Abstract
The plant growth promoting bacteria Herbaspirillum seropedicae is an important model to study biological nitrogen fixation and it is proposed as crop inoculants for grasses. In this study we developed new tools to evaluate the abundance of H. seropedicae SmR1 and its expression in planta and investigated the association of the plant growth promoting bacteria H. seropedicae with maize grown in sterile and nonsterile conditions. Maize seedlings (P30F53) were inoculated with H. seropedicae SmR1 and grown in vitro and in soil. The plants were sampled at 4, 7 and 10 (in vitro) or 14, 21 and 28 days after inoculation (soil). Using qPCR we quantified H. seropedicae DNA and measured nifH, rpoC and hrcN levels of bacterial transcripts. In vitro assay inoculated plants presented highest amount of DNA and transcript contents, not detected in control plants. nifH and rpoC gene expression were detected on roots of inoculated maize cultivated in both growth conditions. However, it was not possible to detect hrcN gene expression in maize roots cultivated in soil. TaqMan assay is species-specific for H. seropedicae DNA and nifH and rpoC transcript levels could be used to monitor H. seropedicae gene expression in planta in sterile and nonsterile growth conditions. We developed specific, reliable and efficient tools to monitor H. seropedicae DNA abundance and H. seropedicae nifH and rpoC expression in planta.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diazotrophic bacteria are able to fix atmospheric nitrogen by biological nitrogen fixation (BNF) process (Bhattacharjee et al. 2008). Some diazothrophs live in close association with plants and have the capacity to transfer the fixed nitrogen to the host (Chubatsu et al. 2012; Carvalho et al. 2014). Furthermore, they can be used to improve crop yield and quality (Chebotar et al. 2015; Berg 2009). Future biotechnological applications, such as an alternative for chemical fertilizers (Babalola 2010), increasing crop production or even conserving biodiversity (Germida et al. 1998; Stets et al. 2013), are processes that can be driven based on the advance of plant-diazotrophic bacteria association’s knowledge. Plant response to inoculation depends on the plant genotype and the plant growth bacterial inoculant (Amaral et al. 2016; Alves et al. 2015). During plant–beneficial bacteria interaction, microorganisms modulate plant molecular responses (Vargas et al. 2012, 2014; Brusamarello-Santos et al. 2012; Berg 2009), and both partners are altered in response to interaction (Cordeiro et al. 2013; Tadra-Sfeir et al. 2015; Balsanelli et al. 2015).
Herbaspirillum seropedicae is an endophytic diazotrophic bacterium, which is able to colonize internal tissues of plants (Chubatsu et al. 2012). It belongs to β-proteobacteria class and colonizes mainly roots, stems and leaves of Poacea (Olivares et al. 1996; James et al. 1997; Baldani et al. 1986). Assessing 21 strains of Herbaspirillum in two maize varieties, Alves et al. (2015) observed that H. seropedicae ZAE94 contributed to plant growth in greenhouse assay and its application as field inoculant increased maize yield up to 34% and can provide 37% of nitrogen plant demand by BNF. One of the best studied strain, H. seropedicae SmR1 (Monteiro et al. 2012) has its complete genome sequenced (Pedrosa et al. 2011). Recent work using Setaria viridis co-inoculated with two nitrogen-fixing bacteria, H. seropedicae SmR1 and Azospirillum brasilense FP2 revealed that the plant could incorporate the fixed nitrogen by BNF in its proteins. Additionally, the nitrogen fixed by the bacteria directly affects overall plant metabolism (Pankievicz et al. 2015). BNF process is mediated by nitrogenase enzymatic complex (Rees and Howard 2000), which is highly regulated at transcriptional and post-translational levels in H. seropedicae (Chubatsu et al. 2012). Nitrogenase metalloenzime is composed by two structural components and the smaller one, Fe-protein, is encoded by nifH gene (Rees and Howard 2000; Dixon and Kahn 2004; Peters et al. 1995; Machado et al. 1996). It was showed that H. seropedicae nif expression occur on and inside roots, leaves and stems of maize, rice, wheat and sorghum plants and nif gene expression in colonized plants suggests that infected tissues provide a suitable environment for BNF (Roncato-Maccari et al. 2003).
In bacteria that interact with eukaryotic hosts, the T3SS is the most common structural apparatus used to translocate effector proteins into the host cytoplasm (He et al. 2004). Analyses of the H. seropedicae SmR1 genome have indicated the presence of genes homologous to the T3SS, suggesting that it is present in this bacterium, and raising the possibility that it may be involved in its interaction with host plants (Monteiro et al. 2012). T3SS is one of the pathways by which some secreted proteins are exported across the inner and outer membranes in Gram-negative bacteria and its machinery is termed injectiosome. Therefore, injectiosome function is to deliver effectors across the bacteria and host membranes into the cytosol of host cells, where they may modulate a large variety of host cell functions, including immune and defense responses (Tseng et al. 2009; Desvaux et al. 2004; He et al. 2004). In a previous study, H. seropedicae dinitrogenase reductase (nif) and T3SS ATP synthase were identified by peptide mass fingerprint among 18 differentially accumulated proteins on proteome of inoculated maize roots 7 days after inoculation (DAI) (Ferrari et al. 2014). DNA-directed RNA polymerase subunit beta’ (rpoC) is a constitutive gene that has been used as a reference gene for normalization in relative gene expression by RT-qPCR in H. seropedicae pure culture in the presence of sugarcane extract in the bacterial culture medium (Cordeiro et al. 2013).
A major problem related to inoculants is the survival of inoculated bacteria in the rhizosphere and the effective plant colonization (Stets et al. 2015). Regarding bacterial DNA presence in planta, quantitative PCR (qPCR) has been the method of choice to quantify plant growth promoting bacteria because it is fast and specific. PCR and qPCR have been used to identify Azospirillum isolates from rhizosphere and soil (Shime-Hattori et al. 2011; Baudoin et al. 2010) and to monitor Azospirillum colonization in wheat and maize (Stets et al. 2015; Couillerot et al. 2010, 2013). qPCR assay was developed using SYBR Green and HERBAS1 species-specific primers to quantify H. seropedicae DNA in inoculated maize roots (Pereira et al. 2014), however a hydrolysis probe assay present higher specificity comparing to a SYBR Green assay.
Herein we developed a hydrolysis probe qPCR tool to quantify H. seropedicae DNA in planta and RT-qPCR assays to measure H. seropedicae transcript levels in plant tissues to evaluate gene expression of a constitutive gene, DNA-directed RNA polymerase subunit beta’ (rpoC), and two regulated genes, related to injectiosome apparatus T3SS (hrcN) and nitrogen fixation (nifH). Our aim was to develop new tools to address the abundance of H. seropedicae SmR1 and its expression (nifH, rpoC and hrcN) in planta in order to monitor the effective plant colonization by this bacterium in two maize growth conditions.
