Abstract
Croton grewioides Baill. is a medicinal and aromatic species with proven biological activity. Considering the economic potential of the species, the use of molecular markers for genetic diversity studies is of utmost importance to develop viable strategies for its management, conservation, and use. Thus, the aim of this study was to evaluate the genetic diversity and structure of natural populations of C. grewioides using SNP markers, and select genotypes to compose the species collection in an Active Germplasm Bank. Genotypes were sampled from two populations, the first in the state of Sergipe (SE) and the second in the state of Bahia (BA). A total of 6,942 high quality SNPs were identified through genotyping-by-sequencing (GBS). Genetic diversity was estimated by observed and expected heterozygosities (Ho: 0.211 < He: 0.225), the Shannon Index (I: 0.357), inbreeding coefficient (f: 0.056), and number of exclusive alleles (BA: 1296 and SE: 799). Analysis of molecular variance (AMOVA) showed greater genetic variation within the populations. The multivariate analyses performed (Discriminant analysis of principal components—DAPC; bayesiana—Structure; Principal Components Analysis—PCA; and the dendrogram based on Rogers’ genetic distance) allowed separation of the genotypes according to the population of origin. Thirteen (13) genotypes from the SE population and 17 from the BA population were selected to compose the species collection. The selection of genotypes coming from the two populations studied is fundamental for effective conservation of the genetic variability detected.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diverse sectors of the economy have intensified their search for products derived from plants. In the medicinal sector alone, 25% of the global production of medicinal drugs is (directly or indirectly) plant based (Newman and Cragg 2020). The increasing search for alternative products is spurred by rising resistance in pathogenic microorganisms, reducing or nullifying the efficiency of medications (Medeiros et al. 2017). Similarly, in the agricultural sector, the development of resistance in plant pathogens, as well as the concern for food safety through the absence and/or reduction of residues in food, has intensified the search for bioproducts (Chojnacka 2015; Li et al. 2019).
The increase in the use of plants with bioactive properties, together with overexploitation, the pressure of anthropization, and the modification of natural environments, compromises the sustainable use and conservation of these species (Costa et al. 2015; Chen et al. 2016; Kadam and Pawar 2020; Silva-Júnior et al. 2020). Under these conditions, they are subject to reduction in their genetic base, to genetic imbalance of their populations, and to variations in effective population size (Santos et al. 2017), making adoption of conservation strategies essential. The implementation of Active Germplasm Banks (AGBs), conservation units of genetic material, is an alternative for the conservation of the genetic variability of a species outside of its natural location of occurrence (Engel and Visser 2003; Gulati 2018).
The selection of genotypes of interest that represent the genetic variability of a species should be performed based on parameters that estimate the genetic diversity and structure of its populations (Almeida-Pereira et al. 2017; White et al. 2018; Motta et al. 2019; Niu et al. 2019). For that purpose, single nucleotide polymorphism (SNP) markers have been successfully applied in various plant species (Otto et al. 2017). The SNPs result from a specific modification due to interchange of a single base (Grover and Sharma 2016). As they are abundant and widely distributed in the genomes (Ganal et al. 2009; Kethom et al. 2019), they allow more accurate estimation of genetic diversity in natural populations (Chung et al. 2017; Nadeem et al. 2018; Kethom et al. 2019), even in non-model species, such as medicinal and aromatic plants (Wang et al. 2016; Otto et al. 2017; Marakli 2018; Do et al. 2019; Brito et al. 2021).
Medicinal and aromatic plants produce secondary metabolites, such as essential oils, that can be composed of an immense variety of chemical compounds of considerable agronomic and medicinal interest (Trindade and Lameira 2014; Santos et al. 2016). Euphorbiaceae is one of the largest and most diverse families of angiosperms and, in Brazil, it is made up of more than 1000 species and 64 genera. The Croton genus is prominent with about 1.200 species, distributed predominantly in the Americas (Silva et al. 2009), and with more than 300 species in Brazil (Caruzo et al. 2020), some with proven biological activities, such as anti-inflammatory (Kim et al. 2015), anticarcinogenic (Sampath et al. 2018), antioxidant (Embuscado 2015), antiprotozoal (Sharma et al. 2017), antifungal (Monteiro et al. 2017; Hailu et al. 2017), and antibacterial (Marchese et al. 2017) properties.
Croton grewioides Baill., popularly known as “alecrim-de-cabocla” or “canelinha” (SiBBr, 2020), is a plant native of Bolivia, Brazil and Peru (Riina et al. 2021). In Brazil, it occours in Caatinga, Cerrado, and Atlantic Forest biomes, with distribution in the states of Alagoas, Bahia, Ceará, Goiás, Minas Gerais, Paraíba, Pernambuco, Piauí, Rio Grande do Norte e Sergipe (Silva et al. 2009; Caruzo et al. 2020; SiBBr 2020). It is a diploid species (2n = 20) (Santos, 2016), monoecious, and has a racemiform inflorescence measuring between 1.3 and 5 cm in length with staminate and pistillate flowers, which are often visited by bees (Silva et al. 2009). It presents shrub growth habit (0.7–2 m), frequently found on rocky and sandy-clay outcrops in shrub and tree Caatinga ecoregions. It is widely used in popular medicine for relief of gastric and stomach problems and intestinal disorders, for cancer treatment, and for fever-reducing baths (Trindade and Lameira 2014). Thus, it has significant biological potential, with some antioxidant (Oliveira et al. 2021), acaricidal (Castro et al. 2019), antibacterial (Medeiros et al. 2017), and insecticidal (Camara 2008; Silva et al. 2008) activities already proven.
Considering the importance of the species and its agronomic and medicinal potential, due to the diversity of compounds it contains, the expectation for this study is to contribute information for designing strategies for conservation and use of this genetic resource. This study was conducted with the aim of studying the genetic diversity and structure of natural populations of C. grewioides in the Brazilian states of Bahia and Sergipe and selecting genotypes to compose the species collection in the AGB of Medicinal and Aromatic Plants of the Universidade Federal de Sergipe. To that end, SNP markers identified through the genotyping-by-sequencing (GBS) technology were used.
Materials and methods
Collection of plant material
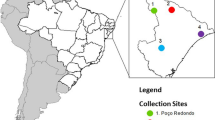
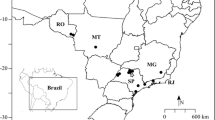
Plant material was surveyed and collection was made of 40 genotypes of C. grewioides Baill. from the states of Bahia (23 genotypes—BA population), in the municipality of Conceição do Coité, and Sergipe (17 genotypes—SE population), in the municipalities of Poço Verde and Poço Redondo.
