Abstract
The objective of this paper was to measure resistance to Puccinia coronata f.sp. avenae (Pca) in a set of 63 previously uncharacterised new and historical oat cultivars representing 120 years of Polish oat breeding. The distinctiveness of the obsolete and modern gene pools led to the expectation that the earlier cultivars could be a valuable source of useful resistance genes discarded during subsequent breeding. The plant material was tested at the seedling stage using 14 Pca pathotypes and at the adult plant stage under natural infection conditions in 3 years of field observations. In all, 23 genotypes displayed an immune response to at least one Pca pathotype in the test at the seedling stage, however, most of them were susceptible to crown rust infection in the field. The most resistant, presenting immunity to 9 out of 14 P. coronata isolates, was Celer. The infection profile of this cultivar corresponded to the pattern of reference line Pc39. An allelism test confirmed that the resistance of both the Pc39 line and Celer is conditioned by either the same locus or by tightly linked genes. The infection analysis also demonstrated that some of the newest cultivars (Elegant, Komfort and Harnaś) showed a relatively high level of resistance in each year of field trials, despite being completely susceptible at the seedling stage. Further research will be required to evaluate the source, durability and suitability of this resistance. The study did not find any novel resistance to crown rust in the historic cultivars. However, results provide a new insight into adult plant stage resistance in Polish modern cultivars. The resistance of these cultivars is the result of many years of selection in breeding programs and provides an excellent base for future work.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
At the beginning of the last century, oat was one of the most widely grown cereals in the world. Currently, it is cultivated on a much smaller acreage of approx. 10.2 million ha, which is 4.7% of the acreage occupied by wheat (218 million ha) (http://www.fao.org/faostat/en/#data/QC). In the past, it owed its popularity mainly to horse feeding, but the mechanization of agriculture and transport significantly reduced the number of these animals, which also resulted in a lower oat production (Spiss 2003). Before World War II, oat in Poland was cultivated on 2 million ha.; in the 1960s it was grown on over 1.5 million ha, but currently, despite its valuable nutritional attributes, oat is grown on less than 0.5 million ha (Spiss 2003). Nevertheless, as of 2018 Poland is the fourth ranked oat producer in the world by harvested grain, after Russian Federation, Canada and United States (http://www.fao.org/faostat/en/#data/QC).
Oat breeding in Poland is of a similar vintage to that of Germany, United Kingdom and Sweden, and dates back to the end of the nineteenth century. Until World War II, during which all breeding materials were lost, the quality of Polish cultivars were comparable to other European oats (Świerczewski and Mazaraki 1993). After World War II Polish oat breeding had to be thoroughly rebuilt. The bases for further breeding programs were German cultivars introduced as compensation for destroyed breeding materials and oat genotypes from all over Europe (Świerczewski and Mazaraki 1993). Studies of genetic variation within Polish oat cultivars shows that diversity within obsolete as well as modern gene pools is quite narrow. However, there are large differences between these two pools, meaning that the earlier oat varieties could be a valuable source of useful genes. Such germplasm may be much more easily introduced into breeding programs than wild-related species. Significantly, both pedigree studies and genetic analyses indicate that only few of them have been used in the creation of modern cultivars (Paczos-Grzęda 2004, 2007; Boczkowska et al. 2015; Boczkowska and Onyśk 2016).
Rusts are known as the most destructive of cereal diseases and historically have caused devastating yield losses worldwide (Chaves et al. 2008; Fetch et al. 2011). In oats, crown rust (Puccinia coronata Cda. f. sp. avenae P. Syd. & Syd., Pca) is considered the most important foliar disease leading to great yield and grain quality losses (Simons 1985). The most common disease control method in Poland is the use of chemical plant protection products. However, fungicides are a serious threat to the natural environment. Moreover, the high cost of chemical protection in comparison to low price of oat grain on the market make such treatment unprofitable (McCallum et al. 2007). Thus plant breeding directed at genetic resistance is an alternative and highly promising approach (Gnanesh et al. 2014).
