Abstract
Annona deceptrix is an endemic species found on the Ecuadorian coast, and its vulnerable status is attributed to the degradation of its ecological niches caused by human activities. This species exhibits desirable agronomic traits with potential use for genetic improvement. The purpose of this study was to unveil the genetic diversity and population structure of 106 individuals from 11 natural populations of A. deceptrix in the province of Manabí using 18 specific simple sequence repeat (SSR) markers. Overall, a moderate level of genetic diversity (He = 0.445) was found. The AD8, AD5, AD6, AD10 and AD1 markers were the most informative for A. deceptrix. Analysis of molecular variance (AMOVA) revealed that 65% of the total genetic diversity was within individuals and 11% of genetic variation was attributable to the differentiation among populations. Low fixation index (Fst = 0) and moderate gene flow (Nm = 1.159), thereby indicating no genetic differentiation among populations. In addition, a null correlation between geographic and genetic distances among populations (R2 = 0.007; p = 0.02) was detected by the Mantel test, indicating no significant isolation by distance. These findings are novel and have a great impact to begin planning conservation strategies for A. deceptrix considering that it is an endemic species, threatened by anthropogenic activity and has received very little research attention. In this sense, we suggest that the strategies should focus on populations such as Humedad, Agua Blanca, and Tachina, since they were those with the greatest diversity and number of private alleles.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Annonaceae is one of the most diverse families of tropical forest species containing between 110 and 200 genera and from 2300 to 2500 species (Leal and Paull 2023). This family comprises woody trees, shrubs and vines which are originally from tropical lowland Central and South America and the West Indies (Cautín and Agustí 2005). Within the Annonaceae family, the Annona genus presents species of commercial interest such as A. cherimola Mill. (cherimoya), A. glabra L. (pond apple), A. muricata L. (soursop), A. reticulata L. (custard apple), atemoya (a hybrid of A. squamosa and A. cherimola) and A. squamosa L. (sweetsop or sugar apple), which are the main species cultivated due to their edible fruits (Leal and Paull 2023). Additionally, species of this genus has excellent nutritional and pharmacological properties for human health through direct consumption as fresh fruit or in processed foods (Al Kazman et al. 2022). However, a large number of threatened Annona species have not been genetically characterized (Awachare et al. 2018).
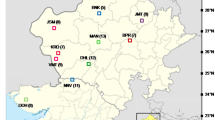
Ecuador is recognized as one of the countries with the highest biodiversity and density of endemic plant species worldwide (Joppa et al. 2013). In this sense, the country has around 113 species belonging to the Annonaceae family, among which 63 species have different uses (De la Torre et al. 2008). Annona deceptrix (Westra) H. Rainer commonly known as ‘chirimoya de monte’ or ‘anonilla’ is an endemic tree of the province of Manabí and other nearby cities in Ecuador (Pico-Mendoza et al. 2020). This species is distributed throughout the humid remnants in the coastal zone of the country. There are populations within the ‘Machalilla’ National Park and in the deciduous forests of the ‘Chongón Colonche’ Mountain range, located between Guayas and Esmeraldas provinces at an altitude of up to 850 m above sea level (Muriel 2011). Moreover, isolated populations have been observed in the northern area of the province of Manabí, specifically, at the ‘Pata de Pájaro’ hill located in the Pedernales canton (Fig. 1).
Annona deceptrix presents interesting botanical and agronomic traits such as the shape and size of its leaves, white to yellowish flowers, and fruits with different shapes (elongated and conical), texture (smooth and rough) and sweetness (Pico-Mendoza et al. 2024). However, due to anthropogenic activities associated with deforestation and agricultural expansion (Muriel 2011), and some intrinsic characteristics such as a low germination capacity of its seeds under natural conditions (Pico-Mendoza et al. 2020), this species has been reported as Vulnerable according to the IUCN Red List of Threatened Species in 2018 (Erkens 2021). Therefore, it is necessary to evaluate its genetic diversity and structure to establish management plans that will allow the species and its diversity to be conserved.
