Abstract
The assessment of genetic variability is indispensable for the development of high-yielding and nutritionally enriched soybean varieties. In this study, fifty-nine soybean genotypes (56 test entries and three standard checks) from different origins were characterized for eight qualitative and 20 quantitative traits. The experiment was laid out in a complete augmented block design for two consecutive seasons under natural rainfed conditions. Results revealed that among qualitative traits, flower colour was found to be most dynamic morphological marker trait for genotypic distribution. Mean square variances for twenty agro-morphological, seed yield and quality traits showed significant differences among tested genotypes for almost all studies traits. Non-hierarchical clustering classified entire set of germplasm into five groups, with the recognition of cluster IV for selection potential. Correlation analysis indicated that seed yield of soybean was positively associated with all yielding components, while a highly negative association was observed between seed quality traits such as protein content (%) and oil content (%). The multivariate principal component analysis (PCA) extracted five essential PCs, which explained 81.22% of the total accumulative variation. Moreover, PCA unveiled the most discriminatory traits as well as superior genotypes which participated intensely in phenotypic variability. Most diverse genotypes identified during study were Jhunghwang, K-D, 24,598, G-35, Brazil-3, 24,560, Ajmeri-1, NARC-2 and 24,608 for improved productivity and enhanced nutritional quality of soybean. The essential traits, including 100-seed weight, seed yield plant−1, protein content and oil content depicted influential effects in identifying these desired genotypes. Conclusively, the hybridization of divergent parents in cross-breeding programs may have successful chances to get transgressive segregants with higher seed yield potential along with improved nutritional quality for developing new soybean varieties.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean is regarded as a miracle crop due to its numerous beneficial properties and versatile usages in human food and animal diet (Ali et al. 2022; Hartman et al. 2011). Being main source of edible oil, soybean is a protein-rich crop mainly used to feed livestock, poultry and aquaculture (Guo et al. 2022; Medic et al. 2014; Selle et al. 2020). It is one of the most widely cultivated oilseed crop, accounting for more than 60% of the global oilseed production (Soy Stats 2022), and holding a premium position in terms of area and production among the oilseed crops (Rai et al. 2016). Globally, the growing area engaged under soybean cultivation is about 122 million hectares, with a total annual production of 385.524 million tons and an estimated average grain yield of 2.8 tons ha−1 (USDA 2019). Brazil is the world's leading producer and exporter of soybean accounting for 36% of global production, followed by the United States (28%), while Argentina, China, and Paraguay each contributing about 15%, 5% and 3% of global production, respectively (FAOSTAT 2023). The demand of soybean has increased manifolds during the recent years, owing to its extensive utility in human food and animal feed as well as multiple industrial applications (Dei 2011; Wilson 2004).
Natural genetic diversity is the basis for crop improvement as well as plant survival that should be exploited to cope with food security demand for growing world population (Breseghello and Coelho 2013; Dong et al. 2014; Salgotra and Chauhan 2023). The presence of genetic variability would help plants in adaptation to a wide range of ecological conditions; however, low variability responses to crop susceptibility to various environmental stresses (Ali et al. 2023; Maxted et al. 1997). Exploration of genetic diversity in initial breeding material provides more opportunities to select potential lines for direct cultivation or efficiently utilized in cross-breeding programs (Govindaraj et al. 2015; Misganaw et al. 2023; Yadava et al. 2022). Thus, the knowledge of genetic diversity would assist plant breeders in broadening the genetic base of adaptable cultivars (Bhandari et al. 2017; Fu 2015; Tester and Langridge 2010; Žulj Mihaljević et al. 2020).
The ultimate goal of a soybean breeder is to develop new cultivars with improved seed yield and acceptable nutritional quality to meet the growing demand for edible oil and protein meal (Dornbos and Mullen 1992; Ghanbari et al. 2018; Sobko et al. 2020). Large-seeded soybean varieties with high protein content coupled with significant amount of oil concentration are foremost demand by the food and feed industries (Stobaugh et al. 2017; Xu et al. 2022). Therefore, novel genes or alleles for these targeted traits must be searched from diverse sources and recombined through conventional breeding procedures to maximize soybean productivity along-with nutritional quality (Mello-Filho et al. 2004; Sharma et al. 2014; Zhao et al. 2021). Nevertheless, selecting promising parents with multiple features is a monumental task in soybean breeding because of the narrow genetic bottleneck of modern cultivars and unknown pedigrees of available genetic resources (Hyten et al. 2006; Kumar et al. 2022; Grainger and Rajcan 2014). Thus developing improved varieties with desired attributes including high seed yield and enhanced nutritional quality is the central focus of soybean breeders (Ali et al. 2022; Sudarić et al. 2019).
Despite being an agricultural country, Pakistan is facing severe shortage of edible oils and overwhelmingly depends on the import of edible oil to meet domestic needs (Asad et al. 2020; Tariq et al. 2022). The total domestic production of edible oil from all sources is about 0.496 million tons, accounting for 12% of the entire necessity. However, 3.177 (88%) million tons would acquire from foreign sources with annual spending of $ 3.562 billion, resulting in a massive trade deficit and a huge burden on the national exchequer (Government of Pakistan 2022–2023). Moreover, Pakistan is unable to produce enough soybeans mainly due to the lack of improved quality varieties with desired characters (Asad et al. 2020; Nasir et al. 2023). Currently, very few varieties are available for general cultivation but most of them are deficient in desirable attributes (Iqbal et al. 2008). Even though the climatic conditions and soil composition of the country are ideal for soybean cultivation, but very little effort has been made to improve the genetic make-up of this valuable crop (Iqbal et al. 2010; Malik et al. 2007). As a result, the imports of soybean has been increased manifolds in recent years and almost reached up to 2.5 million metric tons (USDA 2019), owing to urban sprawl and the steadily expanding poultry industry of the country (Habib et al. 2016).