The municipality of Conceição do Coité (11°35′78.4″ S, 39°18′99.6″ W) is in the northeast region of the state of Bahia, at 420 m above sea level. It has a semi-arid climate, with sandy-clay soils and mean temperature of 22.3 °C. The rainy period is from November to April, with mean annual rainfall of 585 mm (Souza et al. 2003). Poço Verde (10°55′17.8″ S, 37°06′04.1″ W) is in the extreme southwest of the state of Sergipe, at 273 m above sea level. It has a semi-arid climate, shallow sandy-clay soils with a rocky outcropping base, and high salinity. The rainy period is from March to July, with mean annual temperature between 25–26 °C and mean annual rainfall of 646.1 mm (Oliveira et al. 2016). The municipality of Poço Redondo (09°58′06.5″ S, 37°51′48.4″ W) is in the northwest of the state of Sergipe, at 225 m above sea level. It has a semi-arid climate, with shallow, rocky, and dry soils and rocky outcrops. The rainy season is from March to July, with mean temperature of 25.2 °C, and mean annual rainfall of 552.0 mm (Diniz et al. 2014; Bitencurti et al. 2017).
New fresh leaves were collected of each genotype, wrapped in sterile gauze, and placed in ice to prevent oxidation. The samples were stored at -80 °C, then freeze dried in a Terroni LS3000 series bench-top freeze dryer, and stored in a freezer until DNA extraction. The project is registered in the National System of Management of Genetic Patrimony and Associated Traditional Knowledge (Sistema Nacional de Gestão do Patrimônio Genético e Conhecimentos Tradicionais Associados—SISGEN) under number A8CCB3B.
DNA isolation, preparation, and sequencing of the genomic library
Genomic DNA was extracted from C. grewioides leaves through the extraction protocol using the 2% CTAB (cetrimonium bromide) buffer (Ferreira and Grattapaglia, 1998) in the Genetic Diversity and Plant Breeding Laboratory, Genetics Department of ESALQ-USP, Piracicaba, São Paulo, Brazil.
The genomic DNA of each genotype was quantified through electrophoresis in 1% (w/v) agarose gel and stained with SYBR Safe™ DNA (Invitrogen), comparing the DNA with the phage lambda molecular size standards (Invitrogen). After that, the DNA samples were standardized to a concentration of 50 ng/µl for preparation of the GBS (genotyping-by-sequencing) library, following the double digestion protocol, as described by Poland et al. (2012).
The genomic DNA of each genotype was initially digested with two restriction enzymes (Pst1 and Mse1), selected after tests with different combinations of restriction enzymes. The fragments generated of each sample were linked to individual adaptors (bar code) (Sigma-Aldrich, St. Louis, MO, USA) and to specific adaptors for the Illumina technology (Illumina, Certification Program, CSPro). The ligation products were joined in a single pool (multiplex), which was purified using the QIAquick PCR® purification kit, and amplified by PCR, for enrichment of the fragments with both the restriction sites. After that step, the library was once more purified, validated, and quantified in an Invitrogen™ Qubit v 3.0 fluorometer (Thermo Fisher Scientific, Carlsbad, CA, USA). After that, the samples were sequenced on the platform HiSeq3000 (Illumina, Inc., San Diego, CA, USA) with single-end configuration (1 × 100 bp).
Processing of raw sequences and filtering for identification of SNP markers
The sequences obtained were processed on the Stacks program (Catchen et al. 2013) using a pipeline for species that do not have a reference genome. First, the sequences were demultiplexed, using the process_radtags module, and low-quality sequences were removed at the same time. The ustacks module was used to align the sequences and to form stacks of similar sequences for identification of loci with polymorphic nucleotides. The parameters used in this process were minimum depth (− m) = 3 and the maximum distance allowed between stacks to include them in the same locus (− M) = 2. The loci of each sample were grouped in a catalog using the cstacks module, allowing a maximum distance of two nucleotides (− n = 2) between the loci. The sstacks module was used to compare the groups of stacks of each genotype with the catalog. After that, the rxstacks module was used to perform corrections of the individual genotypes through removal of loci with low probability (− ln lim − 10). Finally, the populations module was used to filter the SNP markers using the following parameters: minimum allele frequency (MAF ≥ 0.05), minimum depth of the stack (m ≥ 3), and retention of a single marker per tag, occurring in at least 75% of the individuals in each population.
Genome diversity and structure
The estimate of genetic diversity of the C. grewioides populations was obtained through the observed (Ho) and expected (He) heterozygosities, the Shannon index (I), the inbreeding coefficient (f) of Wright (1965), and the number of exclusive alleles.
The analysis of molecular variance (AMOVA) was used to verify the distribution of genetic variance within and between the populations sampled, and its significance was tested with 10,000 permutations. The genetic structure between the populations was evaluated by the FST statistic of Weir and Cockerham (1984) and its significance was tested through 10,000 bootstraps. Analyses were performed through the GenAlEx program (Peakall and Smouse 2012).
The genomic structure, based on variation of the SNP markers, was evaluated by Discriminant Analysis of Principal Components (DAPC, Jombart et al. 2010) and by Bayesian analysis (Structure software). The DAPC was carried out with the adegenet 2.0.0 package (Jombart and Ahmed 2011) for R (R Core Team 2016). This analysis is free of genetic assumptions, such as the Hardy–Weinberg linkage equilibrium within the groups (Jombart et al. 2010). The DAPC maximizes the difference between the groups and minimizes the difference within them (Roullier et al. 2013). From the information on discriminant functions, a bar graph (Compoplot) was generated, with the probabilities of attribution of each individual to the clusters formed (Jombart and Ahmed 2011).
Bayesian analysis was performed using Structure v.2.3.4 software (Pritchard et al. 2000). Values of genetic clusters (k) ranging from 1 to 5 were tested and, for each k, 10 independent replications were performed. Each iteration consisted of a burn-in period of 10,000 iterations, followed by 50,000 MCMC (Markov Chain Monte Carlo) iterations, assuming the admixture ancestry model and alleles with uncorrelated frequency. The number of genetic groups (k) was identified by the ΔK method described by Evanno et al. (2005), implemented in the STRUCTURE harvester software (Earl and Vonholdt, 2011). Accessions that presented membership values below 0.8 were considered to have mixed ancestry.
The genetic divergence among the genotypes was evaluated by means of the genetic distance of Rogers (1972) and visualized through construction of a dendrogram using the UPGMA (Unweighted Pair Group Method with Arithmetic Means) algorithm. Analysis was performed with the assistance of the poppr package (Kamvar et al. 2014) for R (R Core Team 2016). Ten thousand bootstraps were performed to draw inferences regarding the reliability of the clusters. The final dendrogram was formatted through the program FigTree 1.4.1 (http://tree.bio.ed.ac.uk/software/figtree/). Principal component analysis (PCA) was performed using the dudipca function of the ade4 package (Dray and Dufour 2007) for R (R Core Team 2020).