Most information available for the Annona genus is developed for commercial species such as A. cherimolla, A. muricata, and A. squamosa, which have been widely evaluated in terms of their morphological, agronomic and genetic diversity (Brisibe et al. 2017; Leal and Paull 2023). The lack of information on A. deceptrix limits our knowledge about it agronomic and nutritional properties. Furthermore, genetic research on this species would improve the evaluation of the risk of extinction and develop management strategies that guarantee its conservation (Garner et al. 2020). These investigations would allow the identification of populations with high genetic diversity and genotypes that have valuable nutritional and agronomic traits for genetic improvement of Annona species.
Polymerase chain reaction (PCR)-based molecular markers such as microsatellites or simple sequence repeats (SSR) have been widely used in many crops because they are considered a powerful molecular tool for the characterization of genetic variability and population structure of the species (Väli et al. 2008). Recently, a total of 22 SSR markers were developed for A. deceptrix, which proved to be reproducible and suitable as molecular markers for genetic research in Annona species (Pico-Mendoza et al. 2024). Therefore, the objective of the current study was to evaluate the diversity and genetic structure of natural populations of A. deceptrix, distributed at Manabí province, using specific SSR markers for conservation purposes of this valuable endangered endemic species.
Materials and methods
Plant material
A total of 116 tree of A. deceptrix were collected from 11 sites distributed at Manabí province, including the administrative Regions of Pacoche (9) in the Canton of Manta; Agua Blanca (11), Ayampe (8), and Cerro El Mate (17) in the canton of Puerto Lopez; El Barro in the canton of Jipijapa; Tachina (23), Tachina Alto (6), Nalpe (11), Humedad (13), Vite (8) and Guacucal (6) in the canton of Pedernales. Each collection site was geographically using the GPS information, except for the location of Guacucal whose information was lost (Table 1).
Genomic DNA extraction
A sample of ~ 200 mg of young leaves of A. deceptrix was placed in 2.0 mL tubes and frozen in liquid nitrogen. Leaf samples were macerated with the Precellis and Cryollys systems (Bertin Technologies, Montigny-le-Bretonneux, France), and centrifuged at 6000 rpm in three cycles of 30 s each. Total genomic DNA extraction of all samples was performed using the cetyl trimethyl ammonium bromide (CTAB) protocol (Doyle 1990). The quantity and quality of isolated DNA were determined using a Nanodrop and agarose gel electrophoresis. Eighteen SSR loci were evaluated according to Pico-Mendoza et al. (2024) and the PCR was carried out using the following steps: denaturation at 94 °C for 5 min; specific annealing temperature (Ta) for each primer pair (Table 2) for 45 s, extension at 72 °C for 45 s. This cycle was repeated 30 times, then a final extension at 72 °C. PCR products were separated by capillary electrophoresis in an ABI 3130xl Prism Genetic Analyzer with POP-7 polymer (Life Technologies, Foster City, California, USA). Allele sizes were calculated with GeneMapper software v 4.0 (Applied Biosystems, Foster City, California, USA).
Statistical analysis
Monomorphic SSR, and those SSR loci and genotypes containing 20% or more of missing values were discarded from the subsequent analyses using the ‘poppr’ (Kamvar et al. 2014) and ‘adegenet’ (Jombart 2008) packages implemented in R Studio software. The genetic diversity parameters such as mean number of alleles (Na), the mean effective number of alleles (Ne), expected heterozygosity (He), observed heterozygosity (Ho), number of private allele (Npa), percentage of polymorphic loci (PPL), fixation index (Fst), inbreeding coefficient (Fis), and gene flow (Nm) were estimated for each locus and population using GenAlEx Version 6.5 (Peakall and Smouse 2012) and ‘hierfstat’ package (Goudet 2005).
