Abstract
Understanding the genetic diversity among accessions and germplasm is an important requirement for crop development as it allows for the selection of diverse parental combinations for enhancing genetic gain in varietal selection, advancement and release. The study aimed to characterize 183 provitamin A cassava (Manihot esculenta Crantz) accessions and five Sierra Leonean varieties using morphological traits, total carotenoid content and SNP markers to develop a collection for conservation and further use in the cassava breeding program. Both morphological parameters and 5634 SNP markers were used to assess the diversity among the provitamin-A cassava accessions and varieties. Significant differences were observed among the accessions for most of the traits measured. The first five PCs together accounted for 70.44% of the total phenotypic variation based on yield and yield components among the 183 provitamin-A cassava accessions and five Sierra Leonean varieties. The present study showed that provitamin-A cassava accessions in Sierra Leone have moderate to high diversity based on morphological and molecular assessment studies. The similarity index among the 187 and 185 cassava accessions grouped them into 6 and 9 distinct clusters based on morphological and molecular analyses, respectively. A significant positive, but low correlation (r = 0.104; p < 0.034), was observed between the two dendrograms. The results obtained will serve as a guide and basis of germplasm management and improvement for total carotenoid content, yield and African cassava mosaic disease resistance in Sierra Leone.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic diversity provides species with the ability to adapt to changing environments. Several studies have been reported on the use of morphological descriptors to determine the genetic diversity among cassava genotypes (Rimoldi et al. 2010; Asare et al. 2011; Thompson 2013). Recent advances in molecular biology techniques have led to the development of important tools for genetic diversity study in several plant species. The accuracy in accession characterization may therefore, be enhanced/achieved with the use of molecular markers associated with morphological traits.
Previous studies in plant genetic diversity used DNA molecular markers for beta carotene improvement in cassava (Ferreira et al. 2008; Rimoldi et al. 2010), and included amplified fragment length polymorphism (Benesi et al. 2010), simple sequence repeats (Alves et al. 2011; Parkes 2009; Oliveria et al. 2012; Costa et al. 2013) and single nucleotide polymorphism (Kizito et al. 2005; Tangphatsornruang et al. 2008; Ferguson et al. 2011; Thompson 2013; Rabbi et al. 2015). With recent advances in high throughput genotyping technologies, single nucleotide polymorphism markers (SNPs) are increasingly becoming markers of preference for plant genetic studies and breeding.
SNPs are the most common types of genetic variation among species, involving just a change in a single nucleotide. Expressed Sequence Tags (ESTs) have been exploited to explain and detect SNPs in maize (Zea mays L.) (Ching et al. 2002) and soybean (Glycine max L. Merr.) (Zhu et al. 2003). Lopez et al. (2005) and Rabbi et al. (2014, 2015) have also reported SNPs detection from ESTs in cassava. Cassava being an outbreeding and highly heterogeneous crop, possesses an extreme level of phenotypic plasticity, and thereby, lacks the potential for unified classification system for cultivars (Kawano 1978). Consequently, characterization of agronomic traits becomes a challenge. To conduct a successful genetic diversity study on cassava germplasm in Sierra Leone, there is a need to unravel the genetic potential existing among Sierra Leone’s cassava breeding program, which consists of fourteen released varieties and provitamin-A cassava accessions induction from Institute of International Tropical Agriculture, Nigeria. Thus, the need for assessing and understanding the genetic diversity among the provitamin-A cassava accessions and identifying gaps to be filled within the breeding program in Sierra Leone is required.
The objectives of the study, therefore, were to characterize, quantify and exploit the diversity of 183 provitamin-A cassava accessions and five Sierra Leonean varieties using morphological traits, SNP markers and total carotene content and to develop a collection for conservation and future use in the breeding programmes.