Materials and methods
Bacterial growth and experimental conditions
Herbaspirillum seropedicae strain SmR1, a spontaneous streptomycin resistant mutant of strain Z78 (ATCC 35893), was routinely grown in orbital shaker (120 rpm) at 30 °C in 30 mL NFbHPN medium supplemented with 5 mg L−1 malic acid (Klassen et al. 1997). The bacterial strains used for qPCR probe specificity assay were grown as previously described (Pereira et al. 2014). The optical density (OD) of bacterial cell cultures was measured at 600 nm using Hitachi U2910 Spectrophotometer (Tokyo, Japan). H. seropedicae SmR1 was inoculated in maize seedlings (Pioneer 30F53) cultivated in two distinct conditions:in vitro and in soil. Maize seeds were surface-sterilized in laminar flow by immersion in 70% ethanol for 5 min, followed by submersion in 2% sodium hypochlorite plus 2.5% Tween-20 solution for 30 min. Seeds were then rinsed 3 times with sterile distilled water, germinated on plates containing water-agar 0.8% (in vitro) or water moist filter paper (in soil) and stored at 25 °C in a dark chamber for 3 days. Three days old seedlings were inoculated with washed H. seropedicae SmR1 culture (OD600 = 1; 108 cells of H. seropedicae SmR1 mL−1) after dilution to 105 cells mL−1 (in vitro) or 107 cells mL−1 (in soil) in sterile NFb malate medium without nitrogen source, for 30 min with constant agitation (80 rpm) (Balsanelli et al. 2010). For seedlings grown in vitro, control seedlings were mock-inoculated under the same conditions as described above. For cultivation, seedlings were washed in autoclaved 0.9% saline buffer for 1 min and placed in the glass tubes containing plant medium solution. The seedlings were placed side-by-side in a controlled-environmental chamber with 16 h photoperiod, photosynthetic active radiation of 150 µmol m−2 s−1, 25 °C light/23 °C dark and 40% humidity. The experiment was conducted in completed randomized blocks in three biological replicates. Plants were randomly collected 4, 7 and 10 DAI. For seedling grown in soil, the mock inoculum was prepared in the same conditions as described, but using a bacterial culture boiled at 100 °C for 15 min in water bath until bacteria complete inactivation (Pankievicz et al. 2015). After inoculation, the seedlings were placed in 2 L pots containing 1.5 kg soil (Cerro Negro, Santa Catarina, Brazil, 27°47′43″S, 50°52′33″W) previously supplemented with 100 mL nutrient solution (Rodriguez-Salazar et al. 2009) with reduced nitrogen (0.5 mM KNO3). Plants were grown in greenhouse under natural light without temperature control (average temperature about 29 °C), watered daily and every 2 days was added 100 mL of nutrient solution per pot. The experiment was conducted in completed randomized blocks in three biological replicates. Plants were randomly collected 14, 21 and 28 DAI. Plant tissues were immediately frozen in liquid nitrogen and stored at −80 °C until analysis.
Herbaspirillum seropedicae DNA quantification
Total DNA was isolated from maize tissue using DNeasy Plant Mini Kit (Qiagen) according to the manufacturer’s protocol. Each sample was a pool of three roots or leaves, obtained by manual crushing with mortar and pestle in the presence of liquid nitrogen. DNA extraction from bacterial cultures was performed using Wizard® Genomic DNA purification kit (Promega™, Madison, WI, USA) with modifications (Pereira et al. 2014). DNA concentration was estimated from measurements at 260 and 280 nm on a Thermo Scientific NanoDrop 2000 spectrophotometer (Willmington, DE, USA). H. seropedicae DNA quantification in maize plants grown in vitro and in soil was performed by qPCR using HERBAS1 species-specific primers, targeting Bacteriophage Tail Fiber Protein (GenBank accession CP002039.1| 216,307–217,431), and SYBR Green (Pereira et al. 2014). Additionally, a TaqMan probe (HERBAS1) was designed using Primer Express 3.0 (Applied Biosystems) (Table S1). The HERBAS1 probe specificity test was performed using bacterial DNA isolated from H. seropedicae SmR1, Herbaspirillum hiltneri, Herbaspirillum huttiense, Herbaspirillum lusitanum, Herbaspirillum rubrisubalbicans, Herbaspirillum frisingense, Azospirillum brasilense, Bacillus cereus, Bacillus subtilis, Escherichia coli, Rhizobium sp., Microbacterium sp., and Pseudomonas sp. The amplification reactions contained 12.5 µL of 2× SYBR Green Master Mix or 2× TaqMan Universal PCR Master Mix (Applied Biosystems, Foster City, CA, USA), 100 nM of HERBAS1 forward and reverse primers, 100 nM HERBAS1 probe (TaqMan assay), water and 20 ng of template DNA in a final volume of 25 µL. All control samples reactions were carried out in duplicate and the inoculated samples in triplicate following the cycling conditions: an initial incubation step at 50 °C for 2 min, 95 °C incubation for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. For SYBR Green reactions, an additional dissociation curve with linear increase from 60 to 95 °C was performed immediately after each run. All real-time PCR runs were analyzed using automatic software settings. The reactions were performed in ABI PRISM 7500 Detection System (Applied Biosystems, Foster City, CA, USA). In order to quantify the genomic DNA amount in a pure culture of H. seropedicae SmR1, standard curves were prepared by serial dilution. Genomic DNA was 10-fold serially diluted in ultra-pure water to final concentrations ranging from 106 to 100 DNA copies, equivalent to concentrations of 6.03 ng to 6.03 fg. The number of genome copies (m) was calculated based on H. seropedicae SmR1 genome size (5.51 Mbp) (Pedrosa et al. 2011) as described previously (Pereira et al. 2014). A standard curve was designed for each growth condition as well as plant tissue (roots and leaves) for both SYBR Green and TaqMan assays. The qPCR reactions were carried out separately, in triplicate. Amplification efficiency values were calculated from the equation E = (10−1/S − 1) × 100, where E is the efficiency (percent) and s is the slope obtained from the standard curve.
