Abstract
The origin of diploid C-genome oat Avena bruhnsiana, which is a rare endemic species from Azerbaijan (Apsheron Peninsula) has been clarified. This diploid oat was found only in Apsheron Peninsula in Azerbaijan and is closely related to another diploid species, A. ventricosa that belongs to C-genome oat group. In culture these species form fertile progeny but do not cross with other C-genome oats, A. clauda and A. pilosa. For more precise phylogenetic picture we used the next-generation sequence method that allows to obtain the whole pool of marker sequences (in our case, 18S rDNA(fragment)–ITS1–5.8S rDNA). Based on NGS counts, we revealed that at least 67% of A. bruhnsiana rDNA were received from A. ventricosa. The second ancestor of A. bruhnsiana is probably A. clauda. Avena clauda itself appears to be a homoploid hybrid: one of its main ribotypes is identical to A. pilosa and one of them is separate. Also, one of the minor ribotypes of A. pilosa is related to A-genomes (probably, ancestral state?). The only tetraploid in the genus, perennial A. macrostachya, has CmCm-genome and was also studied in our work. It takes distant position among other C-genome oats having two closely related main ribotypes. Avena macrostachya ribotypes is connected only with A. clauda/A. pilosa complex via minor ribotype fraction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A comparative analysis of mitotic chromosomes morphology in species of the genus Avena L. showed that diploid oat species have A-type or C-type genomes, and it was proposed to distinguish between two variants of C-genomes, Cv and Cp, and five variants of A-genomes (Ac, Ad, Al, Ap and As) (Rajhathy 1991; Loskutov and Rines 2011). To date, there are known one tetraploid Avena species with C-genome, A. macrostachya Balansa and Durieu (Rodionov et al. 2005; Loskutov and Rines, 2011) and four C-genome diploid species—A. ventricosa Balansa, A. pilosa Scop. (= A. eriantha Durieu), A. clauda Durieu and A. bruhnsiana Gruner (Loskutov and Rines 2011). The Cv genome of A. ventricosa consists of only subacrocentric chromosomes (Rajhathy 1971; Loskutov and Abramova 2006; Badaeva et al. 2010). The genomes of A. pilosa and A. clauda, called as Cp genome, are more complicated. There are two large submetacentrics, one medium size metacentrics, two medium size acrocentrics, and two small subtelocentrics (Fominaya et al. 1988; Shelukhina et al. 2008). There is a single nucleolar organizer region (NOR) in Cv genome but two NORs in both Cp genomes of diploid species (Fominaya et al. 1988; Loskutov and Abramova 2006; Shelukhina et al. 2008; Badaeva et al. 2010). As for A. bruhnsiana, the fourth diploid species with C-genome, it has a karyotype 2n = 14 with one or two pairs of submetacentrics (Emme 1930; Rajhathy 1971; Loskutov and Abramova 2006).
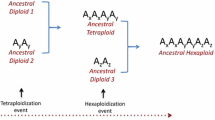
The initial stages of evolution of A- and C-versions of oats karyotypes can be represented as follows: the common ancestor of Avena had a seven pairs submetacentric chromosome set with two pair of NORs, similar in this respect to the Arrhenatherum P.Beauv. karyotype (Mitchell et al. 2003), the chromosome set of A. macrostachya, and the karyotypes of diploid oat species with the A genomes (Rodionov et al. 2005; Winterfeld et al. 2009; Badaeva et al. 2010). Then the separation of phylogenetic lines with A and C genomes occurred, accompanied by the accumulation of differences in scattered repeats (apparently determining the results of GISH hybridization) and the accumulation of chromosome rearrangements specific for each branch. In particular, amplification of some tandem repeats occurred in the phylogenetic branch of C-genome species, which led to the appearance of both few large C-heterochromatic bands and the characteristic "diffuse heterochromatin" on the chromosome arms (Rodionov et al. 2005; Winterfeld et al. 2009; Badaeva et al. 2010).
In the branch of C genome species, diploid ancestor of A. macrostachya was the first species that diverged from the common C-genome group ancestor (Rodionov et al. 2005; Winterfeld et al. 2009). Then several translocations or inversions took place in the common ancestor of current diploid C-genome species that led to significant changes in karyotype morphology C-genome and A-genome + A. macrostachya (Rajhathy 1991; Winterfeld et al. 2009; Badaeva et al. 2010). Avena ventricosa, the species which genome has lost one of NORs, was the first C-genome diploid species that diverged in this branch. C-banding and rDNA mapping showed that A. clauda and A. pilosa are closer relatives (Shelukhina et al. 2008; Badaeva et al. 2010).
