Abstract
The pearly razorfish Xyrichtys novacula (Linnaeus, 1758) is a sedentary benthic species distributed in both sides of the Atlantic Ocean and in the Mediterranean Sea. Previous cytogenetic analysis reported different diploid numbers in samples from Italy, Venezuela and Brazil. This research aims to test the hypothesis that samples from American Atlantic coast and Mediterranean Sea belong to the same single evolutionary lineage, characterized by intra-specific chromosome polymorphism. To this purpose a cytogenetic and molecular (mitochondrial COI sequences) survey was undertaken. Results revealed the existence of three different pearly razorfish molecular lineages: one present in Mediterranean Sea and two in the central and south American area, which are characterized by different karyotypes. One of these lineages shows substantial intra-population chromosomal polymorphism (2n = 45–48) determined by Robertsonian fusions that produce large metacentric chromosomes. On the whole data suggest that specimens morphologically identified as X. novacula correspond to three cryptic species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cytogenetic analyses have proved to be a useful tool in inferring evolutionary and phylogenetic paths within fish taxa, from intraspecific comparisons to higher taxonomic ranks (Artoni et al. 2009; Phillips 2013; Pansonato-Alves et al. 2014). In the last decade the combinations of the cytogenetic traditional methods with the mapping of repetitive sequences (Martins 2007; Cioffi et al. 2011; Cioffi and Bertollo 2012; Parise-Maltempi et al. 2013), and the analysis of sequences commonly used in phylogenetic and phylogeographic reconstruction (Nirchio et al. 2013, 2016) allowed to gain new insight in the process of diversification and in the speciation mechanisms of different groups of fishes.
The Labridae (the wrasses) is the second largest marine fish family and one of the most diversified in shape, color, and size. This mostly reef associated group is distributed in the Atlantic, Pacific and Indian Ocean and includes 71 genera and 525 valid species (Fricke et al. 2018a). The genera are traditionally grouped in subfamilies/tribes, whose number is 9 according to the last molecular phylogenetic analysis (Westneat and Alfaro 2005). The current high biodiversity of this family has been linked to Miocene events of coral reefs expansion that promoted high diversification rates and thus species diversity in fishes (Cowman and Bellwood 2011).
The cytogenetic data on Labridae species at global level are quite limited. Arai (2011) listed 59 species, and very recently the total number of cytogenetically analyzed species was raised to 74, mainly from the tribes Julidini and Labrini (Almeida et al. 2017). The diploid number in this family ranges from 22 to 48, but 56 species (75.7%) show 2n = 48. Some authors consider their karyotype evolution rate as moderate (Artoni et al. 2015), while others report high rates of chromosome evolution (Almeida et al. 2017). The detected macrostructural rearrangements include mainly Robertsonian fusions and pericentric inversions, whose relative recurrence depends on the subfamily (Molina et al. 2014; Almeida et al. 2017). Within this family, Xyrichtys Cuvier 1814 (tribe Novaculini) includes 15 species (Parenti and Randall 2011), commonly known as razor fishes. The taxonomy of these fishes is unclear due to incomplete species descriptions, and because juveniles, females, and males often show different color patterns and morphologies (Victor et al. 2001). Within the genus only one species, the pearly razorfish X. novacula (L., 1758) has been cytogenetically analyzed, since the other three species so far reported (Arai 2011), i.e. X. dea (Temminck and Schlegel, 1845), X. pavo (Valenciennes, 1840), and X. twistii (Bleeker, 1856) have been relocated in Iniistius (Randall and Earle 2002).
Xyrichtys novacula, is a protogynous hermaphrodite (Candi et al. 2004) benthic species that inhabits shallow sandy or muddy bottoms, usually in the vicinity of seagrass beds and corals, at a depth ranging from 5 to 80 m (Westneat 2002). It is distributed in warm latitudes of the Atlantic Ocean, from North Carolina to Brazil through the Caribbean, along the Eastern Atlantic coast from France to Angola including the main archipelagos and the Mediterranean Sea (Froese and Pauly 2018) (Fig. 1a). Previous cytogenetic data on pearly razor fishes, based only on Giemsa and Ag staining, reported different chromosome numbers and karyotype compositions in this species (Fig. 1a). Indeed, both Italian (Vitturi et al. 1986, 1989), and Brazilian (Almeida et al. 2017) samples show 2n = 48 (8 sm + 40 st/a), while preliminary data on samples from Venezuela indicated a reduction in the chromosome number (2n = 46) and the presence of two big metacentric chromosomes, (2 m + 4 sm + 40 a) (Nirchio et al. 2014).