Agro-morphological characterization is a conventional breeding procedure which used to determine the yield potential, genomic selection and maintaining genetic purity of cultivated varieties (Farahani et al. 2019; Li et al. 2020; Malek et al. 2014). The phenotyping of germplasm provides a valuable information for sustainable gene-bank conservation, dynamic management and optimal utilization of elite genetic resources in initial breeding programs (Dong et al. 2004; Gautam et al. 2004; Govindaraj et al. 2015). Traditionally, genetic variability among local and exotic genetic pools of soybean has been evaluated on the basis of pheno-morphic and agronomic traits to undertake selection of elite breeding lines focusing to improve soybean productivity and nutritional composition (Assefa et al. 2019; Sharma et al. 2014; Kumar et al. 2015; Ghanbari et al. 2018). Thus, the current study aims to provide comprehensive information on agro-morphological characterization and selection strategy of available soybean germplasm which might be utilized in future breeding programs to develop new varieties with high yield potential and improved nutritional quality in order to ensure national food security.
Materials and methods
Experimental site discription, plant materials and layout of design
The field experimental trial was conducted at the Knoot Research Farm of PMAS-Arid Agriculture University Rawalpindi, Pakistan, during two Kharif seasons (2019 and 2020). The topsoil (0–15 cm) profiling of research site was examined as sandy clay loam with 54.2% sand, 23.4% silt and 22.4% clay. The geographic position of research station is 33°06′ N latitude, 73°01′ E longitudes and an altitude of 474 feet above sea level. A mini core collection, comprising fifty-nine soybean genotypes (56 test entries and three checks) originated from different countries were explored for qualitative and quantitative traits of soybean. Information of genetic material regarding passport code, variety name, geographic origin and collection source are presented briefly (Table S1).
The germplasm was grown under natural field condition for two cropping seasons using augmented block design (Federer 1956). At the start of rainy seasons, sowing was done on 14th July during the first year and 10th July during second year, depending on the soil moisture availability. The meteorological environment data of the experimental site during two cropping cycles is summarized (Table S2). Eight blocks were assigned for studied soybean genotypes and each block contained seven test entries and three standard checks. All genotypes were planted in a single row of 5-m length and space between row-to-row and plant-to-plant was kept 45 cm and 10 cm, respectively. The test entries were planted in a single replicate, whereas commercial checks were repeated randomly once in each block. The seeds were sown manually in a depth of 3–4 cm and thinning was done after germination in order to maintain a density of 20 plants per linear meter. To ensure proper crop growth, recommended cultural practices were implemented uniformly for each block. The agro-morphological measurements were recorded according to the standard descriptors for soybean (IBPGR 1984). However, seed quality parameters such as protein and oil content were estimated separately for all tested entries and replicated checks following standard procedures (AOAC 1995; Latimer 2016).
Data analysis
The transformed numerical codes data derived from each qualitative trait were subjected to cluster analysis using Ward’s distance matrix (Ward 1963). The Shannon–Weaver diversity index (H′) for all qualitative traits was calculated as per equation (Shannon 1948).
Pi = frequency of total number of genotypes; n = total number of classes for a trait; i = total number of individuals; log = maximum value obtained from all groups.
The mean data of twenty quantitative agro-morphological traits were subjected to Analysis of Variance (ANOVA) and using statistical R software (ver. 3.6.1) for augmented block design analysis. The descriptive statistics was summarized for the arithmetic mean, range (minimum and maximum), variance and coefficient of variation using computer software IBM SPSS version 23 (Kirkpatrick 2015). Two-year mean data (Tables S3–S22) were adjusted by ignoring treatment and block effects and computed for multivariate statistical analyses. Non-hierarchical K-means clustering was calculated based on Euclidian distances to discriminate the trait-specific groups using XLSTAT. The correlation matrices were calculated at a significance level of α = 0.05 (Kwon and Torrie 1964), whereas Principal Component Analysis (PCA) was performed by using R-Studio software.
Results and discussion
Phenotypic characterization of qualitative traits
Seed morphometry is a vital feature for genomic classification and maintaining the seed quality of soybean. The consumers’ preference and global market demand are greatly inspired by soybean seeds quality and purity (Sudarić et al. 2019). In present study, frequency distribution of eight qualitative traits was visually assessed for all soybean genotypes (Table 1). The tested germplasm exhibited remarkable variation for flower colour and pod pubescence, while stem growth, pod colour, seed coat colour, hilum colour, seed shape and seed lustre displayed relatively moderate variation across the genotypes. The wide range of variation for qualitative traits assist soybean breeders to select promising lines with distant features (Kachare et al. 2020; Shrestha et al. 2023).