The 30 C. grewioides genotypes selected to compose the species collection in the Active Germplasm Bank of Medicinal and Aromatic Plants were selected with the assistance of the maximum length sub tree function of the DARwin 6.0.010 software (Perrier and Jacquemoud-Collet 2006). This function removes the redundant genotypes and allows selection of those that retain maximum genetic variability. The calculation of genetic distance among the genotypes, necessary for application of the function, was calculated from the simple matching dissimilarity index: \(d_{ij} = 1 - \frac{1}{L}\mathop \sum \limits_{i = 1}^{L} \frac{{m_{i} }}{\pi }\), where \(d_{ij}\) is the dissimilarity between the i and j units; \(L\) is the number of loci; \(m_{i}\) is the number of corresponding alleles for the \(l\) locus; and \(\pi\) is the ploidy. A total of 10,000 bootstraps were performed.
Results
Sequencing of the GBS library resulted in a total of 272,866,599 reads. After quality control, demultiplexing, and filtering out the low-quality sequences, 6942 high-quality SNP markers were identified. These markers were used for characterization of the genetic diversity and structure of the 40 C. grewioides genotypes sampled from the states of Bahia (23 genotypes) and Sergipe (17 genotypes). The mean sequencing depth was 41.39X (± 17.71 sd), and within the SNPs detected, a greater proportion of transitions (55.75%) than transversions (44.25%) was observed. The most frequent mutations were A-G (28.35%) and C-T (27.40%).
The estimates of genetic diversity are shown in Table 1. The observed heterozygosity (\({H}_{o}\)) was 0.207 for the BA population and 0.215 for the SE population. These results were lower than that of expected heterozygosity for the two populations (0.230—BA and 0.219—SE). The inbreeding coefficient of the BA population was higher (0.085) than that of the SE population (0.025), and the Shannon Index was 0.346 for SE and 0.367 for BA. The presence of exclusive Ae alleles was detected in the two populations: 1296 in the BA population and 799 in the SE population.
Separation of genetic variation into two levels, within and between the populations, was performed through AMOVA (Table 2). The largest proportion of the total genetic variation was observed within the populations, 93% and 7% of this variation was detected between the populations analyzed. The estimate of the fixation index (FST) was 0.071, indicating that there is moderate genetic differentiation between the BA and SE populations.
Discriminant Analysis of Principal Components (DAPC) allowed the 40 genotypes analyzed to be divided into two different groups, without overlap between them (Fig. 1a). The same color in different genotypes indicates that they belong to the same group. The first group was composed by the genotypes of the BA population, and the second by the genotypes of the SE population. In addition, the division of the 40 genotypes into two groups could be visualized by the probability of association graph (Compoplot) (Fig. 1b), based on DAPC analysis.
Discriminant Analysis of Principal Components (DAPC) of the 40 Croton grewioides Baill. genotypes sampled in the states of Bahia and Sergipe, Brazil. (a) Density plot considering the first discriminant function. Each bar represents an individual; (b) Bar graph illustrating the probabilities of assignment of each genotype. Each color represents a state (green: Bahia—BA; lilac: Sergipe—SE)
The Bayesian analysis (Fig. 2), performed using the Structure software, also allowed the division of the 40 genotypes studied into two groups. Group I (green) was composed of individuals from the BA population, and group II (purple) was composed of individuals from the SE population, with the exception of individual Cg 33 who presented mixed ancestry (membership values less than 0.8).
The clustering of the 40 genotypes according to their population of origin was also observed with construction of the dendrogram by the UPGMA method, based on Rogers’ genetic distance (Fig. 3). The CGR18 and CGR21 genotypes exhibited greater genetic distance from the other genotypes sampled in the BA population. For the SE population, the CGR32 and CGR37 genotypes exhibited greater genetic distance from the other genotypes sampled. The Principal Component Analysis explained 16.61% of the total variation (Fig. 4) and allowed separating the genotypes according to the population of origin.
Dendrogram with 40 Croton grewioides Baill. genotypes obtained by the UPGMA clustering method based on Rogers’ genetic distance (Rogers, 1972), with 10,000 bootstraps. Each color represents a state (green: Bahia—BA; lilac: Sergipe—SE)
Use of the maximum length sub tree function of the DARwin 6.0.14 software allowed selection of 30 genotypes to compose the C. grewioides collection (Table 3). This function was used to remove the redundant genotypes, with the aim of retaining the largest proportion of the genetic variability detected in the populations analyzed in a smaller number of genotypes. Seventeen (17) individuals from the BA population and 13 individuals from the SE population were selected, which retained 99.38% of the total genetic variability detected in the two populations.
Discussion
The study of the genetic diversity and structure of natural populations is of utmost importance for designing strategies for conservation and use of species that have economic potential, such as C. grewioides Baill. SNP molecular markers have been an ally for carrying out these studies. SNPs are abundant in the genomes of plant species and, together with reduction in costs and increase in the ease of access to genotyping technologies, their use in genetic studies of natural populations has become frequent (Grattapaglia et al. 2011). In addition, advances in molecular techniques and in bioinformatics have allowed large-scale genotyping of species that do not have a reference genome. Thus, this is the first study performed with SNP markers for C. grewioides, based on GBS genomic libraries. Moreover, this is the first study carried out to analyze the genetic diversity and structure through molecular markers for the species.
Transition bias was detected through analysis of the sequences of the C. grewioides genome; that is, the percentage of transition (55.75%) was greater than that of transversion (54.25%). Transition refers to DNA mutation in which there is interchange between the two purines (A and G) or between the two pyrimidines (C and T), whereas transversion is the interchange of a purine for a pyrimidine, or vice-versa (Guo et al. 2017). The transition bias frequently occurs in plants (Stern and Orgogozo 2008), as was observed in the present study and for the species Jatropha curcas L. (Gupta et al. 2012) and Croton tetradenius Baill. (Brito et al. 2021). Two sources of origin have been reported to explain the transition bias. The first refers to the fact that for transition to occur, distortion of the double strand of DNA is less than in transversion; consequently, transition should occur with greater frequency in DNA replication. In addition, deamination, chemical alteration of the nucleotides, leads to transition (Zou and Zhang 2021).
The C. grewioides populations analyzed showed lower genetic diversity than that expected by Hardy–Weinberg equilibrium (population under random crosses), since the observed heterozygosity was lower than the expected heterozygosity. This result is an indication of deficit of heterozygotes in the populations (Frankham 2003; Hartl and Clark 2010), as seen by the estimated value of the inbreeding coefficient (f: 0.056). The deficit in the frequency of heterozygotes may have been brought about by non-random crosses, as a result of crosses between related individuals and/or self-fertilization, or even as a result of the reproductive biology of the species. Monoecious species, such as C. grewioides (Silva et al. 2009), can self-fertilize, resulting in reduction in the heterozygotes in the population (Awad and Billiard 2017; Islam et al. 2020).