a Distribution range of X. novacula. Sampling sites and karyotypes detected from previous cytogenetic investigation are reported. b–c Sampling origin of specimen collected in this study (stars) or of sequences retrieved from GenBank and BOLD databases and included in the molecular analysis (circles): Brazil (BR), Israel (IS), Italy (IT with the three sites PO, GA and SY), Mexico (MX), Panama (PA), Turkey (TR), Venezuela (VE), Virgin Islands (VI)
On this base, this study aims to test the hypothesis that (a) samples from different sites of the species distribution range belong to the same single evolutionary lineage and (b) differences in the karyotypes represent an intra-specific chromosome polymorphism related to the occurrence of frequent and temporary chromosome rearrangements. To this purpose, a cytogenetic and molecular (COI sequences) survey of pearly razor fishes from Italy and Venezuela has been undertaken. In order to provide a more comprehensive picture, sequences of the species and of other Xyrichtys available in GenBank and BOLD databases were included in the molecular analysis (Fig. 1b, c).
Materials and methods
Specimens collection
A total of 48 specimens of X. novacula were collected in the Caribbean Sea from Margarita Island (n = 41) Venezuela, and in the Mediterranean Sea from Ponza Island (Tyrrhenian Sea) and Gallipoli (Ionian Sea), Italy (n = 6) (Online Resource 1, Fig. 1b, c).
Adults specimens, both exhibiting males or female traits, were morphologically identified according to Cervigón (1993) and Tortonese (1970). After collection the fishes were transported to the lab and kept alive in well aired aquarium with controlled temperature until the sacrifice. The procedures were conducted in accordance with Institutional Authorization Research Project CI-6- 030601-1793/ 2012, Consejo de Investigación, Universidad de Oriente and Protocol number 1027 for experiments with animals of Universidade Estadual Paulista “Julio de Mesquita Filho”. Fish were sacrificed by a numbing overdose of benzocaine (250 mg/L) as recommended by the American Veterinary Medical Association (Leary et al. 2013). Voucher specimens were deposited in the Ichthyology Collection of the Escuela de Ciencias Aplicadas del Mar, Universidad de Oriente, Venezuela (ECAM) and at the Department of Biology and Biotechnology of Sapienza University of Rome, Italy (see Online Resource 1 for details).
Molecular analysis
Genomic DNA of all the specimens was extracted from muscle tissue fragments that were preserved in 95% ethanol (Aljanabi and Martinez 1997). A 510 bp fragment of the mitochondrial cytochrome C oxidase subunit I (COI) region was amplified by PCR and sequenced using primers and procedures previously reported (Milana et al. 2011). The sequences obtained were deposited in GenBank (see Online Resource 1 for Accession Numbers). The basic local alignment search tool (BLAST; https://blast.ncbi.nlm.nih.gov/Blast/) was used to search for similarity of the DNA sequences found in GenBank and in the Barcode of Life Database (BOLD; http://www.boldsystems.org/) to confirm the morphological species identification.
To investigate genealogical relationships among mitochondrial COI haplotypes, a parsimony network was constructed using the TCS 1.21 software (Clement et al. 2000). Genetic distances (D) were calculated using MEGA7 using the Kimura-2-parameters (K2P) substitution model (Kimura 1980).
Phylogenetic tree reconstructions were conducted using neighbour-joining (NJ), maximum-likelihood (ML) and Bayesian inference (BI) analyses. NJ and ML analyses (1000 bootstrap replicates) were performed using MEGA7 (Kumar et al. 2016) and PhyML v3.0 (Guindon et al. 2010) software, respectively. Bayesian analyses were carried out as implemented in MrBayes v3.1.2 (Huelsenbeck and Ronquist 2001). Two independent runs of four Markov chains each for 1,000,000 generations were performed. Modeltest 3.7 (Posada and Crandall 1998) and MrModeltest 2.3 (Nylander 2004) were used to select the evolutionary models that best fit the data set for the ML and the BI analyses, respectively. The Akaike information criterion was used.
In phylogenetic analyses the COI sequences of X. novacula from other sampling sites (Online Resource 1, Fig. 1b, c) and of other species of the genus (Online Resource 2) available in the GenBank and BOLD databases (consulted 31 October 2017) were also included. Cheilio inermis (Forsskål, 1775) was used as outgroup (JF434863).
Cytogenetic analysis
Chromosome preparations were obtained from kidney cells by conventional air-drying techniques (Nirchio and Oliveira 2006). Silver-staining of active nucleolar organizer regions (Ag-NORs) was obtained by argentic impregnation (Howell and Black 1980), and C-banding was performed following the method of denaturation with barium hydroxide (Sumner 1972).
