Abstract
Genetic diversity of 45 genotypes were shown in the list of genotypes Azerbaijani durum wheat (Triticum durum Desf.) genotypes were screened using simple sequence repeats (SSRs). These accessions were collected from various bioclimatic regions of Azerbaijan. Out of the used 22 primers, 13 primers showed polymorphism and were selected for the analyses. Among the genotypes under study, 31 alleles were detected. The highest number of alleles was detected in locus gwm 335 (on chromosome 5B) and on locus gwm 445 (on chromosome 2A) with 5 and 4 alleles, respectively. The lowest number of alleles was found in locus gwm 617 with only 1 allele. For A, B, and D genomes, the total number of alleles detected was 14, 15, and 2, respectively. PIC value between studied SSR markers was 0.912 and this result shows high genetic diversity between Azerbaijani durum wheat genotypes. Therefore, these primers can be recommended for studying the genetic diversity of Azerbaijani durum wheat accessions. The genetic structure of the genotypes was analyzed and the Unweighted Pair Group Method with Arithmetic Mean (UPGMA) dendrogram revealed five major clusters using Nei genetic distance index. The results revealed that SSR markers can efficiently evaluate genetic variation in the wheat samples. Based on the previous characterization of the drought response of these genotypes, links could be established between the SSR markers and drought tolerance. If some of the SSR markers are confirmed for their association with drought tolerance, then, they can be used as markers for the identification of drought-tolerant cultivars needed to enhance wheat productivity for farmers dealing with harsh conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Durum wheat is a high-value cereal crop primarily used for pasta production. Expansion of durum wheat crop production is currently limited, largely due to many biotic and abiotic stress factors, including heat, drought, mineral deficiency, soil salinity etc. Drought is one of the major threats to agricultural crop production including durum wheat in dry land. Plant growth, development, and yield are markedly reduced by the adverse effects of water deficiency. The growth reduction of durum wheat is also believed to be caused by impairing the physiological activities of mitochondria (Flagella et al. 2006). Natural genetic variation exists for drought tolerance within tetraploid wheat species which could be exploited for the improvement of drought tolerance (Munnis and Richards 2007). Breeding for drought tolerance includes screening of wheat germplasm to identify superior genotypes that could be used for the improvement of existing cultivars. The wild relatives and landraces offer the potential of additional sources of drought tolerance due to their adaptation to a wide range of environments and their rich genetic diversity (Colmer et al. 2006).
Conventional breeding relying on phenotypic selection is limited in its ability to accurately and effectively select for drought tolerance due to the complex nature of the trait. Molecular techniques such as molecular markers can be used to reveal sites of variation in DNA between individuals (Langridge and Chalmers 2005). A number of molecular marker technologies are routinely used in generating genetic linkage maps for both tetraploid and hexaploid wheat such as, simple sequence repeats (SSRs) (Roder et al. 1998). SSRs are short tandem repeats of nucleotide units (2–5 bp) that are codominant, abundant, and well dispersed in the genome (Lapitanz 1992; Tautz and Schlotterer 1994). SSR polymorphism can be detected due to variation in the number of these repeat units between the DNA samples. High-throughput detection methods have been established including Multiplex Ready Technology (MRT) (Hayden et al. 2008), which enables simultaneous detection of multiple SSR loci in an economical and efficient manner. To raise the density of microsatellite markers available on a wheat genetic map, Somers et al. (2004) constructed a high-density microsatellite consensus map by mapping common microsatellites on each chromosome in 4 different mapping populations. Microsatellites can be exchanged between laboratories, and are highly transferable between populations (Song et al. 2005; Sun et al. 1998, 2003).
The assessment of genetic diversity is an essential component in the characterization and conservation of germplasm. Within the collections, some of the resources may be genetically redundant or duplicated and therefore not of significance. As conservation and maintenance of the germplasm is an expensive exercise, it is very important to accurately characterise and catalogue germplasm material in Gene bank. Information system of the Plant Genetic Resources (PGR) of Azerbaijan Gene bank plays an important role in the management of genetic resources and in the ongoing activities of the national PGR Network.