Among the studied soybean population, majority of genotypes attended determinate stem growth habit. The white flower colour was more common in population than purple flower colour. There were three different colours were recognized for pods appearance. Hairy structures (pubescence) on the upper surface of pods were present on 39 genotypes and absent on 20 genotypes (Table 1). The evaluated genotypes showed discernible variation for other seed traits, such as seed coat colour, hilum colour, seed shape and seed lustre (Fig. 1). Furthermore, the Shannon’s Diversity Index (H′) was highest (1.704) for hilum colour and seed shape (1.268) while it was found lowest (0.364) for flower colour because 88% of the entire population retained white flower colour. Several previous studies highlighted the importance of qualitative traits in germplasm characterization and varietal identification of soybean (Tripathi and Khare 2016; Khanande et al. 2016; Shrestha et al. 2023; Arteaga et al. 2019).
Cluster analysis based on qualitative traits
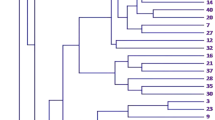
The cluster analysis distributed the soybean genotypes into three main groups and six distinct clusters. The highest number of genotypes (13) were retained in cluster-III, followed by two similar clusters i.e., cluster-IV and cluster-V with 12 genotypes in each. Minimum six genotypes retained by cluster-II, chased by cluster-VI and cluster-I with 7 and 9 genotypes, respectively (Fig. 2). The clustering patterns were found to be very logical based on flower colour because first five clusters (I–V) having 52 genotypes exhibited white flowers, while cluster-VI exhibited only seven genotypes with purple colour. This indicated that the flower colour would be an important marker-trait in soybean breeding in order to identify the desired cross combinations.
Analysis of variance for augmented block design
Augmented block design is an ideal model which primarily used to evaluate large sets of test entries in pre-breeding programs to identify the new selections (Federer 1963; Kempton and Gleeson 1997). It is a cost-efficient design and feasible in situations where the experimental seed is limited in quantity for replication, or the plant breeder is unable to maintain the differences between experimental units (Federer 1956). The information obtained from standard checks can be used to adjust the means of tested entries and provide an appropriate error mean square for various source comparisons (Federer et al. 2001; Saba et al. 2017). In this study, the variability range was measured for 20 agro-morphological, seed yield and quality traits of soybean and observed significant differences across genotypes and other source for all studied traits, except number of nodes plant−1 (Table 2). This indicates that evaluated germplasm penal exhibited high level of variation, which favoured the selection process (Li et al. 2020; Ullah et al. 2021). The high degree of genetic variability plays a vital role in selective breeding and exploring the genetic potential of soybean materials for varietal development (Anderson et al. 2019; Bailey-Serres et al. 2019). Several earlier studies have also reported significant variation for various agronomic traits of soybean (Iqbal et al. 2010; Khurshid et al. 2020; Ullah et al. 2021). Thus, substantial variability observed among tested soybean germplasm could be exploited to maximize the genetic gain of target traits.
Descriptive statistics for quantitative traits
The basic summary statistics (mean, range, variance and coefficient of variation) was computed for each studied trait (Table 3). A remarkable variation was observed in range of all studied traits except number of branches, pod length, pod width, seeds per pod, seed dimension-related features and stem diameter. The presence of sufficient variability indicating that the evaluated soybean materials were highly diverse and could be utilized in various breeding programs to increase soybean grain yield and nutritional quality (Aditya et al. 2011; Sharma et al. 2014). Importantly, high values of variance were noticed for DFF, DFC, DM, PH, NFP and DMWP, which indicating that these traits are of greater importance in soybean selection and genetic improvement (Reni and Rao 2013). However, moderate to low variances observed for SYP, NUFP, PC, NNP, HSW, NBP, OC, SD, ST, SW, SL, PW, PL and NSP subsequently which indicating that selection and improvement of these traits might be limited due to narrow genetic variability. Ghafoor et al. (2001) suggested that conventional breeding procedures would be helpful in improving characters having low magnitude of variance. Moreover, the coefficient of variation (CV) is a crucial indicator for measuring the precision and accuracy of field experiments. During both years, high CV (%) was observed for number of unfilled pods (NUFP), while the lowest CV (%) was recorded for pod width (Table 3). According to Poehlman (2013), the coefficient of variation less than 20% is considered to be reliable for field experiments.
Non-hierarchical cluster analysis
Non-hierarchical clustering technique was used to categorize the soybean genotypes into five distinct clusters (I–V) based on the Euclidean distances. Among the clusters, first cluster retained highest number of genotypes (18) with cumulative contribution of 30.50% (Table 4). The remaining genotypes were grouped in such a way that 12 genotypes in cluster II, 15 were assembled in cluster III, 08 in cluster IV and 06 were grouped in cluster V.
The genotypes concerning to cluster V exhibited the highest mean values for days to flowering (57.42 ± 1.86), days to flower completion (64.25 ± 1.78), maturity duration (117.50 ± 7.40), nodes number (13.78 ± 0.81), branches per plant (6.70 ± 2.43), highest unfilled pods (9.60 ± 4.47), stem diameter (6.78 ± 0.88). This indicated that late maturing genotypes with higher number of empty pods were gathered in cluster V. The genotypes with longest plant height (75.11 ± 13.01) and filled pods (69.83 ± 9.18) were retained in cluster II (Table 5). Likewise, the genotypes assembled in cluster IV were found superior for majority of seed yield related traits such as pod length (3.89 ± 0.40), pod width (8.84 ± 0.53), seeds number per pod (2.42 ± 0.24), seed length (7.23 ± 0.72), seed width (6.39 ± 0.74), seed thickness (5.27 ± 0.69), hundred seed weight (11.95 ± 2.69), dry matter (49.53 ± 7.89) and seed yield per plant (15.23 ± 4.26). For seed quality traits, relatively minor differences were observed between the clusters. Non-hierarchical cluster analysis classified the germplasm by using multiple variables simultaneously and provides distinct classes based on dissimilarity matrices (Anuradha et al. 2011). Our results are consistent with the earlier findings in which soybean germplasm were collected from different countries and classified on the basis of phenotypic variability (Getnet 2018; Malik et al. 2011). Thus selecting potential genotypes from clusters II and IV would be an efficient breeding strategy for improving soybean seed yield accompanied with nutritional composition through conventional breeding procedures.