Metaphases were analyzed under an Olympus BX61 microscope and were photographed using a digital camera (Olympus DP70). Images were processed using the Adobe Photoshop CC ver. 2015.0.0 software. Chromosomes were classified on the base of their arm ratio (Levan et al. 1964).
Results
Molecular analysis
BLAST searching in the GenBank and BOLD databases confirmed the morphological species identification, as all the obtained COI sequences show from 97 to 100% similarity with those of specimens of X. novacula. Similarity values found with other Xyrichtys species reached a maximum of 91%.
The 48 COI sequences of X. novacula obtained in this study correspond to 15 different haplotypes; 4 additional haplotypes were identified after the inclusion of the 18 sequences retrieved from the GenBank and BOLD databases (Online Resource 1). Unique haplotypes correspond to 72.2% of the samples analyzed and the remaining 27.8% are shared among individuals from the same (40%) or different sampling sites (60%). The parsimony network (Online Resource 3) analysis identified the presence of three different haplogroups (Hg), that are characterized by fixed diagnostic substitutions (Table 2). Haplogroup 1 includes 14 haplotypes, and haplogroups 2 and 3 include 2 and 3 haplotypes, respectively.
Additional 52 sequences of other Xyrichtys species were included in the alignment and the phylogenetic trees obtained by NJ, ML and BI analyses revealed that samples of each species are clustered within well supported clades; X. wellingtoni Allen and Robertson, 1995 is very close to X. victori Wellington, 1992 /Xyrichtys sp. D (Fig. 2). The X. novacula monophyletic clade is composed of three different clusters (Fig. 3a), that correspond to the three haplogroups identified by the haplotype network. Cluster 1 (Hg1) includes 33 out of the 36 sequences from Venezuela and six sequences retrieved from the GenBank database, corresponding to specimens collected in other localities within the Caribbean Sea. Cluster 2 (Hg2) groups three sequences from Venezuela and four additional sequences retrieved from the BOLD database, corresponding to specimens from the Brazilian coast. Cluster 3 (Hg3) includes the 12 sequences from Italy collected in this study plus eight sequences retrieved from both GenBank and BOLD database, corresponding to specimens from other localities within the Mediterranean Sea. The genetic divergence between these three clusters (0.016–0.028) is similar to that observed between X. wellingtoni and X. victori/Xyrichtys sp. D (0.022), but substantially lower than that calculated between other Xyrichtys species (Table 1).
a Phylogenetic relationships among X. novacula specimens inferred from COI sequences black circles indicate nodes showing bootstrap values > 70% (NJ and ML) and posterior probabilities > 0.90 (BI). Sequences retrieved from GenBank and BOLD are in italics. Specimens cytogenetically examined in this study are indicated by stars (for site code see Fig. 1). b Idiograms of the cytotypes and NOR localization (in box) obtained in the different haplogroups (Hg) (see below in the text)
Cytogenetic analysis
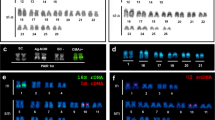
The 25 cytogenetically analyzed individuals show differences in diploid number and karyotype composition (Table 3). All the specimens collected in Italy (6 individuals) show 2n = 48 (6sm + 42 st/a) (Figs. 4a, 3b). Samples collected in Venezuela show a pronounced inter-individual variability in chromosome number, with diploid number ranging from 2n = 45 to 2n = 48, thus allowing the definition of different cytotypes (Figs. 4b–e, 3b). These different numbers are directly related to the presence and number of very large metacentric chromosomes (from 0 to 3): absence of metacentrics when 2n = 48, Cytotype A (1 individual) (Fig. 4b); 1 metacentric when 2n = 47, Cytotype B (8 individuals) (Fig. 4c); 2 metacentrics when 2n = 46, Cytotype C (7 individuals) (Fig. 4d); 3 metacentrics when 2n = 45, Cytotype D (3 individuals) (Fig. 4e) (Table 3). In all the karyotypes 6 submetacentric chromosomes were always clearly identifiable. The remaining chromosomes were not always easily assigned to a category, from a metaphase to another, as morphological differences between submetacentric/subtelocentric and between subtelocentric/acrocentric were not always clear. For this reason and using a parsimony criterion we decided to put together these chromosomes in the subtelocentric/acrocentric category, whose number in the individuals varies from 36 to 42 (Table 2).
