize (Zea mays) plants. New phytol 173:27–38
Huang X, Zhang X, Yang S (2009) A novel chloroplast-localized protein EMB1303 is required for chloroplast development in Arabidopsis. Cell Res 19:1205–1216
Inaba T, Schnell DJ (2008) Protein trafficking to plastids: one theme, many variations. Biochem J 413:15–28
Jiang M, Zhang J (2001) Effect of abscisic acid on active oxygen species, antioxidative defence system and oxidative damage in leaves of maize seedlings. Plant Cell Physiol 42:1265–1273
Jiang M, Zhang J (2002) Role of abscisic acid in water stress induced antioxidant defense in leaves of maize seedlings. Free Rad Res 36:1001–10l0l5
Kaminaka H, Morita S, Tokumoto M, Masumura T, Tanaka K (1999) Differential gene expression of rice superoxide dismutase isoforms to oxidative and environmental stresses. Free Rad Res 31:S219–S225
Kikuchi S, Oishi M, Hirabayashi Y, Lee DW, Hwang I, Nakai M (2009) A 1-megadalton translocation complex containing Tic20 and Tic21 mediates chloroplast protein import at the inner envelope membrane. Plant Cell 21:1781–1797
Kleine T, Kindgren P, Benedict C, Hendrickson L, Strand A (2007) Genome-wide gene expression analysis reveals a critical role for CRYPTOCHROME1 in the response of Arabidopsis to high irradiance. Plant Physiol 144:1391–1406
La Rocca N, Rascio N, Oster U, Rudiger W (2001) Amitrole treatment of etiolated barley seedlings leads to deregulation of tetrapyrrole synthesis and to reduced expression of Lhc and RbcS genes. Planta 213:101–108
Larkindale J, Knight MR (2002) Protection against heat stress induced oxidative damage in Arabidopsis involves calcium, abscisic acid, ethylene, and salicylic acid. Plant Physiol 128:682–695
Lee J, He K, Stolc V, Lee H, Figueroa P, Gao Y, Tongprasit W, Zhao H, Lee I, Deng XW (2007) Analysis of transcription factor HY5 genomic binding sites revealed its hierarchical role in light regulation of development. Plant Cell 19:731–749
Leister D (2005) Genomics-based dissection of the cross-talk of chloroplasts with the nucleus and mitochondria in Arabidopsis. Gene 354:110–116
Lichtenthaler HK, Wellburn AR (1983) Determination of total carotenoids and chlorophylls a and b of leaf extracts in different solvents. Biochem Soc Trans 11:591–592
Liu D, Li W, Cheng J (2016) The novel protein DELAYED PALE-GREENING1 is required for early chloroplast biogenesis in Arabidopsis thaliana. Sci Rep 6:srep25742
Maruta T, Noshi M, Nakamura M, Matsuda S, Tamoi M, Ishikawa T, Shigeoka S (2014) Ferulic acid 5-hydroxylase 1 is essential for expression of anthocyanin biosynthesis-associated genes and anthocyanin accumulation underphotooxidative stress in Arabidopsis. Plant Sci 219–220:61–68
Matsui K, Umemura Y, Ohme-Takagi M (2008) AtMYBL2, a protein with a single MYB domain, acts as a negative regulator of anthocyanin biosynthesis in Arabidopsis. Plant J 55:945–967
Mehrtens F, Kranz H, Bednarek P, Weisshaar B (2005) The Arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiol 138:1083–1096
Miura E, Kato Y, Sakamoto W (2010) Comparative transcriptome analysis of green/white variegated sectors in Arabidopsis yellow variegated2: responses to oxidative and other stresses in white sectors. J Exp Bot 61:2433–2445
Myouga F, Hosoda C, Umezawa T, Iizumi H, Kuromori T, Motohashi R, Shono Y, Nagata N, Ikeuchi M, Shinozaki K (2008) A heterocomplex of iron superoxide dismutases defends chloroplast nucleoids against oxidative stress and is essential for chloroplast development in Arabidopsis. Plant Cell 20:3148–3162
Nott A, Jung HS, Koussevitzky S, Chory J (2006) Plastid-to-nucleus retrograde signaling. Annu Rev Plant Biol 57:739–759
Page M, Sultana N, Paszkiewicz K, Florance H, Smirnoff N (2012) The influence of ascorbate on anthocyanin accumulation during high light acclimation in Arabidopsis thaliana: further evidence for redox control of anthocyanin synthesis. Plant Cell Environ 35:388–404
Pelletier MK, Murrell JR, Shirley BW (1997) Characterization of flavonol synthase and leucoanthocyanidin dioxygenase genes in Arabidopsis. Further evidence for differential regulation of “early” and “late” genes. Plant Physiol 113:1437–1445