The values observed for the Shannon Index (I: 0.346 Bahia; 0.367: Sergipe) in this study were similar to those obtained for C. tetradenius (0.39–0.50), using ISSR markers (Almeida-Pereira et al. 2017); for Croton florisbundus (0.31–0.43), using SSR markers (Sivestrini et al. 2015); and for Croton urucurana Baill. (0.322–0.342) using ISSR markers (Costa et al. 2020). This diversity index shows variation from 0 to 1, and the nearer the value to zero, the lower the genetic diversity of the population under analysis (Botrel et al. 2006; Gois et al. 2014; Silva et al. 2015).
The pattern of distribution of genetic variability within and among populations is the result of the interaction among different factors in the evolution of this distribution. Selection, effective population size, and pollen and seed dispersal are of central importance for the distribution observed (Hamrick 1989). Characteristics intrinsic to the species, such as the reproductive system and floral morphology allow understanding of the genetic structure in natural populations. Allogamous species tend to have high genetic diversity within populations and little interpopulation variation (Hamrick 1989); 10–20% of the genetic variation for these species is found among populations. However, autogamous species can show up to 50% genetic variation among populations (Hamrick and Godt 1989).
From the results of AMOVA for the C. grewioides populations studied, it can be inferred that the species can cross fertilize, given the lower genetic variation between the populations (7%). Similar results were observed for the species C. urucarana (21.72%; Costa et al. 2020) and C. tetradenius (5%; Brito et al. 2021), in which lower genetic variation was observed among the populations than within the populations. The value observed for the FST statistic (0.071) indicates that the populations studied have moderate genetic differentiation (Silva-Júnior et al. 2020apud Wright 1978; Leviyang and Hamilton 2011).
The pollen and seed dispersal pattern (gene flow) of the species is another factor that determines the population genetic structure (Zanella et al. 2012; Gulati 2018; Costa et al. 2020). The presence of exclusive alleles is an indication of restricted gene flow (Seoane et al. 2000) and was observed in the two populations analyzed. Of the total SNP loci evaluated, 18.67% were observed only in the BA population and 11.51% only in the SE population.
The pistillate flowers of the species of the Croton genus have nectar-producing glands around the ovary that attract insects (Thaowetsuwan et al. 2020). Thus, in general, species of the Croton genus are associated with dispersal agents and pollinizers that move in restricted distances, such as ants, flies, wasps, bees, and butterflies (Passos and Ferreira 1996; Lôbo et al. 2011; Nascimento et al. 2014; Silva et al. 2020). In addition, the Croton seeds have a structure called a caruncle, which is rich in lipids (Leal et al. 2007). Ants consume this structure and leave the seeds exposed, generally near the mother plant, collaborating with the spatial genetic structure observed (Vander Wall and Longland 2004), which, for its part, can contribute to crosses between related individuals.
Furthermore, the C. grewioides populations studied are in areas that have marked climatic seasonality and low rainfall, characteristics of arid and semi-arid regions. These conditions favor abiotic vectors (such as autochory and anemochory), very representative in dry periods (Silva and Rodal 2009; Silva et al. 2013; Souza et al. 2013; Quitino and Machado 2014). Pollination through wind, insects, or both have been reported (Dominguez and Bullock 1989; Pires et al. 2004; Quirino and Machado 2014; Costa et al. 2020) for the Croton genus. Autochory was also observed for the species Croton blanchetianus Baill. (Silva and Rodal 2009; Silva et al. 2013), C. florisbundus (Silvestrini et al. 2015), C. heliotropiifolius Kunt (Silva et al. 2013), and Croton rhamnifolioides Pax & K.Hoffin. (Silva and Rodal 2009). Consequently, species of the Croton genus that have this type of dispersal tend to have limited seed spatial dispersal (Passos and Ferreira 1996; Narbona et al. 2005; Silva et al. 2013), and this factor may have contributed to the genetic variability distribution pattern observed among the populations.
The multivariate analyses performed (DAPC, Bayesiana, dendrogram based on Rogers’ genetic distance and PCA) followed the same pattern of division of the 40 genotypes, which were separated according to population of origin. The bar graph, based on DAPC analysis, indicates the probability of association of the genotypes to the different groups (Jombart et al. 2010), suggesting that there is wide agreement between the genetic structure and the locations of origin of the genotypes. Thus, the results of differentiation observed by the multivariate analyses for the populations studied may have been affected by the geographic distance (325 km). In addition, forest fragmentation and unsustainable exploitation of plant resources contribute to reduction in population and, consequently, this can reduce the effectiveness of pollination and seed dispersal by insects and wind (Brito et al. 2021).
The presence of exclusive alleles in both populations corroborates these observations, since populations isolated for a long time can accumulate exclusive alleles, and this is reflected in the genetic divergence observed (Prentice et al. 2003). The moderate genetic differentiation, and the observation of genotypes with mixed ancestry in Bayesian analysis, may portray the historic gene flow between the populations under study, and this therefore suggests that these populations may become more genetically distant over the generations (Bohonak 1999; Pence 2017; Silva-Júnior et al. 2020).
Due to the economic potential of the species, the results obtained in the present study were essential for delineating collection of genotypes to compose the collection of the species in the active germplasm bank, with allelic diversity that represents the populations of origin (Blank et al. 2019). Considering that the genetic variability present in the populations can reflect the chemical variation of the genotypes (Costa et al. 2020), the use of the maximum length sub tree function of the DARwin 6.0.14 software allowed the selection of 30 C. grewioides genotypes that contained the largest proportion of genetic variability detected in the populations analyzed (99.38%).
The conservation of samples that represent the genetic variability of the species is essential for identification and selection of genotypes of interest, which may be used directly or in breeding programs aiming at development of cultivars (Blank 2013; Blank et al. 2015; Souza et al. 2015; Brito et al. 2016). Combining molecular, morphological, and chemical data is of utmost importance for efficient use of genetic resources selected to compose the species collection in the AGB.
Conclusion
Considering the economic potential of C. grewioides, the results of genetic diversity and structure obtained in this study provide essential information for designing strategies for conservation and use of the species. The greater proportion of genetic variation detected within the populations and moderate genetic structuration between the populations show the need for the conservation of genotypes surveyed in the two states. In addition, given the amplitude of natural occurrence of the species, populations should be surveyed in other locations to expand the genetic base to be conserved in the collection.