Genetic diversity in cultivated wheat has been narrowed due to extensive modern breeding approaches using a limited genetic base. In wheat landraces, diversity has been lost by the reduction in population size. To assist with enlarging the genetic base in breeding and conservation, it is important to assess the diversity present within these wheat landraces. The national breeding program, meanwhile, developed over 100 wheat cultivars, most of which had a significant impact on the Azerbaijan economy. Information on germplasm diversity and genetic relationships among cultivars are critical in wheat improvement. Genetic similarities might be evaluated using pedigree analysis (Barrett et al. 1998; Manifesto et al. 2005) or by assessing morphological traits (Schut et al. 1997) as well as biochemical (Metakovsky and Branlard 1998; Blanco et al. 2012) or, more recently, DNA markers (Plaschke et al. 1995; Roy et al. 2002; Burkhamer et al. 1997; Oman et al. 2001; Ogbonnaya et al. 2006; Rizza et al. 2012).
Germplasm characterization in the Gene bank of Genetic Resources Institute of Azerbaijan has been based mostly on morphological descriptors. However, the use of morphological descriptors, presents some limitations, such as limited polymorphism which would require a high number of descriptors, and potential environmental influence on the phenotype. Limited germplasm characterization of plant accessions conserved in Gene bank is a major cause for the limited use of accessions in breeding programs. High throughput genotyping of wheat germplasm accessions allows examination of genetic relationships and sampling of core collections representatives and reveals allelic richness of the Gene bank accessions.
So far, the genetic diversity of Azerbaijan-grown wheat cultivars are not completely studied with genetic markers. The objectives of this work were therefore to (i) characterize by SSR markers durum wheat cultivars, landraces, and cultivars grown in Azerbaijan; (ii) determine the existence of cultivar-specific amplicons; and (iii) assess the relative informativeness of SSRs.
Materials and methods
Plant material
A total of 45 durum wheat genotypes including drought-tolerant, semi-tolerant, and non-tolerant cultivars were used for molecular analysis. All used genotypes are tetraploids (T. durum ssp. durum) and include local cultivars and accesions collected during scientific expeditions. The name/pedigree of cultivars and all information about the genotypes used are presented in Table 1.
Microsatellite assay
A total of 22 wheat microsatellite (SSR) primers originated from two different groups were screened for polymorphism: Gwm primers from Institut fur Pflanzengenetik und Kulturpflanzenforschung (Roder et al. 1997; Pestsova et al. 2000) (Table 2).
DNA extraction and PCR amplification
10 seeds of each genotype were planted in the greenhouse of the Biotechnology laboratory of ICARDA with a mean temperature of 18 °C. After ten days 5–7 cm long piece of fresh leaf material from each plant was cut and the leaf tissues were freeze-dried. Genomic DNA was extracted from fresh leaves with the PVP extraction protocol (F. Ogbonaya group 2006).
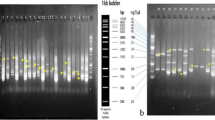
Ten µl of the isolated DNA and 2 µl of 10 × loading dye were loaded in a lane of 1.5% (w/v) agarose gel containing 0.05 µg ml–1 ethidium bromide, for checking the quality of DNA.
For quantitative measurements, a charge-coupled device camera imaging system UVI soft analysis, and Quantity one software were used to capture the image and to calculate the band intensities.
DNA quantification was studied as well with comparison of lambda DNA (10 ng/µl) and different dilution type of DNA (10 µl DNA:2 µl loading dye; 1/2DNA:2 µl loading dye; 1/4DNA:2 µl loading dye; 1/5DNA:2 µl loading dye).
PCR reaction mix was prepared corresponding to the protocol: 4.8 µl H2O, 1 µl 10× buffer, 1 µl dntps, 0.5 µl Primer F and 0.5 µl Primer R, 0.1 µl Taq, and 2 µl DNA per reaction.
Amplifications were performed for each primer pair separately in microtiter plates using the following programs: 5 min at 94 °C, followed by 30 cycles of 30 s at 94 °C, 1 min annealing (between 50 and 62 °C, depending on the published optimal annealing temperature of the primer), and 30 s at 72 °C, and a final extension of 10 min at 72 °C.