Bacterial transcript content quantification
Total RNA was isolated from approximately 100 mg of a pool of three maize roots using TRI Reagent Protocol (Sigma-Aldrich, St. Louis, USA). RNA samples were quantified by Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Wilmington, DE, USA) and treated with DNase I Amplification Grade (Sigma-Aldrich) to eliminate DNA contamination. cDNA synthesis was obtained using High Capacity cDNA Reverse Transcription Kit (Applied Biosystems), with 2 µg of purified RNA and random primer. For absolute quantification of bacterial transcripts in roots of maize plants grown in vitro and in soil, H. seropedicae SmR1 DNA was used as standard and three standard curves were performed using three distinct H. seropedicae SmR1 DNA extracts for each bacterial target gene nifH, hrcN and rpoC (Table S2), in triplicate. Primer Express 3.0 (Applied Biosystems) software was used to design nifH and hrcN primers. Reactions for in vitro and in soil samples were performed in triplicate, with 40 ng of cDNA template and nifH, hrcN and rpoC primer pairs separately. Primer concentrations were described in Table S2. The qRT-PCR reactions were carried out in ABI PRISM 7500 Detection System (Applied Biosystems), following the amplifications conditions as described above using 2× SYBR Green Master Mix (Applied Biosystems) in a final volume of 20 µL. Standard curves (Ct versus log DNA copy number) were used to estimate nifH, rpoC and hrcN gene expression in roots of control and inoculated plants. Gene expression was expressed as nifH, rpoC and hrcN cDNA copy number per 40 ng of total cDNA template. In order to verify the interference of the maize DNA presence in amplification efficiency values, standard curves were constructed for each gene using cDNA from an inoculated maize root sample grown in vitro. The cDNA was fivefold serially diluted in ultra-pure water, with concentrations ranging from 80 to 0.3125 ng of cDNA template, in duplicate and the amplification efficiencies were determined as mentioned above.
Results
qPCR parameters and probe specificity test
Standard curves were prepared for each experimental condition (in vitro and in soil) and plant tissue (maize roots and leaves), using SYBR Green or TaqMan reagents to obtain qPCR parameters such as efficiency, slope and correlation coefficient (R2). SYBR Green assay (Table 1) showed efficiency values varying from 89 to 100%. The qPCR parameters obtained through the standard curves using TaqMan probe (Table 2) showed efficiency values ranging from 84 and 99%. Using six H. seropedicae SmR1 DNA standard curves constructed for the TaqMan assay (six different runs), the general efficiency value obtained was 91% and slope value −3.55. The limit of detection (LOD) using TaqMan assay was established as 101 copies, which corresponds to mean Ct = 32.8 (Fig. S1). Probe specificity test was conducted for TaqMan assay, which was able to detect selectively H. seropedicae SmR1 DNA (Table 3). Even though it was detected late Ct value for different bacteria species but same genera (H. hiltneri, H. huttiense, H. lusitanum, H. rubrisubalbicans and H. frisingense), H. seropedicae SmR1 presented early Ct value (10.21) related to the others (Ct > 32.30).
Quantification of H. seropedicae DNA in maize cultivated in vitro and in soil
Total DNA isolated from roots and leaves of maize plants (control and inoculated) grown in two conditions were used as template to qPCR in order to quantify H. seropedicae DNA. Considering plants grown in vitro (Table 4), inoculated samples presented around 109 H. seropedicae DNA copy number/g of fresh root tissue (4, 7 and 10 DAI) and ranged from 107 (4 DAI) to 109 (7 and 10 DAI) in leaf tissues using SYBR Green. H. seropedicae DNA contents determined by TaqMan assay were similar to SYBR Green results. As expected, bacterial DNA was not detected in control samples by both qPCR assays, either roots or leaves of seedlings grown in vitro. Considering plants grown in soil (Table 5), H. seropedicae DNA was detected using SYBR Green in 9 out of 9 inoculated root samples 14 DAI, 6 out of 9 samples 21 DAI and 3 out of 9 samples 28 DAI. On the other hand, H. seropedicae DNA was detected in only 2 out of 6 control root samples (14 and 21 DAI). H. seropedicae DNA was not detected in leaf samples. Similarly, few control root samples showed H. seropedicae DNA presence comparing to inoculated in TaqMan assay. H. seropedicae DNA was not detected in control leaf samples, while two inoculated leaf samples (14 DAI) showed positive values for DNA presence.
Transcript quantification of H. seropedicae in maize cultivated in two conditions
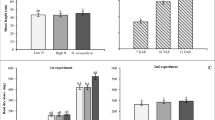
In this study, the cDNA quantity of bacterial transcripts nifH, hrcN and rpoC in maize root samples were determined by RT-qPCR using serial dilutions of DNA from H. seropedicae SmR1 pure culture as standard. Therefore, the standard curves were constructed by plotting Ct versus log DNA copy number to obtain efficiency and slope values (Fig. 1) and to estimate nifH, rpoC and hrcN gene expression in roots of control and inoculated plants. The bacterial gene expression (nifH, rpoC and hrcN) was quantified in maize root samples grown in vitro (Fig. 2) and in soil (Fig. 3) by RT-qPCR. In vitro, gene expression for nifH, rpoC and hcrN was not observed on control roots, whereas on inoculated roots, these genes were expressed 4, 7 and 10 DAI, in different patterns (Fig. 2). The nifH gene expression decreased from 4 to 10 DAI in inoculated samples. rpoC gene expression in inoculated samples slightly decreased from 4 to 10 DAI. Concerning plants grown in soil (Fig. 3), the cDNA bacterial quantification showed highest nifH bacterial gene expression in inoculated samples comparing to control. rpoC gene expression decreased in inoculated samples from 14 to 28 DAI. It was not possible to quantify hrcN gene in control or inoculated samples even 14, 21 or 28 DAI. Standard curves were also constructed for each gene using cDNA from inoculated root as template, which generate curve equations determining qPCR parameters such as efficiency and R2 (Table S3).
Discussion
Considering that there are few studies involving H. seropedicae response inside the plant, and assuming that different experimental settings would be required for a better comprehension about the plant–endophyte interaction (Hardoim et al. 2015), we developed qPCR assays to evaluate Zea mays colonization by H. seropedicae SmR1 in two growth conditions, in vitro and in soil. Herein we quantified H. seropedicae DNA in maize roots and leaves using HERBAS1 species-specific primer set previously designed for SYBR Green qPCR assay (Pereira et al. 2014). Besides, we aimed to validate a more specific tool for H. seropedicae DNA quantification, designing TaqMan HERBAS1 probe. TaqMan assay is the choice for qPCR targeting specific phytopathogens (Oliveira et al. 2002; Li et al. 2013). The labeled fluorogenic hybridization probe (TaqMan) is cleaved by 5′ nuclease activity of Taq DNA polymerase during the extension phase of PCR, yielding a real time measurable fluorescence emission directly proportional to the concentration of the target sequence (Heid et al. 1996). TaqMan probe with conjugated minor-groove-binding (MGB) groups form extremely stable duplexes with single-stranded DNA targets, allowing the design of shorter probes with high specificity of the hybridization (Kutyavin et al. 2000).