The question is what position on the phylogenetic tree is occupied by the rare endemic species A. bruhnsiana. For the first time, this species was collected by pharmacist Alexander Bruhns on the Apsheron Peninsula and Swyatoi Island (now Pirallakhi Isl.) in the Caspian Sea in 1863–1864 and was published as a new species by Gruner (1867). According to its morphological features, A. bruhnsiana is very close to A. ventricosa, however A. bruhnsiana, as a rule, is larger in size characteristics (Musaev and Isaev 1971; Rajhathy 1971), especially in larger spikelets (Tzvelev 1983). The species occurs on the Apsheron Peninsula near coastal sands in undisturbed plant communities with A. ventricosa on gray-brown soils. Apparently, the species differs from A. ventricosa in the morphology of at least one pair of chromosomes (Emme 1930; Rajhathy 1971; Loskutov and Abramova 2006). Unlike A. ventricosa, the species has only one pair of NORs (Loskutov and Abramova 2006). The hybrids between A. bruhnsiana and A. ventricosa appear to be not differ from the parental species in fertility (Rajhathy 1971). On the other hand, A. bruhnsiana, when crossed with A. clauda and A. pilosa, did not produce fertile offspring (Nishiyama and Yabuno 1975). Malzew (1930) united A. ventricosa, which also occurs in the same Apsheron Peninsula, and A. bruhnsiana, into one species A. ventricosa Balansa, with two subspecies: subsp. ventricosa and subsp. bruhnsiana. Tzvelev (1983) considered A. bruhnsiana as a synonym of more widely distributed A. ventricosa. Rajhathy (1971) believed that karyologic data and the absence of reproductive isolation support Malzew (1930) for combining A. ventricosa Balansa and A. bruhnsiana Gruner into one species. At the same time, Azerbaijan botanists Musaev and Isaev (1971) suggested that A. bruhnsiana is possibly an interspecific hybrid.
The hybrid origin of plants can be revealed by examining its rDNA (Kovařik et al. 2005; Punina et al. 2012; Lunerova et al. 2017; Belyakov et al. 2019). In the plant genome, these cistrons are organized in the form of several repetitive transcription units encoding pre-rRNA (45S pre-rRNA in animals, 35-37S pre-rRNA in yeast and plants) (Kovařík et al. 2005; Garcia et al. 2012; Lunerova et al. 2017; Sochorová et al. 2018). In the studied plant species, the 35S rDNA cistron number varied from 150 to 26,048 per haploid genome (Prokopowich et al. 2003). In particular, among cereals, Secale cereale L. has 2900, Hordeum vulgare L. 2900–4200, Aegilops umbellulata Zhuk. 2500 gene copies. Note that S. cereale, like A. bruhnsiana, has single NOR per haploid genome, while both H. vulgare and A. umbellulata have two NORs per haploid genome (Rogers and Bendich 1987).
To study the origin of Avena bruhnsiana and probable relationships between this species and A. ventricosa, we investigated the intragenomic polymorphism of the 18S rDNA (partial sequence)-internal transcribed spacers ITS1-5.8S rDNA (partial sequence) loci using locus-specific sequencing of this region on the platform Illumina.
Materials and methods
Oat samples for our analysis were obtained from the collections of the Federal Research Center N. I. Vavilov All-Russian Institute of Plant Genetic Resources (VIR). Oats in VIR collections were gathered in their native habitats and then maintained via replanting. The following plant materials were used in our research: Avena bruhnsiana (k-212, Azerbaijan, Apsheron Peninsula, Coll. V.N. Soldatov; Genbank accession numbers OK303012–OK303068); Avena ventricosa (k-2056, Algeria, 3 km to the east of Oran, Coll. M. Leggett; Genbank accession numbers OK301935–OK302014); Avena clauda (k-269, Azerbaijan, Agsu District, Agsu, Coll. V.N. Soldatov; Genbank accession numbers OK273905–OK274031); Avena pilosa (k-1890, Syria, 40 km to the north from Damascus, Coll. M. Leggett; Genbank accession numbers OK273862–OK273899); Avena macrostachya (k-1856, Algeria, Atlas Mts, Djurdjura Mt., Coll. M. Leggett; Genbank accession numbers OK256902–OK256954).