Gel documentation and Data analysis
PCR products were separated on 8% PAGE gel, stained in ethidium bromide and vizualized, UVI soft analysis was used to capture the image and to calculate the band intensities. Cluster analysis demonstrating the genetic relationship between genotypes was generated using a simple matching coefficient and Unweighted Pair Group Method Using Arithmetic Averages (UPGMA) and Nei genetic distance. The results are presented graphically in dendrograms.
After gel documentation, results were scored for the construction of cluster analysis using the Power Marker software program.
Polymorphism informative content (PIC)—PIC value is one of the basic steps in determining genetic diversity of genotypes by markers and helps to distinguish molecular differences. The PIC concept is used to measure the effectiveness of a marker and its informativeness in a given population. PIC value was calculated using formula:
where Pij is the frequency of the jth allele.
Results and discussion
The research aimed to investigate the genetic polymorphism of a mapping population of durum wheat obtained from Gene bank accessions of The Genetic Resources Institute of the Ministry of Science and Education and included land races, research material, unreleased breeder material, and local cultivars. To assess genetic distance among 45 durum wheat genotypes 17 microsatellite markers were used out of which 13 makers were polymorphic. Markers gwm 6, gwm 368, gwm 369, gwm 518 didn’t show polymorphism in our experiments. The number of alleles per locus ranged from 1 (gwm 617) to 5 (gwm 335) (Table 3).
Collection Sites of Genebank accessions and Climatic zones of Azerbaijan
Seed material was collected by expeditions from different regions of Azerbaijan. To analyze the relationship between cluster groups we need to collect information about collection sites. As it is known Azerbaijan has multiform climatic conditions, 9 out of the 11 climate patterns in the Koppel climate classification can be found in Azerbaijan (Fig. 1).
Drought response of genotypes are presented in Table 4 and are derived from the Catalogue of Azerbaijan Research Institute of Crop Husbandry (2013).
Polymorphism analysis of durum wheat accessions
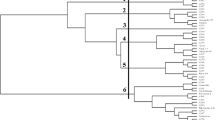
The dendrogram is based on the similarity of genotypes and divides the genotypes into five large groups. The first group is homogenous and consists of genotypes (6112, 6108, and 6104), all belonging to the var. hordeiforme subtaxa and collected from the same collection site 1 which has a dry climate - half deserts and dry steppes with soft winter.
The second group consists of 16 genotypes combining Barakatli-95, Terter, Shirvan, Sharg and, Shiraslan cultivars and 11 land races. Barakatli-95 stood apart from other subgroups but the common feature for all cultivars is their drought response. As shown in Table 2, all cultivars of this cluster group are moderately tolerant to drought tolerant including the cultivar Barakatli-95 (Fig. 2).
The third group can be divided into sub-clusters totaling 16 genotypes: Ag bugda, Vugar, Mirbashir-50, and Garagilchiq-2 cultivars, genotypes 6096, 6097 and 6095 (unreleased research material) and 9 land races. This group included both highly tolerant cultivars Mirbashir-50 and Garagilchig-2 and moderately tolerant Vugar. Research germplasm having similar electrophoresis profile with drought tolerant cultivars and therefore can be characterized as genotypes with high drought tolerance. The land races of the third group were collected from collection sites 1 and 3 (half deserts and dry steppes with cold winter and dry hot climate). High genetic similarity between genotypes in this group can be related to the climate features of their collection sites. Most of the genotypes originated from collection site-1 and site-3. These two collection sites are geographically distant but belong to the same climate classification zone with moderately warm climate and dry winter. The 4th cluster group consists of five land races and the cultivar Mugam which has low drought tolerance. Land races of this cluster group are polymorphic according to their collection sites as collected from sites 3, 4, and 5, all characterized for their moderately warm climate with dry summer. These sites are in between of sites with dry hot climates (sites 1 and 2) and sites with cold climates (5, 6, 7, 8). This can explain the genetic similarity of land races from these sites and the cultivar Mugan. The 5th group is combining research material 6089 and 6105 and three landraces collected from sites 1 and 2.
This study determined the genetic diversity among 45 durum wheat genotypes. The thirteen markers amplified a total 31 allels. The highest numbers of alleles were detected for locus gwm 335 (on chromosome 5B) and locus gwm 445 (on chromosome 2A) with 5 and 4 alleles respectively. The lowest number of alleles was determined in locus gwm 617 with only 1 allele. For A, B, and D genomes, the total number of alleles detected was 14, 15, and 2, respectively.