References
Almeida-Pereira CS, Silva AVC, Alves RP, Feitosa-Alcantara RB, Alvares-Carvalho SV, Costa TS, White LAS, Pinto VS, Sampaio TS, Blank AF (2017) Genetic diversity of native populations of Croton tetradenius Baill. using ISSR markers. Genet and Mol Res 16:1–12. https://doi.org/10.4238/gmr16029602
Awad AD, Billiard MC (2017) The double-edged sword: the demographic consequences of the Evolution of self-fertilization. Evolution 75(5):1178–1190. https://doi.org/10.1111/evo.13222
Bitencurti DP, Melo FP, Gois DV, Ruiz-Esparza J, Ribeiro AS, Ferrari SF, Souza RM (2017) Análise multitemporal do desmatamento nos municípios de Canindé de São Francisco e Poço Redondo. SE Geosul 32(63):117–137. https://doi.org/10.5007/2177-5230.2017v32n63p117
Blank AF (2013) Transformação de recursos genéticos de plantas aromáticas nativas em riqueza: o potencial do alecrim-de-tabuleiro (Lippia gracilis). Hort Bras 31(3):512–512. https://doi.org/10.1590/S0102-05362013000300029
Blank AF, Camêlo LCA, Arrigoni-Blank MF, Pinheiro JB, Andrade TM, Niculau ES, Alves PB (2015) Chemical diversity in Lippia alba (Mill.) N. E. Brown germplasm. Sci World J 2015:321924. https://doi.org/10.1155/2015/321924
Blank AF, Arrigoni-Blank MF, Bacci L, Costa Junior L, Nizio DAC (2019) Chemical diversity and insecticidal and anti-tick activities of essential oils of plants from Northeast Brazil. In: Malik S (ed) Essential Oil Research: Trends in Biosynthesis, Analytics, Industrial Applications and Biotechnological Production, 1ª. Springer International Publishing, Switzerland, pp 235–258
Bohonak AJ (1999) Dispersal, gene flow and population structure. Q Ver Biol 74(4):21–45. https://doi.org/10.1086/392950
Botrel MCG, Souza AMS, Carvalho D, Pinto SIC, Moura MCO, Estopa RA (2006) Caracterização genética de Calophyllum brasiliense camb. em duas populações de mata ciliar. Rev Árvore 30(5):821–827. https://doi.org/10.1590/S0100-67622006000500016
Brito FA, Nizio DAC, Silva AVC, Diniz LEC, Rabbani ARC, Montanari-Júnior I, Blank AF (2016) Genetic diversity analysis of Varronia curassavica Jacq. Accessions using ISSR markers. Genet Mol Res 15(3):1–10. https://doi.org/10.4238/gmr.15038681
Brito FA, Zucci MI, Pinheiro JB, Arrigoni-Blank MF, Dequigiovanni G, Grando C, Batista CE, Alves-Pereira A, Blank AF (2021) Low diversity in the native populations of Croton tetradenius Baill. when using SNP markers: a future crop with an insecticidal activity. Genet Resour Crop Evol. https://doi.org/10.1007/s10722-021-01176-7
Camara CA, Silva CG, Zago HB, Júnior HJ, De Oliveira JV, Barros R, Schwartz MOE, Lucena MFA (2008) Composition and insecticidal activity of the essential oil of Croton grewioides Baill. against Mexican bean weevil (Zabrotes subfasciatus Boheman). J Essent Oil Res 20(2):179–182. https://doi.org/10.1080/10412905.2008.9699985
Caruzo MBR, Secco RA, Medeiros D, Riina R, Torres DSC, Santos RFD, Pereira APN, Rossine Y, Lima LR, Muniz-Filho E, Valduga E (2020) Croton in Flora do Brasil. Jardim Botânico do Rio de Janeiro. http://floradobrasil.jbrj.gov.br/reflora/floradobrasil/FB17516. Access in: 19 nov, 2021.
Castro KNC, Chavas ACS, Costa-Júnior LM, Canuto KM, Brito ES, Rodrigues THS, Andrade IM (2019) Acaricidal potential of volatile oils from Croton species on Rhipicephalus microplus. Rev Bras Farmacogn 29(6):811–815. https://doi.org/10.1016/j.bjp.2019.09.001
Catchen JM, Hohenlohe P, Bassham S, Amores A, Cresko WA (2013) Stacks: an analysis tool set for population genomics. Mol Ecol 22(11):3124–3140. https://doi.org/10.1111/mec.12354
Chen SL, Luo HM, Wu Q, Steinmetz A (2016) Conservation and sustainable use of medicinal plants: problems, progress and prospects. Chin Med 11(37):1–10. https://doi.org/10.1186/s13020-016-0108-7
Chojnacka K (2015) Innovated Bio-Products for Agriculture. Open Chem 13(1):932–937. https://doi.org/10.1515/chem-2015-0111
Chung YS, Choi SC, Jun T-H, Kim C (2017) Genotyping-by-sequencing: a promising tool for plant genetics research and breeding. Hortic Environ Biotechnol 58(5):425–431. https://doi.org/10.1007/s13580-017-0297-8
Costa AS, Arrigoni-Blank MF, Carvalho-Filho JLS, Santana ADD, Santos DA, Alves PB, Blank AF (2015) Chemical diversity in basil (Ocimum sp.) germplasm. Sci World J 2015:352638. https://doi.org/10.1155/2015/352638
Costa RB, Moraes PM, Skowronski L, Oliveira CE, Nogueira MC, Yul RMS, Lorenz AP, Fava WS (2020) Genetic diversity and population structure of Croton urucarana Baill. (Euphorbiaceae) in central Brazil by ISSR markers. Braz J Bot 43:831–838. https://doi.org/10.1007/s40415-020-00657-w
Diniz MTM, Medeiros SC, Cunha CJ (2014) Sistemas atmosféricos atuantes e diversidade pluviométrica em Sergipe. Bol Goiano Geogr 34(1):17–34. https://doi.org/10.5216/bgg.v34i.29313
Do HDK, Jung J, Hyun J, Yoon SJ, Lim C, Park K, Kim JH (2019) The newly developed single nucleotide polymorphism (SNP) markers for a potentially medicinal plant, Crepidiastrum denticulatum (Asteraceae), inferred from complete chloroplast genome data. Mol Biol Rep 46(3):3287–3297. https://doi.org/10.1007/s11033-019-04789-5
Dominguéz CA, Bullock SH (1989) La reproduccíon de Croton suberosus (Euphorbiaceae) em luz y sombra. Rev Biol Trop 37(1):1–10
Dray S, Dufour AB (2007) The ade4 package: implementing the dualy diagram for ecologists. J Stat Softw 22(4):1–20
Earl DA, Vonholdt BM (2011) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361
Embuscado ME (2015) Spices and herbs: natural sources of antioxidants – a mini review. J Funct Foods 18:811–819. https://doi.org/10.1016/j.jff.2015.03.005
Engels JMM, Visser L (2003) A guide to effective management of germplasm collections. International Plant Genetic Resources Institute, Rome
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Ferreira ME, Grattapaglia D (1998) Introdução ao uso de marcadores moleculares em análise genética. Embrapa, Brasília