DNA from leaf material was extracted according to the modified protocol by Doyle and Doyle (1987) and using Qiagen Plant Mini Kit (Qiagen inc., Germany) following the product manual.
Sequencing of the 18S rDNA (fragment)–internal spacer ITS1–5.8S rDNA (fragment) by the NGS method was carried out at the Center for Shared Use "Genomic Technologies, Proteomics and Cell Biology" of the All-Russian Research Institute of Agricultural Microbiology. The resulting sequences were trimmed using the Trimmomatic PE software (Bolger et al. 2014). The sequences (reads) were combined using the fastq-join program (Aronesty 2013). Then all the sequences were sorted using the "the bubble sorting" algorithm. We analyzed related sequences using MEGA X (Kumar et al. 2016). Then we processed the results of targeted-sequencing of the "population" ITS-sequences of studied Avena using the TCS program, which was used to build a network of haplotypes (Clement et al. 2000). The TCS algorithm is based on the probabilistic method of statistical parsimony and allows one to determine the probability of a relationship between all haplotypes with an indication of the number of mutations by which the studied haplotypes differ. The results of TCS calculations were visualized in TCS BU (Múrias dos Santos et al. 2016). The analysis included sequences with at least 10 reads per genome. The sequences of C-genome oats obtained by NGS method were also used in neighbor-net analysis. Neighbor-Net analysis was conducted with the aid of SplitsTree4 (Huson and Bryant 2006).
Results
The read and analyzed region of rDNA in our experiments included the 3 'end of 18S rDNA (71 b.p.), the ITS1 region (224 b.p.) and partial sequence of 5.8S rDNA (54 b.p.). The study of the intragenomic polymorphism of this sequence in the C-genome species showed that the species with one NOR—A. ventricosa demonstrates one dominant rDNA variant ribotype Cm1(12,560 reads, 81% of all reads) (Table 1, Figs. 1 and 2). 19% of the reads in this genome are minor variants of the major version of rDNA (Fig. 1). In the tetraploid species A. macrostachya, we see two closely related major ribotypes (Cm1A—4971 reads, 52% and Cm1B 3033 reads, 32%) (Table 1). In a diploid species with two NORs, A. pilosa, we found 2 main ribotypes—Cp1A (1366 reads, 82%) and Cp1B (1748 reads, 11%). Minor variants are close to these two major rDNA versions (Fig. 1). The level of differences between different ribotypes is small—one or several SNPs and oligonucleotide deletions (Table 2).
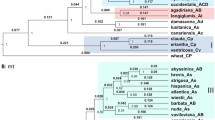
A completely different ribotype pattern is in A. clauda, the rDNA of this species is surprisingly diverse. First, it is the Cc1A ribotype, which is identical to the Cp1B ribotype (21% reads) and the closely related Cc1B ribotype (10% reads) (Table 1). Then a series of ribotypes Cc2A, Cc2B and Cc2C, the sequence of which is close to the rDNA of the species with type A genomes. The rDNA pattern of this species is characterized by a large number of minor variants, one group of which forms a “cloud” around the major variants Cc1B–Cp1B/Cc1A, and the second group is close to variants of ribotypes from the Cc2 family (Figs. 1 and 2).
The pattern of ribotypes of A. bruhnsiana shows that this is a species of hybrid origin, in fact nothospecies, in the genome of which the main part of rDNA (67%) is represented by the ribotype C1v, but 11% of the genome is the ribotype Cc1B (11%) and 2% of reads is the ribotype Cp1B/Cc1A (Table 1, Figs. 1 and 2). Note that the close relationship between A. bruhnsiana and A. ventricosa is confirmed by the fact that the minor rDNA variant with an extended deletion in the ITS1 region with a length of 70 b.p. (107 and 105 reads in A. ventricosa and A. bruhnsiana, correspondently—0.7% of reads in both genomes) was found only in the genomes of these two species. In Fig. 1 these sequences are marked with a four-pointed star.
The question of the second ancestor of this species, A. pilosa or A. clauda, is not resolved so unambiguously. However, it can be assumed that it was A. clauda because we detected A. clauda ribotypes that are common with the ribotypes of A. bruhnsiana. In the genome of A. bruhnsiana, in addition to rDNA of the Cc1B variant, we can see minor variants close to ribotypes of the Cc2B—1.7% of such reads in the genome.