In order to establish a reliable DNA quantification, qPCR parameters for SYBR Green and TaqMan assays were defined using standard curves, once the use of a substantial standard curve based on a known DNA concentration makes it theoretically possible to quantify DNA from any source (Timmusk et al. 2009). Considering that a consistent standard curve should have a R2 value of more than 0.95 and a slope between −3.0 and −3.9 (corresponding to PCR efficiencies of 80 and 115%) (Zhang and Fang 2006), all curves presented reliable qPCR parameters (Tables 1, 2). Using a specific standard curve for each plate run, H. seropedicae DNA quantity on roots and leaves of maize grown in vitro and in soil could be properly estimated using SYBR Green and TaqMan probe assay. Aiming to verify if the TaqMan probe assay could differentiate H. seropedicae from other bacteria (even other bacteria from the same genera), we used DNA samples isolated from pure cultures of H. seropedicae and other bacteria species. Our results confirm the specificity of the HERBAS1 probe (Table 3) based on the Ct values presented by H. seropedicae (Ct = 10.21). In contrast, other species belonging to Herbaspirillum genera presented positive signals, but with late Ct values (>32.30). Furthermore, other bacteria species did not shown amplification signals. The difference of Ct values between the target and other bacteria species (at least 22.9) is more than enough to validate the use of this experimental condition, since the unspecific amplification will not influence the analysis (Martin-Sanchez et al. 2013). In a previous work using SYBR Green chemistry to detect and quantify H. seropedicae DNA in maize plants (Pereira et al. 2014), HERBAS1 primers were tested for H. seropedicae specificity, proving that this pair of primers is very specific for H. seropedicae DNA detection and quantification in planta. Comparing the unspecific Ct values using SYBR Green (25.59 for H. rubrisubalbicans and 29.62 for H. frisingense) with the unspecific Ct values using TaqMan probe (32.68 for H. rubrisubalbicans and 32.3 for H. frisingense), the unspecific amplification was reduced using TaqMan assay as expected. Taken together, these results showed that H. seropedicae DNA can be properly detected and quantified by SYBR Green assay or alternatively using more specific TaqMan assay.
The sensitivity of qPCR quantification experiments is given by the LOD, which is defined as the lowest DNA concentration in a sample that can be reliable detected. In qPCR assays, it is accepted that the LOD is the DNA amount at which 95% of the positive samples are detected (or no more than 5% failed reactions should occur) (Bustin et al. 2009). For the SYBR Green and TaqMan assays, the amplifications below the LOD were discarded, while those above the LOD were accounted and Ct values were used to calculate the mean value of the bacterial DNA copy number/g fresh tissue (Tables 4, 5). For H. seropedicae DNA quantification in maize roots and leaves using SYBR Green assay, the LOD was previously established as 101 genome copies, corresponding to a mean Ct value of 32.15 (Pereira et al. 2014). Using TaqMan probe assay, LOD also corresponded to 101 genome copies (Ct 32.8), permitting a reliable DNA quantification from 106 to 101 genome copies, similar to SYBR Green assay. Despite the difficulties in comparing LOD values between studies due to the differences in experimental designs, such as number of replicates, differences in master mix constituents, or conditions of DNA quantification (Price et al. 2012), the LOD found in the present study (101 genome copy number, corresponding to 60.3 fg) is in accordance with other studies using TaqMan PCR assays to detect and quantify microbial genomic DNA. For example, the genomic DNA of the pathogenic fungus Magnaporthe oryzae in rice, could be detected as low as 1 pg (Su’udi et al. 2013), as well as to detect Paenibacillus polymyxa DNA in wild barley (Timmusk et al. 2009). Also, for Bacillus subtilis and Pseudomonas fluorescens detection in soil, it was set a LOD of 10 pg (Kim et al. 2010).
Using the established set of primers HERBAS1 and HERBAS1 probe, the present study quantified H. seropedicae presence on roots and leaves of maize (P30F53 variety) cultivated in vitro and in soil employing SYBR Green and TaqMan chemicals in qPCR assays. There is a single copy of the HERBAS1 target sequence in H. seropedicae genome (Pedrosa et al. 2011), which allows us to estimate the bacterial cell number in maize tissues (Pereira et al. 2014). H. seropedicae DNA quantification in plants cultivated in vitro, using both SYBR Green and TaqMan qPCR assays (Table 4), indicates an increase or at least maintenance of the genome copy number per gram of root in inoculated maize roots and leaves along the time course (4, 7 and 10 DAI). Our results are consistent with those found previously in maize root plants (DKB240 variety) grown in the same conditions from 1 to 10 DAI (Pereira et al. 2014). Under laboratory conditions, H. seropedicae invade and colonize very quickly maize tissues, and 24 h after inoculation, roots (including xylem vessels) are completely colonized by the bacteria. The plant vascular system colonization allows the bacteria distribution throughout the host, reaching aerial parts (Monteiro et al. 2008, 2012). Our results strengthen these findings, once H. seropedicae DNA was detected in both belowground and aboveground tissues. Likewise qPCR assays were performed to quantify A. brasilense FP2 on roots of maize (Faleiro et al. 2013) and wheat (Stets et al. 2015). For both maize P30F53 and DKB240 varieties, A. brasilense DNA copy number per gram of root increased from 106 to 109 (Faleiro et al. 2013), confirming the possibility to detect and quantify diazotrophic bacteria in maize tissue and the rapid colonization of the plant tissue by plant growth promoting bacteria. Regarding to H. seropedicae DNA quantification of plants grown in nonsterile soil (Table 5), inoculated plants usually presented higher rates of H. seropedicae presence, comparing to control plants. Inoculated root samples showed bacteria presence about 105 DNA copy number per gram of fresh root for SYBR Green and TaqMan assays at 14, 21 and 28 DAI. Otherwise, H. seropedicae DNA was only detected in 2 out of 27 leaf samples of inoculated plants. Even if it is not possible to directly compare different analyzed times after inoculation among the different substrates, it is important to note that under axenic conditions there is absence of competing bacteria and the introduction of competition may substantially change the pattern of the plant colonization observed. Regarding that bacteria (even benefic or pathogenic) penetrate the plant tissue by the same attachment points, competing bacteria may secrete anti-bacterial substances and the plant may elaborate a defense against the mixed bacterial population (Monteiro et al. 2008). H. seropedicae attach and colonize maize root surface progressively from 3 to 15 days after inoculation (Roncato-Maccari et al. 2003). In contrast, a longer time study, which examined the population dynamics of H. seropedicae strain HRC 54 associated with maize roots, showed a continuous decrease in number of bacterial cells per gram of root (fresh weight) from 7 to 14, 21 and 30 DAI (Silva et al. 2014). Similar to our soil experiment, in which H. seropedicae DNA presence was lower 28 DAI (3 positive/9 total) comparing to 14 and 21 DAI. The same study also detected H. seropedicae HRC 54 presence in control plants, but fewer than in inoculated, as our results from plants grown in soil.
It is noteworthy that H. seropedicae DNA quantification stands in the higher bacteria genome copy number found on roots comparing to leaves in maize plants in both growth conditions. This may be a common behavior in diazotrophic bacteria, once other studies also found a greater bacteria presence in rhizosphere of grasses comparing to aerial parts (Rodriguez-Blanco et al. 2015; Pariona-Llanos et al. 2010). In most plants, the endophytes number in belowground is higher comparing to aboveground tissues (Rosenblueth and Martinez-Romero 2006). Higher Herbaspirillum spp. amounts were also observed on roots than stems in sugarcane field experiment (Reis et al. 2000). Additionally, Bacillus megaterium, a Gram-positive nitrogen-fixing bacteria which has similar colonization pattern to some Gram-negative endophytic diazotrophs, such as A. brasilense, migrates slowly from roots to leaves (Liu et al. 2006). This might explain the lower amount of bacteria found in shoots in these studies comparing to roots.