Correlation analysis of quantitative traits
Correlation analysis is a powerful statistical tool for determining the interrelationships between multiple traits simultaneously. Results revealed that out of all possible 190 combinations, 106 pairs were found statistically significant at a probability level of ≥ 95% with a regression coefficient not less than 0.26 (Fig. 3). The phenological traits like DFF, DFC and DM showed a positive correlation with SD and DMWP but a negative association was observed with seed yielding traits i.e., PL, PW, SL, SW, ST and HSW. This indicated that late flowering and delayed maturity would adversely affect pod features and seed related-traits under certain edapho-climatic conditions (Jiang et al. 2014). According to Arshad and Ghafoor (2006), traditional breeding approaches might be productive in overcoming these undesirable linkages for developing short-duration cultivars with improved grain yield in soybean. The correlation among DFF, DFC, DM, NNP, NBP and NUFP were found significant and positive direction which indicating that late-flowering genotypes also delayed maturity and produce higher number of branches and empty pods. Thus selection based on these traits may not be a sound breeding strategy for improving soybean productivity.
Seed yield of soybean is a complex trait which significantly influenced by environmental changes and is primarily determined by the other contributing traits (Malek et al. 2014). In this study, seed yield (SYP) showed positive association with PH, NNP, NFP, PL, PW, NSP, SL, SW, ST, HSW, SD and DMWP but a weak association was observed for seed quality traits including protein and oil content. This indicated that simultaneous selection of elite genotypes with higher protein concentration and oil content accompanied with improved seed yield is a major challenge in soybean improvement (Cober and Voldeng 2000; Vollmann et al. 2000). The major contributing traits such as number of filled pods, seed vigour and hundred seed weight should be prioritized in the genomic selection of soybean (Liu et al. 2019; Malek et al. 2014; Malik et al. 2011). Thus, selecting promising genotypes with a set of positively correlated traits with seed yield may be favoured in indirect selection (Leite et al. 2018; Ghanbari et al. 2018).
The correlation of seed quality traits including protein content (%) and oil content (%) was found to be strongly negative (Fig. 3). This indicating that an increase in oil content would significantly reduce the protein content in soybean seed. Although soybean is a good source of edible oil and harbours a significant amount of seed protein, but the negative association between these two traits poses a real challenge for soybean breeders (Kwanyuen et al. 1997; Qin et al. 2014). This inverse relationship might be due to the distribution of carbon chains that synthesize protein and oil content in soybean (Hernández-Sebastià et al. 2005; Guo et al. 2022). It is noteworthy both seed quality traits are quantitatively inherited and determined by the interaction of multiple genes (Hwang et al. 2014). Several previous studies have also confirmed the strong negative correlation between protein content and oil content in soybean seed composition (Mello-Filho et al. 2004; Popovic et al. 2012). Thus knowledge of characters’ association in soybean would assist breeders in formulating long-term breeding strategies (Ghanbari et al. 2018; Machado et al. 2017).
Principal component analysis (PCA)
The principal component analysis was performed in order to estimate the phenotypic diversity among soybean genotypes. Scree plot indicated that out of twenty reserved principal components, only five initial PCs were declared significant as depicted eigenvalues greater than one (Fig. 4). The extracted five-component axes such as PC1, PC2, PC3, PC4 and PC5 having eigen roots ranging from maximum to minimum of 7.42, 4.73, 1.81, 1.27 and 1.01, respectively explained 81.22% of the total variability across the tested soybean genotypes. The rest of the variance (19%) was cumulatively described by the remaining 15 PCs with lower (< 1) eigenvalues, which were not considered for further interpretation.
PCA variable plot
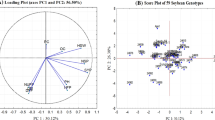
The PCA variable plot revealed that initial two principal components i.e., PC1 and PC2 estimated more than 60% of variation with the contribution of 37.11% and 23.67%, respectively to the total phenotypic variability (Fig. 5a). The projection of quantitative traits with positive loading vectors on PC1 were SW (0.92), HSW (0.88), PL (0.87), PW (0.87), ST (0.86), SL (0.85), SYP (0.82), DMWP (0.81) and NSP (0.69) subsequently (Table 6). On the other hand, traits with short to medium loading vectors were DFF (− 0.441), DFC (− 0.421), DM (− 0.412), NBP (− 0.513) and NUFP (− 0.443) stacked negatively in the first principal component (Table 6). Likewise, PC2 was mainly related to traits were PC (0.73), DFC (0.35), DM (0.32) and HSW (0.31), whereas traits with negative loadings on second principal component were OC, PH, NBP, NFP, NUFP, DMWP and SYP (Fig. 5a). The traits which described PC3 were PC and OC with positive loading weights, while other traits like PL, PW, ST and HSW exhibited very low variances with minor effects. In the case of PC4 and PC5, none of the traits contributed significantly towards assorted variation.