Discussion
The nuclear RNA genes (rDNA) in higher plants are arranged in long tandem repeating units, much like those of other higher eukaryotes (Rogers and Bendich 1987). Plants generally have more rRNA genes than do other groups of organisms. In grasses (Poaceae) ribosomal genes are usually ca. 2000 per haploid genome. The 18S, 5.8S and 25S genes are clustered and transcribed as one unit (Rogers and Bendich 1987) and we need to note that this unit is repeated several times. Thus, our number of reads exceeding 10,000 per genome reflects not the gene quantity but the pool of all marker sequences including repeats of this transcription unit.
Homoploid hybrid speciation, where new hybridogenous species originate without changes in chromosome number has traditionally been considered as rare in relation to more common allopolyploid hybrid speciation (Abbott et al. 2010). Our results show that the diploid A. bruhnsiana is a taxon of hybrid origin: (nothospecies), one of its ancestors was A. ventricosa, and the second, apparently, A. clauda. Hence, we conclude that its name ought to be accepted more correctly as Avena × bruhnsiana. Since the karyotype of A. × bruhnsiana is diploid (2n = 14) (Emme 1930; Rajhathy 1971; Loskutov and Abramova 2006), it is a homoploid hybrid. According to the diversity of rDNA, one of its ancestral species, A. clauda, is also a homoploid hybrid, and perhaps one of its ancestors was a species with rDNA close to rDNA of diploid A-genome Avena species. Thus, of the four studied diploid species in the genus Avena, at least two are homoploid hybrids.
Another observation made in this work requires further research. Species with two nucleolar organizers in their genome on different chromosomes have at least two ribotypes, while A. ventricosa, which has a single NOR, has only one ribotype. This may indicate that homogenization of rDNA occurs within one NOR more efficiently than homogenization of rDNA at loci lying on different chromosomes. This may be because one of the mechanisms of rDNA homogenization is associated with the conjugation of homologous chromosomes and, therefore, it occurs more efficiently within one chromosome than between different chromosomes (Eickbush and Eickbush 2007; Sochorová et al. 2018).
Among studied C-genome oats there is one tetraploid (2n = 28), A. macrostachya. It is considered to be the most ancient species in the genus Avena (Nikoloudakis and Katsiotis 2008; Peng et al. 2008). Avena macrostachya, a perennial, cross-pollinated narrowly endemic species, was first collected in 1971 in the form of individual shoots (clones) in high altitude (1500 m) at the very edge of the snow in the mountainous region of Djurdjura of the Atlas Mts, in the northeastern Algeria (Baum and Rajhathy 1976). On mountain stony slopes covered by meadows and pastures, this species can rise up to 2000 m a.s.l. (Loskutov 2007). According to its morphological features, this perennial appears to be a primitive member of the genus Avena (Malzew 1930). Some researchers even attributed it to the genus Helictotrichon (Holub 1958). Avena macrostachya differs from diploid oat species with the C-genome in symmetric karyotype with a predominance of equal-armed chromosomes, absence of diffuse heterochromatin, predominantly pericentromeric arrangement of C-positive bands, as well as the size and morphology of satellite chromosomes (Badaeva et al. 2010). As it was found, the symmetric karyotype is not characteristic for diploid species with the C-genome. At the same time, large blocks of C-heterochromatin in the pericentromeric regions of chromosomes of this species indicate its affinity to species with C-genome. This confirms that A. macrostachya possesses a special C-genome type designated as CmCm (Rodionov et al. 2005). It was also considered that A. macrostachya could have previously undescribed genome EE (Loskutov 2007). Our analysis of NGS data on 18S–ITS1-5.8S rDNA sequences revealed that A. macrostachya ribotypes are comparatively distant from other existing C-genome oats (Fig. 1, Table 2).
As a conclusion, we can see that the Next-Generation Sequence analysis of rDNA intragenomic polymorphism gives much more complicated but informative results than the results of “traditional” sequence analysis. Here we see that even well-established diploid species have the tracks of ancient hybridization in their NORs.