Materials and methods
Germplasm sources and experimental design
The plant materials used in the study consisted of 183 provitamin-A cassava accessions known for their varying levels of provitamin- A properties, obtained from the International Institute of Tropical Agriculture (IITA, Ibadan, Nigeria) and established at the Taiama experimental site in Sierra Leone, in 2014 (Table 1) and five Sierra Leonean cassava varieties. The trial was established and evaluated during the cropping season of 2015–2016 at the Njala Agricultural Research Institute (NARC), Foya crop site, Njala, representing the transitional rain forest agro-climatic zone (Van Vuure et al. 1972; Odell et al. 1974). The trial was laid out in an Alpha lattice design with two replications, and each replication had four blocks with 47 entries per block. The blocks were separated by 1 m and 2 m alleys between and within blocks to reduce intra and inter block plant competition, respectively. Each entry was grown on 10 m row ridge at a spacing of 1 m × 1 m between and within ridges, respectively. Cassava cuttings of 20–25 cm length were obtained from healthy stem cuttings and horizontally planted.
Morphological traits
Agro-morphological data was collected at 1, 3, 6 and 9 months after planting (MAP) on the parameters listed below using the IITA cassava descriptor (Fukuda et al. 2010) (Table 2).
Harvesting was done at 12 MAP (August–September). The following parameters were taken at harvest: number of marketable roots (expressed as count numbers), number of non-marketable roots (expressed as count numbers), total number of storage roots (expressed as count numbers), roots weight/tuber (kg), inner skin color, and outer skin color, ease of peel, root shape, marketable weight (kg), and non- marketable weight (kg). Dry matter content, expressed in percentage was determined by selecting three representative storage roots. Slices of the fresh root were randomly selected and weighed to obtain a 100 g fresh mass sample per genotype, before being dried for 48 h in an oven at 80 °C. The dried samples were then re-weighed to obtain the dry mass. Disease occurrence and intensity were mostly measured in the 1st, 3rd, 6th and 9th month after planting.
Molecular characterization
The Dellaporta method of DNA extraction (Dellaporta et al. 1983) was carried out at the International Institute of Tropical Agriculture (IITA), Ibadan, Nigeria. For genotyping-by-sequencing library preparation, the ApekI restriction enzyme (recognition site: G|CWCG) that produces less variable distributions of read depth, and therefore, a larger number of scorable SNPs in cassava (Hamblin and Rabbi 2014) was used. Two 96-plex GBS libraries were constructed as described by Elshire et al. (2011) and sequenced at the Institute of Genomic Diversity at Cornell University, using the Illumina HiSeq 2500. Raw read sequences were processed through cassava GBS production pipelines developed using TASSEL 5.0V2. The GBS-derived SNPs were further filtered using the TASSEL software (Bradbury et al. 2007) to retain only polymorphic SNPs. Initially, filtered for minor allele frequency (MAF < 0.05), the generated 5634 SNPs were processed under the Next Generation Cassava project. The resulting SNP dataset was used for the diversity analysis study among the 188 cassava accessions already phenotyped and analyzed. Results from both the phenotype and genotype analyses were compared to check the correspondence between the two.
Data analysis
Agro-morphological data sets from this study were subjected to selected statistical packages for analysis. Analytical procedures comprised the following softwares and statistical procedures: descriptive statistics using XLSTAT (2010), MINITAB 15 and STATA 13. Principal Component Analysis (PCA) were performed using Princomp software to examine the structure of the correlations between the variables using SAS 9.3. Cluster analyses, based on Agro-morphological and SNP markers data sets, were performed to group observations together using the method of Ward’s minimum variance distance using SAS 9.4. A dendrogram was plotted from the computed similarity values for each Agro-morphological traits and SNP markers to show the relationship among the accessions. The provitamin-A studied accessions and varieties were grouped based on the varying levels of total carotenoid content.
Basic diversity indices for the 183 provitamin-A studied accessions and varieties were calculated using Power marker (Liu and Muse 2005) and GenAlex version 6.41 (Peakall and Mouse 2006). The Power maker software was used to generate the following statistics: number of alleles per locus, major allele frequency, observed heterozygosity (Ho), expected heterozygosity (He) and polymorphic information content (PIC) (Bostein et al. 1980). PIC values were calculated with the equation:
where: ΣP2i = sum of each squared ith haplotype frequency.
