Abstract
Some strains of the soil bacterium Rhodococcus fascians maintain an epiphytic life style while others become endophytic. Virulent, endophytic strains cause multiple shoot growth and inhibit root growth of seed-inoculated Pisum sativum L. We were interested in assessing, at the molecular level, the impact of strains of contrasting niche on the emerging shoots and roots of inoculated seeds. The presence of R. fascians was monitored microscopically, endogenous cytokinin and chlorophyll levels were measured, and the expression of genes monitored by RT-qPCR. The expression of the pea sugar transporter genes (SWEET and SUT), amino acid (AAP) transporters and cell wall invertase gene family members, as well as expression of plant and bacterial cytokinin biosynthesis (IPT), activation (LOG) and degradation (CKX) genes were monitored. Both the virulent strain and the epiphytic strain affected the expression of the transporter genes, with less obvious differences between the strains on the shoot compared with the effect on the root. Strong expression of the R. fascians genes, RfIPT, RfLOG and RfCKX, in pea seedlings at 15 days post inoculation was mirrored by increased expression of transporter gene family members in the plant. However, the elevated levels of isopentenyl adenine-type and zeatin-type cytokinins were not consistently associated with the virulent strain. In conclusion, while both the virulent strain and the epiphytic strain impacted the expression of transporter genes in the shoots and roots, only the virulent strain affected morphology. The inhibited root growth, the greening of the roots, and the expression of the pea response regulators in the infected roots are indicative of a response to cytokinin, but a role for the ‘classical’ cytokinins as virulence determinants was not established.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multiple strains of the soil bacterium Rhodococcus fascians maintain an epiphytic life style while others become endophytic and pathogenic (Savory et al. 2017). Infection by pathogenic strains of R. fascians causes leaf deformation, fasciation, leafy galls and the formation of witches’ broom in a wide range of monocot and dicot plants (Vereecke et al. 2000, 2003), as well as the inhibition of root growth as shown in peas (Eason et al. 1995; Dhandapani et al. 2017) and arabidopsis (Francis et al. 2016).
To maintain either an epiphytic or endophytic life style, bacteria need to source both carbon- and nitrogen-containing metabolites from the plant (Bezrutczyk et al. 2018). Both an epiphytic and a pathogenic strain of R. fascians were shown to affect expression of sugar and amino acid transporters in inoculated pea cotyledons (Dhandapani et al. 2017). The effect of the pathogenic strain of R. fascians on the cotyledons of germinating peas was to retain these as a sink for the pathogen as opposed to being used to exhaustion as a source for the growing seedling (Dhandapani et al. 2017). Additionally, Dhandapani et al. (2017) suggested that the interaction between cytokinins, cell wall invertases (CWINV) and the sugar transporters known as Sugar Will Eventually be Exported Transporters (SWEET) may lead to the axillary bud outgrowth observed in germinating peas infected with the pathogenic strain.
That the cytokinins have a role in R. fascians-induced symptoms is strongly supported by the positive correlation in multiple strains between the presence of a plasmid and the genes for cytokinin biosynthesis (isopentenyl transferase-IPT), cytokinin activation (LOG), and cytokinin degradation (CKX) and virulence. Avirulent strains lack such a plasmid (Gális et al. 2005a; Stange et al. 1996; Dhandapani et al. 2017; Savory et al. 2017). Further, Depuydt et al. (2009) suggested that the over-representation of genes involved in cytokinin perception, signal transduction and homeostasis in microarray data from Arabidopsis thaliana infected with R. fascians supported the central role of cytokinin as a virulence determinant.
Crespi et al. (1992) showed that the fas operon of the linear virulence plasmid pFid188 from R. fascians strain 188 consisted of six genes most of which have since been shown to code for enzymes involved in cytokinin biosynthesis and metabolism (Crespi et al. 1992, 1994; Pertry et al. 2010). Isopentenyl transferase (IPT) is encoded by the fasD gene of R. fascians; fasE is homologous to the plant CKX gene which acts to degrade cytokinins; fasF is homologous to the plant Lonely Guy (LOG) which directly converts the products of IPT to active free bases (Kurakawa et al. 2007; Kuroha et al. 2009). Additionally, two SAM-dependent methyl transferases (MT1 and MT2) are located on the fas locus upstream of the fas genes: the products of MT2 are converted to methylated cytokinins by Fas4 (FasD) (Radhika et al. 2015).
