Abstract
Genetic diversity within germplasm holds significant importance in the context of okra breeding programs. However, the limited knowledge regarding the genetic diversity of okra in Palestine poses a challenging obstacle to enhancing the local cultivar. Characterization of Palestinian okra landraces using phenotypic or molecular approaches is limited. Crop characterization usually provides knowledge on diversity within and among crop accessions, which is crucial for breeding efforts and the conservation of germplasm. This study was carried out to evaluate genetic variation in seven okra local landraces. Seven Palestinian okra landraces were grown. DNA was extracted and subjected to polymerase chain reaction (PCR) using 27 random amplified polymorphic DNA (RAPD) markers. The banding pattern for each primer was scored. The data matrix was created to facilitate cluster analysis employing the unweighted pair group method with arithmetic mean (UPGMA) method. From the 15 RAPD markers tested, 92 DNA fragments were identified in total, of these fragments, 76% were polymorphic and 24% were monomorphic, in addition, at least two and no more than eleven DNA fragments were identified. The average resolving power recorded was 2.8. The dendrogram derived from RAPD data categorized the genotypes into two primary clusters. Cluster-I consisted of a single genotype (PARC-OKRA-1). In contrast, Cluster-II further subdivided the genotypes into two subclusters, sub-cluster-II.a represented one genotype accession (UB 405-19), and sub-cluster-II.b included five genotypes (UB 457-19, UB 280-15, UB 457-14, UB 280-12, and UB 457-20). The generated similarity matrix indicated that the similarity coefficients among okra genotypes, as determined by 15 RAPD markers, ranged from 0.226 to 0.678. The present study represents the first application of molecular markers to assess genetic diversity within Palestinian okra landraces. The detection of a high level of polymorphism among RAPD markers demonstrated the efficacy of RAPD markers for identifying polymorphism in Palestinian okra genotypes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Malvaceae, also known as mallows, is a family of flowering plants containing an approximately 240 genera and 4225 species. Okra, cotton, and cocoa are well-known members with great economic importance (Judd et al. 2008). Okra (Abelmoschus esculentus) is an annual vegetable crop grown globally in tropical and subtropical regions (Patil et al. 2015).
Okra is considered a native crop in Africa, however, contradicting evidence exists on its geographical origin in West and Central Africa. So far, most reports indicate that the geographical origin of okra can be either India or Ethiopia (Elkhalifa et al. 2021). Later, the crop probably spread to the Middle East and other parts of the world.
Okra is an economically important crop cultivated in diverse cropping systems and environments. Globally, approximately 10 million tons of okra are harvested annually (Schafleitner et al. 2021). In Palestine, okra is considered one of the favorite crops, with increasing demands due to its valuable benefits for human health. Palestinians consume okra in the form of immature pods, which can be eaten fresh, fried or cooked. Unfortunately, there is no precise data about the cultivated area and productivity of okra in Palestine. However, the Food and Agriculture Organization (FAO) states that in 2023, more than 5340 dunums have been cultivated with okra in Palestine, which produced about 2496 tones (2023).
The characterization of genetic resources could be described as the process of identifying, differentiating, or distinguishing accessions based on their characteristics or qualities (traits). The process of crop characterization usually offer information on crop collection diversity. This acquired knowledge is essential for gene bank curators as it allows for the identification of individual accessions. On top of that, information acquired on the genetic similarity of agricultural plant genetic resources is not only valuable for breeding as well as for germplasm conservation (International 2007).
Identification and characterization are sometimes difficult by conventional methods as the morphological markers are environment dependent, sensitive and time consuming. Additionally, assessment of morphological traits necessitates complete maturation of plants prior to identifying different genotypes (Massucato et al. 2019). To address these constraints, molecular approaches have been extensively implemented. Contrary to morphological markers, DNA expression is highly stable and the expression of DNA markers or molecular markers is not confounded by the environment and are more numerous than morphological characters (Massucato et al. 2019), hence, these markers are a robust and valuable tool to elucidate the hidden relationship among different plant populations.
Molecular markers approaches have become as a viable method for assessing genetic variations. Random Amplified Polymorphic DNA (RAPD), Simple Sequence Repeat (SSR), and Amplified Fragment Length Polymorphism (AFLP) are the most commonly employed markers for genetic research and marker-assisted breeding programs (Sawadogo et al. 2009). However, information on molecular markers for okra is limited (Aladele et al. 2008).