A Mantel matrix test (Mantel 1967) was carried out to compare the extent of agreement between dendrograms derived from morphological and molecular data using the distance matrices. The pairwise genetic distance (identity-by-state, IBS) matrix was calculated among all individuals using PLINK (Purcell et al. 2007). A Ward’s minimum variance hierarchical cluster dendrogram was built from the IBS matrix, using the analyses of phylogenetic and evolution (ape) package in R.
Results and discussion
Summary statistics of morpho-agronomic traits of 183 provitamin-A studied accession and varieties
Table 3 shows summary statistics of some morpho-agronomic traits of 183 provitamin-A studied accessions and varieties. Sprouting was only recorded in the first month after planting (MAP) and ranged from 65 to 100% among the 183 provitamin-A studied accessions and varieties with an average of 9.56 seeds sprouted in the first month. Severity scores for African Cassava Mosaic Disease Cassava Bacterial Blight and Cassava Green Mite variably ranged from 0 to nine in the studied collection consisting of the 183 provitamin-A cassava collection and the five varieties. Percent incidence for African Cassava Mosaic Disease, Cassava Bacterial Blight and Cassava Green Mite variably ranged from 0 to 9. Most of the morphological characters both quantitative and qualitative were taken in the 3rd, 6th, 9th and 12th MAP. Color of apical lobe ranged from 3 to 9 about a mean of 6.8 ± 1.61 3 MAP; whereas the same traits scored ranged from 0 to 9 about a mean of 6.71 ± 1.74 9 MAP. Plant height ranged from 65.5 to 284.5 cm at 6 MAP about a mean of 155.69 ± 26.12 cm. Leaf area ranged from 10.24 to 73.93 cm2 at 6 MAP; whereas leaf retention ranged from 1.75 to 4.5 at the same time. All yield related traits were recorded at 12 MAP. Yield per hectare ranged from 0.2 to 42.5 t/ha; while dry matter content ranged from 4.0 to 44.5% (Table 3). These parameters which were good indicators of growth showed considerable variation for the morpho-agronomic traits evaluated in the study, and the findings were in concordance with previous studies by Mbah et al. (2019) who reported that agro morphological parameters exert strong influence on cassava root yield. In the present study, descriptive analysis of the 183 provitamin-A studied accessions and varieties based on various traits showed high variability among the accessions. The significant variation observed among the 183 provitamin-A studied accessions and varieties studied for these economically important traits, such as African cassava mosaic disease, yield and dry matter content (DMC) offers a prospect for progress in cassava breeding program in Sierra Leone. Diversity studies of cassava germplasm has been widely undertaken worldwide (Bolanos 2001; Chavez et al. 2005; Morillo 2009; Fregene 2007; Parkes 2011; Njoku 2012; Thompson 2013) with little or no attention in Sierra Leone. These findings agree with the findings by Carvalho and Schaal (2001) who reported, in Brazil, a high degree of variability among 94 cassava accessions of Brazilian origin. Raghu et al. (2007) in a similar study, in India, also identified a high level of diversity among 58 cassava accessions from South Indian cassava germplasm based on 29 morphological traits. Lyimo et al. (2012) reported significant variability among 39 cassava accessions of Tanzanian origin using 14 morphological traits. Thompson (2013) observed a moderate to high diversity among 150 Ghanaian landraces and introduced accessions from IITA, Ibadan, Nigeria using 25 morphological traits in Ghana.