There are conflicting hypotheses regarding the efficacy of R. fascians-produced cytokinins and disease symptoms, as the leafy symptoms would normally indicate elevated cytokinin levels (Morris 1987), as would the inhibition of root growth (Werner et al. 2003). Pertry et al. (2010) suggested the continuous presence of cZ and 2MeScZ in Arabidopsis infected with R. fascians virulent strain D188 enabled continuous tissue proliferation, due to the inefficient metabolism of these compounds by CKX, although neither of these cytokinins was detected in tobacco leafy galls (Pertry et al. 2009). Creason et al. (2014), on the other hand, suggested that only one cytokinin type (the iP-type) was necessary for disease symptoms, a statement somewhat supported by Dhandapani et al. (2017) in Rhodococcus-infected cotyledons, while Radhika et al. (2015) suggest that R. fascians-produced methylated cytokinins are the key virulence determinants.
Here we report the impact of strains of contrasting niche on the emerging shoots and roots of inoculated seeds. We were interested if the effect of R. fascians on the expression of both sugar and amino acid transporters continued as the roots and shoots of inoculated peas developed, and whether the levels of the iP-type cytokinins were elevated in both the shoots and roots of peas inoculated with the pathogenic strain, as would be expected if only the iP-type of cytokinin was needed for infection (Creason et al. 2014).
Materials and methods
Plant material, Rhodococcus fascians inoculation and sampling
Pea (Pisum sativum) var. Bohatyr seeds were obtained from the Institute for Plant and Food Research, Christchurch, New Zealand. The pea seeds were surface sterilised and inoculated with R. fascians pathogenic strain 602, which contains a linear virulence plasmid, and a plasmid-free avirulent strain, 589, as described in Dhandapani et al. (2017).
Six seeds were placed in sterilised 500 ml containers with 0.6% (w/v) agar and 10% (w/v) Hoagland’s mineral salts solution (Lawson et al. 1982) and germinated at 22 °C with a 16 h photoperiod in a growth room. Samples of separated shoots and roots from 5 to 35 days post inoculation (dpi) were either immediately immersed in liquid nitrogen and then stored at − 80 °C for gene expression studies, or fixed in FAA (10% formaldehyde:5% acetic acid:50% alcohol) for light microscopy, and cryopreserved for scanning electron microscopy. For each sampling, tissues from five plant samples were collected from each of the three treatments: the avirulent and virulent strain-inoculated peas, and the mock-inoculated controls.
Chlorophyll estimation
The chlorophyll content of shoot and root samples was measured from 5 to 35 dpi by immersing 0.1 g FW in 1 ml dimethylformamide overnight at 4 °C, and using a Nanodrop spectrophotometer to determine the absorbance at 664 and 667 nm as described in Evans et al. (2012).
Light microscopy
A modified procedure of Carletom and Druvy (1957) for fixation, dehydration, embedding and microtoming was followed. The detailed protocol is described in Dhandapani et al. (2017).
RNA isolation and cDNA synthesis
Total RNA was extracted from pea tissues (shoot and root) with the TRIzol® Reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol and as explained by Dhandapani et al. (2017). The concentration and integrity of isolated total RNA were assessed using a NanoDrop™ Spectrophotometer (Thermo Fisher Scientific Inc.) and 1% (w/v) agarose gel electrophoresis. The extracted RNA was converted to cDNA by reverse transcription, diluted fivefold with TE buffer and stored at − 20 °C. The cDNA quality was validated by RT-qPCR assay by using two reference genes and a target gene (Dhandapani et al. 2017).
Genes of interest and real-time reverse transcription quantitative PCR (RT-qPCR)
Gene isolation, sequence analysis and identification are described in Dhandapani et al. (2017). The detailed phylogenetic analysis of the cytokinin (PsIPTs, PsLOGs, PsCKXs and PsRRs) gene family members and transporter (PsSUTs, PsAAPs, PsCWINVs and PsSWEETs) gene family members has been published in Dhandapani et al. (2017). Previously these were aligned using the ClustalX program (Thompson et al. 1997) and MEGA4 (Tamura et al. 2007).
Primers were designed which were unique and specific for the genes of interest based on P. sativum transcriptome analysis results and R. fascians BLAST search results using Primer Premier TM 5.00. The expression studies were conducted following the RT-qPCR MIQE guidelines (Bustin et al. 2009), with three technical replicates for each of two biological replicates. The RT-qPCR assay was run using the KAPA SYBR® FAST qPCR Kits (Kapa Biosytems, Boston, USA). The temporal expression of each gene of interest and selected reference genes were quantified using a Qiagen Rotor-Gene Q.
To achieve accurate normalisation in the RT-qPCR, four reference genes U18S, PsEF, PsGAP and PsACT, were used as internal controls as described by Song et al. (2012) and Dhandapani et al. (2017). RT-qPCR was used to determine the relative expression of the genes of interest in pea shoots and roots at different growth stages following seed inoculation with the virulent strain (602), the avirulent strain (589) and the mock inoculated control. The expression data are presented as heat maps with fold differences calculated relative to the mock-inoculated control.