Bacterial gene expression (nifH, rpoC and hrcN) was analyzed on maize roots by RT-qPCR absolute quantification using standard curves performed with serial dilutions of H. seropedicae SmR1 DNA. Similar approach has already been employed in study of environmental diazotrophic diversity using nifH gene expression as a molecular marker (Martensson et al. 2009). The accuracy of the bacterial transcript quantification assay on roots of maize inoculated with H. seropedicae SmR1 in the present study was verified by qPCR standard curves parameters using bacterial DNA as template. According to Zhang and Fang (2006) the efficiencies between 80 and 115% are consistent with reliable standard curves. In this way, the efficiency values found in this study for nifH (105%), rpoC (103%) and hrcN (97%) bacterial genes (Fig. 1) are in accordance with the established parameters and hence the bacterial absolute transcript quantification could be properly measured. In this study, H. seropedicae nifH, rpoC and hrcN transcripts were quantified in maize roots grown in two growth conditions. We were able to detect nifH gene expression on inoculated maize roots, cultivated in two growth conditions. The highest amounts of nifH transcripts was found in inoculated samples grown in vitro, remaining about 103 bacterial cDNA copy number at 4, 7 and 10 DAI, although lower amounts were observed in soil plants. Our results are in accordance with previous studies showing that genera Herbaspirillum were able to express nif genes and presented nitrogenase activity when associated with other Poaceae, such as sorghum, rice and sugarcane (James et al. 1997, 2002; Olivares et al. 1997; Gyaneshwar et al. 2002; Elbeltagy et al. 2001).
Inoculated maize samples cultivated in axenic conditions (in vitro) had a relatively constant rpoC gene expression, between 5.7 and 4.2 × 103 cDNA copy number per 40 ng of total cDNA. Our results suggest that rpoC transcript level is a possible tool to monitor H. seropedicae activity in planta. rpoC gene expression was tested as a reference gene for Gluconacetobacter diazotrophicus grown in different carbon sources, however it was highly variable (Galisa et al. 2012). In contrast, rpoC presented constant expression for H. seropedicae culture in the presence of sugarcane extract, and it has been used as a reference gene for normalization of relative gene expression by RT-qPCR (Cordeiro et al. 2013). However our report is the first that successfully measure rpoC transcript levels in planta. Herein, transcripts of hrcN gene, coding for T3SS ATP synthase, was also measured. In vitro inoculated roots presented higher rates of hrcN transcripts at 10 DAI, comparing to 4 and 7 DAI. In contrast, hrcN transcript was not detected on roots in soil experiment at 14, 21 or 28 DAI. It was previously suggested that specific physiological conditions might be required for expression and activity of T3SS and synthesis of effector proteins in H. seropedicae (Pedrosa et al. 2011; Monteiro et al. 2012). It is possible that the necessary conditions were achieved in vitro plant growth condition 4, 7 and 10 DAI, but not in soil 14, 21 and 28 DAI. Despite transcriptomic analysis of in vitro H. seropedicae and maize interaction did not reveal T3SS gene expression (Balsanelli et al. 2015), T3SS was identified by mass spectrometry when investigating the root proteome of H. seropedicae and maize interaction 7 DAI (Ferrari et al. 2014). We observed that H. seropedicae DNA copy number slightly increased in roots and leaves of plants grown in vitro from 4 to 10 DAI, while nifH transcript level decreased over time in roots, but the same was not observed for rpoC or hrcN transcript levels, suggesting that nifH expression was regulated in the beginning of this plant–bacteria interaction.
In conclusion, we quantified H. seropedicae DNA presence on roots and leaves of maize plants (P30F53 variety) inoculated with diazotrophic bacteria H. seropedicae, and also measured the transcripts levels of bacterial genes on maize roots of plants grown in sterile and nonsterile conditions. We developed a specific, reliable and efficient TaqMan probe for qPCR assay, to quantify H. seropedicae DNA in plant tissues. H. seropedicae nifH and rpoC gene expression were detected on roots of inoculated maize grown in both growth conditions, however it was not possible to verify hrcN gene expression in plants grown in soil. Our results suggest that it is possible to use nifH and rpoC transcript levels to monitor H. seropedicae gene expression inside plant tissue.
References
Alves GC, Videira SS, Urquiaga S, Reis VM (2015) Differential plant growth promotion and nitrogen fixation in two genotypes of maize by several Herbaspirillum inoculants. Plant Soil 387(1–2):307–321. doi:10.1007/s11104-014-2295-2
Amaral FP, Pankievicz VCS, Arisi ACM, Souza EM, Pedrosa F, Stacey G (2016) Differential growth responses of Brachypodium distachyon genotypes to inoculation with plant growth promoting rhizobacteria. Plant Mol Biol 90(6):689–697. doi:10.1007/s11103-016-0449-8
Babalola OO (2010) Beneficial bacteria of agricultural importance. Biotechnol Lett 32(11):1559–1570. doi:10.1007/s10529-010-0347-0
Baldani JI, Baldani VLD, Seldin L, Dobereiner J (1986) Characterization of Herbaspirillum seropedicae Gen-Nov, Sp-Nov, a root-associated nitrogen-fixing bacterium. Int J Syst Bacteriol 36(1):86–93
Balsanelli E, Serrato RV, de Baura VA, Sassaki G, Yates MG, Rigo LU, Pedrosa FO, de Souza EM, Monteiro RA (2010) Herbaspirillum seropedicae rfbB and rfbC genes are required for maize colonization. Environ Microbiol 12(8):2233–2244. doi:10.1111/j.1462-2920.2010.02187.x