Summary statistics of the genetic variation among the 183 provitamin-A studied accessions and varieties using SNP markers
Summary statistics for number of alleles observed, expected heterozygosity and polymorphic information content are presented in Table 4. The number of observed alleles ranged from 1.30 to 1.47, with an average of 1.38 alleles per locus. The expected heterozygosity was the lowest for TR 1233 (0.15) and SLICASS 6 (0.15) and highest in TR 1525 (0.23), with a mean of 0.19. The observed heterozygosity per individual observation ranged from 0.30 (TR 1233) to 0.47 (TR 1525) with a mean of 0.38. The mean of observed heterozygosity (0.38) was moderately higher than the expected heterozygosity (0.19). This substantiates the difference in the relatedness of most of the provitamin-A studied accessions which were developed from varieties of half sib families with different female know parental sources (Female plants) been pollinated by different sources. However, the major allele frequency (MAF) of all the ‘markers used in the observations was generally, below 0.95, indicating that they were all polymorphic. PIC values ranged from 0.11 in TR 1233 to 0.18 in TR 1199 and TR 1525 with a PIC mean of 0.14. The higher the PIC value the more informative is the marker. Since morphological traits are influenced by the environment, molecular markers which are not influenced or controlled by the environment are preferable in genetic diversity studies (Kaemmer et al. 1992; Gepts 1993; Njoku 2012; Thompson 2013). The study carried out by Kawuki et al. (2009) was the first published report where SNPs were used for genetic diversity studies in cassava. They characterized and identified some SNP markers and assessed their utilization in cassava genetic diversity analysis assessment. The present study seems to be the first reported case in Sierra Leone, where SNP markers were used in cassava diversity study of provitamin-A cassava accessions. Using the 5634 SNP markers, 95% of them were polymorphic. The informativeness of a genetic marker is measured by the polymorphic information content (PIC). The mean PIC value observed in this study (0.14) is relatively lower than previously reported. Indeed, Kawuki et al. (2009) reported a PIC value of 0.29 in 74 cassava accessions using 26 SNP: while Thompson (2013) also reported PIC value of 0.29 using 150 cassava accessions. PIC values for SNP markers in cassava are generally lower than observed in genetic diversity studies in other crops. For instance, Yang et al. (2011) reported PIC value of 0.34 in maize genotypes using 884 SNP markers.
Principal component analysis among yield and yield related traits of 183 provitamin-A cassava studied accessions and varieties
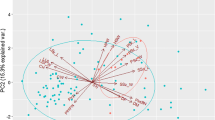
The first five PCs together accounted for 70.44% of the total phenotypic variation among the 183 provitamin-A cassava studied accessions and five varieites (Table 5). PC1 axis had an eigenvalue of 4.44 and acounted for 27.74% of the total variation, whereas PC, PC3, PC4 and PC5 axes had eigenvalues of 3.1, 1.45, 1.11% and 1.09% acounted for 19.8%, 9.03%, 6.95% and 6.82% of the total variation, respectively. Marketable root, marketable weight and yield had positive loadings on PC1. Non-marketable weight, storage root weight and ease of peel had positive loadings on PC2. Unmarketable root and total number of storage roots had negative loadings in PC3. Root Size had a positive loading in PC4 and Inner color had a positive loading in PC5.
Principal Component Analysis is a technique which identifies plant traits that contribute most to the observed variation within a group of 183 provitamin-A studied accessions and five varieties. The tool has a practical application in the selection of parent lines for breeding purposes and varietal development. The cumulative variance of 70.44% by the first five axes with eigen values > 1.0 indicates that the identified traits within these axes exhibited great influence on the phenotype of these accessions, and could effectively be used for selection among them. This study agrees with findings of Afuape and Nwachukwu (2005; Afuape et al. 2010), who reported a cumulative variance of 70.09% for the first three axes in the dry evaluation of nine sweetpotato genotypes, weight of total roots, weight of biomass, and dry matter as the important traits that distinguished the elite materials been researched on.
Cluster groupings of the studied accessions and varieties based on morpho-agronomic traits using ward’s minimum variance and SNP markers
Agro-morphological traits diversity analysis: The dendrogram constructed based on the data generated from the agro-morphological traits divided the provitamin-A studied accessions and five varieties into six major clusters (A to F), and at a genetic distance of 0.30, and each had sub clusters apart from Cluster A (Table 6). Cluster A consisted of only two cassava accession germplasm with no sub clusters. Cluster B, had two sub cluster, Cluster D recorded the highest number of accessions, 57 in total, followed by Cluster E and F, grouping 53 and 34 accessions, respectively. In general, most of the accessions in this study were grouped according to their morpho-agronomic traits and geographical location. For example, the accessions in major Cluster E scored similar values for most of the morph-agronomic traits studied. Three out the five Sierra Leonean varieties developed in Sierra Leone were grouped into cluster F: while cluster B and D contained only provitamin-A studied accessions introduced to Sierra Leone in the form of seeds from IITA, Nigeria, and had a discrete pattern of clustering, which have been grouped more or less per their state, geographical distribution or country.