Frankham R (2003) Genetics and conservation biology. Biologies 326:22–29. https://doi.org/10.1016/S1631-0691(03)00023-4
Ganal MW, Altmann T, Röder M (2009) SNP identification in crop plants. Curr Opin Plant Biol 12(2):211–217. https://doi.org/10.1016/j.pbi.2008.12.009
Gois IB, Ferreira AF, Silva-Mann R, Pantaleão SM, Gois CB, Oliveira RSC (2014) Variabilidade genética em populações naturais de Ziziphus joazeiro Mart., por meio de marcadores moleculares RAPD. Rev Árvore 38(4):621–630. https://doi.org/10.1590/S0100-67622014000400005
Grattapaglia D, Silva-Junior OB, Kirt M, Lima BM, Faria DA, Pappas-Junior GJ (2011) High-throughput SNP genotyping in the highly heterozygous genome of Eucalyptus: assay success, polymorphism and transferability across species. Plant Biol 11(65) http://www.biomedcentral.com/1471-2229/11/65
Grover A, Sharma PC (2016) Development and use of molecular markers: past and present. Crit Revin Biotechnol 36(2):290–302. https://doi.org/10.3109/07388551.2014.959891
Gulati R (2018) Strategies for sustaining plant germplasm evaluation and conservation a review. Res J Life Sci 4(5):313–321
Guo C, McDowell IC, Nodzenski M, Scholtens DM, Allen AS, Lowe WL, Reddy TE (2017) Transversions have larger regulatory effects than transitions. BMC Genomics 19(1):394. https://doi.org/10.1186/s12864-017-3785-4
Gupta F, Idris A, Mantri S, Asif MH, Yada HK, Roy JK, Tuli R, Mohanty CS, Sawant SV (2012) Discovery and use of single nucleotide polymorphic (SNP) markers in Jatropha curcas L. Mol Breeding 30:1325–1335. https://doi.org/10.1007/s11032-012-9719-6
Hailu G, Bitew M, Temesgen M (2017) In vitro antifungal activity of Croton macrostachys and Allium sativum extracts against Candida albicans and Trichophyton mentagrophytes isolates. J Int J Biotechnol Food Sci 5(3):42–47
Hamrick JL (1989) Isozymes and the analysis of genetic structure in plant populations. In: Soltis DE, Soltis PS (eds) Isozymes in plant biology, 1ª. Chapman e Hall, London, pp 87–105
Hamrick JL, Godt MJW (1989) Allozyme diversity in plant species. In: Broen ADG, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding, and genetic resources, 1ª. Sinauer Associates, Sunderland, pp 43–63
Hartl DL, Clark AG (2010) Princípios de Genética de Populações. Artmed, Porto Alegre
Islam MR, Zhanf Y, Li ZZ, Liu H, Chen JM, Yang XY (2020) Genetic diversity, population structure and historical gene flow of Nelumbo lutea in USA using microsatellite markers. Aquat Bot 160:1–8. https://doi.org/10.1016/j.aquabot.2019.103162
Jombart T, Ahmed I (2011) Adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics 27:2070–3071. https://doi.org/10.1093/bioinformatics/btr52
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of the principal components: a new method for the analysis of genetically structured populations. Genetics 11(94):1–15. https://doi.org/10.1186/1471-2156-11-94
Kadam ST, Pawar AD (2020) Conservation of medicinal plants a review. Int J Ayurvedic Med. https://doi.org/10.46607/iamj0807112020
Kamvar ZN, Tabina JF, Gruenwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal and/or sexual reproduction. PeerJ. https://doi.org/10.7717/peerj.281
Kethom W, Togyoo P, Mongokollpo O (2019) Genetic diversity and capsaicinoids content association of Thai chili landraces analyzed by whole genome sequencing-based SNPs. Sci Hort 249:401–406. https://doi.org/10.1016/j.scienta.2019.02.022
Kim DS, Lee HJ, Jeon YD, Han YH, Kee JY, Kim HJ, Kim S (2015) Alpha-pinene exhibits anti-inflammatory activity through the suppression of MAPKs and the NF-κB pathway in mouse peritoneal macrophages. Am J Chin Med 43(4):731–742. https://doi.org/10.1142/S0192415X15500457
Leal IR, Wirth R, Tabarelli M (2007) Seed dispersal by ants in the semi-arid Caatinga of north-east Brazil. Ann Bot 99(5):885–894. https://doi.org/10.1093/aob/mcm017
Leviyang LS, Hamilton MB (2011) Properties of Weir and Cockerham’s FST estimators and associated bootstrap confidence intervals. Theor Popul Biol 79(1–2):39–52. https://doi.org/10.1016/j.tpb.2010.11.001
Li HX, Nuckols TA, Harris T, Stevenson KL, Brewer MT (2019) Differences in fungicide resistance profiles and multiple resistance to a quinone-outside inhibitor (QoI), two succinate dehydrogenase inhibitors (SDHI), and a demathylation inhibitor (DMI) for two Stagonosporopsis species causing gummy stem blight of cucurbits. Pest Manag Sci 75(11):3093–3101. https://doi.org/10.1002/ps.5426
Lôbo D, Tabarelli M, LeaL IR (2011) Relocation of Croton sonderianus (Euphorbiaceae) seeds by Pheidole fallax Mayr (Formicidae): a case of post-dispersal seed protection by ants? Neotrop Entomol 40(4):440–444. https://doi.org/10.1590/S1519-566X2011000400005
Marakli S (2018) A brief review of molecular markers to analyse medicinally important plants. Int J Life Sci Biotech 1(1):29–36
Marchese A, Barbieri R, Coppo E, Orhan IE, Daglia M, Nabavi SN, Abdollahi M, Nabavi SM, Ajami M (2017) Antimicrobial activity of eugenol and essential oils containing eugenol: A mechanistic viewpoint. Crit Rev Microbiol 3(6):668–689. https://doi.org/10.1080/1040841X.2017.1295225
Medeiros VM, Nascimento YM, Souto AL, Madeiro SAL, Costa VCO, SilVA SMPM, Silva VSF, Agra MF, Siqueira-Júnior JP, Tavares JF (2017) Chemical composition and modulation of bacterial drug resistance of the essential oil from leaves of Croton grewioides. Microb Pathog 11:468–471. https://doi.org/10.1016/j.micpath.2017.09.034
Monteiro PA, Zelioli IAM, Sousa IMO, Ruiz ALTG, Vendramini-Costa DB, Foglio MA, Carvalho JE (2017) Chemical composition and proliferative activity of Croton campestris A. St.-Hil. essential oil. Nat Prod Res 33(4):1–4. https://doi.org/10.1080/14786419.2017.1399377
Motta FGA, Skowronski L, Pinto JVC, Silvestrini M, Oliveira CE, Moraes PM, Costa RB (2019) Transferability of microsatellite loci from Croton floribundus Spreng to Croton urucarana Baill. (Euhporbiaceae). Afr Biotechnol 18(18):383–389. https://doi.org/10.5897/AJB2019.16810