Cytokinin analyses
Pea shoots and roots from four individual plants from each of the three treatments, were ground under liquid nitrogen and freeze-dried. The four biological replicates were extracted and purified using the method described in Antoniadi et al. (2015), and Dhandapani et al. (2017). The samples were analysed by LC–MS/MS (Svačinová et al. 2012).
Results
Morphological and microscopic differences in the shoot and root of pea following inoculation with avirulent and virulent strains of Rhodococcus fascians
The morphological variations in the pea plants inoculated with the avirulent strain of R. fascians (avir-plants) and with the virulent strain of R. fascians (vir-plants) and mock inoculated (con-plants) are described in detail in Dhandapani et al. (2017). The characteristic multiple shoot symptoms were visible from 5 dpi in vir-plants and other symptoms such as reduced leaf, stunted shoot growth, shortened and thickened roots were evident from 11 to 45 dpi in vir-plants. Both avir-plants and con-plants had similar morphological characteristics throughout their growth period.
The chlorophyll content of vir-roots was low but increased with time, while the chlorophyll content of the vir-shoots was less relative to control and avir-shoots from 11 to 20 dpi and was only greater by 35 dpi (Fig. 1). The total chlorophyll content in vir-roots was greater than control and avir-roots especially at the later developmental stages (Fig. 1).
Total chlorophyll content in a shoot and b root tissues of P. sativum infected with R. fascians. The pea seeds were imbibed for 4 h (hpi) with R. fascians avirulent strain 589 (avirulent), virulent strain 602 (virulent) and medium as mock inoculation (control) and grown in sterile agar containers until 35 days post inoculation (dpi). The error bars are ± SE of two biological replicates and four technical replicates
The multiple shoots emerging at 5 dpi were colonised by the virulent strain (Fig. 2a: C); by 15 dpi there was noticeable bacterial colonisation on the surface of the multiple shoots and on the single shoot of avir-plants (Fig. 2a: F). By 35 dpi, both strains of R. fascians appeared to colonise the shoot tissue in a similar manner (Fig. 2a: H, I). Through scanning electron microscopy, it was noticed that the colonisation of both avirulent and virulent strains of R. fascians on the surface of the pea tissues (root and shoot) increased with time from 5 to 35 dpi (Dhandapani 2014).
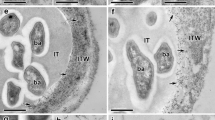
Light micrographs of sections of P. sativum shoot and root following seed inoculation with R. fascians. a Light micrographs of sections of P. sativum shoot tissues following inoculation with R. fascians at 5 days post inoculation (dpi) (A–C), 15 dpi (D–F) and 35 dpi (G–I). b Light micrographs of sections of P. sativum root tissues following inoculation with R. fascians at 2 days post inoculation (dpi) (A–C), 5 dpi (D–F), 15 dpi (G–I) and 35 dpi (J–L)
Both strains of R. fascians (avirulent and virulent) were detected on the radicle tissue of the pea at 2 dpi (Fig. 2b: B, C). The presence of mucilage-like material was seen near the surface of radicle root hairs of the roots infected by the avirulent strain (Fig. 2b: E) and the bacteria were mainly clumped together (Fig. 2b: B, E, H). The radicle infected by the virulent strain exhibited a layer of bacteria on the root epidermis and amongst the root hairs (Fig. 2b: C, F, I) which appeared as bacterial rods on enlargement (Fig. 2b: L). By 5 dpi, clumps of avirulent bacteria were evident on the root epidermis and root hairs (Fig. 2b: E), whereas the virulent bacteria were spread across the epidermal surface of the root amongst the root hairs (Fig. 2b: F). By 15 dpi, profuse colonies of virulent and avirulent bacteria were evident on the surface of the root (Fig. 2b: H, I). The avir-roots showed small rods clumped together on the root epidermis (Fig. 2b: E, H), whereas the virulent strain exhibited more diffuse spread of bacteria on roots (Fig. 2b: I, L).
Expression of RfIPT, RfCKX, and RfLOG during infection of shoots and roots
In peas inoculated with the virulent strain of R. fascians, RfIPT, RfLOG and RfCKX showed markedly increased expression in shoots and roots by 15 dpi, and more so in roots (Fig. 3). No expression of RfIPT, RfLOG and RfCKX in control (con-shoots and con-roots) and shoots and roots inoculated with the avirulent strain (avir-shoots and avir-roots) was detected, confirming that the designed primers discriminated between the R. fascians genes and the pea genes as shown in Dhandapani et al. (2017).