SNP markers diversity analysis: The 181 Provitamin-A cassava accession germplasm and 4 Sierra Leonean varieties were grouped into nine clusters based on the 5643 SNP markers (Fig. 1). Clusters A, B, C, D and E, had 21, 7, 11, 8, and 16 accessions, respectively; while cluster F, G, H and I consisted of 10, 47, 50 and 17 accessions, respectively (Table 7). Clusters A, B, C, E, G, H and I had 3, 1, 2, 4, 9, 10, and 1 accessions with varying levels of total carotenoid content. Cluster I consisted of only one provitamin-A studied accessions.
Correlation Analysis between Clusters from Agro-Morphological Traits and SNP Makers: A comparison of the two dendrogram based on Mantel matrix test showed a significant positive, but weak correlation between the morphological and molecular data sets (r = 0.104, p < 0.034). In a similar study, Raghu et al. (2007) mentioned that 24 morphological traits out of 28, contributed to the total variation observed. Here, our clustering study showed six and nine distinct clusters based on morphological and molecular analyses, respectively, indicating a large variability in the collection. In a similar study, Carvalho and Schaal (2001) identified 22 distinct clusters using 94 cassava accessions in Brazil, whereas Raghu et al. (2007) identified six distinct groups using 58 accessions. Our study is, therefore, in agreement with all these studies. Although the morphological and SNP data grouped the accessions into six and nine distinct clusters, respectively, some similarities were observed. Accessions TR 0747 and TR 0365 which were selected as provitamin-A studied accessions were found to be closely similar using both morphological and genetic markers. This could explain why the morphological and molecular analysis showed similar accessions between the two clusters. There are no reports on the genetic diversity of provitamin-A cassava accessions using morphological traits, molecular markers and total carotenoid content so far. This remains the first study using morphological, genetic diversity characterization and total carotenoid content levels of our provitamin-A cassava accessions in Sierra Leone.
The study reveals a moderate degree of diversity among the provitamin-A cassava accessions and varieties which can be further used for crop improvement. This may provide an opportunity to enhance and boost the breeding strategy.
Thirty provitamin-A studied accessions with varying levels of total carotenoid content, yield and dry matter content
The 30 accessions grouped in the different clusters were selected as provitamin-A studied accessions for formation of core collection, conservation and improvement in the breeding program. These accessions were selected based on the higher levels of total carotene content after laboratory analysis using color chat and the i-check device. The core selected provitamin-A cassava accessions across different clusters revealed significant variation of total carotenoid content, yield and dry matter. These provitamin-A cassava accessions TR 0998, TR 0222, TR 1337 and TR 0461 contained higher levels of total carotenoid content with TR 0365 been the lowest. Dry matter content ranged from 12.5 (TR 0696) to 39.5 (TR 1208) with yield ranging from 2.0 (TR 0461) to 22.8 (TR 0232) in the study provitamin-A accessions. TR 0747, TR 1337, TR 0232, TR 0998 and TR1755 clustered similarly morphologically and genetically (B, D, E, E and E). The wide range of total carotenoid content, dry matter content, yield, and distribution of morphological variability revealed in the study might provide a broader scope for the crop’s improvement through hybridization and selection. The higher dry matter content and significant variability observed in some provitamin-A cassava accessions in this study contradict findings reported by Esuma et al. (2012) who reported high DMC and low total carotenoid content for local white cassava root varieties using the Ugandan landraces.
Conclusion
The present morphological and molecular assessment studies reported that provitamin-A cassava accessions in Sierra Leone have moderate to high diversity based on total carotenoid content, morphological, and molecular assessment (Table 8).