Balsanelli E, Tadra-Sfeir MZ, Faoro H, Pankievicz VCS, de Baura VA, Pedrosa FO, de Souza EM, Dixon R, Monteiro RA (2015) Molecular adaptations of Herbaspirillum seropedicae during colonization of the maize rhizosphere. Environ Microbiol. doi:10.1111/1462-2920.12887
Baudoin E, Couillerot O, Spaepen S, Moenne-Loccoz Y, Nazaret S (2010) Applicability of the 16S–23S rDNA internal spacer for PCR detection of the phytostimulatory PGPR inoculant Azospirillum lipoferum CRT1 in field soil. J Appl Microbiol 108(1):25–38. doi:10.1111/j.1365-2672.2009.04393.x
Berg G (2009) Plant–microbe interactions promoting plant growth and health: perspectives for controlled use of microorganisms in agriculture. Appl Microbiol Biotechnol 84(1):11–18. doi:10.1007/s00253-009-2092-7
Bhattacharjee RB, Singh A, Mukhopadhyay SN (2008) Use of nitrogen-fixing bacteria as biofertiliser for non-legumes: prospects and challenges. Appl Microbiol Biotechnol 80(2):199–209. doi:10.1007/s00253-008-1567-2
Brusamarello-Santos LCC, Pacheco F, Aljanabi SMM, Monteiro RA, Cruz LM, Baura VA, Pedrosa FO, Souza EM, Wassem R (2012) Differential gene expression of rice roots inoculated with the diazotroph Herbaspirillum seropedicae. Plant Soil 356(1–2):113–125. doi:10.1007/s11104-011-1044-z
Bustin S, Benes V, Garson J, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl M, Shipley G, Vandesompele J, Wittwer C (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55(4):611–622. doi:10.1373/clinchem.2008.112797
Carvalho TLG, Balsemao-Pires E, Saraiva RM, Ferreira PCG, Hemerly AS (2014) Nitrogen signalling in plant interactions with associative and endophytic diazotrophic bacteria. J Exp Bot 65(19):5631–5642. doi:10.1093/jxb/eru319
Chebotar VK, Malfanova NV, Shcherbakov AV, Ahtemova GA, Borisov AY, Lugtenberg B, Tikhonovich IA (2015) Endophytic bacteria in microbial preparations that improve plant development (review). Appl Biochem Microbiol 51(3):271–277. doi:10.1134/s0003683815030059
Chubatsu LS, Monteiro RA, de Souza EM, de Oliveira MAS, Yates MG, Wassem R, Bonatto AC, Huergo LF, Steffens MBR, Rigo LU, Pedrosa FD (2012) Nitrogen fixation control in Herbaspirillum seropedicae. Plant Soil 356(1–2):197–207. doi:10.1007/s11104-011-0819-6
Cordeiro FA, Tadra-Sfeir MZ, Huergo LF, Pedrosa FD, Monteiro RA, de Souza EM (2013) Proteomic analysis of Herbaspirillum seropedicae cultivated in the presence of sugar cane extract. J Proteome Res 12(3):1142–1150. doi:10.1021/pr300746j
Couillerot O, Bouffaud M-L, Baudoin E, Muller D, Caballero-Mellado J, Moenne-Loccoz Y (2010) Development of a real-time PCR method to quantify the PGPR strain Azospirillum lipoferum CRT1 on maize seedlings. Soil Biol Biochem 42(12):2298–2305. doi:10.1016/j.soilbio.2010.09.003
Couillerot O, Ramirez-Trujillo A, Walker V, von Felten A, Jansa J, Maurhofer M, Defago G, Prigent-Combaret C, Comte G, Caballero-Mellado J, Moenne-Loccoz Y (2013) Comparison of prominent Azospirillum strains in Azospirillum–Pseudomonas–Glomus consortia for promotion of maize growth. Appl Microbiol Biotechnol 97(10):4639–4649. doi:10.1007/s00253-012-4249-z
da Silva Lima L, Olivares F, de Rodrigues Oliveira R, Vega M, Aguiar N, Canellas L (2014) Root exudate profiling of maize seedlings inoculated with Herbaspirillum seropedicae and humic acids. Chem Biol Technol Agric C7–C231(1):1–18. doi:10.1186/s40538-014-0023-z
Desvaux M, Parham NJ, Scott-Tucker A, Henderson IR (2004) The general secretory pathway: a general misnomer? Trends Microbiol 12(7):306–309. doi:10.1016/j.tim.2004.05.002
Dixon R, Kahn D (2004) Genetic regulation of biological nitrogen fixation. Nat Rev Microbiol 2(8):621–631. doi:10.1038/nrmicro954
dos Reis FB Jr, Reis VM, Urquiaga S, Dobereiner J (2000) Influence of nitrogen fertilisation on the population of diazotrophic bacteria Herbaspirillum spp. and Acetobacter diazotrophicus in sugar cane (Saccharum spp.). Plant Soil 219(1–2):153–159
Elbeltagy A, Nishioka K, Sato T, Suzuki H, Ye B, Hamada T, Isawa T, Mitsui H, Minamisawa K (2001) Endophytic colonization and in planta nitrogen fixation by a Herbaspirillum sp. isolated from wild rice species. Appl Environ Microbiol 67(11):5285–5293. doi:10.1128/aem.67.11.5285-5293.2001
Faleiro AC, Pereira TP, Espindula E, Brod FCA, Arisi ACM (2013) Real time PCR detection targeting nifA gene of plant growth promoting bacteria Azospirillum brasilense strain FP2 in maize roots. Symbiosis 61(3):125–133. doi:10.1007/s13199-013-0262-y
Ferrari CS, Amaral FP, Ferreira Bueno JC, Scariot MC, Valentim-Neto PA, Maisonnave Arisi AC (2014) Expressed proteins of Herbaspirillum seropedicae in maize (DKB240) roots–bacteria interaction revealed using proteomics. Appl Biochem Biotechnol 174(6):2267–2277. doi:10.1007/s12010-014-1197-3
Galisa PS, da Silva HAP, Macedo AVM, Reis VM, Vidal MS, Baldani JI, Simoes-Araujo JL (2012) Identification and validation of reference genes to study the gene expression in Gluconacetobacter diazotrophicus grown in different carbon sources using RT-qPCR. J Microbiol Methods 91(1):1–7. doi:10.1016/j.mimet.2012.07.005
Germida JJ, Siciliano SD, de Freitas JR, Seib AM (1998) Diversity of root-associated bacteria associated with held-grown canola (Brassica napus L.) and wheat (Triticum aestivum L.). FEMS Microbiol Ecol 26(1):43–50. doi:10.1111/j.1574-6941.1998.tb01560.x
Gyaneshwar P, James EK, Reddy PM, Ladha JK (2002) Herbaspirillum colonization increases growth and nitrogen accumulation in aluminium-tolerant rice varieties. New Phytol 154(1):131–145. doi:10.1046/j.1469-8137.2002.00371.x
Hardoim PR, van Overbeek LS, Berg G, Pirttilä AM, Compant S, Campisano A, Döring M, Sessitsch A (2015) The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79(3):293–320. doi:10.1128/MMBR.00050-14
