Abstract
The collection and characterization of apricot germplasm is an important step for its utilization, conservation, and breeding programs. In the present study, the genetic diversity and population structure of 106 accessions of apricot including 82 indigenous and 24 exotic accessions collected from various geographical locations of Jammu and Kashmir, India were evaluated using 14 inter simple sequence repeat markers. The PCR amplification produced 365 loci of which 356 (97.53%) were polymorphic. The average number of alleles, effective number of alleles, expected heterozygosity, Shannon’s information index indicated high level of genetic diversity in the collected apricot accessions, and the polymorphism information content (0.82) revealed that these markers were highly polymorphic. The results of UPGMA dendrogram and model-based STRUCTURE analysis clearly divided the 106 apricot accessions into two main groups; one group included the 78 accessions of Kashmir province and the other included 28 accessions of Jammu province indicating the distinction of two genetic pools of apricot in the region. Nonetheless, PCoA also revealed a similar grouping of accessions except few exotic accessions that formed a small separate group. Moreover, analysis of molecular variance showed high genetic variation within the population (67%) and low among the population (33%). To our knowledge, this study represents the first comprehensive report on the ISSR based genetic diversity, relationship, and the structure of apricot accessions of Jammu and Kashmir. Further, the results of the present study will assist in efficient utilization, conservation, and MAS based breeding programs of apricot.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Apricot (Prunus armeniaca L.) of the family Rosaceae is one of the most important plants of the genus Prunus, commonly cultivated for its high nutritional value (Shangguan et al. 2012). It is one of the four economically important fruit crops of the genus Prunus, such as P. persica L., P. domestica L., and P. avium L. (Burgos et al. 2007). It is an interfertile diploid species with 8 chromosome pairs (2n = 16) and has an intermediate genome between that of peach and cherry i.e. (5.9 × 108 bp/2n) (Arumuganathan and Earle 1991). Apricot is a temperate zone fruit that can be grown under climatically variable conditions in many regions of the world such as Central Asia, Japan, Siberia, and China (Asma 2007). In North–Western Himalayan regions, wild apricot is found growing in the southern part of China, cold deserts of Tibet and in northern India it is found growing in temperate regions of Jammu and Kashmir, Himachal Pradesh and Uttarakhand at 2000–2500 m above average sea level (Sharma 2000). Moreover, in northern India, Jammu & Kashmir is the leading producer of apricots contributing a total of 20456 MT in the year 2018–2019 (http://hortikashmir.gov.in/).
Jammu and Kashmir particularly Kashmir valley holds a rich diversity of both indigenous as well as exotic cultivars. Indigenous accessions originated in the region, are locally grown, seedling origin trees (wild), or locally grafted (cultivated) accession, while the exotic accessions are those which have been introduced from other countries mostly of European origin (Bhat et al. 2013). The main purpose of apricot cultivation is fruit production for fresh consumption that offers high profit to the growers, while a part of the produce is destined for the processing industry. However, lack of cultivation practices, poor shelf life, lack of awareness in perceiving apricot quality, and poor market opportunities has led to erosion of genetic diversity of landraces in apricot. This depletion of apricot germplasm in the region may lead to the loss of many elite landraces which represent a natural repository of certain novel genotypes in terms of color, flavor, and taste of fruit, disease resistance, and high yield. Information on genetic diversity and population structure is therefore desired for this crop to assist the conservation, determine genetic relationships between accessions and characterization of germplasm for breeding purposes.
A variety of methods and techniques such as morphological, molecular, biochemical have been developed to expedite work on population genetic diversity, among which molecular tools play a vital role in genetic resource management and use (Chen et al. 2020). Such molecular tools increase our possibilities to study genetic diversity and to advance genome-wide association studies of complex characters and select suitable donors for breeding purposes (Aliyev et al. 2007; Hajiyev et al. 2015). Furthermore, these molecular markers have been in use for more than 30 years to estimate genetic diversity values due to their ability to produce more accurate information (Reddy et al. 2002; Madhumati 2014).
ISSR is one of the DNA marker systems involving the amplification of a DNA segment between two (SSR sequences) identical microsatellites in the opposite direction (Gebrehiwet et al. 2019). They provide a detailed genome coverage, high performance, time efficiency, and cost-efficiency (Ganopoulos et al. 2011; Yilmaz et al. 2012). Moreover, ISSR markers are highly reproducible and are one of the most widely used markers, revealing a variety of information bands in a single amplification (Gelotar et al. 2019). Several studies have been undertaken to study the genetic diversity of apricot germplasm using different molecular markers in the other parts of the world such as RAPD, AFLP, SSR, ISSR, and SRAP (Ercisli et al. 2009; Lamia et al. 2010; Yilmaz et al. 2012; Ai et al. 2011). However, in Jammu and Kashmir including Ladakh province, morphological tools have been mostly used to study the variability of apricot germplasm (Bhat et al. 2013; Malik et al. 2010; Girish et al. 2012; Abdul et al. 2016; Angmo et al. 2017; Kumar et al. 2015; Wani et al. 2017) and to the best of our knowledge, very limited work is available using molecular markers such as RAPD (Kumar et al. 2009a; Mir et al. 2012) and except for the study of Kumar et al. (2009b) from Ladakh region, no report is available on the evaluation of genetic diversity of apricot germplasm of Jammu and Kashmir using ISSR markers. Furthermore, these studies were often restricted by the limited number of accessions and specific locations. Therefore, genetic diversity data on the major apricot germplasm of the Jammu and Kashmir region are by and large lacking. Hence, the present study was conducted on 106 accessions of apricot collected from different areas of Jammu and Kashmir using 14 ISSR markers. This is the first report on genetic diversity analysis of apricot germplasm using ISSR markers that include indigenous (wild and cultivated) accessions as well as exotic accessions grown throughout the Jammu and Kashmir region. The generated information could be used as a baseline in the future to support conservation, marker-assisted breeding, crop improvement, and comparative genomic studies.