Conventional Giemsa-stained karyotypes of Xyrichtys novacula from Italy (a) and Venezuela (b–e). The different cytotypes are indicated. Enlargement of NOR bearing chromosomes, after silver staining, and idiograms are shown in the right side. In samples from Venezuela different NOR patterns, not related to cytotypes, are observed
In all the specimens, regardless of the sampling site and chromosome number/cytotype, two positive signals were observed after silver nitrate staining (Ag-NOR), localized always on the antepenultimate chromosome pair (Fig. 4). However, while in Italian samples positive silver signals were observed in interstitial position on a small st/a pair (Fig. 4a, right side), in samples from Venezuela a noticeable polymorphism in the localization of these chromosome regions was observed (Fig. 4b–e; Table 3): NOR signals could be in interstitial position, or close to the centromere, or evenly, as a putative heterozygote combination, one interstitial and the other in terminal position. The C-banding revealed the presence of conspicuous blocks of constitutive heterochromatin in the centromeric region of almost all chromosomes, regardless of their geographic origin; in samples from Venezuela these blocks characterize also the centromere of the large metacentric chromosomes (Fig. 5), present in cytotypes B, C and D.
Discussion
Our results indicate that individuals morphologically identified as X. novacula show different karyotypes and also belong to different molecular lineages that partly reflect their geographic origin. Pearly razor fish sampled from different Mediterranean areas as Sicily channel, Tyrrhenian Sea, Ionian Sea, Levant Sea, all belong to the same molecular cluster (Hg3). Previous cytogenetic data (Vitturi et al. 1986, 1989) and results herein presented show that specimens belonging to this lineage share the same 2n = 48 karyotype and the same chromosome composition. Indeed, looking at the karyotypes of samples (10 individuals) from Sicily examined by Vitturi et al. (1986, 1989) and those from Ponza collected in this survey, the differences in the number of submetacentric and subtelocentric chromosome can be attributed to the use of different criteria in chromosome classification.
Pearly razor fishes from central and south America are split into two molecular lineages, that share samples from Venezuela and show different karyotypes (Fig. 3b). Indeed, the first lineage (Hg1) includes samples spread around the Caribbean Sea (analyzed in the present study or from the databases). Cytogenetic data are available only from Venezuela and reveal an extensive chromosomal polymorphism not previously detected in a preliminary analysis (Nirchio et al. 2014), that allows the recognition of four different cytotypes. The second lineage (Hg2) is quite rare in the Caribbean area (includes only three out of the 36 samples from Venezuela) but extends much more southward, to central-eastern Brazil. Unfortunately, cytogenetic data are available only for a single individual from Venezuela, which shows 2n = 46.
The main question is how to interpret obtained data. Floyd et al. (2002) defined the concept of molecular operational taxonomic units (MOTUs) to identify molecular clusters that do not correspond to well-accepted criteria for species recognition, and Ryberg (2015) underlined that to be of biological relevance these units should correspond as well as possible to species. Thus is the molecular divergence observed among the three pearly razor fish lineages sufficient for their attribution to different cryptic species? Or in other words do these MOTUs correspond to species? According to Ward (2009) 2% divergence in COI represents the reasonably threshold value sufficient to attribute unknown samples to different fish species, and values here observed are comparable and close to this limit. Furthermore, although the genetic divergence generally observed between other Xyricthys species is much higher (Table 2), a similar value is reported between X. wellingtoni from Clipperton island (off Pacific Mexican costs) and X. victori/Xyricthys sp. D from Galapagos islands, two well recognized species (Fricke et al. 2018b). In addition, such divergence is associated to differences in the karyotypes. Thus, although the presence of co-occurring lineages in the same area (like in Venezuela), per se, could represent the footprint of ancestral events of isolation and secondary contact within conspecific populations (Angiulli et al. 2016), more likely these lineages represent cryptic species, as already detected within the Caribbean Sea in other fishes (Nirchio et al. 2016, 2017).