Relative expression of RfIPT, RfLOG and RfCKX from P. sativum shoots and roots following seed inoculation with R. fascians strain 589 (avirulent), strain 602 (virulent) and mock inoculation (control) from 5 to 35 dpi. a Shoot and b root. Data are means of relative mRNA levels in fold changes (log) detected using RT-qPCR with three technical replicates for each of two biological replicates. PsEF, U18S, PsGAP and PsACT were used as internal controls. Error bars represent ± SD calculated for the combined technical and biological replicates
Cytokinin biosynthesis and metabolism in shoot and root of pea following inoculation with avirulent and virulent strains of Rhodococcus fascians
In general, the expression of the cytokinin biosynthetic (PsIPT) and activation (PsLOG) gene family members in both vir- and avir-roots and shoots at 5 dpi was reduced relative to the controls (Figs. 4, 5). Expression of all PsIPT and LOG gene family members was elevated relative to controls from 15 to 35 dpi in vir-shoots and at 15 dpi in vir-roots. In the latter stages PsIPT and LOG expression was elevated in avir-shoots but not in avir-roots.
Relative expression of cytokinin biosynthesis (PsIPT), activation (PsLOG), degradation (PsCKX) and response regulator (PsRR) gene family members along with transporter (PsAAP, PsSUT, PsCWINV and PsSW) gene family members in P. sativum shoot tissues following seed inoculation with R. fascians strain 589 (AVIR) and virulent strain 602 (VIR) from 5 to 35 days post-inoculation. Values are fold changes relative to the expression of the mock-inoculated control. Initial fold change values were calculated using PsEF, Ps18S, PsGAP and PsACT as internal controls using three technical replicates for each of two biological replicates in the RT-qPCR. The relative expression level of each gene was then compared with the expression in pea shoots inoculated with medium as the mock-inoculated control. The colour scale indicates upregulated expression (red scale), similar (white) and downregulated expression (blue scale) relative to the mock-inoculated control. (Color figure online)
Relative expression of cytokinin biosynthesis (PsIPT), activation (PsLOG), degradation (PsCKX) and response regulator (PsRR) gene family members along with transporter (PsAAP, PsSUT, PsCWINV and PsSW) gene family members in P. sativum root tissues following seed inoculation with R. fascians strain 589 (AVIR) and virulent strain 602 (VIR) from 5 to 35 days post-inoculation. Values are fold changes relative to the expression of the mock-inoculated control. Initial fold change values were calculated using PsEF, Ps18S, PsGAP and PsACT as internal controls using three technical replicates for each of two biological replicates in the RT-qPCR. The relative expression level of each gene was then compared with the expression in pea shoots inoculated with medium as the mock-inoculated control. The colour scale indicates upregulated expression (red scale), similar (white) and downregulated expression (blue scale) relative to the mock-inoculated control. (Color figure online)
Expression patterns of PsCKX generally mirrored those of PsIPT and LOG (Figs. 4, 5). The expression of the PsCKX family members in both vir- and avir-shoots was elevated from 9 to 35 dpi (Fig. 4). PsCKX2 expression was elevated at all time points, especially in vir-shoots. Expression of the PsCKX gene family members was increased in vir-and avir-roots relative to control, particularly at 15 dpi.
Expression of the cytokinin response regulator (PsRR) gene family members in both vir- and avir-shoots was generally elevated across all stages of development relative to control (Fig. 4). However, in vir-roots expression of the PsRRs was suppressed at 5 dpi but increased at 9 and 15 dpi. In contrast, in avir-roots PsRR expression was frequently less than the relevant control.
The total cytokinin content of the vir-shoots was reduced significantly relative to control and avir-shoots at 5 and 11 dpi, but then increased at 15 and 25 dpi relative to con-shoots. At these latter time points the cytokinin content was similar in avir- and vir-shoots (Table 1). The total cytokinin content of avir-roots was significantly greater than controls at 5 and 15 dpi, whereas that in vir-roots was significantly less at 11 dpi, but greater at 15 dpi. Cytokinin levels were similar in avir and vir-roots at 15 and 25 dpi (Table 1).
Considering the individual cytokinins, there was significantly less tZ, tZRMP, iPR and iPRMP in vir-shoots 5 and 11 dpi relative to control. iP levels were consistently elevated relative to controls and avir-shoots. However, the total iP-type cytokinins were less in the vir-shoots relative to controls and avir-shoots at 5 and 11 dpi, but greater at 15 and 25 dpi.
Initially, there was no difference in the total iP-type cytokinins in the roots at 5 dpi. Subsequently, the iP-type cytokinins in the vir-roots only exceeded those in the controls at 25 dpi. The iP levels were elevated relative to control at the four time points in vir-roots. However, at both 11 and 15 dpi the iP levels in both vir- and avir-roots were significantly elevated relative to control. At 5 dpi, tZ-type cytokinins were elevated in vir-roots relative to con-roots, particularly tZR. However, at 11 dpi most cytokinin forms, including tZ, were decreased in vir-roots, while at 15 and 25 dpi free bases and nucleotides were generally elevated.