To determine the genetic structure, analysis of molecular variance (AMOVA) was applied to estimate the variance components within and between populations in GenAlEx Version 6.5 (Peakall and Smouse 2012). The population structure of the A. deceptrix genotypes was evaluated by principal component analysis (PCA) implemented in ‘ade4’ (Chessel et al. 2004) and ‘adegenet’ (Jombart 2008) and visualized with ‘ggplot2’ (Wickham 2010) packages. Additionally, an unrooted tree was constructed to show the genetic relationship among the A. deceptrix genotypes. It was implemented using the function aboot implemented in the ‘poppr’ package (Kamvar et al. 2014) based on Nei’s genetic distance with 1,000 bootstrap replicates. The resulting tree was visualized with function plot.phylo in the ‘ape’ package (Paradis and Schliep 2019). Moreover, the population structure was inferred using a Bayesian-based clustering method in Structure v2.3.4 (Pritchard et al. 2000). An admixture ancestry model with correlated allele frequencies assuming no prior information of population origin was used. Ten independent runs for each putative number of subpopulations (K), ranged from K = 1 to 11, were performed with a burn-in period of 100,000 steps followed by 1,000,000 MCMC iterations. The optimal K value was determined in Structure Harvester (Earl and vonHoldt 2012) using the ad hoc statistic ΔK (Evanno et al. 2005). Finally, a Mantel test was performed to determine isolation by distance (IBD) between populations of A. deceptrix in GenAlEx Version 6.5 (Peakall and Smouse 2012).
Results
Screening of SSR primers in A. deceptrix
In this study, a total of 18 SSR markers were successfully amplified in 116 samples of A. deceptrix. After applying quality filters on raw data, ten samples were discarded because they presented more than 20% missing data (Fig. 2). Therefore, a total of 106 A. deceptrix individuals were evaluated with the 18 SSR markers in the subsequent analyses.
Genetic diversity
The 18 SSR markers were polymorphic and generated a total of 186 alleles across the 106 individuals of A. deceptrix. The number of total alleles (Ta) per locus ranged from 5 to 25 in the markers AD18 and AD8, respectively, with an average of 10.3 alleles per locus. In general, the AD8 locus was the one that presented the highest diversity parameters (Ho = 0.777; He = 0.810; Ne = 4.423), while the AD18 locus was the one that presented the lowest diversity (Ho = 0.070; He = 0.190; Ne = 1.324). Among the SSR markers evaluated the AD8, AD5, AD6, AD10 and AD1 were the most informative for A. deceptrix. The genetic differentiation coefficient (Fst) varied from 0.102 to 0.354 in the loci AD11 and AD9, respectively, with an average value of 0.207. On contrary, the gene flow (Nm) at loci AD11 (Nm = 2.196) and AD9 (Nm = 0.455) presented the highest and lowest value, respectively. All diversity parameters evaluated at the locus level are shown in Table 3.
At population level, El Barro was the population with the lowest number of alleles per locus (Na = 1.444) and number of effective alleles (Ne = 1.404). While Tachina was the population with the highest Na (7.500) and Ne (3.577). Overall, the number of alleles per locus and number of effective alleles was of 3.654 and 2.367, respectively. A total of 89 private alleles (Npa) were identified in the 11 populations by 18 SSR markers, except for Pacoche and El Barro where no Npa were identified. The highest number of private alleles was found in the Tachina (Npa = 36), followed by Agua Blanca (Npa = 14), and Humedad (Npa = 13) populations. The observed heterozygosity (Ho) varied from 0.352 to 0.519 in the Ayampe and Guacucal, respectively. The expected heterozygosity (He) was highest in Humedad with a value of 0.649, followed by Agua Blanca (He = 0.623), and Tachina (He = 0.616). The lowest He was observed in El Barro with a value of 0.194. In general, the level of genetic diversity in the 106 A. deceptrix sample was moderate (He = 0.445). The percentage of polymorphic loci (PPL) was of 100% in Agua Blanca, Tachina, Nalpe, and Humedad, while El Barro presented the lowest number of PPL with a value of 38.89%. Finally, El Barro was the population with the lowest inbreeding (Fis = − 0.667) and genetic differentiation (Fst = − 0.848) coefficients, while the population Tachina presented the highest values of Fis = 0.391 and Fst = 0.373. All diversity parameters evaluated at the population level are shown in Table 4.