The PIC value between studied SSR markers was 0.912 and shows high genetic diversity between studied durum wheat genotypes. Therefore, these primers can be recommended for the genetic evaluation of Azerbaijan wheat accessions. Khavarinejad and Karimov (2012) detected 33 and 17 polymorphic alleles for spring wheat genotypes by RAPD and SSR markers, respectively. Xgwm 120-2B had the most PIC with 57%. The obtained results showed an average of 3.4 polymorphic bands per primer for SSR.
Research conducted to assess in formativeness and efficiency of 26 different molecular markers for genetic diversity among 40 durum wheat determined 74% of the diversity. A high level of polymorphism was recorded for SSR markers used in the study. A total of 136 fragments were obtained from the 26 SSR primers and all the bands were polymorphic across all the genotypes screened. The PIC values ranged from 38 to 94% with an average of 74% (Ratiba et al. 2012). In the study conducted by Khavarnejad (2013) polymorphism percentage was detected by UBC 350 and UBC 109 markers with values 0.53 and 0.50 respectively using RAPD and concluded that RAPD could detect more polymorphism alleles than SSRs.
In our experiments more differentiations were observed between accessions in populations originating from different climatic regions of Azerbaijan. Similar results were observed in experiments conducted to study the genetic diversity of Ethiopian durum wheat landrace collections. The genetic diversity of 141 Ethiopian durum wheat landrace collections and 19 cultivars was analyzed using 12 SSR primers. Based on the analysis of molecular variance, a lower level (19%) of variation was observed among populations and a higher level (81%) within the population (Asmamaw et al. 2019).
Breeding programs focused on morphological traits also can affect diversity among wheat accessions. morphological characterization and genetic diversity analysis of Tunisian durum wheat accessions revealed high genetic variation within regions (81%) and within genetic groups (41%), reflecting a considerable amount of admixture between landraces. The moderate (19%) and high (59%) levels of genetic variation detected among regions and genetic groups, respectively, highlighted the selection practices of local farmers (Ouaja et al. 2021).
Christov et al. (2022) studied of population structure in combination with genetic variation has big importance for the improvement of breeding programs under drought-prone environmental conditions. The genetic variation and population structure of a panel of 90 durum wheat consisting of 62 cultivars and breeding lines originating from two agro-ecological zones in Bulgaria (Northern and Southern Bulgaria) and 28 introduced cultivars from Southwestern, Central, and Eastern Europe, and the USA showed a good relationship between the genetic and phenotypic population structures according to the division of the genotypes by their agro-geographical origin.
Dagnaw et al. (2023) studied 104 Ethiopian durum wheat genotypes representing thirteen populations, three regions, and four altitudinal classes for their genetic diversity, using 10 grain quality- and grain yield-related phenotypic traits and 14 simple sequence repeat (SSR) makers. Analysis of molecular variance (AMOVA) revealed a high level of variation within populations, regions, and altitudinal classes, accounting for 88%, 97%, and 97% of the total variation, respectively.
Summarizing the characterization results of 45 durum wheat genotypes we conclude that genotypes having similar profiles with drought-tolerant cultivars Mirbashir-50 and Garagilchig-2 can be characterized as genotypes with high tolerance to drought. Most of the land races of this cluster group were collected from half deserts and dry steppes prevailing in the collection site 1, which could explain the importance of natural selection of landraces and their adaptation to drought. Genetic similarity of genotypes can be explained by the presence of common alleles probably related to drought resistance. More SSR markers will be needed to further confirm the tolerance to drought of the genotypes in this group.
The research findings may be useful for the improvement of drought tolerance in durum wheat crop cultivars.
Data availability
The datasets analyzed during the current study are available from the corresponding author – Ruhangiz Mammadova upon request.