Materials and methods
Collection site and sample collection
In the present study, 106 apricot accessions were collected from 22 geographical areas of Jammu and Kashmir, India (Table 1, Fig. 1). The young leaf samples were collected and stored at − 80 °C after snap freezing in liquid nitrogen until subjected to DNA extraction.
DNA extraction
Genomic DNA extraction from the young and healthy leaves of collected apricot accessions was carried out using the DNeasy Plant Mini Kit (Qiagen, Hilden GmbH, Germany), according to the manufacturer’s instructions. In order to check the quality of extracted genomic DNA, 0.8% w/v agarose gel with ethidium bromide staining and 1X TAE as running buffer were used and the quantity was calculated by spectrophotometer at 260 nm.
PCR amplification
A total of 14 ISSR markers chosen from the literature (Yılmaz et al. 2009) obtained from IDT (Integrated DNA Technologies, USA) were used to access the genetic diversity of 106 apricot accessions. Details of 14 ISSR markers with their sequence and annealing temperature (°C) are presented in Table 2. For each amplification, the reaction mixture was prepared in a final volume of 20 µl which consists of 2 µl 10× reaction buffer, 1.5 µl of 25 mM MgCl2, 0.3 µl 25 mM of dNTP mix (dATP, dGTP, dCTP, and dTTP, 3 µl of primer (10 µM), 0.3 µl Taq polymerase (5U/µl) (Thermo Scientific, USA), 11.9 µl of sterilized distilled water and 1.0 µl of template DNA (50 ng). The amplification was carried out in Applied Biosystems thermocycler, USA using the following PCR conditions: preheating and initial denaturation at 94 °C for 5 min followed by 35 cycles of denaturation at 94 °C for 1 min; primer annealing at specific annealing temperature for each primer (Tm) 45 s; extension at 72 °C for 2 min and a final extension at 72 °C 10 min and 4 °C hold. After the completion of the PCR, 2.5 µl of 6× loading dye (Thermo Scientific, USA) was added to the amplified products and separated in 1.8% (w/v) agarose gel with ethidium bromide staining using 1× TAE (Tris-acetate EDTA) as running buffer. Finally, the size of the fragments was measured against the 100 bp DNA ladder (Thermo Fisher Scientific, USA).
Data scoring and analysis
The well separated and intense bands were scored in binary code with 1 and 0, indicating the presence and absence of bands respectively. The informativeness of the ISSR marker system in defining, differentiating, and evaluating the apricot diversity was evaluated by the following parameters for each assay unit; polymorphic information content (PIC), resolving power (RP), and marker index (MI). PIC was calculated according to Smith et al. (1997); PIC = 1 − ΣPi2, RP was calculated according to Prevost and Wilkinson (1999); RP = ΣIb, similarly marker index (MI) according to Powell et al. (1996); MI = PIC × β × α. Furthermore, band frequency, observed number of alleles (Na), effective number of alleles (Ne), Shannon’s information index (I), expected heterozygosity (He), and unbiased heterozygosity (uHe) were estimated by using GenAlEx version 6.5 (Peakall and Smouse 2012). Based on Jaccard’s genetic similarity coefficient (Rohlf 2000), a dendrogram of 106 accessions of apricot was constructed using the NTSYS-pc Version 2.1 program with the UPGMA (unweighted pair-group method with arithmetic means) approach. Besides, PAST software was used to calculate the principal coordinate analysis (PCoA) (Hammer et al. 2001). A model-based Bayesian clustering analysis program STRUCTURE (Pritchard et al. 2000) determined the genetic structure and the number of clusters in the data set of apricot accessions. The number of possible populations (K) was estimated between 1 and 10 and the analysis was repeated twice. For each run, the burn-in and Markov Chain Monte Carlo (MCMC) were set to 50,000 each, and iterations were set to 5. The run with the highest likelihood was used to designate individual accessions into groups and true K was estimated according to the method defined by Evanno et al. (2005). Within a group, accessions with inferred ancestry based on probability values P ≥ 70% were assigned to a different group, and those with < 70% were treated as ‘‘admixture’’, i.e., these accessions tend to have mixed ancestry from the parents belonging to various geographical origins or gene pools. The results were further processed and the accessions were assigned to different clusters using CLUMPP software v1.1.2 (Jakobsson and Rosenberg 2007). The output of CLUMPP was directly used as input for STRUCTURE software (Pritchard et al. 2000) to obtain results in graphical format. Further, the analysis of molecular variation (AMOVA) among and within populations was performed by GenAlEx version 6.5 (Peakall and Smouse 2012).