Transporter gene expression in Rhodococcus fascians-inoculated pea tissues
The expression of sucrose transporter (PsSUT) gene family members was similar in both vir- and avir-shoots at 5 dpi and was reduced relative to con-shoots. Subsequently, expression was increased or was similar to controls in both vir-and avir-shoots (Fig. 4). Likewise, the expression of the SUT gene family members was similar in both vir- and avir-roots: decreased expression at 5 and 9 dpi and elevated expression particularly at 15 dpi (Fig. 5).
Generally, the PsAAP gene family members showed decreased expression relative to controls at 5 dpi. At 15 dpi expression relative to controls was greater in the vir-shoots than the avir-shoots. Cluster 1 PsAAP 7a expression was high at 9 dpi in vir-shoots and at 15 dpi in avir-shoots (Fig. 4). The relative expression of Cluster 4B AAP6a and 8 was greater at 15 dpi in vir-shoots and at 25 dpi in avir-shoots. In general, for many PsAAP gene family members, the expression in vir-and avir-roots was reduced relative to control at 5, 9 and 35 dpi but elevated at 15 dpi and more so in the vir-roots than the avir-roots (Fig. 5).
Generally, expression of the PsCWINVs was only modestly increased in both vir- and avir-shoots relative to the controls, with the exception of PsCWINV1 at 15 dpi in vir-shoots (Fig. 4). Expression in vir-roots was generally decreased relative to controls, with the marked exception at 15 dpi where expression was elevated for the four family members, and more strongly in the vir-roots than the avir-roots (Fig. 5).
Expression of Clades I, II and III SWEET gene family members was usually decreased in vir-shoots at 5 and 9 dpi, but increased at 15 dpi (Fig. 4). SWEET expression in the avir-shoots was more variable and lacked the increase at 15 dpi seen in the vir-shoots. Clade III SW15c was markedly decreased in both vir- and avir-shoots. In vir-roots, SWEETs were noticeably elevated relative to controls at 15 dpi, more so in the vir-roots than the avir-roots (Fig. 5). Clade II PsSW7 was strongly elevated relative to controls across the experiment in vir-roots, as was Clade III SW15b in both avir- and vir-roots. In contrast, expression of most SWEETS in vir-and avir-roots was decreased relative to controls at 5 and 9 dpi.
Discussion
Rhodococcus fascians is a soil bacterium. We were interested in the effect of seed inoculation, as most previous research has focused on infection of above ground tissues. The virulent strain of R. fascians transformed the cotyledon from being a sole source for the germinating seed to a competitive source of nutrients for the pathogen (Dhandapani et al. 2017). As the integrity of the cotyledon was maintained by the virulent strain, we investigated whether there was an ongoing impact from the presence of the bacterium on the emerging roots and shoots of seeds that were inoculated with either the virulent or the avirulent strain.
We previously showed that there was a noticeably greater impact of the virulent strain on gene expression in the cotyledons compared to the avirulent strain, especially between 2 and 15 dpi (Dhandapani et al. 2017). However, as the shoot developed, on first examination of the heat map (Fig. 4) both strains led to increased expression of the genes of interest, but the differences in expression between the virulent and avirulent strains are not particularly pronounced, even though morphologically the inoculated plants were extremely different. This supports the notion of the avirulent epiphytic strain of R. fascians also affecting the metabolism of the plant (Depuydt et al. 2009; Dhandapani et al. 2017). Whether this is due to cytokinin production by the bacteria is not known, but both virulent and avirulent strains of R. fascians produce substantial quantities of non-hydroxylated cytokinins in vitro (Eason et al. 1996) as well as methylthio derivatives (Francis et al. 2016)—potentially from the turnover of tRNA (Matsubara et al. 1968; Jameson 2000).
However, at 5 dpi, when symptoms are first becoming apparent, there is reduced expression of PsIPT and PsLOG in vir-shoots compared with avir-shoots, relative to controls. This matches the reduced levels of the Z-type and iP-type endogenous cytokinins in the vir-shoots relative to controls and avir-shoots, indicating that the pathogen continues to impact the homeostatic mechanisms of the germinating plant. Expression of PsCKX gene family members is apparent in both the vir- and avir-shoots, but with noticeably greater PsCKX2 expression occurring in vir-shoots. PsCKX expression is particularly evident in the vir-roots at 15 dpi, as is PsIPT and LOG expression. CKX expression and activity frequently parallels enhanced IPT expression and increased endogenous cytokinin levels (see references cited in Jameson and Song 2016), and the virulent strain (but not the avirulent strain) of R. fascians has been shown to specifically up-regulate AtCKX3 in transgenic tobacco (Gális et al. 2005b).