References
Asmamaw M, Keneni G, Tesfaye K (2019) Genetic diversity of Ethiopian durum wheat (Triticum durum Desf.) landrace collections as revealed by SSR markers. Adv Crop Sci Technol 7:413
Barrett BA, Kidwell KK, Fox PN (1998) Comparison of AFLP and pedigree-based genetic diversity assessment methods using wheat cultivar from the Pacific Northwest. Crop Sci 38:1271–1278
Blanco A, Mangini G, Giancaspro A, Giove S, Colasuonno P, Simeone R, Signorile A, Vita P, Mastrangelo M, Cattivelli L, Gadaleta A (2012) Relationships between grain protein content and grain yield components through quantitative trait locus analyses in a recombinant inbred line population derived from two elite durum wheat cultivars. Mol Breed 30:79–92. https://doi.org/10.1007/s11032-011-9600-z
Burkhamer RL, Lanning SP, Martens RJ, Martin JM, Kim HS, Ward RW (1997) Genetic diversity in Eastern U.S. soft winter wheat (Triticum aestivum L. em. Thell.) based on RFLPs and coefficient of parentage. Theor Appl Genet 94:472–479
Catalogue of Cereal and Legume Varieties (2013) Ministry of Agriculture of the Republic of Azerbaijan Agrarian Science Center. Research Institute of Crop Husbandry, Baku, p 295
Christov NK, Tsonev S, Dragov R, Taneva K, Violeta Bozhanova & Elena Georgieva Todorovska (2022) Genetic diversity and population structure of modern Bulgarian and foreign durum wheat based on microsatellite and agronomic data. Biotechnol Biotechnol Equip 36(1):637–652
Colmer TD, Flowers TJ, Munns R (2006) Use of wild relatives to improve salt tolerance in wheat. J Exp Bot 57:1059–1078
Dagnaw T, Mulugeta B, Haileselassie T, Geleta M, Ortiz R, Tesfaye K (2023) Genetic diversity of durum wheat (Triticum turgidum L. ssp. durum, desf) germplasm as revealed by morphological and SSR markers. Genes 14:1155
Flagella Z, Trono D, Pompa M, Di Fonzo N, Pastore D (2006) Seawater stress applied at germination affects mitochondrial function in durum wheat (Triticum durum) early seedlings. Funct Plant Biol 33:357–366
Hayden MJ, Nguyen TM, Waterman A, McMichael GL, Chalmers KJ (2008) Application of multiplex-ready PCR for fluorescence based SSR genotyping in barley and wheat. Mol Breed 21:271–278
Khavarinejad MS, Karimov M (2012) Assessment of genetic diversity in wheat spring genotypes by molecular markers in Northern Iran. Afr J Biotechnol 11(82):14724–14731
Khavarinejad MS, Karimov M, Samadi M (2013) Evaluation of RAPD and SSR molecular markers in Bread wheat genotypes. Int J Plant Anim Environ Sci 3(4):131–139
Langridge P, Chalmers K (2005) The principle: identification and application of molecular markers. In: Lorz H, Wenzel (eds) Molecular marker systems in plant breeding and crop improvement. Springer, Berlin, pp 3–22
Lapitanz NLV (1992) Organization and evolution of higher plant nuclear genomes. Genome 35:171–181
Manifesto MM, Schlatter AR, Hopp HE, Suarez EY, Pagnotta M, Mondini L, Atallah M (2005) Morphological and molecular characterization of Italian emmer wheat accessions. Euphytica 146:29–37
Metakovsky EV, Branlard G (1998) Genetic diversity of french common wheat germplasm based on gliadin alleles. Theor Appl Genet 96:209–218
Munnis R, Richards RA (2007) Recent advances in breeding wheat for drought and salt stresses. In: Jenks MA, Hasegawa PM, Jain SM (eds) Advances in molecular breeding toward drought and salt tolerant crops. Springer, Netherlands, pp 565–585
Ogbonnaya FC, Imtiaz M, Hearnden P, Wilson J, Eastwood RF (2006) Identification of novel gene for seed dormancy in wheat. In: Proceedings of the 13th Australasian plant breeding conference, Christchurch, New Zealand
Oman J, Ogbonnaya FC, Matassa V, Eastwood RF (2001) Comparison of dormancy between a synthetic hexaploid bread wheat and its tetraploid and diploid progenitors. In: Proceedings of the 10th wheat breeding assembly, Mildura, Australia, pp 205–208
Ouaja M et al (2021) Morphological characterization and genetic diversity analysis of tunisian durum wheat (Triticum turgidum var. durum) accessions. BMC Genomic Data 22:1–7