PCA score plot
Two-dimension PCA score plot was drawn between PC1 and PC2 for identifying the diverse genotypes. The genotypes dispersed at the extreme positions away from the origin point, demonstrated high genetic variability (Fig. 5b). Certain genotypes namely, Jhunghwang, K-D, 24,598, G-35, Brazil-3, 24,562, 24,592, 24,578, 24,560, Aga1, Ajmeri-1, NARC-2 and 24,608 were found to be more diverse in the studied materials. It is worth mentioning that the genotypes located at the right side of plot exhibited high variances, and may be given preference in the selection process (Bartual et al. 1985). Thus genotypes including G-35, K-D, Jhunghwang, and 24,598 with higher variances for one or more traits identified as valuable genetic sources for introgressing desired genes into adaptable local varieties. Several former studies have utilized the principal component analysis for determining the genetic variability and identification of diverse genotypes in soybean (Iqbal et al. 2008; Mannan et al. 2010).
Conclusion
This study comprehensively described a suitable strategy for the identification and selection of desirable superior progenitors with broad genetic spectrum based on phenotypic variability. The evaluated soybean genetic materials displayed a considerable divergence for all agronomic, yield and seed quality traits. Various agro-morphological characters were recognized as key indices based on multivariate approach, which might be effective in future soybean breeding programs to improve seed yield and nutritional quality simultaneously. Overall, three exotic lines viz., K-D, Jhunghwang and 24,598 were identified for improved productivity and may be approved for general cultivation or even conserved as elite resources for future breeding programs. Additionally, two local varieties such as NARC-2 and Ajmari-1 performed better for oil composition, hence could be utilized as potential parents for increasing oil content in other commercial cultivars. Taken together, the prominent genotypes identified during this study would be served as benchmark for developing high-yielding and nutritionally enriched soybean cultivars for local ecologies.
Data availability
The research data is available in main text and/or in supplementary material. Raw data will be made available by corresponding author upon the reasonable request.
References
Aditya JP, Bhartiya P, Bhartiya A (2011) Genetic variability, heritability and character association for yield and component characters in soybean (G. max (L.) Merrill). J Cent Eur Agric 12:27–34
Ali A, Ahmed Z, Maqbool R, Shahzad K, Hameed A, Awan FS, Khan AI, Shah ZH, Ali Z, Alsamadany H, Bilal M (2022) Advances in molecular markers to develop soybean cultivars with increased protein and oil content. Springer, Cham, pp 125–136
Ali Z, Maryam H, Saddique MAB, Ikram RM (2023) Exploiting genetic diversity in enhancing phenotypic plasticity to develop climate-resilient cotton. Genet Resour Crop Evol 70:1305–1320
Anderson EJ, Ali ML, Beavis WD, Chen P, Clemente TE, Diers BW, Graef GL, Grassini P, Hyten DL, McHale LK, Nelson RL (2019) Soybean [Glycine max (L.) Merr.] breeding: history, improvement, production and future opportunities. Adv Plant Breed Strateg Legumes 7:431–516
Anuradha B, Aditya JP, Singh G, Gupta A, Agarwal PK, Bhat JC (2011) Assessment of genetic diversity in indigenous and exotic collections of black soybean (Glycine max (L.) Merrill.). SABRAO J Breed Genet 43:81–90
AOAC (1995) Association of Official Analytical Chemists. Official methods of analysis, 16th edn. AOAC International, Washington
Arshad M, Ali N, Ghafoor A (2006) Character correlation and path coefficient in soybean Glycine max (L.) Merrill. Pak J Bot 38(1):121–130
Arteaga S, Yabor L, Torres J, Solbes E, Muñoz E, Díez MJ, Vicente O, Boscaiu M (2019) Morphological and agronomic characterization of Spanish landraces of Phaseolus vulgaris L. Agriculture 9:149
Asad SA, Wahid MA, Farina S, Ali R, Muhammad F (2020) Soybean production in Pakistan: experiences, challenges and prospects. Int J Agric Biol 24:995–1005
Assefa Y, Purcell LC, Salmeron M, Naeve S, Casteel SN, Kovács P, Archontoulis S, Licht M, Below F, Kandel H, Lindsey LE (2019) Assessing variation in US soybean seed composition (protein and oil). Front Plant Sci 10:298
Bailey-Serres J, Parker JE, Ainsworth EA, Oldroyd GE, Schroeder JI (2019) Genetic strategies for improving crop yields. Nature 575:109–118
Bartual R, Carbonell EA, Green DE (1985) Multivariate analysis of a collection of soybean cultivars for southwestern Spain. Euphytica 34:113–123
Bhandari HR, Bhanu AN, Srivastava K, Singh MN, Shreya HA (2017) Assessment of genetic diversity in crop plants-an overview. Adv Plants Agric Res 7:279–286