Increased expression of the RR gene family members is indicative of enhanced cytokinin levels and/or perception (Hwang et al. 2012). Increased PsRR expression relative to the controls in the inoculated shoots and the vir-roots indicates that they are responding to more cytokinin than the control shoots. The lack of PsRR gene expression is particularly noticeable in the avir-roots. A similar differential for AtIPT, AtCKX and AtRR response has been reported for arabidopsis leaves inoculated with either a virulent (D188) strain or its avirulent equivalent (D188-5) (Depuydt et al. 2008, 2009).
Detection by the plant of the bacteria is also evidenced in early shoot growth with down-regulation of several SWEETS, but particularly a Clade III SWEET. SWEETs in this clade move sucrose across the plasma membrane to the apoplast (Chen 2014). Down-regulation would remove a source of carbohydrate from the bacteria. However at 15 dpi, the expression of PsCWINVs, PsSWEETS, PsSUTs and PsAAP gene family members are all upregulated in the vir-shoots but not the avir-shoots. At this time the expression of the R. fascians genes was strongly detected in the shoots and roots (Figs. 4, 5) and bacterial growth was prolific (Fig. 2). Clearly, the virulent strain was having a marked impact on the plant shoot as a source of nutrients. However, subsequently, both epiphytic and endophytic strains continued to affect nutrient transporters in the shoot, as noted also by Depuydt et al. (2009). The ability of an epiphytic strain to utilise plant-derived carbon sources was recently reported by Francis et al. (2016).
The impact of both the virulent and avirulent strains on transporters in the roots was clearly evident by 15 dpi, again when expression of RfIPT, LOG and CKX was detected and when bacterial colony growth was profuse. However, while the patterns of expression are remarkably similar, there is a stronger upregulation of gene expression in the vir-roots compared with the avir-roots at 15 dpi. Generally, there is a greater level of down-regulation of the gene families of interest by the avir-roots compared to the avir-shoots. The one exception is the noticeable up-regulation of PsSW2b in the avir-roots. SWEETS in Clade I are associated with the transport of hexoses across either the plasmalemma or the tonoplast (Chen 2014). Transport across the tonoplast would sequester the glucose away from the bacteria (Chen et al. 2015). We suggest that the plant has actively combated the demands of the epiphytic strain more successfully than the virulent strain, and more successfully in the roots than the shoots.
Cytokinin is strongly implicated in the development of chloroplasts and the accumulation of chlorophyll (Cortleven and Schmülling 2015), and in the transition of etioplasts to chloroplasts (Cortleven et al. 2016). Chlorophyll accumulated in the cotyledons of the peas infected by the virulent strain, and Dhandapani et al. (2017) suggested that this was an indicator of enhanced cytokinin levels in the infected cotyledons. Likewise, the vir-roots accumulated more chlorophyll than the avir-roots or controls, which again could be caused by enhanced cytokinin levels. Recently, Kobayashi et al. (2017) showed chloroplast development in arabidopsis roots occurred through cytokinin response regulator signalling. We show here that expression of PsRRs was elevated in vir-roots, supporting the suggestion that the vir-roots, but not the avir-roots, were responding to an elevated cytokinin content.
Root growth is inhibited by the virulent R. fascians (Eason et al. 1995; Dhandapani et al. 2017). As endogenous cytokinin levels in roots are considered to be supraoptimal (Werner et al. 2003), and applied cytokinin inhibits root growth (Guo et al. 2017), further inhibition could be expected if R. fascians was increasing endogenous cytokinin levels. The inference of greater cytokinin levels in the vir-roots is described above with respect to chlorophyll accumulation. However, in terms of the individual cytokinins measured, there is a lack of correlation between the cytokinin content of the vir-roots (growth inhibition) and the similar or greater cytokinin content of the avir-roots (no growth inhibition). Neither do the data support the contention of the iP-type cytokinins being responsible for the root inhibition caused by the virulent strain. As the methylated cytokinins are reported to inhibit root growth in arabidopsis (Radhika et al. 2015), we are currently investigating the effect of these cytokinins in pea.
In conclusion, both the virulent and avirulent strains influenced transporter gene expression in the developing root and shoot of inoculated pea plants, with a more generally positive manipulation of transporter expression in the shoots over time, but with a lesser effect on the roots until a significant mass of bacteria had accumulated. Both strains are capable of releasing cytokinin (Eason et al. 1996) and, while this may affect transporter gene expression, it appears that the ‘classical’ cytokinins, the iP- and Z-types, are unlikely to be the causative molecules of the extreme symptoms invoked by the virulent strain of R. fascians.