Breeding disease resistant cultivars is a long and laborious process. Usually recurrent back-crosses are required to eliminate transmission of unfavourable traits from the resistant donor into the new cultivar (Pradesh et al. 2013). Selection of resistant genotypes can be carried out using either physiological tests or marker-assisted selection (MAS) (Miedaner and Korzun 2012; Miedaner 2016). There are close to 100 P. coronata resistance genes identified (CDL 2018) however, the number of molecular markers useful in breeding is very limited. Most of the reported markers (RAPDs, AFLPs, and DArTs) are dominant and are not ideal for high-throughput genotyping, nevertheless, both major and minor R genes have been utilised in breeding for resistance to crown rust worldwide (Cabral et al. 2014). The highly variable nature of P. coronata (Chong and Zegeye 2004; Carson 2008, 2011; Chong et al. 2008, 2011; Menzies et al. 2015; Paczos-Grzęda and Sowa 2019) allows it to eventually overcome major gene-for-gene resistance, so constant replacement of these genes is required.
Characterizing historic breeding programme material may provide valuable information about pedigrees and lead to recovery of germplasm of use to current programmes. In this paper the crown rust resistance spectrum of oat cultivars developed throughout 120 years of Polish oat breeding was studied. The resistance of a diverse set of 63 new and historical Polish oat cultivars was tested at seedling as well as adult plant stage.
Materials and methods
Plant material
Plant material consisted of 63 Polish oat cultivars (Avena sativa L.) of which 21 were cultivars registered in Research Centre for Cultivar Testing (COBORU) (http://www.coboru.pl). The remaining genotypes were historical oat cultivars from the collection of Institute of Genetics, Plant Breeding and Biotechnology, University of Life Sciences in Lublin, Poland as well as the National Centre for Plant Genetic Resources (Radzików, Poland). Oat cultivars registered in COBORU after 2012 were tested as lines obtained directly from breeding companies.
The differential near-isogenic lines Pc14, Pc36, Pc51, Pc52, Pc70, Pc71 were developed at Iowa State University, USA, and the lines Pc35, Pc38, Pc39, Pc40, Pc45, Pc46, Pc48, Pc50, Pc54, Pc55, Pc56, Pc57, Pc62, Pc63, Pc64, Pc68, Pc91, Pc94, Pc96, Pc97, Pc98, Pc101, Pc103-1 and Pc104 were created at the Cereal Research Centre AAFC Winnipeg, Canada (Carson 2011; Chong et al. 2011; Menzies et al. 2015). Pc58, Pc59, Pc60 and Pc61 were represented by TAM-O-301, TAM-O-312, Coker227, and Coker234, respectively (Simons et al. 1978).
Pathogen isolates
Crown rust resistance of Avena genotypes was tested using 14 P. coronata isolates (Table S1). Isolates were selected from a wide collection of single-pustule isolates derived from populations collected in central, southern and south-eastern Poland in the years 2013–2015 (Fig. 1) according to the method described by Sowa et al. (2016). The virulence of each isolate was determined on a set of 35 differential oat lines with different resistance genes (below).