Results
ISSR polymorphism
In the present study, the ISSR markers showed a high polymorphism and reproducible fragments on the 106 accessions of apricot. A representative image illustrating the banding pattern of 106 apricot accessions using ISSR markers BC840 is shown in Fig. 2. A total of 365 alleles were amplified ranging from 3 for BC825 to 36 for BC827 with an average of 26.07 alleles per locus among which 356 alleles were polymorphic (97.53%) and 9 alleles were monomorphic. Furthermore, among 14 markers, most of the markers showed 100% polymorphism except BC825, BC835, BC873, BC880, and BC888. The allelic size of these 14 ISSR markers ranged from 150 bp in BC835, BC841, BC843, and BC888 to 3500 bp in BC807. The PIC value recorded revealed the lowest PIC value 0.67 in marker BC825 and the highest PIC value 0.92 in marker BC835 and BC843. Other parameters like RP, MI, and EMR were also studied for all 14 ISSR markers. RP ranged from 2.01 in BC825 to 23.24 in BC827, MI was recorded 1.35 in BC825 to 31.09 in BC827 and EMR ranged from 2 in BC825 to 36 in BC827 (Table 2).
Genetic diversity analysis
Band frequency, observed number of alleles (Na), effective number of alleles (Ne), Shannon’s information index (I), expected heterozygosity (He), and unbiased heterozygosity (uHe) were used to evaluate the genetic diversity of collected apricot accessions (Table 3). The highest band frequency was observed for BC880 (0.958) followed by BC888 (0.844) and BC868 (0.806). The average value of observed number of alleles was 2.960, ranging from 2.500 (BC873) to 3.448 (BC812) while the effective number of alleles ranged from 2.252 (BC825) to 2.830 (BC841) with an average value of 2.646. In addition highest, Shannon’s information index (I) was recorded for marker BC827 (0.793) followed by BC812 (0.762) and BC888 (0.749). The highest expected heterozygosity (He = 0.515) and unbiased heterozygosity (uHe = 0.521) were recorded for BC827 followed by BC888 respectively. BC825 showed the least expected heterozygosity (He = 0.189) as well as unbiased heterozygosity (uHe = 0.192) while 0.403 and 0.408 were the average values of expected heterozygosity and unbiased heterozygosity recorded for all the 14 ISSR markers (Table 3).
Genetic relationship and population structure
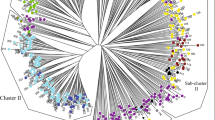
ISSR data was used to calculate pairwise distances between all possible pairs of 106 accessions to identify the genetic relationship among the apricot accessions. The similarity coefficients among the collected accessions ranged from 0.16 (RJC20 and BPW06) to 0.90 (SNC01and SNC03). The UPGMA dendrogram obtained divided apricot accessions into two main clusters, namely cluster I and cluster II. Most of the accessions grouped in individual clusters belonged to a particular geographical location (Fig. 3). Cluster I includes all accessions collected from different locations of Kashmir province while cluster II comprises accessions collected from different locations of Jammu province. Cluster I which includes most of the accessions (78) was divided into two main subgroups IA and IB at a genetic similarity coefficient level of 0.37. IA formed a smaller cluster consisting of 16 accessions from Srinagar, Baramulla, and Shopian districts of Kashmir province. These all 16 accessions are commonly grown exotic apricot accessions in Kashmir province which are mostly of European origin. However, IB formed the largest group of 62 accessions consisting of both wild and cultivated accessions of apricot, and was further subdivided into two sub-clusters (IB1 and IB2) at a genetic similarity coefficient level of 0.52. Subgroup IB1 was further divided into two clusters (IB1a and IB1b); IB1a includes 16 cultivated accessions of Srinagar, Shopian, and Budgam districts of Kashmir province while IB1b again formed two sub-clusters with one sub-cluster containing cultivated accessions of Srinagar, Ganderbal, Bandipora, and Kupwara, two wild accessions (BGW06 and SNW20) of Budgam and Srinagar and another sub-cluster containing 15 wild accessions of Bandipora and one wild accession from Ganderbal district. Similarly, the main sub-cluster IB2 was further divided into two subgroups in which one group comprises 09 wild accessions from the Ganderbal district and 04 accessions from Bandipora district while its 2nd subgroup contains only one wild accession BPW06 from Bandipora district.
All the 28 accessions collected from different locations of Jammu province formed the main cluster II, which further separated into two subgroups IIA and IIB. Subgroup IIA contained 10 wild and 06 cultivated accessions of Rajouri district of Jammu province while IIB formed two sub-clusters in which one sub-cluster comprised of 11 accessions from Rajouri district and 2nd sub-cluster contains only one accession of Poonch district of Jammu province (Fig. 3).
Principal coordinate analysis (PCoA) was performed by PAST software and graphically showed two distinct clusters except for a few exotic accessions that formed a small separate cluster (Fig. 4). The PCoA revealed the first two most informative principal components with a percentage of variability of 18.21% and 15.33% respectively, which together accounted for 33.54% of the total genetic variation. Figure 4 showed similar patterns of clustering with that of the UPGMA dendrogram except for the sub-cluster IB which formed a separate cluster. As seen in Fig. 4, three groups were obtained and all the three groups clustered according to their geographical location. Group I include accessions collected from Kashmir province which all are exotic accessions while group II consists of 65 wild and cultivated accessions of Kashmir province among which most of the accessions are indigenous except few exotic accessions. Similarly, group III includes 28 accessions collected from different locations of Jammu province.