The inter-relationships of morpho- agronomic factors in determining cassava fresh root yield based on provitamin-A cassava accessions require additional research to fully understand concept of improving total carotenoid content and yield on provitamin-A cassava accession germplasm. Even though the agro-morphological traits are generally employed to estimate genetic diversity in crop plants, such a method has its own limitations as the traits are heavily influenced by the environmental conditions and climate being the main factor influencing the growth and development of the species (Cadena Iniguez and Arevalo Galarza 2011). This also confirms the importance of molecular techniques and markers on Provitamin-A cassava accession germplasm to carry out successful research and improvement studies. The present study has revealed that during provitamin-A cassava variety development, high dry matter content (quality trait) is a priority trait that should be considered at both primary and advance (yield evaluation) stages with good root qualities to facilitate adoption after varietal release.
Finally, the genetic diversity revealed from this study would provide the cassava breeding program in Sierra Leone an opportunity to boost the breeding strategy on crop genetic improvement for Provitamin-A cassava varieties with end-use preferred traits (total carotenoid content, dry matter, yield and African cassava mosaic disease resistance).
References
Afuape SO, Nwachukwu EC (2005) Variability and correlation studies in some quantitative characters in selected sweetpotato (Ipomea batatas (L.) Lam) genotypes. Genetics and Sustainable Agriculture. In: Proceedings of the 30th annual conference of genetic society of Nigeria held at the University of Nigeria, Nsukka, 5th–8th September, 2005, pp 124–129
Afuape SO, Okocha PI, Njoku D (2010) Multivariate assessment of the agro-morphological variability and yield components among sweetpotato (Ipomoea batatas (L.) Lam) landraces. Afr J Plant Sci 5(2):123–132.
Alves PA, Peroni N, Abreu AG, Gribel R, Clement CR (2011) Genetic structure of traditional varieties of bitter manioc in three soils in central Amazonia. Genetica 139:1259–1271
Asare PA, Labuschagne MT, Herselman L, Mahungu N (2011) Morphological and molecular based diversity studies of some cassava (Manihot esculenta Crantz) germplasm in Ghana. Afr J Biotechnol 10:13900–13908
Benesi IRM, Labuschagne MT, Herselman L, Mahungu N (2010) Ethnobotany, morphology and genotyping of cassava germplasm from Malawi. J Biol Sci 10:616–623
Bolanos EA (2001) Caracterización de la diversidad genética en cuanto a contenido de carotenos deraíces y hojas de 682 Genotipos de yuca (Manihot esculenta Crantz) de la colección de CIAT y 158de 411 Genotipos en cuanto a minerales. Universidad Nacional de Colombia, Sede Palmira, Tesis de grado
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. J Hum Genet 32(3):314–331
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler S (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Cadena Iniguez J, Arevalo Galarza ML (2011) Las Variedades del Chayote Mexicano, Recurso Ancestral con Potencial de Commercialization. College of Postgraduates Mexico, Grupo Interdisciplinario de Investigacion en Sechium edule en Mexico, AC
Carvalho LJ, Schaal BA (2001) Assessing genetic diversity in the cassava (Manihot esculenta Crantz) germplasm collection in Brazil using PCR-based markers. Euphytica 120:133–142
Chavez AL, Sanchez T, Jaramillo G, Bediya JMI, Echeverry J, Bolanos EA, Ceballos H, Iglesias CA (2005) Variation of quality traits in cassava roots evaluated in landraces and improved clones. Euphytica 143:125–133
Ching AKS, Caldwell M, Jung M, Dolan O, Smith S et al (2002) SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet 3:19