The origin and presence of different pearly razorfish lineages/species is congruent with known phylogeographic barriers (Hodge and Bellwood 2016). Indeed, X. novacula of the Mediterranean Sea are separated from those from the West Atlantic area by the Mid-Atlantic barrier and by an oceanic gap of thousands of kilometers; the two groups present in the Caribbean area and south America are divided by the Amazon and Orinoco barriers, about 2300 km of coastline that are characterized by freshwater outflow. These latter barriers between marine faunas, that changed dramatically during Pleistocene, are considered responsible for the high endemism of each Province (Floeter and Gasparini 2000), as well as for the existence of numerous sister species pairs between Brazilian and Caribbean coral and reef associated faunas (Rocha 2003). The presence of rare samples from Venezuela in the South America lineage can be due to a secondary contact due to a huge and rich reef corridor (about 200 kilometers, as calculated from Fig. 1 of Moura et al. 2016), that is present between the Caribbean and the South Atlantic Ocean, from the mouth of the Amazon River (Collette and Rützler 1977). This biogeographic corridor could allow the cross of the Amazon barrier and explain the presence of fish species that are widely distributed in Brazilian province but can be recorded only in few sites in the southern region of the Caribbean Province (Rocha 2003). This suggestion is strengthened by the checklist of reef-associated organisms by Moura et al. (2016, supplementary material) which shows an inventory dominated by wide depth–ranging species that are broadly distributed in the Atlantic Ocean and include, among the Labridae, Halichoeres dimidiatus (Agassiz, 1831) and Xyrichtys splendens Castelnau, 1855.
In this picture whether chromosomal rearrangements represent a consequence or a cause (reinforced by biogeographic barriers) of the observed divergence between sympatric lineages is not clear. Chromosome rearrangements can prevent crossing-over and play a role in speciation (Faria and Navarro 2010), being the cause of gene flow restriction and population split in different vertebrate taxa (Mezzasalma et al. 2017 and references therein), so that different local cytotypes correspond to cryptic species (Bertollo 2007; Utsunomia et al. 2014). Therefore, high karyotypic variability could reveal complete speciation or parapatric chromosomal speciation in progress (do Nascimento et al. 2018).
In marine fishes chromosomal speciation has been suggested in several groups as in the family Gobiidae (Lima-Filho et al., 2016 and references therein), but usually chromosome rearrangements and variability are not common. Few exceptions due to Robertsonian events are known (Vitturi et al. 1984, 1989; Thode et al. 1985, 1988; Amores et al. 1990; Vitturi and Lafargue 1992; Caputo et al. 1999; Vasil’ev et al. 2012; Artoni et al. 2015). On the other hand, in freshwater environment, the small population size and the lower dispersion are associated to a much greater chromosomal diversity (Nirchio et al. 2014; Cioffi et al. 2015).
Xyrichtys novacula is a protogynous sequential hermaphrodite with an haremic social structure characterized by groups of 4 to 6 females occupying small territories defended by a dominant male that mates with these females (Marconato et al. 1995; Cardinale et al. 1998; Candi et al. 2004). According to Alós et al. (2012) the species is considered sedentary and occupies a reduced ‘home range’ (< 0.5 km2). Protoginy favors a strong sexual selection on males that monopolize multiple females increasing the reproductive success of a small number of larger males (Benvenuto et al. 2017). These biological traits could affect dispersal potential and gene flow and favor the maintenance of chromosome polymorphism, according to what observed in tropical reef fishes (Martinez et al. 2015; de Sena and Molina 2007). Thus, the simplest way to explain the persistence of the extensive chromosomal polymorphism detected in the Caribbean lineage in X. novacula is to assume that the basal karyotype corresponds to cytotype A, present also in the Mediterranean lineage, and composed of 2n = 48 chromosomes. The other cytotypes are produced by fusion of some acrocentric elements, that originated the large metacentric chromosomes, without alteration of the genome (because all the chromosome arms are present), and therefore without effect on the viability of the carrier individuals. The combination of gametes that are expected to be produced from the existing cytotypes would originate a proportion of non-viable zygotes, due to the loss or gain of one or more chromosome arms (Online Resource 4). The Chi-Square Goodness of Fit test allowed to accept (P = 0.877) the null hypothesis of non-significant differences between the frequency of the observed cytotypes and the expected frequency of the putative viable cytotypes, thus supporting the maintenance of a balanced chromosome polymorphism.
In conclusion our data unveiled the presence of different cryptic species across X. novacula species range, and the presence of parapatric divergence in progress in Venezuela. This latter could be related to a combination of factors that include local environmental condition. Indeed, higher diversification rates (and higher rates of speciation) exist between tropical and extra-tropical regions in different fish families including Labridae (Siqueira et al. 2016), according to the idea that the Caribbean region is a marine biodiversity hotspot (Miloslavich et al. 2010).