References
Antoniadi I, Plačková L, Simonovik B, Doležal K, Turnbull C, Ljung K, Novák O (2015) Cell-type-specific cytokinin distribution within the Arabidopsis primary root apex. Plant Cell 27:1955–1967
Bezrutczyk M, Yang J, Eom J-S et al (2018) Sugar flux and signalling in plant-microbe interactions. Plant J. https://doi.org/10.1111/tpj.13775
Bustin SA, Benes V, Garson JA et al (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622
Carletom HM, Druvy RAB (1957) Histological technique: for normal pathological tissues and the identification of parasites. Oxford University Press, London
Chen L-Q (2014) SWEET sugar transporters for phloem transport and pathogen nutrition. New Phytol 201(4):1150–1155
Chen H-Y, Huh J-H, Yu Y-Chi, Ho L-H, Chen L-Q, Tholl D, Frommer WB, Guo W-J (2015) The Arabidopsis vacuolar sugar transporter SWEET2 limits carbon sequestration from roots and restricts Pythium infection. Plant J 83:1046–1058
Cortleven A, Schmülling T (2015) Regulation of chloroplast development and function by cytokinin. J Exp Bot 66:4999–5013
Cortleven A, Marg I, Yamburenko MV, Schlicke H, Hill K, Grimm B, Schaller GE, Schmülling T (2016) Cytokinin regulates etioplast-chloroplast transition through activation of chloroplast-related genes. Plant Physiol 172:464–478
Creason AL, Vandeputte OM, Savory EA, Davis EW II, Putnam ML, Hu E, Swader-Hines D, Mol A, Baucher M, Prinsen E, Zdanowska M, Givan SA, Jaziri ME, Loper JE, Mahmud T, Chang JH (2014) Analysis of genome sequences from plant pathogenic Rhodococcus reveals genetic novelties in virulence loci. PLoS ONE 9(7):e101996. https://doi.org/10.1371/journal.pone.0101996
Crespi M, Messens E, Caplan AB, Van Montagu M, Desomer J (1992) Fasciation induction by the phytopathogen Rhodococcus fascians depends upon a linear plasmid encoding a cytokinin synthase gene. EMBO J 11:795–804
Crespi M, Vereecke D, Temmerman W, Van Montagu M, Desomer J (1994) The fas operon of Rhodococcus fascians encodes new genes required for efficient fasciation of host plants. J Bacteriol 176:2492–2501
Depuydt S, Doležal K, Van Lijsebettens M, Moritz T, Holsters M, Vereecke D (2008) Modulation of the hormone setting by Rhodococcus fascians results in ectopic KNOX activation in Arabidopsis. Plant Physiol 146:1267–1281
Depuydt S, Trenkamp S, Fernie AR, Elftieh S, Renou J-P, Vuylsteke M, Holsters M, Vereecke D (2009) An integrated genomics approach to define niche establishment by Rhodococcus fascians. Plant Physiol 149:1366–1386
Dhandapani P (2014) Rhodococcus fascians-plant interactions: microbiological and molecular aspects. Unpublished PhD thesis, University of Canterbury, Christchurch, New Zealand
Dhandapani P, Song J, Novak O, Jameson PE (2017) Infection by Rhodococcus fascians maintains cotyledons as a sink tissue for the pathogen. Ann Bot 119:841–852
Eason JR, Jameson PE, Bannister P (1995) Virulence assessment of Rhodococcus fascians strains on pea cultivars. Plant Pathol 44:141–147
Eason JR, Morris RO, Jameson PE (1996) The relationship between virulence and cytokinin production by Rhodococcus fascians (Tilford 1936) Goodfellow 1984. Plant Pathol 45:323–331
Evans T, Song J, Jameson PE (2012) Micro-scale chlorophyll analysis and developmental expression of a cytokinin oxidase/dehydrogenase gene during leaf development and senescence. Plant Growth Regul 66:95–99
Francis IM, Stes E, Zhang Y, Rangel D, Audenaert K, Vereecke D (2016) Mining the genome of Rhodococcus fascians, a plant growth-promoting bacterium gone astray. New Biotechnol. https://doi.org/10.1016/j.nbt.2016.01.009
Gális I, Bilyeu K, Wood G, Jameson PE (2005a) Rhodococcus fascians: shoot proliferation without elevated cytokinins? Plant Growth Regul 46:109–115
Gális I, Bilyeu KD, Godinho MJG, Jameson PE (2005b) Expression of three Arabidopsis cytokinin oxidase/dehydrogenase promoter::GUS chimeric constructs in tobacco: response to developmental and biotic factors. Plant Growth Regul 45:173–182
Guo Q, Love J, Song J, Roche J, Turnbull MH, Jameson PE (2017) Insights into the functional relationship between cytokinin-induced root system phenotypes and nitrate uptake in Brassica napus L. Funct Plant Biol 44:832–844
Hwang I, Sheen J, Muller B (2012) Cytokinin signaling networks. Annu Rev Plant Biol 63:353–380
Jameson PE (2000) Cytokinins and auxins in plant-pathogen interactions—an overview. Plant Growth Regul 32:369–380