Breseghello F, Coelho AS (2013) Traditional and modern plant breeding methods with examples in rice (Oryza sativa L.). Chem Biol Technol Agric 61:8277–8286
Cober ER, Voldeng DH (2000) Developing high-protein, high-yield soybean populations and lines. Crop Sci 40:39–42
Dei HK (2011) Soybean as a feed ingredient for livestock and poultry. IntechOpen, London, pp 215–226
Dong D, Fu X, Yuan F, Chen P, Zhu S, Li B, Yang Q, Yu X, Zhu D (2014) Genetic diversity and population structure of vegetable soybean (Glycine max (L.) Merr.) in China as revealed by SSR markers. Genet Resour Crop Evol 61:173–183
Dong YS, Zhao LM, Liu B, Wang ZW, Jin ZQ, Sun H (2004) The genetic diversity of cultivated soybean grown in China. Theor Appl Genet 108:931–936
Dornbos DL, Mullen RE (1992) Soybean seed protein and oil contents and fatty acid composition adjustments by drought and temperature. J Am Oil Chem Soc 69:228–231
FAOSTAT (2023) Food and Agriculture Organization of the United Nations. http://Faostat.fao.org. Accessed 25 April 2023
Farahani M, Salehi-Arjmand H, Khadivi A, Akramian M (2019) Phenotypic diversity among Morus alba var. nigra genotypes as revealed by multivariate analysis. Sci Hortic 248:41–49
Federer WT (1956) Augmented (or hoonuiaku) designs. Hawaiian Pl Rec 55:191–208
Federer WT (1963) Procedures and designs useful for screening material in selection and allocation, with a bibliography. Biometrics 1:553–587
Federer WT, Reynolds M, Crossa J (2001) Combining results from augmented designs over sites. Agron J 93:389–395
Fu YB (2015) Understanding crop genetic diversity under modern plant breeding. Theor Appl Genet 128:2131–2142
Gautam PL, Singh BB, Saxena S, Sharma RK (2004) Collection, conservation and utilization of plant genetic resources. In: Plant breeding: mendelian to molecular approaches, pp 657–690. https://doi.org/10.1007/978-94-007-1040-5_28
Getnet BE (2018) Genetic variability, heritability and expected genetic advance as indices for selection in soybean [Glycine max (L.) Merrill] varieties. Am J Life Sci 6:52–56
Ghafoor A, Sharif A, Ahmad Z, Zahid MA, Rabbani MA (2001) Genetic diversity in blackgram (Vigna mungo L. Hepper). Field Crops Res 69:183–190
Ghanbari S, Nooshkam A, Fakheri BA, Mahdinezhad N (2018) Assessment of yield and yield component of soybean genotypes (Glycine max L.) in north of Khuzestan. J Crop Sci Biotechnol 21:435–441
Government of Pakistan (2022–2023) Government of Pakistan, Ministry of Finance. Economic Advisor’s Wing, Islamabad. http://www.mnfsr.gov.pk. Accessed 12 Aug 2023
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Genet Res Int 1:1–14
Grainger CM, Rajcan I (2014) Characterization of the genetic changes in a multi-generational pedigree of an elite Canadian soybean cultivar. Theor Appl Genet 127:211–229
Guo B, Sun L, Jiang S, Ren H, Sun R, Wei Z, Hong H, Luan X, Wang J, Wang X, Xu D (2022) Soybean genetic resources contributing to sustainable protein production. Theor Appl Genet 135:4095–41121
Habib G, Khan MF, Javaid S, Saleem M (2016) Assessment of feed supply and demand for livestock in Pakistan. J Agric Sci Technol 6:191–202
Hartman GL, West ED, Herman TK (2011) Crops that feed the World 2. Soybean—worldwide production, use, and constraints caused by pathogens and pests. Food Secur 3:5–17
Hernández-Sebastià C, Marsolais F, Saravitz C, Israel D, Dewey RE, Huber SC (2005) Free amino acid profiles suggest a possible role for asparagine in the control of storage-product accumulation in developing seeds of low-and high-protein soybean lines. J Exp Bot 56:1951–1963
Hwang EY, Song Q, Jia G, Specht JE, Hyten DL, Costa J, Cregan PB (2014) A genome-wide association study of seed protein and oil content in soybean. BMC Genom 15:1–12
Hyten DL, Song Q, Zhu Y, Choi IY, Nelson RL, Costa JM, Specht JE, Shoemaker RC, Cregan PB (2006) Impacts of genetic bottlenecks on soybean genome diversity. Proc Natl Acad Sci 103:16666–16671
IBPGR (1984) Descriptors for Soybean. International Board for Plant Genetic Resources; IBPGR Secretariat: Rome Italy, pp 19‒38
Iqbal Z, Arshad M, Ashraf M, Mahmood T, Waheed A (2008) Evaluation of soybean [Glycine max (L.) Merrill] germplasm for some important morphological traits using multivariate analysis. Pak J Bot 40:2323–2328
Iqbal Z, Arshad M, Ashraf M, Naeem R, Malik MF, Waheed A (2010) Genetic divergence and correlation studies of soybean [Glycine max (L.) Merrill.] genotypes. Pak J Bot 42:971–976
Jiang B, Nan H, Gao Y, Tang L, Yue Y, Lu S, Ma L, Cao D, Sun S, Wang J, Wu C (2014) Allelic combinations of soybean maturity loci E1, E2, E3 and E4 result in diversity of maturity and adaptation to different latitudes. PLoS ONE 9:1–11
Kachare S, Tiwari S, Tripathi N, Thakur VV (2020) Assessment of genetic diversity of soybean (Glycine max) genotypes using qualitative traits and microsatellite markers. Agric Res 9:23–34