There are a few investigations that have employed general DNA markers to evaluate the genetic diversity of okra (Gulsen et al. 2007; Prakash et al. 2011; Kumar et al. 2017). RAPD and AFLP have been employed in previous research comparing the genetic diversity of several okra accessions (Ojo; Martinello et al. 2001; Prakash et al. 2011; Salameh 2014; Japda and Dhruve 2018). Sequence-Related Amplified Polymorphism (SRAP) has been used to study genetic variation within the okra plant as well. (Gulsen et al. 2007). SSR markers are an effective tool for plant genetics and breeding (de Magalhães Bertini et al. 2006; Japda and Dhruve 2018; Jhanji et al. 2021; Mkhabela et al. 2022). However, 402 microsatellites developed by using next-generation sequencing technology for amplification in okra (Ravishankar et al. 2018).
RAPD is a DNA-based marker approach developed by Welsh and Mcclelland (Welsh and Mcclelland 1990). Although RAPD is simple, dominant DNA marker, they may identify many loci ranging from 0.5 to 5 Kb, making them useful for genetic diversity analysis.
Genetic diversity within germplasm holds significant importance in the context of okra breeding programs. However, the limited knowledge regarding the genetic diversity of okra in Palestine poses a challenging obstacle to enhancing the local cultivar. Moreover, as a potentially self-pollinating crop, planted okra has a limited genetic base, consequently, concentrated efforts are required to determine the full potential of the okra germplasm resources in Palestine. To assess the nature and magnitude of genetic divergence among available okra landraces in Palestine, which is essential for selecting genetically diverse parents for an effective breeding program, the existing germplasm accessions must be evaluated in detail for a variety of horticultural and genetic traits.
Therefore, the objective of the current study is to evaluate the genetic diversity and relationships among seven okra landraces preserved in the Palestinian gene banks and cultivated in the West Bank, employing RAPD molecular markers.
Materials and Methods
Plant Material
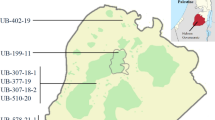
In the current study, seeds of 7 okra accessions of Palestinian local landrace from different places were used (Fig. 1). Among the accessions, six were provided by the Union of Agricultural Work Committees seed bank, and one was provided by Palestinian Agricultural Relief Committees seed bank (Table 1). In December 2022, at least 10 seeds from each accession were cultivated in pots containing a mixture of perlite and peat moss in a 1:2 (v:v) in December 2022. Seedlings were grown in a greenhouse for 2 weeks.
DNA Extraction and Molecular Analysis
Total DNA was extracted from 14-day-old fresh leaves by using the DNeasy Plant Mini Kit (Qiagen) according to the manufacturer's instructions. 27 RAPD primers (Table 2) were obtained from Sigma-Aldrich to amplify random DNA banding patterns. Polymerase chain reaction (PCR) amplifications were performed in a final volume of 25 μl, and the amplification mixture consisted of 12.5 μl of 2X GoTaq master mix, 9.5 μl of nuclease-free water, 2 μl of a 10 pmol primer, and 1 μl of DNA template (50 ng of total DNA). The PCR reactions were conducted in an Eppendorf thermal cycler, following this protocol: an initial denaturation step at 91 °C for 5 min, followed by 40 cycles of 1 min at 91 °C, 1 min at 36 °C, and 2 min at 72 °C, concluding with a final extension step of 5 min at 72 °C. Electrophoresis was used to separate the amplification products on 1.5% agarose gels in 1X TBE buffer. The approximate size of DNA fragments was approximated using either a 100 bp or 1 KB DNA ladder (hylabs) as a standard molecular weight marker. The generated bands were visualized by a gel documentation system under UV light.
Data Analysis
The banding pattern from RAPD analysis was assessed visually for each primer. In each position, the existence of an amplification product (band) was recorded as 1 and its absence as 0. When these bands were absent from some samples, they were assumed polymorphic. The polymorphism proportion for each marker was computed as [number of polymorphic bands divided by the total number of scored bands] multiplied by 100.
Data matrix for cluster analysis using the UPGMA (Unweighted Pair Group Method with Arithmetic Average) was generated using Excel software. (Sneath and Sokal 1973). Cluster analysis was conducted utilizing the Nei and Li distance coefficients (Nei 1978). Using MVSP (Multi Variate Statistical Package), UPGMA was performed (1986).