Costa TR, Vidigal-Filho PS, Goncalves-Vidigal MC, Galvan MZ, Lacanallo GF, Silva LI, Kvitschal MV (2013) Genetic diversity and population structure of sweet cassava using simple sequence repeat (SSR) molecular markers. Euphytica 120(143–157):77
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA mini preparation: version II. Plant Mol Biol Report 4:19–21
Elshire RJ, Glaubitz Q, Sun J, Poland J, Kawamoto K, Buckler E, Mitchell S (2011) A robust, simple genotyping-by sequencing (GBS) approach for high diversity species. PLoS ONE 6:E19379. https://doi.org/10.1371/journalpone.0019379
Esuma W, Rubaihayo P, Pariyo A, Kawuki R, Wanjala B, Nzuki I, Harvey JW, Baguma Y (2012) Genetic diversity of provitamin-A cassava in Uganda. J Plant Sci 1(1):60
Ferguson LC, Green J, Surridge A, Jiggins CD (2011) Evolution of the insect yellow gene family. Mol Biol Evol 28(1):257–272. https://doi.org/10.1093/molbev/msq192
Ferreira CF, Alves E, Pestana KN, Junghans DT, Kobayashi AK, V’de Santos J, Fukudu W (2008) Molecular characterization of cassava (Manihot esculenta Crantz) with yellow orange roots for beta carotene improvement. Crop Breed Appl Biotechnol 8:23–29
Fregene MA, Blair MW, Beebe SE, Ceballos H (2007) Marker-assisted selection in common beans and cassava. In: Marker-assisted selection (MAS) in crops, livestock, forestry and fish: status and the way forward. FAO publishing, pp 471
Fukuda WMG, Guevara CL, Kawuki R, Ferguson ME (2010) Selected morphological and agronomic descriptors for the characterization of cassava. International Institute of Tropical Agriculture (IITA), Ibadan, pp 5–9
Gepts P (1993) The use of molecular and biochemical markers in crop evolution studies. Evolut Biol 27:15–94
Hamblin MT, Rabbi IY (2014) The effects of restriction-enzyme choice on properties of genotyping-by-sequencing libraries: a study in cassava (Manihot esculenta). Crop Sci 54(6):2603–2608
Kaemmer D, Afza R, Weising K, Kahl G, Novak FJ (1992) Oligonucleotide and amplification fingerprinting of wild species and cultivars of banana (Musa spp). Biol Technol 10:1030–1035
Kawano K (1978) Genetic improvement of cassava (Manihot esculenta Crantz) for productivity. Trop Agric Res Ser 11:9–21
Kawuki R, Ferguson S, Labuschagne M, Herselman ML, Kim DJ (2009) Identification characterization and application of single nucleotide polymorphisms for diversity assessment in cassava (Manihot esculenta Crantz). Mol Breed 9:669–9280
Kizito EB, Bua A, Fregene M, Egwang T, Gullberg U, Westerbergh A (2005) The effect of cassava mosaic disease on the genetic diversity of cassava in Uganda. Euphytica 146:45
Liu K, Muse SV (2005) Power marker: integrated analysis environment for genetic marker data. Bioinformatics 221(9):2128–2129
Lopez C, Piegu B, Cooke R, Delseny M, Tohme J, Verdier V (2005) Using cDNA and genomic sequences as tools to develop SNP strategies in cassava (Manihot esculenta Crantz). Theor Appl Genet 110:425–431
Lyimo LD, Tairo FM, Rweyemamu CL (2012) Morphological and molecular characterization of cassava (Manihot esculenta Crantz) landraces using SSR markers. Internal J Integr Biol 13(2):71–77
Mantel N (1967) The detection of disease clustering and generalized regression approach. Cancer Res 27:209–220
Mbah EU, Nwankwo BC, Njoku DN, Gore MA (2019) Genotypic evaluation of twenty-eight high- and low-cyanide cassava in low-land tropics, southeast Nigeria. Heliyon. https://doi.org/10.1016/j.heliyo.e01855
Morillo CY (2009) Inheritance of carotenoids content in cassava roots (Manihot esculenta Crantz). Ph.D. Dissertation. National University of Colombia, Palmira Campus
Njoku D (2012) Improving beta-carotene content in farmers’ preferred cassava cultivars in Nigeria. Ph.D. Thesis. West Africa Centre for Crop Improvement. University of Ghana. Legon. Accra. Ghana
Odell RT, Dijkerman JC, Van Vuure W, Melsted SW, Beavers AH, Sutton PM, Kurtz LT, Miedema R (1974) Characteristics, classification and adaptation of soils in selected areas of Sierra Leone West Africa. Bulletin 748.Agricultural Experiment Station, College of Agriculture, University of Illinois at Urbana-Champaign. Bulletin 4 Njala University College, University of Sierra Leone