References
Abbott RJ, Hegarty MJ, Hiscock SJ, Brennan AC (2010) Homoploid hybrid speciation in action. Taxon 59:1375–1386. https://doi.org/10.2307/20774035
Aronesty E (2013) Comparison of sequencing utility program. Open Bioinform J 7:1–8. https://doi.org/10.2174/1875036201307010001
Badaeva ED, Shelukhina O, Diederichsen A, Loskutov IG, Pukhalskiy VA (2010) Comparative cytogenetic analysis of Avena macrostachya and diploid C-genome Avena species. Genome 53:125–137. https://doi.org/10.1139/G09-089
Baum BR, Rajhathy T (1976) A study of Avena macrostachya. Can J Bot 54:2434–2439. https://doi.org/10.1139/b76-258
Belyakov EA, Machs EM, Mikhailova YV, Rodionov AV (2019) The study of hybridization processes within genus Sparganium L. subgenus Xanthosparganium Holmb. Based on data of next generation sequencing (NGS). Ecol Genet 17:27–35. https://doi.org/10.17816/ecogen17427-35
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660. https://doi.org/10.1046/j.1365-294x.2000.01020.x
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Eickbush TH, Eickbush DG (2007) Finely orchestrated movements: evolution of the ribosomal RNA genes. Genetics 175:477–485. https://doi.org/10.1534/genetics.107.071399
Emme H (1930) Über chromosomen von hafer und haferbastarden. Der Züchter 2:65–72
Fominaya A, Vega C, Ferrer E (1988) Giemsa C-banded karyotypes of Avena species. Genome 30:627–632. https://doi.org/10.1139/g88-106
Garcia S, Garnatje T, Kovařík A (2012) Plant rDNA database: ribosomal DNA loci data including other karyological and cytogenetic information in plants. Chromosoma 121:389–394. https://doi.org/10.1007/s00412-012-0368-7
Gruner L (1867) Plantae Bakuenses Bruhnsii. Verzeichniss der von dem Prov. Alexander Bruhns auf der Insel Swätoi und der Halbinsel Apscheron während der Jahre 1863–1865 gesammelten Pflanzen. Bull Soc Imp Nat Moscou 40:380–463
Holub J (1958) Bemerkungen zur taxonomie der gattung Helictotrichon Bess. In: Klaštersky I (ed) Philipp Maximilian opiz und seine bedeutung für die pflanzentaxonomie. Tschechoslowakische Akademie der Wissenschaften, Prague, pp 101–133
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267. https://doi.org/10.1093/molbev/msj030
Kovařik A, Pires JC, Leitch AR, Lim KY, Sherwood AM, Matyasek R, Rocca J, Soltis DE, Soltis PS (2005) Rapid concerted evolution of nuclear ribosomal DNA in two Tragopogon allopolyploids of recent and recurrent origin. Genetics 169:931–944. https://doi.org/10.1534/genetics.104.032839
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Loskutov IG (2007) Oat (Avena L.). Distribution, taxonomy, evolution and breeding value. VIR Publishers, St. Petersburg
Loskutov IG, Abramova LI (2006) Morphological and karyological study of wild species of the genus Avena L. Proc Appl Bot Genet Breed 162:108–113 (In Russian)
Loskutov IG, Rines HW (2011) Avena L. In: Kole Ch (ed) Wild crop relatives: genomic and breeding resources, vol 1. Springer, Heidelberg, p 77
Lunerova J, Renny-Byfield S, Matyášek R, Leitch A, Kovařík A (2017) Concerted evolution rapidly eliminates sequence variation in rDNA coding regions but not in intergenic spacers in Nicotiana tabacum allotetraploid. Plant Syst Evol 303:1043–1060. https://doi.org/10.1007/s00606-017-1442-7
Malzew AI (1930) Wild and cultivated oat. Sectio Euavena Griseb. Publ. of the All-Union Inst. of Appl. Botany and New Cultures under the Council of People's Commissars of the USSR, Leningrad
Mitchell CC, Parkinson SE, Baker TJ, Jellen EN (2003) C-banding and localization of 18S–5.8S S-26S rDNA in tall oatgrass species. Crop Sci 43:32–36
Múrias dos Santos A, Cabezas MP, Tavares AI, Xavier R, Branco M (2016) tcsBU: a tool to extend TCS network layout and visualization. Bioinformatics 32:627–628. https://doi.org/10.1093/bioinformatics/btv636
Musaev SG, Isaev YM (1971) Bruns’ Oat—an endemic species of the flora of Azerbaijan. Rep AS Azerb SSR 27:64–65 (In Russian)