Genetic differentiation and genetic structure
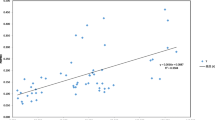
The analysis of molecular variance (AMOVA) revealed that a large part of the genetic variation occurred within individuals (65%), while 11% of genetic variation was attributable to the differentiation among populations (Table 5). The fixation index (Fst) was 0, indicating no genetic differentiation among populations. Pairwise estimates of Fst calculated between pairs of populations are shown in the Fig. 3. El Barro population presented the greatest differentiation (Fst) with the Ayampe (Fst = 0.372), Tachina Alto (Fst = 0.358), and Pacoche (Fst = 0.351) populations, which come from the Puerto López, Pedernales, and Manta cantons, respectively. In general, the least differentiation was found between populations belonging to the canton of Pedernales. In this sense, the pairs of populations Nalpe and Humedad (Fst = 0) and Nalpe and Tachina (Fst = 0.009) were the least differentiated. This result is in accordance with the Mantel test conducted for A. deceptrix populations, where a null correlation between geographic and genetic distances among populations (R2 = 0.007; p = 0.02) was detected, indicating no significant isolation by distance (IBD) between populations (Fig. 4).
The principal component analysis (PCA) also showed a low differentiation between A. deceptrix populations, where the first principal component (PC) explained 25.6% of the variation, while the second PC explained 8.3%. This result indicated that the 11 natural populations can be grouped into two genetic clusters. In one cluster, individuals from the 11 populations were grouped, while in the second cluster, individuals from the populations of Agua Blanca (3), Tachina (5), Nalpe (2), and Humedad (4) were grouped (Fig. 5). Result is in agrees with the Bayesian analysis, where the most probable number of subpopulations was K = 2 (Fig. 6A and B). Finally, the genetic relationship between the A. deceptrix individuals from different populations can be grouped into three cluster based on genetic distance, where some individuals from the Agua Blanca, Tachina, Nalpe and Humedad populations were the most genetically distant in accordance with the structure and PCA results (Fig. 6C).
Genetic structure of 106 genotypes from 11 Annona deceptrix populations inferred by Bayesian clustering and genetic distance. A Plot of mean posterior probability Ln (K) values varying of number of subpopulations (K) from 1 to 11. B The estimated membership probability (Q) for each genotype. C Unrooted neighbor joining tree of 106 genotypes of A. deceptrix. The color of the names is according to the result obtained by the structure analysis, where red and green corresponds to cluster 1 and cluster 2, respectively
Discussion
SSR primers in A. deceptrix
SSR markers have been widely used to carry out research of genetic diversity due to their hyper-variability, multiallelic nature, reproducibility, relative abundance, extensive genome coverage, high polymorphism, and co-dominant inheritance (Kalia et al. 2011). The 18 SSR markers used in this study had a high percentage of polymorphic loci (82.3%) in the A. deceptrix populations, confirming that the SSR markers developed by Pico-Mendoza et al. (2024) are useful to evaluate genetic diversity in A. deceptrix. These results are also in consonance with those reported by Saitwal et al. (2022), where a total of 20 SSR were evaluated in different Annona genotypes from Maharashtra, India, and produced 83.72% of polymorphic loci. In addition, a 95.0% of polymorphic loci was reported in populations of A. senegalensis using 10 SSR markers (Donhouedé et al. 2023). In our current study, the AD5, AD6, AD8, AD10 and AD1 markers were the most informative for A. deceptrix. Consequently, considering the high rate of transferability of SSR markers between related species, these primers could be useful for the evaluation of evaluate genetic diversity in other understudied Annona species, without the need to develop species-specific markers.
Genetic diversity of A. deceptrix
Ecuador contains a unique set of species and ecosystems, many of them endemic to the country (Cuesta et al. 2017). However, due to different sources of anthropogenic threat, many of these endemic species are expected to become extinct or threatened (Brooks et al. 2002) in the future. A. deceptrix is an endemic and endangered tree distributed throughout the humid remnants in the coastal zone of Ecuador (Pico-Mendoza et al. 2020). Ecosystems near coastal areas are considered fragile and with high rates of disturbance, being considered priority conservation areas (Cuesta et al. 2017). In Manabí, a coastal province of Ecuador, there is a high deforestation rate (6159 ha/year), which causes a high degree of threat to its native flora, including A. deceptrix. Despite this reality, there are no programs aimed at the conservation of threatened species. In this sense, to evaluate the genetic diversity of A. deceptrix populations is an important prerequisite to determine the management strategies that would guarantee its conservation.