Kempton RA, Gleeson AC (1997) Unreplicated trials. statistical methods for plant variety evaluation. Springer, Dordrecht, pp 86–100
Khanande AS, Jadhav PV, Kale PB, Madavi SM, Mohari MP, Nichal SS (2016) Genetic diversity in vegetable and grain type soybean genotypes identified using morphological descriptor and EST-SSR markers. Vegetos Int J Plant Res 29:1–16
Khurshid H, Baig D, Nawaz N, Rabbani MA, Shinwari ZK (2020) intraspecific variation in soybean (Glycine max) revealed by multivariate approaches to morphometric traits and soybean mosaic virus resistance. Int J Agric Biol 23:1149–1157
Kirkpatrick LA (2015) A simple guide to IBM SPSS Statistics-Version 23.0. Cengage Learning, Chennai
Kumar A, Pandey A, Aochen C, Pattanayak A (2015) Evaluation of genetic diversity and interrelationships of agro-morphological characters in soybean (Glycine max) genotypes. Proc Natl Acad Sci India Biol Sci 85:397–405
Kumar SJ, Susmita C, Sripathy KV, Agarwal DK, Pal G, Singh AN, Kumar S, Rai AK, Simal-Gandara J (2022) Molecular characterization and genetic diversity studies of Indian soybean (Glycine max (L.) Merr.) cultivars using SSR markers. Mol Biol Rep 1‒12. https://doi.org/10.1007/s11033-021-07030-4
Kwanyuen P, Pantalone VR, Burton JW, Wilson RF (1997) A new approach to genetic alteration of soybean protein composition and quality. J Am Oil Chem Soc 74:983–987
Kwon SH, Torrie JH (1964) Heritability and interrelationships among traits of two soybean populations. Crop Sci 4:196–198
Latimer Jr GW (2016) Official methods of analysis of AOAC International 20th edition, Appendix D, Guidelines for collaborative study procedures to validate characteristics of a method of analysis. Gaithersburg, MD, USA
Leite WD, Unêda-Trevisoli SH, Silva FM, Silva AJ, Mauro AO (2018) Identification of superior genotypes and soybean traits by multivariate analysis and selection index. Rev Cienc Agron 49:491–500
Li M, Liu Y, Wang C, Yang X, Li D, Zhang X, Xu C, Zhang Y, Li W, Zhao L (2020) Identification of traits contributing to high and stable yields in different soybean varieties across three Chinese latitudes. Front Plant Sci 10:1642
Liu S, Xue H, Zhang K, Wang P, Su D, Li W, Xu S, Zhang J, Qi Z, Fang Y, Li X (2019) Mapping QTL affecting the vertical distribution and seed set of soybean [Glycine max (L.) Merr.] pods. Crop J 7:694–706
Machado BQ, Nogueira AP, Hamawaki OT, Rezende GF, Jorge GL, Silveira IC, Medeiros LA, Hamawaki RL, Hamawaki CD (2017) Phenotypic and genotypic correlations between soybean agronomic traits and path analysis. Genet Mol Res 16:1–11
Malek MA, Rafii MY, Afroz SS, Nath UK, Mondal M (2014) Morphological characterization and assessment of genetic variability, character association, and divergence in soybean mutants. Sci World J 1:1–12
Malik MF, Ashraf M, Qureshi AS, Ghafoor A (2007) Assessment of genetic variability, correlation and path analyses for yield and its components in soybean. Pak J Bot 39:405–413
Malik MF, Ashraf MU, Qureshi AS, Khan MR (2011) Investigation and comparison of some morphological traits of the soybean populations using cluster analysis. Pak J Bot 43:1249–1255
Mannan MA, Karim MA, Khaliq QA, Haque MM, Mian MA, Ahmed JU (2010) Assessment of genetic divergence in salt tolerance of soybean (Glycine max L.) genotypes. J Crop Sci Biotechnol 13:33–37
Maxted N, Hawkes JG, Guarino L, Sawkins M (1997) Towards the selection of taxa for plant genetic conservation. Genet Resour Crop Evol 44:337–348
Medic J, Atkinson C, Hurburgh CR (2014) Current knowledge in soybean composition. J Am Oil Chem Soc 91:363–384
Mello-Filho OL, Sediyama CS, Moreira MA, Reis MS, Massoni GA, Piovesan ND (2004) Grain yield and seed quality of soybean selected for high protein content. Pesqui Agropecu Bras 39:445–450
Misganaw A, Feyissa T, Mekonnen T, Desalegne O, Disasa T (2023) Genetic diversity analysis of sorghum genotypes for sustainable genetic resource conservation and its implication for breeding program in ethiopia. Genet Resour Crop Evol 70:1831–1852
Nasir B, Razzaq H, Sadaqat HA, Wahid MA (2023) Selection of soybeans for adaptation through principal component analysis under different climatic factors at seedling stage. Pak J Bot 55:665–678
Poehlman JM (2013) Breeding field crops. Springer, Berlin, pp 76–84
Popović V, Vidić M, Jocković Đ, Ikanović J, Jakšić S, Cvijanović G (2012) Variability and correlations between yield components of soybean [Glycine max (L.) Merr.]. Genetika 44:33–45
Qin P, Song W, Yang X, Sun S, Zhou X, Yang R, Li N, Hou W, Wu C, Han T, Ren G (2014) Regional distribution of protein and oil compositions of soybean cultivars in China. Crop Sci 54:1139–1146
Rai SK, Charak D, Bharat R (2016) Scenario of oilseed crops across the globe. Plant Arch 16:125–132
Reni YP, Rao YK (2013) Genetic variability in soybean [Glycine max (L) Merrill]. Int J Plant Anim Env Sci 3:35–38