Nadeem MA, Nawaz MA, Shahid MQ, Dogan Y, Comertpay G, Yildiz M, Hatipoglu R, Ahmad F, Alsaleh A, Labhane N, Ozkan H, Chung G, Baloch FS (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnol Biotechnol Equip 32(2):261–285. https://doi.org/10.1080/13102818.2017.1400401
Narbona E, Arista M, Ortiz PL (2005) Explosive seed dispersal in two perennial Mediterranean Euphorbia species (Euphorbiaceae). Am J Bot 92(3):510–516. https://doi.org/10.3732/ajb.92.3.510
Nascimento AS, Carvalho CAL, Martins MLL (2014) Plants visited by Apis mellifera (Hymenoptera: Apidae) in Recôncavo Baiano, State Bahia. Brazil Brazilian Journal of Agriculture 89(2):97–116
Newman DJ (2020) Cragg GM (2020) Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J Nat Prod 83(3):770–803. https://doi.org/10.1021/acs.jnatprod.9b01285
Niu S, Song Q, Koiwa H, Qiao D, Zhao D, Chen Z, Liu X, Wen X (2019) Genetic diversity, linkage disequilibrium, and population structure analysis of the tea plant (Camellia sinensis) from an origin center, Guizhou plateau, using genome-wide SNPs developed by genotyping-by-sequencing. BMC Plant Biol 19:328. https://doi.org/10.1186/s12870-019-1917-5
Oliveira AR, Pinto JESS, Mendonça FA (2016) A desertificação do Alto Sertão de Sergipe/Brasil: abordagem na perspectiva das vulnerabilidades socioambientais. Investig Geogr Chile 52(139):139–149. https://doi.org/10.5354/0719-5370.2016.43294
Oliveira SDDS, Silva AMDOE, Blank AF, Nogueira PCDLN, Nizio DADC, Almeida-Pereira CS, Pereira RO, Menezes-S ÁTS, Santana MHDSS, Arrigoni-Blank MF (2021) Radical scavenging activity of essential oils from Croton grewioides Baill accessions and the major compounds eugenol, methyl eugenol and methyl chavicol. J Essent Oil Res 33(1):94–103. https://doi.org/10.1080/10412905.2020.1779139
Otto LG, Mondal P, Brassac J, Preiss S, Degenhardt J, He S, Reif JC, Sharbel TF (2017) Use of genotyping-by-sequencing to determine the genetic structure in the medicinal plant chamomile, and to identify flowering time and alpha-bisabolol associated SNP-loci by genome-wide association mapping. BMC Genomics 18(1):599–617. https://doi.org/10.1186/s12864-017-3991-0
Passos L, Ferreira SO (1996) Ant dispersal of Croton priscus (Euphorbiaceae) seeds in a tropical semideciduous forest in Southeastern Brazil. Biotropica 28:697–700. https://doi.org/10.2307/2389055
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research –an update. Bioinformatics 28(19):2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Pence CH (2017) Is genetic drift a force? Synthese 194:1967–1988. https://doi.org/10.1007/s11229-016-1031-2
Perrier X, Jacquemoud-Collet JP (2006) DARwin Software. http://darwin.cirad.fr/darwin
Pires MMY, Souza LA, Terada Y (2004) Biologia floral de Croton urucarana Baill. (Euphorbiaceae) ocorrente em vegetação riária da ilha Porto Rico, Porto Rico, Estado do Paraná, Brasil. Acta Sci, Biol Sci 26:209–215. https://doi.org/10.4025/actascibiolsci.v26i2.1638
Poland J, Brown PJ, Sorrells ME, Jannink JC (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequence approach. PLoS ONE 7(2):e32253. https://doi.org/10.1371/journal.pone.0032253
Prentice HC, Malm JU, Mateu-Andrés I, Segarra-Moragues G (2003) Allozyme and chloroplast DNA variation in island and mainland populations of the rare Spanish endemic, Silene hifacensis (Caryophyllaceae). Conserv Genet 4:543–555. https://doi.org/10.1023/A:1025603328704
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Quitino ZGM, Machado IC (2014) Pollination syndromes in a Caatinga plant community in northeastern Brazil: seasonal availability of floral resources in different plant growth habitats. Braz J Biol 74(1):62–71. https://doi.org/10.1590/1519-6984.17212
R Development Core Team (2020) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria
Riina R, Van EBW, Caruzo MBR, Carneiro-Torres DS, Santos RF, Berry PE (2021) The Neotropical Croton sect. Geiseleria (Euphorbiaceae): Classification update, phylogenetic framework, and seven new species from South America. Ann Missouri Bot Gard. 106(1):111–166. https://doi.org/10.3417/2021669
Rogers JS (1972) Measures of genetic similarity and genetic distance. University of Texas Publication, Austin
Roullier C, Benoit L, Lebot V (2013) Historical collections reveal patterns of diffusion of Sweet potato in Oceania obscured by modern plant movements and recombination. Proc Natl Acad Sci USA 110(5):2205–2210. https://doi.org/10.1073/pnas.1211049110
Sampath S, Subramani S, Janardhanam S, Subramani P, Yuvaraj A, Chellan R (2018) Bioactive compound 1, 8-Cineole selectively induces G2/M arrest in A431 cells through the upregulation of the p53 signaling pathway and molecular docking studies. Phytomedicine 46:57–68. https://doi.org/10.1016/j.phymed.2018.04.007
Santos CP, Pinto JAO, Santos CA, Cruz EMO, Arrigoni-Blank MF, Andrade TM, Santos DA, Alves PB, Blank AF (2016) Harvest time and geographical origin affect the essential oil of Lippia gracilis Schauer. Ind Crops Prod 79:205–210. https://doi.org/10.1016/j.indcrop.2015.11.015
Santos MO, Almeida BV, Ribeiro DA, Macêdo DG, Macêdo MJF, Macedo JF, De Sousa FFS, Oliveira LGS, Saraiva ME, Araújo TMS, Souza MMA (2017) The conservation of native priority medicinal plants in a Caatinga area in Ceará, northeastern Brazil. An Acad Bras De Ciênc 89(4):2675–2685. https://doi.org/10.1590/0001-3765201720160633
Santos AS (2016) Biologia reprodutiva e citogenética de espécies simpátricas de Croton L. (Euphorbiaceae). Dissertation, University Federal of Paraíba
Seoane CRS, Kageyama PY, Sebbenn AM (2000) Efeitos da fragmentação florestal na estrutura genética de populações de Esenbeckia leiocarpa Enfl (Guarantã). Sci for 57:123–139
Sharma UK, Sharma AK, Gupta A, Kumar R, Pandey A, Pandey AK (2017) Pharmacological activities of cinnamaldehyde and eugenol: antioxidant, cytotoxic and anti-leishmanial studies. Cell Mol Biol 63(3):73–78