To carry out effective conservation strategies, it is essential to understand the spatial patterns of the genetic diversity of the target species (Escudero et al. 2003), allowing the identification of populations with greater and lesser diversity, and thus, creating conservation strategies that optimize the use of available genetic resources (van Zonneveld et al. 2012). Among the populations of A. deceptrix, the greatest expected (He) and observed (Ho) heterozygosity was reported for Humedad (He = 0.649; Ho = 0.474), followed by Agua Blanca (He = 0.623; Ho = 0.472), and Tachina (He = 0.616; Ho = 0.426). In a previous work, individuals of A. deceptrix from Tachina (12), Agua Blanca (10), and Humedad (18) were evaluated with 22 SSR markers, whose He and Ho values were like those obtained in this study (Pico-Mendoza et al. 2024). Our results indicated that the A. deceptrix populations collected have a moderate level of genetic diversity, showing that the expected and observed heterozygosity of A. deceptrix was He = 0.445 and Ho = 0.416, respectively. This is similar to what has been reported in other Annona species, where a moderate level of genetic diversity (He = 0.470) was found in 14 populations of A. squamosa distributed in the Mexican states of Yucatán and Quintana Roo using isozyme markers (Salazar et al. 2010). Moreover, in A. senegalensis populations occurring in Western and Southern Africa a high level of genetic diversity (He = 0.56) was found (Donhouedé et al. 2023). However, these values are higher than those reported in other populations of Annona localized in Brazil. For example, in four natural populations of A. crassiflora from northern Minas Gerais State, the genetic diversity evaluated with RAPD markers was He = 0.31 (Cota et al. 2011). In A. squamosa, populations from the Brazilian states of Maranhão and Piauí had a low genetic diversity (He = 0.222) using ISSR markers (Sá et al. 2022). Our findings suggests that the anthropogenic effect on the reduction in population size has affected the genetic variation of the species to a lesser extent.
Genetic differentiation and structure in A. deceptrix
The analysis of molecular variance (AMOVA) revealed that only 11% of the total genetic variation in A. deceptrix occurred among populations, while 65% resided within individuals. The low differentiation coefficient (Fst = 0) and the moderate gene flow (Nm = 1.159) indicated that there is no genetic differentiation between populations. However, the markers AD9, AD22, AD16, AD1, AD19, and AD2 have a high power of discrimination between populations (Fst > 0.25). The level of population differentiation reported in this study for A. deceptrix was contrary to what has been reported in other Annonas species such as A. squamosa (Nagori et al. 2018; Sá et al. 2022), A. muricata (Brisibe et al. 2017; Lira-Ortiz et al. 2022), A. senegalensis (Donhouedé et al. 2023), where populations are, generally, genetically differentiated according to its distribution and geographic distance. This result may be due to the negative relationship between the number of alleles per locus and the estimates of Fst values, resulting in a decrease in the power to discriminate between populations (O’Reilly et al. 2004). Furthermore, the result may be influenced by the population size used in this study, since a number of ~ 20–25 individuals (Danusevičius et al. 2016) per population or at least 10% of the individuals from populations are recommend for genetic research (Sinclair and Hobbs 2009; Costa et al. 2015).
Although anthropogenic activity has affected tropical forests since ancient times (van der Sande et al. 2019), our results suggest that the fragmentation of the coastal habitat in the province of Manabí has been below the fragmentation threshold which causes loss of genetic diversity and population structure (Cruzan 2001; Bacles and Jump 2011). Part of the data in the current study is supported by the Mantel test where no isolation by distance was detected between A. deceptrix populations, indicating that the reduction in population size due to anthropogenic activity has not greatly affected the genetic variation of the species. However, due to the intensive increase in land use change, especially during recent decades, the conservation of threatened species (mainly endemic) should be a high priority (Gibbs et al. 2010; Cuesta et al. 2017; Erkens et al. 2023). For the purposes of conservation of A. deceptrix, the strategies should focus on populations such as Humedad, Agua Blanca, and Tachina, since they were the populations with the greatest diversity and number of private alleles, which are good candidates for conservation because high levels of genetic variation are expected to increase the potential of populations to respond to selection and maintain the health of individuals (Kalinowski 2004). In addition, these populations are interesting for genetic improvement purposes because there is a greater probability of finding genotypes with characteristics of commercial interest than in populations with low genetic diversity (Frankel et al. 1995; van der Sande et al. 2019).