He SY, Nomura K, Whittam TS (2004) Type III protein secretion mechanism in mammalian and plant pathogens. Biochim Biophys Acta 1694(1–3):181–206. doi:10.1016/j.bbamcr.2004.03.011
Heid CA, Stevens J, Livak KJ, Williams PM (1996) Real time quantitative PCR. Genome Res 6(10):986–994. doi:10.1101/gr.6.10.986
James EK, Olivares FL, Baldani JI, Dobereiner J (1997) Herbaspirillum, an endophytic diazotroph colonizing vascular tissue in leaves of Sorghum bicolor L. Moench. J Exp Bot 48(308):785–797. doi:10.1093/jxb/48.3.785
James EK, Gyaneshwar P, Mathan N, Barraquio QL, Reddy PM, Iannetta PPM, Olivares FL, Ladha JK (2002) Infection and colonization of rice seedlings by the plant growth-promoting bacterium Herbaspirillum seropedicae Z67. Mol Plant Microbe Interact 15(9):894–906. doi:10.1094/mpmi.2002.15.9.894
Kim SE, Moon JS, Choi WS, Lee EN, Lee SH, Kim SU (2010) Qualitative and quantitative detection of agricultural microorganisms expressing iturin and mop cyclase in soils. J Agric Food Chem 58(24):12657–12663. doi:10.1021/jf103216g
Klassen G, Pedrosa FO, Souza EM, Funayama S, Rigo LU (1997) Effect of nitrogen compounds on nitrogenase activity in Herbaspirillum seropedicae SMR1. Can J Microbiol 43(9):887–891
Kutyavin IV, Afonina IA, Mills A, Gorn VV, Lukhtanov EA, Belousov ES, Singer MJ, Walburger DK, Lokhov SG, Gall AA, Dempcy R, Reed MW, Meyer RB, Hedgpeth J (2000) 3′-minor groove binder-DNA probes increase sequence specificity at PCR extension temperatures. Nucleic Acids Res 28(2):655–661. doi:10.1093/nar/28.2.655
Li W, Teixeira DC, Hartung JS, Huang Q, Duan Y, Zhou L, Chen J, Lin H, Lopes S, Juliano Ayres A, Levy L (2013) Development and systematic validation of qPCR assays for rapid and reliable differentiation of Xylella fastidiosa strains causing citrus variegated chlorosis. J Microbiol Methods 92(1):79–89. doi:10.1016/j.mimet.2012.10.008
Liu XM, Zhao HX, Chen SF (2006) Colonization of maize and rice plants by strain Bacillus megaterium C4. Curr Microbiol 52(3):186–190. doi:10.1007/s00284-005-0162-3
Machado IMP, Yates MG, Machado HB, Souza EM, Pedrosa FO (1996) Cloning and sequencing of the nitrogenase structural genes nifHDK of Herbaspirillum seropedicae. Braz J Med Biol Res 29(12):1599–1602
Martensson L, Diez B, Wartiainen I, Zheng WW, El-Shehawy R, Rasmussen U (2009) Diazotrophic diversity, nifH gene expression and nitrogenase activity in a rice paddy field in Fujian, China. Plant Soil 325(1–2):207–218. doi:10.1007/s11104-009-9970-8
Martin-Sanchez PM, Bastian F, Alabouvette C, Saiz-Jimenez C (2013) Real-time PCR detection of Ochroconis lascauxensis involved in the formation of black stains in the Lascaux Cave, France. Sci Total Environ 443:478–484. doi:10.1016/j.scitotenv.2012.11.026
Monteiro RA, Schmidt MA, de Baura VA, Balsanelli E, Wassem R, Yates MG, Randi MAF, Pedrosa FO, de Souza EM (2008) Early colonization pattern of maize (Zea mays L. Poales, Poaceae) roots by Herbaspirillum seropedicae (Burkholderiales, Oxalobacteraceae). Genet Mol Biol 31(4):932–937. doi:10.1590/s1415-47572008005000007
Monteiro RA, Balsanelli E, Wassem R, Marin AM, Brusamarello-Santos LCC, Schmidt MA, Tadra-Sfeir MZ, Pankievicz VCS, Cruz LM, Chubatsu LS, Pedrosa FO, Souza EM (2012) Herbaspirillum–plant interactions: microscopical, histological and molecular aspects. Plant Soil 356(1–2):175–196. doi:10.1007/s11104-012-1125-7
Olivares FL, Baldani VLD, Reis VM, Baldani JI, Dobereiner J (1996) Occurrence of the endophytic diazotrophs Herbaspirillum spp. in roots, stems, and leaves, predominantly of Gramineae. Biol Fertil Soils 21(3):197–200
Olivares FL, James EK, Baldani JI, Dobereiner J (1997) Infection of mottled stripe disease-susceptible and resistant sugar cane varieties by the endophytic diazotroph Herbaspirillum. New Phytol 135(4):723–737. doi:10.1046/j.1469-8137.1997.00684.x
Oliveira AC, Vallim MA, Semighini CP, Araujo WL, Goldman GH, Machado MA (2002) Quantification of Xylella fastidiosa from citrus trees by real-time polymerase chain reaction assay. Phytopathology 92(10):1048–1054. doi:10.1094/phyto.2002.92.10.1048
Pankievicz VCS, do Amaral FP, Santos K, Agtuca B, Xu YW, Schueller MJ, Arisi ACM, Steffens MBR, de Souza EM, Pedrosa FO, Stacey G, Ferrieri RA (2015) Robust biological nitrogen fixation in a model grass-bacterial association. Plant J 81(6):907–919. doi:10.1111/tpj.12777
Pariona-Llanos R, Ferrara FID, Gonzales HHS, Barbosa HR (2010) Influence of organic fertilization on the number of culturable diazotrophic endophytic bacteria isolated from sugarcane. Eur J Soil Biol 46(6):387–393. doi:10.1016/j.ejsobi.2010.08.003
Pedrosa FO, Monteiro RA, Wassem R, Cruz LM, Ayub RA, Colauto NB, Fernandez MA, Fungaro MHP, Grisard EC, Hungria M, Madeira HMF, Nodari RO, Osaku CA, Petzl-Erler ML, Terenzi H, Vieira LGE, Steffens MBR, Weiss VA, Pereira LFP, Almeida MIM, Alves LR, Marin A, Araujo LM, Balsanelli E, Baura VA, Chubatsu LS, Faoro H, Favetti A, Friedermann G, Glienke C, Karp S, Kava-Cordeiro V, Raittz RT, Ramos HJO, Ribeiro E, Rigo LU, Rocha SN, Schwab S, Silva AG, Souza EM, Tadra-Sfeir MZ, Torres RA, Dabul ANG, Soares MAM, Gasques LS, Gimenes CCT, Valle JS, Ciferri RR, Correa LC, Murace NK, Pamphile JA, Patussi EV, Prioli AJ, Prioli SMA, Rocha C, Arantes OMN, Furlaneto MC, Godoy LP, Oliveira CEC, Satori D, Vilas-Boas LA, Watanabe MAE, Dambros BP, Guerra MP, Mathioni SM, Santos KL, Steindel M, Vernal J, Barcellos FG, Campo RJ, Chueire LMO, Nicolas MF, Pereira-Ferrari L, Silva JLD, Gioppo NMR, Margarido VP, Menck-Soares MA, Pinto FGS, Simao RDG, Takahashi EK, Yates MG (2011) Genome of Herbaspirillum seropedicae Strain SmR1, a specialized diazotrophic endophyte of tropical grasses. Plos Genetics 7(5):10. doi:10.1371/journal.pgen.1002064