Rowan DD, Cao M, Lin WK, Cooney JM, Jensen DJ, Austin PT, Hunt MB, Norling C, Hellens RP, Schaffer RJ, Allan AC (2009) Environmental regulation of leaf colour in red 35S:PAP1 Arabidopsis thaliana. New Phytol 182:102–115
Sakamoto W, Miura E, Kaji Y, Okuno T, Nishizono M, Ogura T (2004) Allelic characterization of the leaf-variegated mutation var2 identifies the conserved amino acid residues of FtsH that are important for ATP hydrolysis and proteolysis. Plant Mol Biol 56:705–716
Sarafraz-Ardakani MR, Khavari-Nejad RA, Moradi F, Najafi F (2014) Abscisic acid and cytokinin-induced carbohydrate and antioxidant levels regulation in drought-resistant and -susceptible wheat cultivar during grain filling under field conditions. Int J Biosci 5:11–24
Shin J, Park E, Choi G (2007) PIF3 regulates anthocyanin biosynthesis in an HY5-dependent manner with both factors directly binding anthocyanin biosynthetic gene promoters in Arabidopsis. Plant J 49:981–994
Steyn WJ, Wand SJE, Holcroft DM, Jacobs G (2002) Anthocyanins in vegetative tissues: a proposed unified function in photoprotection. New Phytol 155:349–361
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B (2007) Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J 50:660–667
Teng S, Keurentjes J, Bentsink L, Koornneef M, Smeekens S (2005) Sucrose-specific induction of anthocyanin biosynthesis in Arabidopsis requires the MYB75/PAP1 gene. Plant Physiol 139:1840–1852
Wang Y, Wang Y, Song Z, Zhang H (2016) Repression of MYBL2 by both microRNA858a and HY5 leads to the activation of anthocyanin biosynthetic pathway in Arabidopsis. Mol Plant 9:1395–1405
Wilkinson S, Davies WJ (2002) ABA-based chemical signalling: the co-ordination of responses to stress in plants. Plant Cell Environ 25:195–210
Winkel-Shirley B (2001) Flavonoid biosynthesis: a colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126:485–493
Xie Y, Tan H, Ma Z, Huang J (2016) DELLA proteins promote anthocyanin biosynthesis via sequestering MYBL2 and JAZ suppressors of the MYB/bHLH/WD40 complex in Arabidopsis thaliana. Mol Plant 9:711–721
Yang J, Zhang J, Wang Z, Zhu Q, Wang W (2001) Hormonal changes in the grains of rice subjected to water stress during grain filling. Plant Physiol 127:315–323
Youssef A, Laizet Y, Block MA, Marecha E, Alcaraz JP, Larson TR, Pontier D, Gaffe J, Kuntz M (2010) Plant lipid-associated fibrillin proteins condition jasmonate production under photosynthetic stress. Plant J 61:436–445
Yousuf PY, Ahmad A, Ganie AH, Sareer O, Krishnapriya V, Aref IM, Iqbal M (2017) Antioxidant response and proteomic modulations in Indian mustard grown under salt stress. Plant Growth Regul 81:31–50
Yu HD, Yang XF, Chen ST, Wang YT, Li JK, Shen Q, Liu XL, Guo FQ (2012) Downregulation of chloroplast RPS1 negatively modulates nuclear heat-responsive expression of HsfA2 and its target genes in Arabidopsis. PLoS Genet 8:e1002669
Zeevaart JAD, Creelman RA (1988) Metabolism and physiology of abscisic acid. Annu Rev Plant Physiol Plant Mol Biol 39:439–473
Zhu D, Scandalios JG (1994) Differential accumulationof manganese-superoxide dismutase transcripts in maize in response to abscisic acid and high osmoticum. Plant Physiol 106:173–178
Acknowledgements
We would like to give our great thanks to Mrs. Li-Xia Ma for technical assistance, and the Arabidopsis Biological Resource Center at The Ohio State University for providing the T-DNA insertion line. This work was supported by the National Natural Science Foundation of China (Grant Number 31560077) and the National Key Research and Development Program of China (Grant Number 2017YFD0301605).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, M., Lv, X., Zhou, Y. et al. High anthocyanin accumulation in an Arabidopsis mutant defective in chloroplast biogenesis. Plant Growth Regul 87, 433–444 (2019). https://doi.org/10.1007/s10725-019-00481-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-019-00481-7