Conclusion
The SSR markers developed for A. deceptrix were informative and allowed evaluating the genetic diversity of 11 natural populations distributed in the province of Manabí, Ecuador. A moderate level of genetic diversity and no differentiation between populations was found. However, the large number of private alleles reported could indicate that populations are experiencing significant reductions in size. Therefore, these results have a great impact on the conversation policies of A. deceptrix considering that it is an endemic and threatened species. In this sense, it is suggest that the strategies should focus on populations such as Humedad, Agua Blanca, and Tachina, since they were the populations with the greatest diversity and number of private alleles, doing conservation activities such as habitat conservation, in situ and ex situ conservation, genetic banks, with the intention of safeguarding the genetic variability evaluated.
Data availability
The data sets generated and/or analyzed during the current study are not publicly available due to future work being planned. But they are available from the corresponding author upon reasonable request.
References
Al Kazman BS, Harnett JE, Hanrahan JR (2022) Traditional uses, phytochemistry and pharmacological activities of Annonacae. Molecules 27:3462. https://doi.org/10.3390/molecules27113462
Awachare CM, Kurian RM, Upreti KK, Laxman RH (2018) Morpho-physiological diversity in Annona species. Sci Hortic 234:58–62. https://doi.org/10.1016/j.scienta.2018.02.005
Bacles CF, Jump AS (2011) Taking a tree’s perspective on forest fragmentation genetics. Trends Plant Scie 16:13–18. https://doi.org/10.1016/j.tplants.2010.10.002
Brisibe EA, Ogbonna NC, Chukwurah PN (2017) Characterization and selection of exploitable genetic diversity in soursop (Annona muricata Linn.) accessions based on phenotypic attributes and RAPD markers. Agrofor Syst 91:781–793. https://doi.org/10.1007/s10457-016-9965-4
Brooks TM, Mittermeier RA, Mittermeier CG, Da Fonseca GA, Rylands AB, Konstant WR, Flick P, Pilgrim J, Oldfield S, Magin G, Hilton-Taylor C (2002) Habitat loss and extinction in the hotspots of biodiversity. Conserv Biol 16:909–923. https://doi.org/10.1046/j.1523-1739.2002.00530.x
Cautín R, Agustí M (2005) Phenological growth stages of the cherimoya tree (Annona cherimola Mill.). Sci Hortic 105:491–497. https://doi.org/10.1016/j.scienta.2005.01.035
Chessel D, Dufour AB, Thioulouse J (2004) The R Journal: the ade4 package—I: One-table methods. R News 4:5–10
Costa LSD, Corneleo NS, Stefenon VM (2015) Conservation of forest biodiversity: how sample size affects the estimation of genetic parameters. An Acad Bras Ciênc 87:1095–1100. https://doi.org/10.1590/0001-3765201520140064
Cota LG, Vieira FA, Melo Júnior AF, Brandão MM, Santana KNO, Guedes ML, Oliveira DA (2011) Genetic diversity of Annona crassiflora (Annonaceae) in northern Minas Gerais state. Genet Mol Res 10:2172–2180. https://doi.org/10.4238/vol10-3gmr1188
Cruzan MB (2001) Population size and fragmentation thresholds for the maintenance of genetic diversity in the herbaceous endemic Scutellaria montana (Lamiaceae). Evolution 55:1569–1580. https://doi.org/10.1111/j.0014-3820.2001.tb00676.x
Cuesta F, Peralvo M, Merino-Viteri A, Bustamante M, Baquero F, Freile JF, Muriel P, Torres-Carvajal O (2017) Priority areas for biodiversity conservation in mainland Ecuador. Neotrop Biodivers 3:93–106. https://doi.org/10.1080/23766808.2017.1295705
Danusevičius D, Kavaliauskas D, Fussi B (2016) Optimum sample size for SSR-based estimation of representative allele frequencies and genetic diversity in Scots pine populations. Balt for 22:194–202
Donhouedé JCF, Marques I, Salako KV, Assogbadjo AE, Ribeiro N, Ribeiro-Barros AI (2023) Genetic and morphological diversity in populations of Annona senegalensis Pers. occurring in western (Benin) and southern (Mozambique) Africa. PeerJ 11:e15767. https://doi.org/10.7717/peerj.15767
Doyle JJ (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
De la Torre L, Navarrete H, Muriel MP, Macía MJ, Balslev H (2008) Enciclopedia de las Plantas Útiles del Ecuador. Herbario QCA de la Escuela de Ciencias Biológicas de la Pontificia Universidad Católica del Ecuador and Herbario AAU del Departamento de Ciencias Biológicas de la Universidad de Aarhus. Quito & Aarhus. Quito. https://bibdigital.rjb.csic.es/idurl/1/16016
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Erkens RHJ (2021) Annona deceptrix. The IUCN Red List of Threatened Species 2021: e.T42828A176434191. https://doi.org/10.2305/IUCN.UK.2021-2.RLTS.T42828A176434191.en. Accessed on 29 September 2023.