To find out the number of populations from 106 accessions using 14 ISSR markers, STRUCTURE analysis was carried out. Our results showed a clear peak for ΔK at K = 2 with the next largest peaks at K = 4 and K = 6 (Fig. 5A). For the first level of clustering at K = 2, the apricot accessions were differentiated into two clusters, the first one consisted of accessions from Kashmir province while the second cluster included all accessions of Jammu province (Fig. 5B; Table S1). We used the second level of clustering at K = 4 to define the clusters considered in subsequent analyses. At K = 4, the 78 accessions of Kashmir province got separated into 3 clusters while the accessions from the Jammu region remained as a single cluster. Cluster 1 (16 accessions; in blue) included exotic accessions of those of European origin. Cluster 2 (30 accessions; in yellow) grouped few exotic and indigenous cultivated accessions from Srinagar, Baramulla, Shopian, Budgam and wild accessions from Ganderbal and Bandipora districts. Cluster 3 (32 accessions; in green) grouped cultivated accessions from Ganderbal, Bandipora, Srinagar, Kupwara, and wild accessions from Bandipora districts, and Cluster 4 (28 accessions; in red) grouped all the wild and cultivated accessions of Jammu province (Table S2). At the same time, the third-largest ΔK at K = 6 was much larger than the previous ones. When K = 6, there was a further division of 78 apricot accession collected from Kashmir province while no further clustering was recorded in the accessions collected from Jammu province. Therefore, at K = 6, cluster 1 (16 accessions; in pink) grouped exotic accessions collected from different districts of Kashmir province, Cluster 2 (16 accessions; in yellow) grouped few exotic and cultivated accessions of Kashmir province, Cluster 3 (16 accessions; in turquoise) grouped 14 cultivated accessions from Ganderbal, Bandipora, Srinagar, Kupwara districts and 2 wild accessions from Budgam and Srinagar districts of Kashmir province. Cluster 4 (16 accessions; in green) grouped the wild accessions collected from the Bandipora district of Kashmir province, while cluster 5 (14 accessions; in blue) grouped 9 wild accessions from Ganderbal and 5 wild accessions from Bandipora districts of Kashmir province and the last cluster 6 (28 accessions; in red) grouped all the wild and cultivated accessions collected from Jammu province (Fig. 5B; Table S3).
A Graphical representation of mean likelihood L (K) and variance per K values generated by model based-STRUCTURE on dataset containing 106 apricot accessions across 14 ISSR markers. B STRUCTURE plot of membership coefficients for all the 106 apricot accessions in the study sample sorted in the same order and classified according to successive preset K values ranging from 2 to 10. Each individual is represented by a thin vertical line, which is partitioned into K colored segments that represent the individual’s estimated membership fractions in K clusters
Model-based Bayesian clustering of 106 accessions revealed a similar pattern as inferred with the UPGMA dendrogram and PCoA, with clear genetic discrimination of two major clusters where major Cluster 1 formed several sub-clusters separating the exotic, cultivated (grafted accessions) and wild (seedling origin) accessions and formed the most diversified cluster. Overall, the observed cluster distribution reflected the clear separation of two main genetic clusters in Jammu and Kashmir, India.
Based on the grouping of accessions in two populations defined by the model-based Bayesian clustering algorithm using the STRUCTURE program, the expected heterozygosity for the two populations ranged from 0.202 for population II to 0.256 for population I with an average value of 0.229, and the Fst values ranged from 0.267 for population I to 0.558 for the population II with an average value of 0.412 (Table S4). Besides, the genetic differentiation based on Fst value for both populations identified by STRUCTURE analysis was recorded as 0.1382 (Table S5).
Analysis of molecular variance (AMOVA)
The analysis of molecular variance (AMOVA) was done to study the variation among and within the populations after the grouping of accessions in two populations by STRUCTURE analysis (Table 4). The results demonstrated that 67% of the total variation occurred within populations whereas 33% of the total variance was distributed among the populations. These results suggested that there is a substantial gene flow within the populations and the presence of different combinations of alleles as well the allelic exchange rates within the populations are very high than among the populations.
Discussion
Detailed genetic characterization of plants is a crucial step in the implementation of breeding programs, successful conservation, and usage of plant genetic resources (Tripathi et al. 2012; Izzatullayeva et al. 2014). The most effective and efficient technique for this characterization is the molecular marker analysis. Various properties, particularly non-responsiveness towards environmental, pleiotropic, and epistatic effects, make these molecular markers extremely profitable and useful in comparison with traditional phenotypic or morphological markers (Mondini et al. 2009). The ISSR markers are one of the most frequently used molecular markers for the assessment of genetic diversity and the relationship in the number of crop plants (Reddy et al. 2002). In the present investigation, as many as 14 ISSR markers were used to study the genetic diversity, population structure, and the relationship between 106 accessions of apricot collected from the different locations of Jammu and Kashmir, India.