References
Aljanabi SM, Martinez I (1997) Universal and rapid salt-extraction of high quality genomic DNA for PCR-based techniques. Nucl Acids Res 25:4692–4693
Almeida LAH, Nunes LA, Bitencourt JA, Molina WF, Affonso PRAM (2017) Chromosomal evolution and cytotaxonomy in wrasses (Perciformes; Labridae). J Hered 108:239–253
Alós J, Cabanellas-Reboredo M, Lowerre-Barbieri S (2012) Diel behaviour and habitat utilisation by the pearly razorfish during the spawning season. Mar Ecol Prog Ser 460:207–220
Amores A, Giles V, Thode G, Alvarez MC (1990) Adaptative character of a Robertsonian fusion in chromosomes of the fish Gobius paganellus (Pisces, Perciformes). Heredity 65:151
Angiulli E, Sola L, Ardizzone G, Fassatoui C, Rossi AR (2016) Phylogeography of the common pandora Pagellus erythrinus in the central Mediterranean Sea: sympatric mitochondrial lineages and genetic homogeneity. Mar Biol Res 12:4–15
Arai R (2011) Fish Karyotypes: a check list. Springer, Berlin
Artoni RF, Vicari MR, de Almeida MC, Moreira-Filho O, Bertollo LAC (2009) Karyotype diversity and fish conservation of southern field from South Brazil. Rev Fish Biol Fish 19:393–401
Artoni RF, Castro JP, Jacobina UP, Lima-Filho PA, da Costa GWWF, Molina WF (2015) Inferring diversity and evolution in fish by means of integrative molecular cytogenetics. Sci World J 5:365787
Benvenuto C, Coscia I, Chopelet J, Sala-Bozano M, Mariani S (2017) Ecological and evolutionary consequences of alternative sex-change pathways in fish. Sci Rep 7:9084
Bertollo LAC (2007) Chromosome evolution in the Neotropical Erythrinidae fish family: an overview. In: Pisano E, Ozouf Costaz C, Foresti F, Kapoor BG (eds.) Fish Cytogenetics. Science Publishers, Stevenson Ranch, pp 195–211
Candi G, Castriota L, Andaloro F, Finoia MG, Marino G (2004) Reproductive cycle and sex inversion in razor fish, a protogynous labrid in the southern Mediterranean Sea. J Fish Biol 64:1498–1513
Caputo V, Caniglia ML, Machella N (1999) The chromosomal complement of Aphia minuta, a paedomorphic goby. J Fish Biol 55:455–458
Cardinale M, Colloca F, Ardizzone GD (1998) Growth and reproduction of Xyrichthys novacula (Pisces: Labridae) in the Mediterranean Sea. Sci Mar 62:193–201
Cervigón F (1993) Los peces marinos de Venezuela, vol 2. Fundación Científica Los Roques, Caracas
Cioffi MB, Bertollo LAC (2012) Chromosomal distribution and evolution of repetitive DNAs in fish. Genome Dyn 7:197–221
Cioffi MB, Sánchez A, Marchal JA, Kosyakova N, Liehr T et al (2011) Cross-species chromosome painting tracks the independent origin of multiple sex chromosomes in two cofamiliar Erythrinidae fishes. BMC Evol Biol 11:186
Cioffi MB, Bertollo LAC, Villa MA, de Oliveira EA, Tanomtong A et al (2015) Genomic Organization of Repetitive DNA Elements and Its Implications for the Chromosomal Evolution of Channid Fishes (Actinopterygii, Perciformes). PLoS One 10:e0130199
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Collette BB, Rützler K (1977) Reef fishes over sponge bottoms off the mouth of the Amazon River. In: Proceedings, third international Coral Reef symposium rosenstlel school of Marine and Atmospheric Science University of Miami. Florida, USA. May 1977
Cowman PF, Bellwood DR (2011) Coral reefs as drivers of cladogenesis: expanding coral reefs, cryptic extinction events, and the development of biodiversity hotspots. J Evol Biol 24:2543–2562
de Sena DCS, Molina WF (2007) Chromosomal rearrangements associated with pelagic larval duration in Labridae (Perciformes). J Exp Mar Bio Ecol 353:203–210
do Nascimento VD, Almeida Coelho K, Nogaroto V, de Almeida RB, Ziemniczak K et al (2018) Do multiple karyomorphs and population genetics of freshwaterdarter characines (Apareiodon affinis) indicate chromosomal speciation? Zool Anz 272:93–103
Faria R, Navarro A (2010) Chromosomal speciation revisited: rearranging theory with pieces of evidence. Trends Ecol Evol 25:660–669
Floeter SR, Gasparini JL (2000) The southwestern Atlantic reef fish fauna: composition and zoogeographic patterns. J Fish Biol 56:1099–1114