Erkens RH, Blanpain LM, Carrascosa Jara I, Runge K, Verspagen N, Cosiaux A, Couvreur TL (2023) Spatial distribution of Annonaceae across biomes and anthromes: knowledge gaps in spatial and ecological data. Plants People Planet 5:520–535. https://doi.org/10.1002/ppp3.10321
Escudero A, Iriondo JM, Torres ME (2003) Spatial analysis of genetic diversity as a tool for plant conservation. Biol Conserv 113:351–365. https://doi.org/10.1016/S0006-3207(03)00122-8
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Frankel OH, Brown AHD, Burdon J (1995) The conservation of cultivated plants. The conservation of plant biodiversity. pp. 79–117. Cambridge University Press, UK. First edition.
Garner BA, Hoban S, Luikart G (2020) IUCN Red List and the value of integrating genetics. Conserv Genet 21:795–801. https://doi.org/10.1007/s10592-020-01301-6
Gibbs HK, Ruesch AS, Achard F, Clayton MK, Holmgren P, Ramankutty N, Foley JA (2010) Tropical forests were the primary sources of new agricultural land in the 1980s and 1990s. PNAS 107:16732–16737. https://doi.org/10.1073/pnas.0910275107
Goudet J (2005) Hierfstat, a package for R to compute and test hierarchical F-statistics. Mol Ecol Notes 5:184–186. https://doi.org/10.1111/j.1471-8286.2004.00828.x
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Joppa LN, Visconti P, Jenkins CN, Pimm SL (2013) Achieving the convention on biological diversity’s goals for plant conservation. Science 341:1100–1103. https://doi.org/10.1126/science.124170
Kalia RK, Rai MK, Kalia S, Singh R, Dhawan AK (2011) Microsatellites markers: an overview of the recent progress in plants. Euphytica 177:309–334. https://doi.org/10.1007/s10681-010-0286-9
Kalinowski ST (2004) Counting alleles with rarefaction: private alleles and hierarchical sampling designs. Conserv Genet 5:539–543. https://doi.org/10.1023/B:COGE.0000041021.91777.1a
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281. https://doi.org/10.7717/peerj.281
Leal F, Paull RE (2023) The genus Annona: Botanical characteristics, horticultural requirements and uses. Crop Sci 63:1030–1049. https://doi.org/10.1002/csc2.20833
Lira-Ortiz R, Cortés-Cruz MA, López-Guzmán GG, Palomino-Hermosillo YA, Sandoval-Padilla I, Ochoa-Jiménez VA, Sánchez-Herrera LM, Balois-Morales R, Berumen-Varela G (2022) Diversidad Génetica de poblaciones de guanábana (Annona muricata L.) en Nayarit, México mediante marcadores SSR y SRAP. Acta Biol Colomb 27:104–112. https://doi.org/10.15446/abc.v27n1.88241
Muriel P (2011) Raimondia deceptrix. In: León-Yánez S, Valencia R, Pitmam N, Endara L, Ulloa Ulloa C, Navarrete H (Eds) Libro Rojo de Plantas Endémicas del Ecuador, 2ª edición. Publicaciones del Herbario QCA, Pontificia Universidad Católica del Ecuador, Quito.