The Marker Index (MI) for all markers was determined following the procedure outlined by Powell et al. in 1996 (Powell et al. 1996), which serves as an indicator of a marker's discriminatory power. Additionally, the Polymorphic Information Content (PIC) was calculated for each primer in order to evaluate its capacity for distinguishing between different individuals. This calculation followed to the methodology outlined by Prevost and Wilkinson in (1999), as presented by Prevost, A. (Prevost and Wilkinson 1999). Furthermore, the Resolving Power (RP) or Discriminatory Power, representing a marker's ability to differentiate between genotypes or individuals, was determined for each primer using the approach detailed by Prevost and Wilkinson (Prevost and Wilkinson 1999).
Results
Diversity of alleles in Okra utilizing RAPD markers
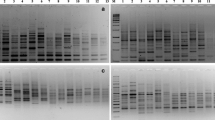
Out of 27 RAPD markers tested, only18 markers generated variable banding patterns and 3 markers shown no polymorphic bands. A total of 92 DNA fragments were detected, resulting in an average of 6.1 loci per primer, spanning a size range from 150 to 2000 base pairs (bp) (Table 2). Among these fragments, 70 (76%) revealed polymorphism, the test units reveal an average of 4.7 polymorphic bands, while the remaining 22 (24%) were monomorphic. The highest percentage of polymorphic markers observed was 100%, exemplified by OPA-4, OPC-6, OPC-8, OPE-8, OPG-4, and OPE-12, while the lowest was 40% for OPD-2 (Fig. 2). Consequently, the use of different primers resulted in the obtaining of at least 2 and up to 11 DNA fragments, observed in the cases of OPG-8, OPB-1, and OPF-1 primers, respectively.
Amplification profiles of random amplified polymorphic DNA (RAPD) markers in DNA of okra genotypes. Lane 1 and 17 (M) 1000 bp DNA ladder; lanes 2–8, Banding pattern as revealed by primers OPD-20; 10–16 Banding pattern as revealed by primers OPD-2; Lane 9(W); negative control (bulk from the two PCR reactions using OPD-20 and OPD-2 primers)
Marker Index (MI), which indicates the discriminatory power of a marker, was an important feature calculated, however, RAPD markers OPG-5 and OPH-7 displayed a higher Marker Index value of 0.61 (Table 2). Average PIC for RAPD markers were 0.52. The primer with the highest PIC was OPG-4 (Table 2). Further resolving power (RP)/discriminatory power was calculated for every primer. The average resolving power observed was 2.8. Primers OPB-1 and OPBI-1 have the highest and lowest resolving powers, respectively, among RAPD markers. Primer OPB-1 has a resolving power of 6.3, while primer OPBI-1 has a resolving power of 1.2.
Genetic relationship among okra genotypes
The RAPD data sets have been analyzed in order to evaluate the genetic relationships among the okra genotypes. The dendrogram generated by the analysis successfully categorized the genotypes into two primary clusters. (Fig. 3). Cluster-I consisted of one genotype (PARC-Okra-1), while Cluster-II encompassed six distinct genotypes. Subsequently, within Cluster-II, the genotypes were further divided into two additional subclusters. Specifically, sub-cluster-II.a was exclusive to one genotype accession (UB 405-19). In contrast, sub-cluster-II.b comprised five genotypes (UB 457-19, UB 280-15, UB 457-14, UB 280-12, and UB 457-20) (Fig. 3).
The constructed similarity matrix shown that the similarity coefficients among okra genotypes determined from 15 RAPD markers, varied between 0.226 and 0.678 (Table 3). The genotypes UB 48-7-20 and UB 214-12, originating from the southern and central regions of the Dura area, respectively, exhibited the highest similarity index, reaching 0.678. The lowest similarity coefficients (0.226) occurred in genotypes PARC-OKRA-1 and UB 214–12 which were collected from Qalqilya (north part of Palestine) and Dura city (south part of Palestine), respectively (Fig. 1).
Discussion
Our finding proved that RAPD marker techniques was highly effective in discriminating the polymorphism between the 7 Palestinian Okra genotypes. In the analysis of detected DNA fragments, each primer produced an average of 6.1 loci. Out of these amplified fragments, 76% exhibited polymorphism, yielding an average of 4.7 polymorphic bands per assay. Consequently, the use of RAPD primers led to the detection of at least 2 and up to 11 DNA fragments, such results was reported by (Aladele et al. 2008). This polymorphism was indicative of the existence of moderate genetic diversity within these seven okra genotypes; hence, the use of RAPD primers revealed the existence of a significant polymorphism, demonstrating genetic variation in the okra genotypes, such result obtained by Punitha and Raveendran (Punitha and Raveendran 2004) and Ikram-ul-haq (Haq et al. 2013).