Floyd R, Abebe E, Papert A, Blaxter M (2002) Molecular barcodes for soil nematode identification. Mol Ecol 11:839–850
Fricke R, Eschmeyer WN, (eds) (2018b). Catalog of fishes: genera, species, references. http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp (accessed on 15 November 2018)
Fricke R, Eschmeyer WN, Fong JD (2018a) Species by family/subfamily. http://researcharchive.calacademy.org/research/ichthyology/catalog/SpeciesByFamily.asp (Accessed 15 November 2018)
Froese R, Pauly D (2018) FishBase. http://www.fishbase.org. accessed on 30 June 2018)
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W et al (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Hodge JR, Bellwood DR (2016) The geography of speciation in coral reef fishes: the relative importance of biogeographical barriers in separating sister-species. J Biogeogr 43:1324–1335
Howell WM, Black DA (1980) Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia 36:1014–1015
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol 33:1870–1874
Leary SL, Underwood W, Anthony R, Cartner S, Corey D et al (2013) AVMA guidelines for the euthanasia of animals: 2013 edition. American Veterinary Medical Association, Schaumburg
Levan A, Fredga K, Sandberg AA, Lima-Filho PA, Rosa RS et al (1964) Nomenclature for centromeric position on chromosomes. Hereditas 52:201–220
Marconato A, And VT, Marin G (1995) The mating system of Xyrichthys novacula: sperm economy and fertilization success. J Fish Biol 47:292–301
Martinez PA, Zurano JP, Amado TF, Penone C, Betancurr RR et al (2015) Chromosomal diversity in tropical reef fishes is related to body size and depth range. Mol Phylogenet Evol 93:1–4
Martins C (2007) Chromosomes and repetitive DNAs: a contribution to the knowledge of fish genome. In: Pisano E, Ozouf Costaz C, Foresti F, Kapoor BG (eds.) Fish Cytogenetics. Science Publishers, Stevenson Ranch, pp 421–453
Mezzasalma M, Andreone F, Aprea G, Glaw F, Odierna G et al (2017) When can chromosomes drive speciation?: The peculiar case of the Malagasytomato frogs (genus Dyscophus). Zool Anz 268:41–46
Milana V, Fusari A, Rossi A, Sola L (2011) Molecular and morphological identification of an uncommon centrolophid fish. Central Europ J Biol 6:445 ·
Miloslavich MP, Díaz JM, Klein E, Alvarado JJ, Díaz C et al (2010) Marine biodiversity in the Caribbean: regional estimates and distribution patterns. PLoS One 5:e11916
Molina WF, Martinez PA, Bertollo LAC, Bidau CJ (2014) Evidence for meiotic drive as an explanation for karyotype changes in fishes. Mar Genom 15:29–34
Moura RL, Amado-Filho GM, Moraes FC, Brasileiro PS, Salomon PS et al (2016) An extensive reef system at the Amazon River mouth. Sci Adv 2:e1501252
Nirchio M, Oliveira C (2006) Citogenética de peces. Editado por Universidad de Oriente, Cumaná, Venezuela, ISBN 980–234
Nirchio M, Mujica A, Oliveira C et al (2013) Pseudoplatystoma metaense and P. orinocoense (Siluriformes: Pimelodidae) from the Orinoco basin, Venezuela: cytogenetic and molecular analyses. Ital J Zool 80:526–535
Nirchio M, Rossi AR, Foresti F, Oliveira C (2014) Chromosome evolution in fishes: a new challenging proposal from Neotropical species. Neotrop Ichthyol 12:761–770
Nirchio M, Oliveira C, Siccha-Ramirez ZR, Sene VF, Sánchez-Romero OR et al (2016) Cryptic Caribbean species of Scorpaena (Actinopterygii: Scorpaeniformes) suggested by cytogenetic and molecular data. J Fish Biol 89:1947–1957
Nirchio M, Oliveira C, Siccha-Ramirez ZR, Sene VF, Sola L et al (2017) The Mugil curema species complex (Pisces, Mugilidae): a new karyotype for the Pacific white mullet mitochondrial lineage. Comp Cytogenet 11:225–237
Nylander JAA (2004) MrModeltest v2. 3 software. Evolutionary Biology Center, University U, Sweden Available from: http://www.abcse/nylander/mrmodeltest2/mrmodeltest2
Pansonato-Alves JC, Serrano ÉA, Utsunomia R, Camacho JPM, da Costa Silva GJ et al (2014) Single origin of sex chromosomes and multiple origins of B chromosomes in fish genus Characidium. PLoS One 9:e107169
Parenti P, Randall JE (2011) Checklist of the species of the families Labridae and Scaridae: an update. Smithiana Bull 29–44