Nagori R, Sharma P, Purohit SD (2018) Molecular characterization of diversity in Annona squamosa L.: a high value tropical medicinal plant. Int J Phytocos Nat Ingred 5:4. https://doi.org/10.15171/ijpni.2018.04
O’Reilly PT, Canino MF, Bailey KM, Bentzen P (2004) Inverse relationship between FST and microsatellite polymorphism in the marine fish, walleye pollock (Theragra chalcogramma): implications for resolving weak population structure. Mol Ecol 13:1799–1814. https://doi.org/10.1111/j.1365-294X.2004.02214.x
Paradis E, Schliep K (2019) ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35:526–528. https://doi.org/10.1093/bioinformatics/bty633
Peakall R, Smouse P (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Pico-Mendoza J, Pinoargote M, Madrid L, Flor J, Álava J, Sancán G, Basilio C, Limongi R, Peña G, Quiroz K (2020) Germination capacity of Annona deceptrix (Westra) H. Rainer (Annonaceae) an endemic and endangered species in Manabí, Ecuador. Indian J Agric Res 54:329–335. https://doi.org/10.18805/IJARe.A-481
Pico-Mendoza J, Madrid L, Pinoargote M, Limongi R, Peña G, Flor J, Vélez A, Quiroz K, Arevalo B, Carrasco B (2024) Simple sequence repeat markers (SSR) in Annona Deceptrix Westra H. Rainer, an endangered species of the Ecuadorian coast. J Trop For Sci 36:132–138. https://doi.org/10.26525/jtfs2024.36.1.132
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Sá GH, Lima FC, Viana JP, Lopes ÂC, Carvalho LC, Valente SE, Lima PS (2022) Genetic diversity and structure of an active germplasm collection of Annona squamosa L. Ecol Genet Genom 25:100143. https://doi.org/10.1016/j.egg.2022.100143
Saitwal YS, Musmade AM, Kale AA, Supe VS, Joshi VR, Ranpise SA, Mehetre SS (2022) Characterization of Annona genotypes by ISSR and SSR markers. Bhartiya Krishi Anusandhan Patrika 37:43–49. https://doi.org/10.18805/BKAP494
Salazar C, Vargas-Mendoza CF, Flores JS (2010) Estructura y diversidad genética de Annona squamosa en huertos familiares mayas de la península de Yucatán. Rev Mex Biodivers 81:759–770
Sinclair EA, Hobbs RJ (2009) Sample size effects on estimates of population genetic structure: Implications for ecological restoration. Restor Ecol 17:837–844. https://doi.org/10.1111/j.1526-100X.2008.00420.x
Väli Ü, Einarsson A, Waits L, Ellegren H (2008) To what extent do microsatellite markers reflect genome-wide genetic diversity in natural populations? Mol Ecol 17:3808–3817. https://doi.org/10.1111/j.1365-294X.2008.03876.x
van der Sande MT, Gosling W, Correa-Metrio A, Prado-Junior J, Poorter L, Oliveira RS, Mazzei L, Bush MB (2019) A 7000-year history of changing plant trait composition in an Amazonian landscape; the role of humans and climate. Ecol Lett 22:925–935. https://doi.org/10.1111/ele.13251
van Zonneveld M, Scheldeman X, Escribano P, Viruel MA, Van Damme P, Garcia W et al (2012) Mapping genetic diversity of cherimoya (Annona cherimola Mill.): Application of spatial analysis for conservation and use of plant genetic resources. PLoS ONE 7:e29845. https://doi.org/10.1371/journal.pone.0029845
Wickham H (2010) A layered grammar of graphics. J Comput Graphical Stat 19:3–28. https://doi.org/10.1198/jcgs.2009.07098
Funding
The authors are grateful to the Endangered Species Conservation Program of the Faculty of Agronomic Engineering of the Technical University of Manabí. Universidad Laica Eloy Alfaro de Manta, extension Pedernales. Centro de Estudios en Alimentos Procesados (CEAP) Talca, Chile; and many others in one way or another supported this research.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by José Pico-Mendoza, Luis Madrid, Eduardo Morillo and Juan Flor. The first draft of the manuscript was written by Basilio Carrasco and Osvin Arriagada and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pico-Mendoza, J., Madrid, L., Morillo, E. et al. Genetic diversity and structure populations in Annona deceptrix (Westra) H. Rainer (Annonaceae), an endangered species from Ecuador. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-02037-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-02037-9