Saba I, Sofi PA, Zeerak NA, Mir RR, Gull M (2017) Using augmented design for evaluation of common bean (Phaseolus vulgaris L.) germplasm. Int J Curr Microbiol Appl Sci 6:246–254
Salgotra RK, Chauhan BS (2023) Genetic diversity, conservation, and utilization of plant genetic resources. Genes 14:174
Selle PH, Dorigam JC, Lemme A, Chrystal PV, Liu SY (2020) Synthetic and crystalline amino acids: alternatives to soybean meal in chicken-meat production. Animals 10:729
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27:379–423
Sharma S, Kaur M, Goyal R, Gill BS (2014) Physical characteristics and nutritional composition of some new soybean (Glycine max (L.) Merrill) genotypes. J Food Sci Technol 51:551–557
Shrestha P, Pandey MP, Dhakal KH, Ghimire SK, Thapa SB, Kandel BP (2023) Morphological characterization and evaluation of soybean genotypes under rainfed ecosystem of Nepal. J Agric Food Res 11:100526
Sobko O, Zikeli S, Claupein W, Gruber S (2020) Seed yield, seed protein, oil content, and agronomic characteristics of soybean (Glycine max L. Merrill) depending on different seeding systems and cultivars in Germany. Agronomy 10:1020
Soy Stats (2022) International: world oilseed production. http://soystats.com/international-world-oilseed-production Accessed 21 Sept 2022
Stobaugh B, Florez-Palacios L, Chen P, Orazaly M (2017) Agronomic evaluation of high-protein and high-oil soybean genotypes for specialty markets. J Crop Improv 31:247–260
Sudarić A, Kočar MM, Duvnjak T, Zdunić Z, Kulundžić AM (2019) Improving seed quality of soybean suitable for growing in Europe. In: Soybean for human consumption and animal feed, pp 416–680. https://doi.org/10.5772/intechopen.89922
Tariq A, Jabeen Z, Farrakh S, Noreen K, Arshad W, Ahmed H, Haider W (2022) Exploring the genetic potential of Pakistani soybean cultivars through RNA-seq based transcriptome analysis. Mol Bio Rep 49:2889–2897
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327:818–822
Tripathi N, Khare D (2016) Molecular approaches for genetic improvement of seed quality and characterization of genetic diversity in soybean: a critical review. Biotechnol Lett 38:1645–1654
Ullah A, Akram Z, Malik SI, Khan KS (2021) Assessment of phenotypic and molecular diversity in soybean [Glycine max (L.) Merr.] germplasm using morpho-biochemical attributes and SSR markers. Genet Resour Crop Evol 68:2827–2847
USDA (2019) Oilseed and products update 2019; Global Agricultural Information Network (GAIN) Report Number: PK1916. US Department of Agriculture (USDA) Foreign Agricultural Service: Washington, DC, USA
Vollmann J, Fritz CN, Wagentristl H, Ruckenbauer P (2000) Environmental and genetic variation of soybean seed protein content under Central European growing conditions. J Sci Food Agric 80:1300–1306
Ward JH Jr (1963) Hierarchical grouping to optimize an objective function. J Am Stat Assoc 58:236–244
Wilson RF (2004) Seed composition. In: Boerma RH, Specht JE (eds) Soybeans: improvement, production, and uses, 3rd edn. American Society of Agronomy, Madison, pp 621–669
Xu C, Wu T, Yuan S, Sun S, Han T, Song W, Wu C (2022) Can soybean cultivars with larger seed size produce more protein, lipids, and seed yield? A meta-analysis. Foods 11:4059
Yadava DK, Dikshit HK, Mishra GP, Tripathi S (2022) Fundamentals of field crop breeding. Springer, Berlin, pp 907–944
Zhao Q, Shi X, Yan L, Yang C, Liu C, Feng Y, Zhang M, Yang Y, Liao H (2021) Characterization of the common genetic basis underlying seed hilum size, yield, and quality traits in soybean. Front Plant Sci 12:610214
Žulj Mihaljević M, Šarčević H, Lovrić A, Andrijanić Z, Sudarić A, Jukić G, Pejić I (2020) Genetic diversity of European commercial soybean [Glycine max (L.) Merr.] germplasm revealed by SSR markers. Genet Resour Crop Evol 18:1587–1600
Funding
This research was supported by Higher Education Commission (HEC) Pakistan under the Indigenous Scholarship Program “MS leading to PhD for the Students of Balochistan”.
Author information
Authors and Affiliations
Contributions
AU designed and performed the field and lab experiments. ZA conceptualized and conceived the experiment. GR and MW analyzed the data and wrote the paper. HK provided the genetic materials and assistance for experiment.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that there are no conflicts of interest.
Ethical approval
Ethical approval is not applicable for this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ullah, A., Akram, Z., Rasool, G. et al. Agro-morphological characterization and genetic variability assessment of soybean [Glycine max (L.) Merr.] germplasm for yield and quality traits. Euphytica 220, 67 (2024). https://doi.org/10.1007/s10681-024-03322-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-024-03322-5