ISSR polymorphism
All the 14 ISSR markers used in the present study were selected based on their high polymorphism, ability to amplify a single locus and their high discriminating power that allows differentiation between the accessions (Yilmaz et al. 2009, 2012; Li et al. 2013, 2014a, b; Liu et al. 2015, 2016). These ISSR markers showed an overall high level of polymorphism (97.53%) which suggests that there is a considerable level of genetic variation present in the accessions of apricot grown in both Jammu and Kashmir provinces. This percentage of polymorphism is higher than reported by Yilmaz et al. (2009) (88%); Li et al. (2014b) (90.37%) in Turkish apricot germplasm and Siberian apricots in China respectively. The ISSR markers used in the present study generated 356 highly informative polymorphic loci with an average PIC value of 0.82 which is considerably higher than found by (Yilmaz et al. 2009, 2012) (PIC = 0.564) in the Turkish apricot germplasm and in the Prunus spp. using 20 ISSR markers. The higher PIC value indicates that the set of ISSR markers used in the present study was highly efficient and suitable for the diversity analysis of apricot accessions from the Jammu and Kashmir region. Similarly, in comparison to other studies, the average MI = 20.95 observed in our study was higher than those reported on apricot by Kumar et al. (2009b) (MI = 3.74) and sweet cherry by Ivanovych et al. (2017) (MI = 2.46). In a similar way, the average values of the other parameters such as R.P (15.95) and EMR (24.90) were also observed to be high as compared to the studies of Yilmaz et al. (2009) (RP = 0.829), Dvin et al. (2020) (R.P = 7.882) and Kumar et al. (2009b) (EMR = 4.81). In the diversity analysis of apricot by ISSR markers, the majority of previous studies did not involve PIC, MI, RP, and EMR calculations, however, very low values were recorded in a few studies. The reason for the high level of polymorphism obtained in the present study is because of a rich genetic diversity of apricot germplasm in both Jammu and Kashmir provinces, the large number of accessions selected (sample size), and the high efficiency of ISSR markers used in the present study. Besides, high allelic diversity in the studied germplasm may be due to the presence of wild apricot accessions that grow locally and provide opportunities for cross-pollination with the cultivated accessions. The presence of allelic richness in any germplasm is a valuable tool for obtaining desirable accessions for commercial cultivation as well as in the breeding programs.
Genetic diversity analysis
There are different types of measures of genetic diversity such as the observed number of alleles, the effective number of alleles, Shannon’s information index, expected heterozygosity, and unbiased heterozygosity. All these parameters were used to study the genetic diversity of collected apricot accessions of Jammu and Kashmir. The average value of the observed number of alleles (Na) per locus was 2.960 while the average value of the effective number of alleles (Ne) per locus was 2.646, which appears to be higher than the previous studies on apricot germplasm. Effectively, Liu et al. (2015) recorded a mean value of 1.73 (Na) and 1.40 (Ne) using 9 ISSR markers for sweet kernel apricots of China and Li et al. (2013) obtained a mean value of 1.62 (Na) and 1.36 (Ne) for 14 wild apricot populations of Ili Valley area of northwestern China using 15 ISSRs. Similarly, the average values of Shannon’s information index (I = 0.628) were higher than the average value reported by Li et al. (2014a), who recorded the average value of Shannon’s information index (I) as 0.414 for 76 accessions using 15 ISSR markers and Li et al. (2013) who obtained the average value as 0.317 for 14 wild apricots. The average value of expected heterozygosity in the present study was 0.403 indicating the substantial genetic diversity in the apricot accessions of Jammu and Kashmir. By comparison, the expected heterozygosity obtained from Prunus avium was lower (He = 0.238) by Ivanovych et al. (2017). Apricot is a long-lived woody tree with a broad geographical range in Jammu and Kashmir. The accessions have most likely acquired major genetic variations in fruit weight and dimensions, fruit color, fruit shape, fruit firmness, etc., due to the adaptation to the environment, selection through domesticating processes, and different pollination mechanisms. All such factors in combination or alone have resulted in a high degree of genetic diversity in the presently studied apricot accessions. Moreover, the high genetic diversity observed in the present study is because the area surveyed falls in the Central Asian region (from the Tianshan mountains to Kashmir) which is one of the primary domestication centers of the commercial apricot and is considered as one of the oldest and richest reservoirs of apricot germplasm (Vavilov 1951; Faust et al. 1998; Hormaza et al. 2007).
Genetic relationship and population structure
The UPGMA method was used for the cluster analysis based on Jaccard’s genetic similarity coefficient (Rohlf 2000). Two main clusters distinguished 78 apricot accessions (73.59%) collected from Kashmir province, from the remaining 28 apricot accessions (26.41%) collected from Jammu province, which could be related to the existence of two genetic stocks (gene pools) in Jammu and Kashmir. The major cluster I contained two subgroups structured according to the geographical origin with cultivated exotic accession in subgroup IA and indigenous accessions (wild and cultivated) in subgroup IB. The obtained genetic relationship was in agreement with the results of Hu et al. (2018) based on their diversity analysis of apricot germplasm collected from the Tien-Shan Mountains of China using SSR markers and Bourguiba et al. (2010a) using mapped SSR markers on the genetic diversity of apricot in Tunisia. The further division of subgroup IB into sub-clusters IB1 and IB2 according to the mode of propagation was also revealed in the dendrogram. Evidence of this structure according to the mode of propagation (grafting vs. seedling origin trees) was also obtained by (Bourguiba et al. 2010b, 2012a) when studying Tunisian apricot germplasm and genetic diversity of grafted and seed propagated apricot in the Maghreb region respectively. However, in the sub-cluster IB1, clustering of a few exotic accession and wild accessions with the cultivated accessions can be explained by the fact that these accessions present large adaptability as well as a substantial gene exchange among materials originating from the different regions as described by Crossa-Raynaud (1960), Carraut and Crossa- Raynaud (1974). Similar results were also shown using PCoA analysis which agreed with the separation of apricot accessions of Kashmir province from Jammu province. Although the grouping corresponded largely to the geographical provinces, there was a notable exception with the 16 accessions of the main cluster I which have formed a subgroup IA in UPGMA tended to form a separate cluster which all are exotic accessions. Besides the PCoA analysis also suggested that apricot accessions of Kashmir province were more diverse as they had a wider distribution in the PCoA plot as compared to the apricot accessions collected from Jammu province. The genetic structure of 106 apricot accessions was studied by model-based Bayesian clustering by STRUCTURE analysis and the results indicated the existence of clear two group-structure. The two major cluster structures revealed the presence of two different apricot genetic stocks present in the apricot germplasm of Jammu and Kashmir. A similar result of the two-group structure was reported in the previous study conducted on Tunisian apricot accessions (Krichen et al. 2008, 2014). Li et al. (2014b), Gürcan et al. (2015) and Bourguiba et al. (2012b) also confirmed this structure and clearly allocated apricot accessions into two groups with the Bayesian method using STRUCTURE software.