The crown rust resistance tests
The host–pathogen test (Hsam et al. 1997) was conducted on three leaf fragments, each from a different seedling of one Avena genotype, as described in Sowa et al. (2016). Seeds were grown for 10 days in plug trays filled with a universal substrate containing peat. 3-cm-long leaf fragments were placed onto 12-well culture plates with agar (0.6%) containing benzimidazole (3.4 mM). In each well a single leaf fragment of susceptible cultivar Marvelous was used as infection control. Inoculations were performed in a settling tower by applying 500–700 spores of P. coronata per 1 cm2. Plates were incubated for 10 days in a phytotron at 18 °C with 70% humidity and light intensity of approximately 4 kLx for a 16-h photoperiod. Assessment of crown rust disease symptoms was performed after 12 days using 0–4 infection type (IT) qualitative scale, which were transformed to S, MS, MR, R, and HR, where S = 4 = susceptible (large to moderately large pustules with little or no chlorosis); MS = 3 = moderately susceptible (moderately large pustules surrounded by extensive chlorosis); MR = 2, 2 N, 12C, ;1C = moderately resistant (small pustule surrounded by chlorosis or necrosis); R = ;-N, ;C, ;+ C, ;1N = resistant (chlorotic or necrotic flecking); and 0 = HR = highly resistant (no visible reaction) (Sowa et al. 2016, see Nazareno et al. 2018 for an illustration). Reactions to the crown rust isolate infections were grouped into two classes: phenotypes described as S and MS were considered as susceptible, and the remainder as resistant. On the bases of these classes, infection profiles (IP) describing distinct patterns of response to crown rust isolates infection were determined. The infection profiles (IP) of the screened cultivars were then compared with the IPs of the differential lines with Pc genes used to evaluate virulence of P. coronata isolates.
Scores of infection were transformed into a 0/1 matrix that recorded the susceptibility (0) or resistance (1) to crown rust in order to perform hierarchical analysis. Data were used to conduct clustering and construct a dissimilarity dendrogram based on the unweighted pair-group method (UPGMA) in XlStat v. 2014.1.01 Excel add-in software. Groups and subgroups were determined using Dice coefficient (Dice 1945).
Allelism test
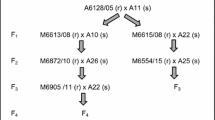
Celer was crossed with the Pc39 reference line. F1 plants were selfed and 96 F2 plants used for allelism tests conducted as above. One leaf from each seedling was cut into four 3-cm-long fragments. Seedling progeny and three seedlings of each parent were tested with four P. coronata pathotypes (4.2, 11.1, 12.2, 13.3).
Field experiments
Field experiments were used to evaluate P. coronata crown rust resistance of tested cultivars in adult plants. Experiments were carried out at the experimental farm of the University of Life Sciences in Lublin (Czesławice 51°18 N, 22°15 E) in 2012, 2013 and 2014 using 1 m rows arranged in a randomised block design, with three replications. About 30 seeds were sown in each row. The plants were cultivated in a crop rotation system after sugar beet, using the recommended fertilizer inputs and allowed to become naturally inoculated with uredospores of P. coronata. Every second plot one row of the susceptible cultivar Marvelous was planted to ensure that each entry had an equal chance to get secondary infection.
Crown rust severity was assessed three times during the growing season by visual inspection of leaves according to the 0–9 scale proposed by McNeal et al. (1971). The highest degree of infection was recorded. Infection score of 0–4 was cosidered as resistant; 5–6, intermediate and 7–9, susceptible. Obtained data were subjected to one-way ANOVA followed by Newman-Keuls test at 0.05 probability level in Statistica Software 13.1 (Tibco Software Inc. 2017).
Results
Fourteen P. coronata isolates (Table S1) were used for the host–pathogen tests and screening of seedling resistance of 63 Polish new and historic oat cultivars. The virulence of each isolate was determined on a set of 35 differentials. These were the most diverse pathotypes selected from a wide collection of single-pustule isolates derived from populations collected in central, southern and south-eastern Poland in the years 2013–2015 where the level of virulence against reference Pc genes ranged between 23 and 66%. The most aggressive isolate (3.2) overcame the resistance of 23 out of 35 tested lines, while the two least aggressive (4.2 and 107.2) were virulent against 8 reference lines. The diverse virulence profile of pathotypes differentiates 25 Pc reference oat lines, each showing a unique infection profile (IP_16–IP_36, IP_38–IP_40, IP_42) (Table S2). Of the remaining lines, three infection patterns can be determined. The first group consists fully resistant lines Pc52, Pc59, Pc60, Pc68, Pc71 and Pc91 (IP_43). The second group containing genes Pc57 and Pc104 were overcome by isolates 94.1, 94.2 and 132.1 (IP_41), and the third group including lines Pc51 and Pc101 were overcome by 3.2, 94.1, 94.2 and 132.1 crown rust pathotypes (IP_37).