Parise-Maltempi PP, da Silva EL, Rens W, Dearden F, O’Brien PCM et al (2013) Comparative analysis of sex chromosomes in Leporinus species (Teleostei, Characiformes) using chromosome painting. BMC Genet 14:60
Phillips RB (2013) Evolution of the sex chromosomes in salmonid fishes. Cytogenet Genome Res 141:177–185
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818
Randall JE, Earle JL (2002) Review of Hawaiian Razorfishes of the Genus Iniistius (Perciformes: Labridae). Pac Sci 56:389–402
Rocha LA (2003) Patterns of distribution and processes of speciation in Brazilian reef fishes. J Biogeogr 30:1161–1171
Ryberg R (2015) Molecular operational taxonomic units as approximations of species in the light of evolutionary models and empirical data from Fungi. Mol Ecol 24:5770–5777
Siqueira AC, Oliveira-Santos LGR, Cowman PF, Floeter SR (2016) Evolutionary processes underlying latitudinal differences in reef fish biodiversity. Glob Ecol Biogeogr 25:1466–1476
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304–306
Thode G, Giles V, Alvarez MC (1985) Multiple Chromosome Polymorphism in Gobius paganellus (Teleostei, Perciformes). Heredity 54:3
Tortonese E (1970) Fauna d’Italia,“Osteichthyes”, pesci ossei, volume 1. Bologna, Edizioni Calderini
Utsunomia R, Pansonato Alves JC, Paiva LRS, Costa Silva GJ, Oliveira C et al (2014) Genetic differentiation among distinct karyomorphs of the wolf fish Hoplias malabaricus species complex (Characiformes, Erythrinidae) and report of unusual hybridization with natural triploidy. J Fish Biol 85:1682–1692
Vasil’ev VP, Prazdnikov DV, Vasil’eva ED (2012) Chromosome polymorphism of stargazer Uranoscopis scaber (Uranoscopidae, Perciformes) from the Black Sea. J Ichthyol 52:296–300
Victor BC, Wellington GM, Caldow C (2001) A review of the razorfishes (Perciformes: Labridae) of the eastern Pacific Ocean. Rev Biol Trop 49(Suppl 1):101–110
Vitturi R, Lafargue F (1992) Karyotype analyses reveal inter-individual polymorphism and association of nucleolus-organizer-carrying chromosomes in Capros aper (Pisces: Zeiformes). Mar Biol 112:37–41
Vitturi R, Carbone P, Catalano E, Macaluso M (1984) Chromosome polymorphism in Gobius paganellus, Linneo 1758 (pisces, gobiidae). Biol Bull 167:658–668
Vitturi R, Catalano E, Macaluso M, Zava B (1986) Karyotypes of nine species of the family Labridae (Pisces, Perciformes). Biol Zent Bl 105:519–530
Vitturi R, Catalano E, Lo Conte MR, Spampinato P (1989) Ag-NORs and C-banding pattern of the labrid species Xyrichthys novacula (L.) (Pisces, Perciformes). Biol Zent Bl 108:263–266
Westneat MW (2002) Labridae. In: Carpenter KE (ed) The living marine resources of the Western Central Atlantic. Bony fishes part 2 (Opistognathidae to Molidae), sea turtles and marine mammals. FAO, Rome, pp 1701–1739
Westneat MW, Alfaro ME (2005) Phylogenetic relationships and evolutionary history of the reef fish family Labridae. Mol Phylogenet Evol 36:370–390
Acknowledgements
Financial support was provided by Consejo de Investigación, Universidad de Oriente-CIUDO, Venezuela (to M.N. and J.I.G.), Grant, no. Cl-6-030601-1793/12, by Fundação de Amparo à Pesquisa do Estado de São Paulo-FAPESP, Brazil and Conselho Nacional de Desenvolvimento Científico e Tecnológico—CNPq, Brazil (to C.O. and F.F.), and by Sapienza University of Rome, Italy, Progetto Università 2014 (to V.M. and A.R.). We are grateful to Elisa Angiulli and Mariangela Coriandri (Sapienza University of Rome, Italy), who kindly helped in the collection of fishes in Italy and to Regulo López and Juan Marcano (Universidad de Oriente) who helped in the collection of fishes in Margarita and Cubagua Islands.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nirchio, M., Gaviria, J.I., Siccha-Ramirez, Z.R. et al. Chromosomal polymorphism and molecular variability in the pearly razorfish Xyrichtys novacula (Labriformes, Labridae): taxonomic and biogeographic implications. Genetica 147, 47–56 (2019). https://doi.org/10.1007/s10709-019-00051-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-019-00051-9