The resulting dendrogram separated the genotypes into two primary clusters. Nevertheless, Cluster-I was represented by a single genotype accession (PARC-OKRA-1), collected from Qalqilya in the northern region of Palestine, this observation agrees with the findings of (Aladele et al. 2008), who employed RAPD markers on 93 okra genotypes and identified one genotype as exhibiting greater variation than the others. In contrast, the majority of genotypes were grouped within Cluster-II, originating from the southern part of Palestine. These findings might provide evidence of an association between genetic diversity and the geographical origins of these landraces. Except for PARC-Okr-1 landrace, all other landraces were collected from Hebron governorate. The results (Fig. 3) showed that this landerace has been located in a separate cluster. This association coincides with the findings presented in the study by Massucato et al. (2019) and Salameh (2014), where they explored genetic diversity among okra landraces using molecular AFLP markers and proved the presence of variability among accessions. Both authors, also found genetic diversity among different Okr landraces with association with geographical region.
Regarding Cluster-II, where the majority of genotypes were grouped together and showed genetic diversity, a possible explanation is that these six genotypes were collected from a relatively small geographical area in Hebron province. Therefore, we predict that the majority of these landraces may have transitioned from one region to another as a result of human activity,, such a result was reported by (Massucato et al. (2019). In fact, the comparable agroclimate that characterizes the area may be the explanation for their high similarity and corresponding clustering.
Conclusion
In the current work, PCR that uses RAPDs is considered the first molecular marker approach used for assessing genetic diversity and relatedness among Palestinian okra landraces. This work revealed the usefulness of RAPD molecular markers for identifying polymorphism in Palestinian okra landraces by obtaining a high degree of polymorphism as well as the identification of particular RAPD markers. The results of our research provide a solid foundation for upcoming breeding efforts, enabling the selection of diverse genetic resources to enhance the quality of locally cultivated okra cultivars.
Ethics Declarations
Not applicable.
Consent for Publication
The author declare that the work has been consented for publication.
Competing Interests
The author declare that they have no competing interests.
Availability of Data and Materials
All data and materials of the study have been presented in the main manuscript.
References
Aladele S, Ariyo O, Lapena R de (2008) Genetic relationships among West African okra (Abelmoschus caillei) and Asian genotypes (Abelmoschus esculentus) using RAPD. African J Biotechnol 7. https://doi.org/10.4314/AJB.V7I10.58689
de Magalhães Bertini CHC, Schuster I, Sediyama T et al (2006) Characterization and genetic diversity analysis of cotton cultivars using microsatellites. Genet Mol Biol 29:321–329. https://doi.org/10.1590/S1415-47572006000200021
Elkhalifa AEO, Alshammari E, Adnan M, et al (2021) Okra (Abelmoschus Esculentus) as a potential dietary medicine with nutraceutical importance for sustainable health applications. Molecules 26. https://doi.org/10.3390/MOLECULES26030696
Gulsen O, Karagul S, Abak K (2007) Diversity and relationships among Turkish okra germplasm by SRAP and phenotypic marker polymorphism. Biologia (bratisl) 62:41–45. https://doi.org/10.2478/S11756-007-0010-Y/MACHINEREADABLECITATION/RIS
Haq I, Khan AA, Azmat MA (2013) Assessment of genetic diversity in okra (Abelmoschus esculentus L.) using rapd markers. Pakistan J Agric Sci 50:655–662
International B (2007) Developing crop descriptor lists: guidelines for developers
Japda AR, Dhruve JJ (2018) Assessment of genetic diversity of okra (Abelmoschus esculentus L.) for yvmv using rapd and ssr markers
Jhanji S, Kha S, Sanjit Bauri DF, et al (2021) SSR marker development in Abelmoschus esculentus (L.) Moench using transcriptome sequencing and genetic diversity studies. J Hortic Sci 16:206–214. https://doi.org/10.24154/jhs.v16i2.1251
Judd WS (Walter S, Campbell CS, Kellogg EA (2008) Plant systematics : a phylogenetic approach. 611