From the analysis of UPGMA dendrogram, PCoA, and Bayesian clustering, two genetic clusters were clearly indicated among the apricot accessions of Jammu and Kashmir. Moreover, the estimate of genetic variation reflected in the AMOVA was higher within the populations than among the populations. These estimates indicate that significant genetic diversity exists in apricot accessions of both Jammu and Kashmir provinces that formed two genetic clusters in which isolation by distance may have played a determining role. Besides, there could be a lot of factors responsible for the separation of these two genetic clusters, such as landscape variability, environmental and human-mediated transport of reproductive material which becomes barriers to the migration and gene flow among the two genetic clusters. However, the geographical topology, acting as an extrinsic barrier between the two provinces may play an important role. On the other hand, existing genetic barriers have possibly arisen as a result of changes in the local climate and vegetation that might cause genetic discontinuity among the two populations (Pacheco-Olvera et al. 2012). Several previous studies also reported the existence and influence of geographical barriers on the genetic structure patterns (Lusini et al. 2014; Meng et al. 2016; Bhandawat et al. 2019; Torokeldiev et al. 2019).
Conclusion
This study represents the first major effort to characterize the apricot germplasm collected from various geographical locations of Jammu and Kashmir, India. The results of the high variability parameters obtained by the ISSR markers indicated that the apricot germplasm of Jammu and Kashmir maintained a high level of genetic diversity. Our study showed that the collected 106 apricot accessions were divided into two main genetic clusters by UPGMA, PCoA, and STRUCTURE analysis according to the geographical location/distribution. The distinction of two genetic clusters may be attributed to human selection as well as the ecological constraints. This study provided a precise picture in understanding the genetic diversity and the relationship between these 106 apricot accessions of Jammu and Kashmir which will be important for identification, utilization, and conservation of germplasm as well as in molecular marker-assisted breeding programs.
References
Abdul K, Ram RB, Dwivedi SK, Dwivedi DH, Meena ML (2016) Genetic variability, heritability and genetic advance studies for physico-chemical characters of apricot germplasm under Ladakh region. Progress Hortic 48(1):22–27
Ai PF, Zhen ZJ, Jin ZZ (2011) Genetic diversity and relationships within sweet kernel apricot and related Armeniaca species based on sequence-related amplified polymorphism markers. Biochem Syst Ecol 39(4–6):694–699
Aliyev RT, Abbasov MA, Mammadov AC (2007) Genetic identification of diploid and tetraploid wheat species with RAPD markers. Turk J Biol 31(3):173–180
Angmo P, Angmo S, Upadhyay SS, Targais K, Kumar B, Stobdan T (2017) Apricots (Prunus armeniaca L.) of trans-Himalayan Ladakh: potential candidate for fruit quality breeding programs. Sci Hortic 218:187–192
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Rep 9(3):208–218
Asma BM (2007) Malatya: world’s capital of apricot culture. Chron Hortic 47(1):20–24
Bhandawat A, Sharma V, Singh P, Seth R, Nag A, Kaur J, Sharma RK (2019) Discovery and utilization of EST-SSR marker resource for genetic diversity and population structure analyses of a subtropical bamboo, Dendrocalamus hamiltonii. Biochem Genet 57(5):652–672
Bhat MY, Padder BA, Wani IA, Banday FA, Ahsan H, Dar MA, Lone AA (2013) Evaluation of apricot cultivars based on physicochemical characteristics observed under temperate conditions. Int J Agric Sci 3(5):535–537
Bourguiba H, Krichen L, Audergon JM, Khadari B, Trifi-Farah N (2010a) Impact of mapped SSR markers on the genetic diversity of apricot (Prunus armeniaca L.) in Tunisia. Plant Mol Biol Rep 28(4):578–587
Bourguiba H, Khadari B, Krichen L, Trifi-Farah N, Santoni S, Audergon JM (2010b) Grafting versus seed propagated apricot populations: two main gene pools in Tunisia evidenced by SSR markers and model-based Bayesian clustering. Genetica 138(9–10):1023–1032
Bourguiba H, Audergon JM, Krichen L, Trifi-Farah N, Mamouni A, Trabelsi S, Khadari B (2012a) Genetic diversity and differentiation of grafted and seed propagated apricot (Prunus armeniaca L.) in the Maghreb region. Sci Hortic 142:7–13
Bourguiba H, Audergon JM, Krichen L, Trifi-Farah N, Mamouni A, Trabelsi S, D’Onofrio C, Asma BM, Santoni S, Khadari B (2012b) Loss of genetic diversity as a signature of apricot domestication and diffusion into the Mediterranean Basin. BMC Plant Biol 12(1):49
Burgos L, Petri C, Badenes ML (2007) Prunus spp. In: Pua EC, Davey MR (eds) Biotechnology in agriculture and forestry, vol 60. Springer, Heidelberg, pp 283–307