The response of the 63 analysed cultivars on 14 Pca isolates infection ranged from S to HR (Table 1, Table S3). In all, 23 genotypes displayed immune response to at least one Pca pathotype. Five cultivars, Bajka, Furman, Rajtar, Szakal and Zuch, showed resistance to one isolate. Two cultivars, Cacko and Deresz, were resistant to two isolates. The resistance of Stoper was not overcome by five isolates. Six varieties Chwat, Haker, Krezus, Paskal and Skrzat showed resistance to six isolates. Arden, Bohun, Grajcar and Koneser were resistant to seven isolates, whereas cultivars Borowiak, Nawigator, Sławko and Sprinter showed resistance to 8 isolates. The highest level of resistance to crown rust was presented by the cultivar Celer, which was resistant to 9 out of 14 P. coronata isolates.
Based on the reaction to 14 Pca isolates inoculations, 16 infection profiles (IPs) of screened genotypes were determined. A highly virulent response to all of the crown rust races, assigned as IP_0, was presented by 40 Avena cultivars. The remaining genotypes could be grouped into some identical IP sets. Szakal and Zuch were resistant to Pca race 124.1 (IP_2). Cacko and Deresz showed resistance to 11.1 and 12.2 (IP_5). Chwat and Krezus were resistant to 11.1, 12.2, 15.1, 23.1, 107.2, 132.1 (IP_8). Haker and Skrzat were resistant to the 12.2, 15.1, 22.1, 23.1, 107.2, 132.1 (IP_10). Arden, Koneser, Bohun and Grajcar displayed resistance to Pca races 11.1, 12.2, 15.1, 22.1, 23.1, 107.2 and 132.1 (IP_11). Cultivar infection profiles were compared with Pc differential line responses to inoculation with the same isolates. Only cultivar Celer (IP_16) presented a profile which corresponded to the pattern of line Pc39 (resistance to 4.2, 13.3, 22.1, 23.1, 26.1, 94.1, 94.2, 107.2 i 132.1). These relationships are displayed in a dissimilarity dendrogram based on the Dice coefficient (Fig. 2). The cluster analysis identified two distinct clusters composed of 40 genotypes of IP_0 and 23 displaying immune response to at least one crown rust pathotype. Within the second cluster two subgroups can be distinguished, the first composed of five cultivars of IP_2, IP_3 and IP_5 and the second including eighteen remaining cultivars and line Pc39. All cultivars with identical infection profile have a dissimilarity value of zero. Additionally, clusters with a dissimilarity coefficient ≤ 0.2 can be identified. Cultivars characterized by IP_11 cluster with Chwat and Krezus (IP_8). Haker and Skrzat (IP_10) group with Stoper (IP_6), Nawigator (IP_12) cluster with Sławko and Celer—line Pc39 group with Borowiak (IP_14) as well as Sprinter (IP_15).
UPGMA dendrogram of 63 polish oat cultivars and Pc39 reference line based on the similarity of infection obtained by 14 P. coronata isolates inoculation. Each cultivars name is followed by infection profile IP at the seedling stage (in brackets) as well as the year of registration. Cultivars currently registered in Research Centre for Cultivar Testing (COBORU) are marked with *
Within the Celer/Pc39 F2 population no phenotype segregation was observed. Both parental forms and F2 progeny plants showed resistance against isolates 4.2 and 13.3 and susceptibility to infection with isolates 11.1 and 12.2.