Kumar S, Parekh MJ, Fougat RS et al (2017) Assessment of genetic diversity among okra genotypes using SSR markers. J Plant Biochem Biotechnol 26:172–178. https://doi.org/10.1007/S13562-016-0378-2
Martinello GE, Leal NR, Amaral AT, et al (2001) Comparison of morphological characteristics and rapd for estimating genetic diversity in abelmoschus spp.: Comparaison entre caracteres morphologiques et marqueurs rapd pour l’evaluation de la diversite genetique dans le genre abelmoschus. Acta Hortic 546:101–104. https://doi.org/10.17660/ACTAHORTIC.2001.546.7
Massucato LR, Nakamura KK, Ruas PM et al (2019) Genetic diversity among Brazilian okra landraces detected by morphoagronomic and molecular descriptors. Acta Sci Agron 42:e43426. https://doi.org/10.4025/ACTASCIAGRON.V42I1.43426
Mkhabela SS, Shimelis H, Gerrano AS, et al (2022) Characterization of okra (Abelmoschus esculentus L.) accessions with variable drought tolerance through simple sequence repeat markers and phenotypic traits. Divers 14:747. https://doi.org/10.3390/D14090747
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590. https://doi.org/10.1093/GENETICS/89.3.583
Ojo DK. Morphological classification of genetic diversity in cultivated okra, Abelmoschus esculentus (L.) Moench using principal component analysis (PCA) and single linkage cluster analysis (SLCA). AFRICAN J Biotechnol
Patil P, Sutar S, John JK et al (2015) A systematic review of the genus Abelmoschus (Malvaceae). Rheedea 25:14–30
Powell W, Morgante M, Andre C et al (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238. https://doi.org/10.1007/BF00564200
Prakash K, Pitchaimuthu M, Ravishankar KV (2011) Assessment of genetic relatedness among okra genotypes Abelmoschus esculentus (L.) Moench using rapd markers. Electron J Plant Breed 2:80–86
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112. https://doi.org/10.1007/S001220051046
Punitha D, Raveendran TS (2004) DNA fingerprinting studies in coloured cotton genotypes. Plant Breed 123:101–103. https://doi.org/10.1046/J.0179-9541.2003.00921.X
Ravishankar KV, Muthaiah G, Mottaiyan P, Gundale S, kumar, (2018) Identification of novel microsatellite markers in okra (Abelmoschus esculentus (L.) Moench) through next-generation sequencing and their utilization in analysis of genetic relatedness studies and cross-species transferability. J Genet 97:39–47. https://doi.org/10.1007/S12041-018-0893-0
Salameh NM (2014) Genetic diversity of okra (Abelmoschus esculentus L.) genotypes from different agro-ecological regions revealed by amplified fragment length polymorphism analysis. Am J Appl Sci 11:1157–1163. https://doi.org/10.3844/ajassp.2014.1157.1163
Sawadogo M, Ouedraogo J, Balma D, et al (2009) The use of cross species SSR primers to study genetic diversity of okra from Burkina Faso. African J Biotechnol 8. https://doi.org/10.4314/AJB.V8I11.60730
Schafleitner R, Lin CY, Lin YP, et al (2021) The world vegetable center okra (Abelmoschus esculentus) core collection as a source for flooding stress tolerance traits for breeding. Agric 11:165. https://doi.org/10.3390/AGRICULTURE11020165
Sneath PHA, Sokal RR (1973) Numerical taxonomy: the principles and practice of numerical classification
Welsh J, Mcclelland M (1990) Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res 18:7213. https://doi.org/10.1093/NAR/18.24.7213
FAOSTAT
(1986) Package statistical variate multi-version 3.1 users’ manual kovach computing services
Acknowledgements
The authors are thankful to the Union of Agricultural Work Committees and the Palestinian Agricultural Relief Committees for providing the Okra landraces seeds.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
Yamen Hamdan, Abdalmenem Hawamda and Mazen Salman designed and carried out the study. Yamen Hamdan, Abdalmenem Hawamda carried out the field and laboratory work. Yamen Hamdan and Rezq Basheer-Salimia carried out the statistical analyses and interpretation of the study. Yamen Hamdan and Abdalmenem Hawamda wrote the first draft of the manuscript. Yamen Hamdan wrote the final draft of manuscript. Yamen Hamdan and Mazen Salman reviewed the final draft. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hamdan, Y.A.S., Hawamda, A.I.M., Basheer-Salimia, R. et al. Genetic Diversity Assessment of Palestinian Okra Landraces (Abelmoschus esculentus L.) Through RAPD Marker. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01859-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01859-x