Oliveira MVC, Baliza DP, Souza GA, Carvalho SP, Assis LHB (2012) Caracterização de clones de mandioca utilizando marcadores microssatélites. Revista Ciência Agronômica 43:170–176
Parkes E (2009) Using molecular markers to identify new sources of CMD resistant genes for cassava improvement. In: 2nd GSS workshop CIAT Cali
Parkes E (2011) Assessment of genetic diversity, combining ability, stability and farmer preference of cassava germplasm in Ghana Ph.D. Thesis. Department of Plant Sciences (Plant Breeding), in the Faculty of Natural and Agricultural Sciences at the University of the Free State. Bloemfontein South Africa
Peakall RS, Mouse PE (2006) GenAlEx 6: genetic analysis in excel. population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a toolset for whole-genome association and population-based linkage analysis. Am J Hum Genet 81:559–575
Rabbi I, Hamblin M, Melaku G, Kulakow P, Ferguson M, Kpan I, Delphine AS, Jean-Luc J (2014) Genetic mapping using genotyping-by-sequencing in the clonally propagated cassava. Int Plant Animal Genomics Conf, XXI, p 2013
Rabbi I, Kulakow P, Manu-Aduening P, Dankyi A, Asibuo J, Parkes EY, Abdoulaye T, Girma G, Gedil M, Ramu P, Reyes B, Maredia M (2015) Tracking crop varieties using genotyping by-sequencing markers: a case study using cassava (Manihot esculenta Crantz). BMC Genet 16:115. https://doi.org/10.1186/s12863-015-0273-1
Raghu D, Senthil N, Saraswathi T, Raveendran M, Gnanam R, Venkatachalam R, Shanmug Asundaram P, Mohan C (2007) Morphological and simple sequence repeats (SSR) based finger printing of South Indian cassava germplasm. Int J Integr Biol 1:141–148
Rimoldi F, Vidigal-Filho PS, Kvitschal MV, Conclaves Vidigal MC, Prioli AJ, Prioli SM, Costa TR (2010) Genetic divergence in sweet cassava cultivars using morphological agronomic traits and RAPD molecular markers. Braz Arch Biol Technol 53:1477–1486
Tangphatsornruang S, Sraphet S, Singh R (2008) Development of polymorphic markers from expressed sequence tags of Manihot esculenta Crantz. Mol Ecol Resour 8:682–685
Thompson R (2013) Genetic analysis of postharvest physiological deterioration in cassava (Manihot esculenta Crantz) storage roots. Ph.D. Thesis. West Africa Centre for Crop Improvement. University of Ghana. Legon. Accra. Ghana
Van Vuure W, Odell RT, Sutton PM (1972) Soil Survey of the Njala Area—Bulletin No. 3. Njala University College, University of Sierra Leone in cooperation with Ministry of Agriculture and Natural Resources, Sierra Leone
Yang X, Xu Y, Shah T, Li H, Han Z, Li J, Ya J (2011) Comparison of SSRs and SNPs in assessment of genetic relatedness in maize. Genetica 11:9606–96015
Zhu YL, Song QJ, Hyten DL, Tasell CP, Matukumali LK, Grim DR, Cregan PB (2003) Single nucleotide polymorphisms in Soybean. Genetics 163:1123–1134
Acknowledgements
Support for this research was provided by the West Africa Agricultural Productivity Programme Sierra Leone (WAAPP-1C) through a Grant acquired from the Word Bank. We are grateful to Sierra Leone Agricultural Research Institute, West Africa Centre for Crop Improvement (WACCI), University of Ghana, Legon and the International Institute of Tropical Agriculture (IITA), Ibadan, Nigeria for providing free access to their facilities.
Funding
West Africa Agricultural Productivity Programme Sierra Leone (WAAPP).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We the authors of this manuscript declare that we have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kamanda, I., Blay, E.T., Asante, I.K. et al. Genetic diversity of provitamin-A cassava (Manihot esculenta Crantz) in Sierra Leone. Genet Resour Crop Evol 67, 1193–1208 (2020). https://doi.org/10.1007/s10722-020-00905-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-020-00905-8