Carraut A, Crossa-Raynaud P (1974) L’amélioration des variétés d’abricotier en Tunisie. Ann Inst Nat Rech Agro Tun 47:3–33
Chen C, Chu Y, Ding C, Su X, Huang Q (2020) Genetic diversity and population structure of black cottonwood (Populus deltoides) revealed using simple sequence repeat markers. BMC Genet 21(1):2
Crossa-Raynaud P (1960) Problèmes d’arboriculture fruitière en Tunisie. Ann Inst Nat Rech Argo Tun 33:39–63
Dvin SR, Gharaghani A, Pourkhaloee A (2020) Genetic diversity, population structure, and relationships among wild and domesticated almond (Prunus spp.) germplasms revealed by ISSR markers. Adv Hortic Sci 34(3):287–300
Ercisli S, Agar G, Yildirim N, Esitken A, Orhan E (2009) Identification of apricot cultivars in Turkey (Prunus armeniaca L.) using RAPD markers. Rom Biotechnol Lett 14(4):4582–4588
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620
Faust M, Surányi D, Nyujtó F (1998) Origin and dissemination of apricot. Hort Rev 22:225–266
Ganopoulos IV, Kazantzis K, Chatzicharisis I, Karayiannis I, Tsaftaris AS (2011) Genetic diversity, structure and fruit trait associations in Greek sweet cherry cultivars using microsatellite based (SSR/ISSR) and morpho-physiological markers. Euphytica 181(2):237–251
Gebrehiwet M, Haileselassie T, Gadissa F, Tesfaye K (2019) Genetic diversity analysis in Plectranthus edulis (Vatke) Agnew populations collected from diverse geographic regions in Ethiopia using inter-simple sequence repeats (ISSRs) DNA marker system. J Biol Res Thessalon Greece 26(1):7
Gelotar MJ, Dharajiya DT, Solanki SD, Prajapati NN, Tiwari KK (2019) Genetic diversity analysis and molecular characterization of grain amaranth genotypes using inter simple sequence repeat (ISSR) markers. Bull Natl Res Cent 43(1):103
Girish K, Tsering S, Dwivedi SK, Ashish Y, Srivastava RB (2012) Pomological and fruit quality characteristics of Halman and Raktsey-Karpo apricot cultivars of trans-Himalayan Ladakh region, India. Progress Hortic 44(2):211–214
Gürcan K, Öcal N, Yılmaz KU, Ullah S, Erdoğan A, Zengin Y (2015) Evaluation of Turkish apricot germplasm using SSR markers: genetic diversity assessment and search for Plum pox virus resistance alleles. Sci Hortic 193:155–164
Hajiyev ES, Akparov ZI, Aliyev RT, Saidova SV, Izzatullayeva VI, Babayeva SM, Abbasov MA (2015) Genetic polymorphism of durum wheat (Triticum durum Desf.) accessions of Azerbaijan. Russ J Genet 51(9):863–870
Hammer Ø, Harper DA, Ryan PD (2001) PAST: Paleontological statistics software package for education and data analysis. Palaeontol Electron 4(1):9
Hormaza JI, Yamane H, Rodrigo J (2007) Apricots. In: Kole C (ed) Genome mapping and molecular breeding in plants, volume 4: fruits and nuts. Springer, Berlin, pp 171–187
Hu X, Zheng P, Ni B, Miao X, Zhao Z, Li M (2018) Population genetic diversity and structure analysis of wild apricot (Prunus armeniaca L.) revealed by SSR markers in the Tien-Shan mountains of China. Pak J Bot 50(2):609–615
Ivanovych YI, Udovychenko KM, Bublyk MO, Volkov RA (2017) ISSR-PCR fingerprinting of Ukrainian sweet cherry (Prunus avium L.) cultivars. Cytol Genet 51(1):40–47
Izzatullayeva V, Akparov Z, Babayeva S, Ojaghi J, Abbasov M (2014) Efficiency of using RAPD and ISSR markers in evaluation of genetic diversity in sugar beet. Turk J Biol 38(4):429–438
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Krichen L, Audergon JM, Trifi-Farah N (2014) Assessing the genetic diversity and population structure of Tunisian apricot germplasm. Sci Hortic 172:86–100
Krichen L, Martins JM, Lambert P, Daaloul A, Trifi-Farah N, Marrakchi M, Audergon JM (2008) Using AFLP markers for the analysis of the genetic diversity of apricot cultivars in Tunisia. J Am Soc Hortic Sci 133(2):204–212
Kumar D, Lal S, Ahmed N (2015) Morphological and pomological diversity among apricot (Prunus armeniaca L.) genotypes grown in India. Indian J Agric Sci 85(10):1349–1355
Kumar M, Mishra GP, Singh R, Naik PK, Dwivedi S, Ahmad Z, Singh SB (2009a) Genetic variability studies among apricot populations from cold arid desert of Ladakh using DNA markers. Indian J Hort 66(2):147–153
Kumar M, Mishra GP, Singh R, Kumar J, Naik PK, Singh SB (2009b) Correspondence of ISSR and RAPD markers for comparative analysis of genetic diversity among different apricot genotypes from cold arid deserts of trans-Himalayas. Physiol Mol Biol Plants 15(3):225
Lamia K, Hedia B, Jean-Marc A, Neila TF (2010) Comparative analysis of genetic diversity in Tunisian apricot germplasm using AFLP and SSR markers. Sci Hortic 127(1):54–63
Li M, Zhao Z, Miao XJ (2013) Genetic variability of wild apricot (Prunus armeniaca L.) populations in the Ili Valley as revealed by ISSR markers. Genet Resour Crop Evol 60(8):2293–2302
Li M, Zhao Z, Miao X (2014a) Genetic diversity and relationships of apricot cultivars in North China revealed by ISSR and SRAP markers. Sci Hortic 173:20–28