Jameson PE, Song J (2016) Cytokinin: a key driver of seed yield. J Exp Bot 67:593–606
Jameson PE, Dhandapani P, Novak O, Song J (2016) Cytokinins and expression of SWEET, SUT, CWINV and AAP genes increase as pea seeds germinate. Int J Mol Sci 17:2013. https://doi.org/10.3390/ijms17122013
Kobayashi K, Ohnishi A, Sasaki D, Fujii S, Iwase A, Sugimoto K, Masuda T, Wada H (2017) Shoot removal induces chloroplast development in roots via cytokinin signalling. Plant Physiol 173:2340–2355
Kurakawa T, Ueda N, Maekawa M, Kobayashi K, Kojima M, Nagato Y, Sakakibara H, Kyozuka J (2007) Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature 445:652–655
Kuroha T, Tokunaga H, Kojima M, Ishida T, Nagawa S, Fukuda H, Sugimoto K, Sakakibara H (2009) Functional analyses of LONELY GUY cytokinin-activating enzymes reveal the importance of the direct activation pathway in Arabidopsis. Plant Cell 21:3152–3169
Lawson E, Gantotti B, Starr M (1982) A 78-megadalton plasmid occurs in avirulent strains as well as virulent strains of Corynebacterium fascians. Curr Microbiol 7:327–332
Matsubara S, Armstrong DJ, Skoog F (1968) Cytokinins in tRNA of Corynebacterium fascians. Plant Physiol 43:451–453
Morris RO (1987) Molecular aspects of hormone synthesis and action genes specifying auxin and cytokinin biosynthesis in prokaryotes. In: Davies PJ (ed) Plant hormones. Kluwer Academic Publishers, Dordrecht, pp 318–339
Pertry I, Vaclavikova K, Depuydt S, Galuszka P, Spichal L, Temmerman W, Stes E, Schmulling T, Kakimoto T, Van Montagu MCE, Strnad M, Holsters M, Tarkowski P, Vereecke D (2009) Identification of Rhodococcus fascians cytokinins and their modus operandi to reshape the plant. Proc Natl Acad Sci 106(3):929–934
Pertry I, Václavíková K, Gemrotová M, Spíchal L, Galuszka P, Depuydt S, Temmerman W, Stes E, De Keyser A, Riefler M, Biondi S, Novák O, Schmülling T, Strnad M, Tarkowski P, Holsters M, Vereecke D (2010) Rhodococcus fascians impacts plant development through the dynamic fas-mediated production of a cytokinin mix. Mol Plant Microbe Interact 23:1164–1174
Radhika V, Ueda N, Tsuboi Y, Kojima M, Kikuchi J, Kudo K, Sakakibara H (2015) Methylated cytokinins from the phytopathogen Rhodococcus fascians mimic plant hormone activity. Plant Physiol 169:1118–1126
Savory EA et al (2017) Evolutionary transitions between beneficial and phytopathogenic Rhodococcus challenge disease management. eLIFE. https://doi.org/10.7554/eLife.30925
Song J, Jiang L, Jameson PE (2012) Co-ordinate regulation of cytokinin gene family members during flag leaf and reproductive development in wheat. BMC Plant Biol 12:78
Stange RR, Jeffares D, Young C, Scott DB, Eason JR, Jameson PE (1996) PCR amplification of the fas-1 gene for the detection of virulent strains of Rhodococcus fascians. Plant Pathol 45:407–417
Svačinová J, Novák O, Plačková L, Lenobel R, Holík J, Strnad M, Doležal K (2012) A new approach for cytokinin isolation from Arabidopsis tissues using miniaturized purification: pipette tip solid-phase extraction. Plant Methods 18:17
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acid Res 25:4876–4882
Vereecke D, Burssens S, Simón-Mateo C et al (2000) The Rhodococcus fascians-plant interaction: morphological traits and biotechnological applications. Planta 210:241–251
Vereecke D, Temmerman W, Jaziri M, Holsters M, Goethals K (2003) Towards an understanding of the Rhodococcus fascians-plant interaction. In: Stacey G, Kean N (eds) Molecular plant microbe interactions, vol 6. American Phytopathological Society, St. Paul
Werner T, Motyka V, Laucou V et al (2003) Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15:2532–2550
Acknowledgements
A UC scholarship to PD is gratefully acknowledged. Thanks to Graeme Bull, Jan McKenzie and Neil Andrews for assistance with microscopy, and to the anonymous referees for their constructive comments. O.N. was funded by the Czech Science Foundation (Nr. 17-06613S).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Dhandapani, P., Song, J., Novak, O. et al. Both epiphytic and endophytic strains of Rhodococcus fascians influence transporter gene expression and cytokinins in infected Pisum sativum L. seedlings. Plant Growth Regul 85, 231–242 (2018). https://doi.org/10.1007/s10725-018-0387-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-018-0387-3