SiBBr (2020) Sistema de Informação sobre a Biodiversidade Brasileira: Croton grewioides Baill. Retrieved from: https://sibbr.gov.br. Accessed on: 24 May 2021
Silva MCNA, RodaL MJN (2009) Padrões das síndromes de dispersão de plantas em áreas com diferentes graus de pluviosidade, PE. Brasil Acta Bot Bras 23(4):1040–1047. https://doi.org/10.1590/S0102-33062009000400014
Silva GV, Zago HB, Júnior HJGS, Camara CAG, De Oliveira JV, Barros R, Schwartz MOE, Lucena MFA (2008) Composition and insecticidal activity of the essential oil of Croton grewioides Baill. against Mexican bean weevil (Zabrotes subfasciatus Boheman). J Essent Oil Res 20(2):179–182. https://doi.org/10.1080/10412905.2008.9699985
Silva JSS, Carneiro-Torres MF, Santos D (2009) O gênero Croton (Euphorbiaceae) na microrregião do Vale do Ipanema, Pernambuco. Brasil Rodriguésia 60(4):879–901. https://doi.org/10.1590/2175-7860200960409
Silva AC, Prata APN, Mello AA, Santos ACAS (2013) Síndromes de dispersão de angiospermas em uma Unidade de Conservação na Caatinga, SE. Brasil Hoehnea 40(4):601–609. https://doi.org/10.1590/S2236-89062013000400003
Silva AVC, Muniz EN, Almeida CS, Vitória MF, Ledo AS, Vasconcelos-Melo MF, Rabbani ARC (2015) Genetic diversity and sex identification in Genipa americana L. Trop Subtrop Agroecosyst 18(1):81–86
Silva TSS, Freitas JS, Santos ESL, Cerqueira-Silva CBM (2020) Diversity and genetic structure in natural populations of Croton linearifolius (Euphorbiaceae) based on molecular markers. Afr J Biotechnol 16(9):613–624. https://doi.org/10.5897/AJB2018.16595
Silva-Júnior AL, Cabral RLR, Sartori L, Souza LC, Miranda FD, Caldeira MVW, Moreira SO, Godinho TO (2020) Evaluation of diversity and genetic structure as strategies for conservation of natural populations of Dalbergia nigra (Vell.) Allemão ex Benth. Cerne 26(4):435–443. https://doi.org/10.1590/01047760202026042754
Silvestrini M, Mccauley DE, Zucci MI, Santos FA (2015) How do gap dynamics and colonization of a human disturbed area affect genetic diversity and structure of a pioneer tropical tree species? Forest Ecol Manag 344:38–52. https://doi.org/10.1016/j.foreco.2015.01.026
Souza DCL (2015) Técnicas moleculares para caracterização e conservação de plantas medicinais e aromáticas: uma revisão. Revi Brasi De Plantas Med 17(3):495–503. https://doi.org/10.1590/1983-084X/13071
Souza JT, Ferraz EMN, Albuquerque UP, Araújo EL (2013) Does proximity to a mature forest contribute to the seed rain and recovery of an abandoned agriculture area in a semiarid climate? Plant Biol 16(4):748–756. https://doi.org/10.1111/plb.12120
Souza JD, Melo KM, Santos RA, Teixeira LR, Sampaio A, Guimarães JR, Vieira T, Bento R, Borges VP, Martins AAM, Arcanjo JB, Loureiro HSC, Angelim LA (2003) A Mapa geológico do Estado da Bahia—Escala 1:1.000.000, Salvador: CRPM https://rigeo.cprm.gov.br/jspui/handle/doc/8665?mode=full. Accessed on: 22 Jan. 2021
Stern DL, Orgogozo V (2008) The loci of evolution: how predictable is genetic evolution? Evolution 62(9):2155–2177. https://doi.org/10.1111/j.1558-5646.2008.00450.x
Thaowetsuwan P, Ritchie S, Riina R, Craene LPR (2020) Divergent developmental pathways among staminate and pistillate flowers of some unusual Croton (Euphorbiaceae). Front Ecol Evol 8:253. https://doi.org/10.3389/fevo.2020.00253
Trindade MT, Lameira AO (2014) Espécies úteis da família Euphorbiaceae no Brasil. Rev Cuba De Plantas Med 19(4):292–309
Vander Wall B, Longland W (2004) Diplochory: are two seed dispersers better than one? Trends Ecol Evol 19(3):155–161. https://doi.org/10.1016/j.tree.2003.12.004
Wang H, Li G, Kwon WS, Yang DC (2016) Development of EST intron-targeting SNP markers for Panax ginseng and their application to cultivar authentication. Int J Mol Sci 17(6):884. https://doi.org/10.3390/ijms17060884
Weir B, Cockerham MC (1984) Estimating F statistics for analysis of- population structure. Evolution 38(6):1358–1370. https://doi.org/10.2307/2408641
White LAS, Silva AVC, Carvalho SVA, Silva-Mann R, Arrigoni-Blank MF, Souza EMS, Almeida-Pereira CS, Nizio DAC, Sampaio TS, Blank AF (2018) Genetic diversity of a native population of Myrcia ovata Cambess using ISSR molecular markers. Genet Mol Res 17:1–11. https://doi.org/10.4238/gmr18022
Zanella CN, Palma-Silva C, Kaltchuk-Santos E, Pinheiro FG, Paggi GM, Soares LES, Goetze M, Büttow MV, Bered F (2012) Genetic evolution and conservation of Bromeliaceae. Genet Mol Biol 35(4):1020–1026. https://doi.org/10.1590/S1415-47572012000600017
Zou Z, Zhang J (2021) Are nonsynonymous transversions generally more deleterious than nonsynonymous transitions? Mol Biol Evol 38(1):181–191. https://doi.org/10.1093/molbev/msaa200
Acknowledgements
The authors thank the Fundação de Apoio a Pesquisa e Inovação Tecnológica do Estado de Sergipe (Fapitec/SE), the Conselho Nacional de Desenvolvimento Científico e Tecnológico—Brasil (CNPq), the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES—Finance code 001), and the Financiadora de Estudos e Projetos—Brasil (FINEP) for financial support. AA-P is grateful to the São Paulo Research Foundation for a post-doctoral scholarship (FAPESP 2018/00036-9).
Not applicable.
Funding
Funding provided by CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior) (code AUXPE 2971/2014).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest. All of the authors have read and approved the manuscript in its final form.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Oliveira, S.D.S., Gois, I.B., Blank, A.F. et al. Genome-wide diversity in native populations of Croton grewioides Baill., a future crop with fungicidal and antioxidant activity, using SNP markers. Genet Resour Crop Evol 69, 1965–1978 (2022). https://doi.org/10.1007/s10722-022-01357-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-022-01357-y