Li M, Zhao Z, Miao X, Zhou J (2014b) Genetic diversity and population structure of Siberian apricot (Prunus sibirica L.) in China. Int J Mol Sci 15(1):377–400
Liu MP, Du HY, Zhu GP, Fu DL, Tana WY (2015) Genetic diversity analysis of sweet kernel apricot in China based on SSR and ISSR markers. Genet Mol Res 14:9722–9729
Liu J, Liao K, Nasir M, Zhao S, Sun Q, Peng X (2016) Analysis of genetic diversity of the apricot germplasm in the southern region of the Tianshan Mountains in Xinjiang, China using the ISSR technique. Eur J Hortic Sci 81(1):37–43
Lusini I, Velichkov I, Pollegioni P, Chiocchini F, Hinkov G, Zlatanov T, Cherubini M, Mattioni C (2014) Estimating the genetic diversity and spatial structure of Bulgarian Castanea sativa populations by SSRs: implications for conservation. Conserv Genet 15(2):283–293
Madhumati B (2014) Potential and application of molecular markers techniques for plant genome analysis. Int J Pure App Biosci 2(1):169–188
Malik SK, Chaudhury R, Dhariwal OP, Mir S (2010) Genetic diversity and traditional uses of wild apricot (Prunus armeniaca L.) in high-altitude north-western Himalayas of India. Plant Genet Resour 8(3):249
Meng J, He SL, Li DZ, Yi TS (2016) Nuclear genetic variation of Rosa odorata var. gigantea (Rosaceae): population structure and conservation implications. Tree Genet Genomes 12(4):1–14
Mir JI, Ahmed N, Rashid R, Wani SH, Sheikh MA, Mir H, Parveen I, Shah S (2012) Genetic diversity analysis in apricot (Prunus armeniaca L.) germplasm using RAPD markers. Indian J Biotechnol 11:187–190
Mondini L, Noorani A, Pagnotta MA (2009) Assessing plant genetic diversity by molecular tools. Diversity 1(1):19–35
Pacheco-Olvera A, Hernandez-Verdugo S, Rocha-Ramirez V, Gonzalez- Rodriguez A, Oyama K (2012) Genetic diversity and structure of pepper (Capsicum Annuum L.) from Northwestern Mexico analyzed by microsatellite markers. Crop Sci 52(1):231–241
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2(3):225–238
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98(1):107–112
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Reddy MP, Sarla N, Siddiq EA (2002) Inter simple sequence repeat (ISSR) polymorphism and its application in plant breeding. Euphytica 128(1):9–17
Rohlf FJ (2000) NTSYSpc: Numerical Taxonomy and Multivariate Analysis System. Version 2.02. Exeter Software, Setauket, New York, USA
Shangguan LF, Wang YZ, Li XY, Wang YJ, Song CN, Fang JG (2012) Identification of selected apricot cultivars using RAPD and EST-SSR markers. Caryologia 65(2):130–139
Sharma JK (2000) Morphological studies on apricot and its wild relatives. J Hill Res 13(1):5–10
Smith FW, Ealing PM, Dong B, Delhaize E (1997) The cloning of two Arabidopsis genes belonging to a phosphate transporter family. Plant J 11(1):83–92
Torokeldiev N, Ziehe M, Gailing O, Finkeldey R (2019) Genetic diversity and structure of natural Juglans regia L. populations in the southern Kyrgyz Republic revealed by nuclear SSR and EST-SSR markers. Tree Genet Genomes 15(1):5
Tripathi N, Saini N, Mehto V, Kumar S, Tiwari S (2012) Assessment of genetic diversity among Withania somnifera collected from central India using RAPD and ISSR analysis. Med Aromatic Plant Sci Biotechnol 6(1):33–39
Vavilov NI (1951) The origin, variation, immunity and breeding of cultivated plants. Soil Sci 72:482
Wani AA, Zargar SA, Malik AH, Kashtwari M, Nazir M, Khuroo AA, Ahmad F, Dar TA (2017) Assessment of variability in morphological characters of apricot germplasm of Kashmir, India. Sci Hortic 225:630–637
Yılmaz KU, Ercişli S, Asma BM, Doğan Y, Kafkas S (2009) Genetic relatedness in Prunus genus revealed by inter-simple sequence repeat markers. HortScience 44(2):293–297
Yilmaz KU, Paydas-Kargi S, Dogan Y, Kafkas S (2012) Genetic diversity analysis based on ISSR, RAPD and SSR among Turkish Apricot Germplasms in Iran Caucasian eco-geographical group. Sci Hortic 138:138–143
Acknowledgements
The authors sincerely acknowledge the support and cooperation from the Head Department of Botany, Punjabi University Patiala, Director Horticulture Kashmir Division, and Director CITH Rangreth, Srinagar. We also sincerely thank all those orchardists who allowed us sample collections during field surveys.
Funding
This work was supported by the Department of Biotechnology (DBT), Govt. of India under Grant No. BT/PR11394/PBD/16/1077/2014.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animals rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zargar, S.A., Saggoo, M.I.S., Wani, A.A. et al. Genetic diversity, population structure and genetic relationships in apricot (Prunus armeniaca L.) germplasm of Jammu and Kashmir, India using ISSR markers. Genet Resour Crop Evol 69, 255–270 (2022). https://doi.org/10.1007/s10722-021-01225-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-021-01225-1