Pestsova E, Ganal MW, Roder MS (2000) Isolation and mapping of microsatellite markers specific for the d-genome of bread wheat. Genome 43(4):689–697
Plaschke J, Ganal MW, Roder MS (1995) Detection of genetic diversity in closely related bread wheat using microsatellite markers. Theor Appl Genet 91:1001–1007
Ratiba B, Baum M, Djekoune A, Labadidi S, Djighly A, Benbelkacem K, Labhilili M, Gaboun F, Ykhlef N (2012) Screening for drought tolerance using molecular markers and phenotypic diversity in durum wheat genotypes. World Appl Sci J 16(9):1219–1226
Rizza F, J.Ghashghaie S, Meyer L, Matteu A, Mastrangelo FB (2012) Constitutive differences in water use efficiency between two durum wheat cultivars. Field Crops Res 120:49–60. https://doi.org/10.1016/j.fcr.2011.09.001
Roder MS, Worland AJ, Borner A (1997) Abundance, variability and chromosomal location chromosomal mapping of the genes for dwarfing (Rht12) and of microsatellites in wheat. Mol Gen Genet 246:327–333
Roder NS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149(4):2007–2023.
Roy JK, Balyan HS, Prasad M, Gupta PK (2002) Use of SAMPL for a study of polymorphism, genetic diversity and possible gene tagging in bread wheat. Theor Appl Genet 104:465–472
Schut JW, Oi X, Stam P (1997) Association between relationship measures based on AFLP markers, pedigree data and morphological traits in barley. Theor Appl Genet 95:1161–1168
Somers JD, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114. https://doi.org/10.1007/s00122-004-1740-7
Song GJ, Shi JR, Singh S, Fickus EW, Costa JM, Lewis J, Gill BS, Ward R, Cregan PB (2005) Development and mapping of microsatellite (SSR) markers in wheat. Theor Appl Genet 110:550–560. https://doi.org/10.1007/s00122-004-1871-x
Sun G, Bond M, Nass H, Martin R, Dong Z (2003) RAPD polymorphisms in spring wheat cultivar and lines with different level of Fusarium resistance. Theor Appl Genet 106:1059–1067
Sun Q, Ni Z, Liu Z, Cao J, Huang T (1998) Genetic relationships and diversity among tibetan wheat, common wheat and european spelt wheat revealed by RAPD markers. Euphytica 99:205–211
Tautz D, Sclotterer C (1994) Simple sequences. Curr Opin Genet Dev 4:832–837
Acknowledgements
We thank the Department of Biotechnology of ICARDA (International Center of Agricultural Research in the Dry Areas) for all support in conducting the experiments. The study presented in this paper is a part of project supported by the Japan International Cooperation Agency (JICA).
Funding
This work was supported by the Japan International Cooperation Agency (JICA) and the International Center for Agricultural Research in the Dry Areas (ICARDA). Author Ruhangiz Mammadova has received travel and research support from JICA during a specialized long-term individualized training course on “Biotechnology Application in Crop Improvement” held under the third country training program (TCTP) at ICARDA headquarters.
The authors declare that no other funds, grants, or other support was received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation was performed by ZA and NA, phenotypic data collection was performed by FS, EH, and MA, experiment design and molecular analysis was performed by RM, FYA and AA. The first draft of the manuscript was written by RM and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Financial interests: author Ruhangiz Mammadova, Ahmed Amri, Zeynal Akparov, Fida Yossef Alo, Fatma Shekhzamanova, Mehraj Abbasov, Nurlan Amrahov, Elchin Hajiyev, Shader Alizade declare they have no financial interests. Author Ruhangiz Mammadova received training and travel support from the Japan international cooperation agency (JICA) in 2011. The experimental part of the work was implemented during the outlined long-term training program.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mammadova, R., Amri, A., Akparov, Z. et al. Evaluation of genetic diversity using SSR markers and link with drought response of Azerbaijani durum wheat (Triticum durum Desf.) genotypes. Genet Resour Crop Evol 71, 2203–2212 (2024). https://doi.org/10.1007/s10722-023-01763-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01763-w