Nikoloudakis N, Katsiotis A (2008) The origin of the C-genome and cytoplasm of Avena polyploids. Theor Appl Genet 117:273–281. https://doi.org/10.1007/s00122-008-0772-9
Nishiyama I, Yabuno T (1975) Meiotic chromosome pairing in two interspecific hybrids and a criticism of the evolutionary relationship of diploid Avena. Jpn J Genet 50:443–451
Peng Y-Y, Wei Y-M, Baum BR, Zheng Y-L (2008) Molecular diversity of the 5S rRNA gene and genomic relationships in the genus Avena (Poaceae: Aveneae). Genome 51:137–154
Prokopowich CD, Gregory TR, Crease TJ (2003) The correlation between rDNA copy number and genome size in eukaryotes. Genome 46:48–50. https://doi.org/10.1139/g02-103
Punina EO, Machs EM, Krapivskaya EE, Kim ES, Mordak EV, Myakoshina YA, Rodionov AV (2012) Interspecific hybridization in the genus Paeonia (Paeoniaceae): polymorphic sites in transcribed spacers of the 45S rRNA genes as indicators of natural and artificial peony hybrids. Rus J Genet 48:684–697. https://doi.org/10.1134/S1022795412070113
Rajhathy T (1971) Chromosome polymorphism in Avena ventricosa. Chromosoma 35:206–216. https://doi.org/10.1007/BF00285737
Rajhathy T (1991) The chromosomes of Avena. In: Gupta PK, Tsuchiya T (eds) Chromosome engineering in plants: genetics, breeding, evolution (Part A), vol 2. Elsevier, Amsterdam, pp 449–467
Rodionov AV, Tyupa NB, Kim ES, Machs EM, Loskutov IG (2005) Genomic configuration of the autotetraploid oat species Avena macrostachya inferred from comparative analysis of ITS1 and ITS2 sequences: on the oat karyotype evolution during the early events of the Avena species divergence. Russ J Genet 41:518–528
Rogers SO, Bendich AJ (1987) Ribosomal RNA genes in plants: variability in copy number and in intergenic spacer. Plant Mol Biol 9:509–520. https://doi.org/10.1007/BF00015882
Shelukhina OY, Badaeva ED, Brezhneva TA, Loskutov IG, Pukhalskiy VA (2008) Comparative analysis of diploid species of Avena L. Using cytogenetic and biochemical markers: Avena pilosa M. B. and A. clauda Dur. Rus J Gen 44:1087–1091. https://doi.org/10.1134/S1022795408090111
Sochorová J, Garcia S, Gálvez F, Symonová R, Kovařík A (2018) Evolutionary trends in animal ribosomal DNA loci: introduction to a new online database. Chromosoma 127:141–150. https://doi.org/10.1007/s00412-017-0651-8
Tzvelev NN (1983) Grasses of the Soviet Union (translated from the Russian edition of 1976). Oxonian Press Pvt. Ltd., New Delhi
Winterfeld G, Döring E, Röser M (2009) Chromosome evolution in wild oat grasses (Aveneae) revealed by molecular phylogeny. Genome 52:361–380. https://doi.org/10.1139/g09-012
Acknowledgements
The authors are grateful to A.G. Pinaev, and all researchers of Center for Shared Use "Genomic Technologies, Proteomics and Cell Biology" of the All-Russian Research Institute of Agricultural Microbiology for Next-generation sequencing.
Funding
This study was supported by Russian Foundation for Basic Research grant 20-516-10002 KO_a, St. Petersburg State University Grant 60256916 (bioinformatics) and within the framework of State Assignment No. 0662-2019-0006, as well as State Assignment No. AAAAA18-118040290161-3.
Author information
Authors and Affiliations
Contributions
AG, NN, EM carried out the experiments. AG, NN, EM and AR analyzed the data. IG, EB provided seed material. AG, NN, AR and NP wrote the manuscript. All authors read, revised and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gnutikov, A.A., Nosov, N.N., Loskutov, I.G. et al. New insights into the genomic structure of the oats (Avena L., Poaceae): intragenomic polymorphism of ITS1 sequences of rare endemic species Avena bruhnsiana Gruner and its relationship to other species with C-genomes. Euphytica 218, 3 (2022). https://doi.org/10.1007/s10681-021-02956-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-021-02956-z