Pereira TP, do Amaral FP, Dall’Asta P, Angonesi Brod FC, Maisonnave Arisi AC (2014) Real-time PCR quantification of the plant growth promoting bacteria Herbaspirillum seropedicae strain SmR1 in maize roots. Mol Biotechnol 56(7):660–670. doi:10.1007/s12033-014-9742-4
Peters JW, Fisher K, Dean DR (1995) Nitrogenase structure and function: a biochemical-genetic perspective. Annu Rev Microbiol 49:335–366
Price EP, Dale JL, Cook JM, Sarovich DS, Seymour ML, Ginther JL, Kaufman EL, Beckstrom-Sternberg SM, Mayo M, Kaestli M, Glass MB, Gee JE, Wuthiekanun V, Warner JM, Baker A, Foster JT, Tan P, Tuanyok A, Limmathurotsakul D, Peacock SJ, Currie BJ, Wagner DM, Keim P, Pearson T (2012) Development and validation of Burkholderia pseudomallei-specific real-time PCR assays for clinical, environmental or forensic detection applications. PLoS ONE 7(5):9. doi:10.1371/journal.pone.0037723
Rees DC, Howard JB (2000) Nitrogenase: standing at the crossroads. Curr Opin Chem Biol 4(5):559–566. doi:10.1016/s1367-5931(00)00132-0
Rodriguez-Blanco A, Sicardi M, Frioni L (2015) Plant genotype and nitrogen fertilization effects on abundance and diversity of diazotrophic bacteria associated with maize (Zea mays L.). Biol Fertil Soils 51(3):391–402. doi:10.1007/s00374-014-0986-8
Rodriguez-Salazar J, Suarez R, Caballero-Mellado J, Iturriaga G (2009) Trehalose accumulation in Azospirillum brasilense improves drought tolerance and biomass in maize plants. FEMS Microbiol Lett 296(1):52–59. doi:10.1111/j.1574-6968.2009.01614.x
Roncato-Maccari LDB, Ramos HJO, Pedrosa FO, Alquini Y, Chubatsu LS, Yates MG, Rigo LU, Steffens MBR, Souza EM (2003) Endophytic Herbaspirillum seropedicae expresses nif genes in gramineous plants. FEMS Microbiol Ecol 45(1):39–47. doi:10.1016/s0168-6496(03)00108-9
Rosenblueth M, Martinez-Romero E (2006) Bacterial endophytes and their interactions with hosts. Mol Plant Microbe Interact 19(8):827–837. doi:10.1094/mpmi-19-0827
Shime-Hattori A, Kobayashi S, Ikeda S, Asano R, Shime H, Shinano T (2011) A rapid and simple PCR method for identifying isolates of the genus Azospirillum within populations of rhizosphere bacteria. J Appl Microbiol 111(4):915–924. doi:10.1111/j.1365-2672.2011.05115.x
Stets MI, Pinto AS, Huergo LF, de Souza EM, Guimaraes VF, Alves AC, Steffens MBR, Monteiro RA, Pedrosa FD, Cruz LM (2013) Rapid identification of bacterial isolates from wheat roots by high resolution whole cell MALDI-TOF MS analysis. J Biotechnol 165(3–4):167–174. doi:10.1016/j.jbiotec.2013.04.001
Stets MI, Alqueres SMC, Souza EM, Pedrosa FD, Schmid M, Hartmann A, Cruz LM (2015) Quantification of Azospirillum brasilense FP2 bacteria in wheat roots by strain-specific quantitative PCR. Appl Environ Microbiol 81(19):6700–6709. doi:10.1128/aem.01351-15
Su’udi M, Kim J, Park JM, Bae SC, Kim D, Kim YH, Ahn IP (2013) Quantification of rice blast disease progressions through Taqman real-time PCR. Mol Biotechnol 55(1):43–48. doi:10.1007/s12033-012-9632-6
Tadra-Sfeir MZ, Faoro H, Camilios-Neto D, Brusamarello-Santos L, Balsanelli E, Weiss V, Baura VA, Wassem R, Cruz LM, Pedrosa FD, Souza EM, Monteiro RA (2015) Genome wide transcriptional profiling of Herbaspirillum seropedicae SmR1 grown in the presence of naringenin. Front Microbiol 6:8. doi:10.3389/fmicb.2015.00491
Timmusk S, Paalme V, Lagercrantz U, Nevo E (2009) Detection and quantification of Paenibacillus polymyxa in the rhizosphere of wild barley (Hordeum spontaneum) with real-time PCR. J Appl Microbiol 107(3):736–745. doi:10.1111/j.1365-2672.2009.04265.x
Tseng TT, Tyler BM, Setubal JC (2009) Protein secretion systems in bacterial-host associations, and their description in the gene ontology. BMC Microbiol 9:9. doi:10.1186/1471-2180-9-s1-s2
Vargas L, Gurjao de Carvalho TL, Gomes Ferreira PC, Divan Baldani VL, Baldani JI, Hemerly AS (2012) Early responses of rice (Oryza sativa L.) seedlings to inoculation with beneficial diazotrophic bacteria are dependent on plant and bacterial genotypes. Plant Soil 356(1–2):127–137. doi:10.1007/s11104-012-1274-8
Vargas L, Brigida ABS, Mota JP, de Carvalho TG, Rojas CA, Vaneechoutte D, Van Bel M, Farrinelli L, Ferreira PCG, Vandepoele K, Hemerly AS (2014) Drought tolerance conferred to sugarcane by association with Gluconacetobacter diazotrophicus: a transcriptomic view of hormone pathways. PLoS ONE 9(12):37. doi:10.1371/journal.pone.0114744
Zhang T, Fang HHP (2006) Applications of real-time polymerase chain reaction for quantification of microorganisms in environmental samples. Appl Microbiol Biotechnol 70(3):281–289. doi:10.1007/s00253-006-0333-6
Acknowledgements
This work was financially supported by the National Institute of Science and Technology-Biological Nitrogen Fixation (INCT-FBN), National Counsel of Technological and Scientific Development (CNPq), Ministry of Science and Technology, Brazil. Pâmela Dall Asta, Tomás Pellizzaro Pereira and Fernanda Plucani Amaral were recipients of PhD fellowships from Coordination of Personnel Improvement of Higher Education (CAPES), Ministry of Education, Brazil and Ana Carolina Maisonnave Arisi is recipient of research fellowship (PQ2) from CNPq.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest declared.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dall’Asta, P., Pereira, T.P., do Amaral, F.P. et al. Tools to evaluate Herbaspirillum seropedicae abundance and nifH and rpoC expression in inoculated maize seedlings grown in vitro and in soil. Plant Growth Regul 83, 397–408 (2017). https://doi.org/10.1007/s10725-017-0306-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-017-0306-z