Every year in field experiments conducted in Czesławice infection of the control cultivar Marvelous was at the highest level of the 10-point scale of McNeal et al. (1971). 75% of tested cultivars were scored as susceptible (Fig. 3). Nine cultivars, Arden, Bohun, Haker, Stoper, Arab, Dukat, Hetman and Romuluspresented intermediate resistance. Eight remaining cultivars, Elegant, Nawigator, Borowiak, Celer, Paskal, Komfort, Sprinter and Harnaś were scored as resistant, with an average infection score not exceeding 4. Newman-Keuls test confirmed statistically significant difference between these resistant cultivars and the rest of genotypes (Table S4).
Characteristics of A. sativa cultivar reactions to natural infection of P. coronata at the adult plant stage according to the 0–9 scale proposed by McNeal et al. (1971). Each cultivars name is followed by infection profile IP at the seedling stage (in brackets) as well as the year of registration. Cultivars currently registered in Research Centre for Cultivar Testing (COBORU) are marked with *
Discussion
The current work presents the characteristics of resistance to P. coronata in a set of 63 new and historical Polish oat cultivars. Material selection was based on the assumption that old varieties might be a rich and underutilized source of diversity. Nine cultivars represented the period of breeding before World War II. Most of them were the selection from Polish (Sobieszyński, Jagiełło) or foreign (Antoniński Żółty, Teodozja) landraces. Despite many efforts of Polish breeders, for many years after the war the Polish market was dominated by foreign cultivars, Flamingsweiss and Romulus (East Germany), Diadem (Czechoslovakia), Leanda and Perona (Netherlands) (Król and Mucha 1977). The first postwar Polish cultivar, which immediately took over 60% of the Polish market, was Dragon, registered in 1982 (Świerczewski and Mazaraki 1993). This cultivar began a new era in Polish breeding, with a few new cultivars registered every year. Currently, about 90% of Avena sativa cultivars on The Polish National List of Agricultural Plant Varieties of COBORU (http://www.coboru.pl) has been developed by Polish breeders. Out of cultivars released after 1982, more than 50 were selected based on their importance to the current studies.
P. coronata isolates used in host–pathogen tests were selected from a wide collection of single-pustule isolates derived from populations collected in Poland in the years 2013–2015 (Paczos-Grzęda and Sowa 2019). To differentiate crown rust resistance conditioned by R and adult plant resistance (APR) genes, genotypes were screened at the seedling and adult plant stage as R genes mostly function at all growing stages, whereas APR function mainly at the adult stage (Periyannan et al. 2017). In all, 23 genotypes displayed immune response to at least one Pca pathotype in the test at the seedling stage, however, most of them were susceptible to crown rust infection in field studies. Such resistance may be determined by race-specific genes or genes effective only at the seedling stage, which are not expressed or are ineffective at the adult stage.
Almost all susceptible genotypes were historic and modern cultivars registered in COBORU before the year 2000. However, some modern cultivars, such as Berdysz, Bingo, Komfort, Siwek and Harnaś were also fully susceptible. In contrast, a very high level of resistance was presented by the recent varieties Arden and Nawigator. The most resistant, presenting immunity to 9 out of 14 P. coronata isolates was Celer. This cultivar was developed in the breeding station in Borowo and one of its parental genotypes is Góral. Our results indicate that this cultivar was not the source of resistant alleles, as it was susceptible in both growing stages. The second parental component of Celer was the KR-KOR, a line derived in the Krukanice Plant Breeding Station. This station was one of the locations of field trials of Pc reference lines conducted in Czech Republic (Šebesta and Harder 1983), so possibly some of these lines were used as a breeding component. Góral along with Santor were also parental forms of Borowiak. Borowiak presented one of the highest levels of resistance in both growing stages, with its parental forms being completely susceptible, so the resistance of Borowiak may be quantitative, conditioned by many genes of minor effect.
A. sativa cultivars were analyzed in terms of the infection patterns similarity. Some varieties were characterized by the same IP, which may indicate a common origin. Szakal and Zuch were resistant to the same pathotype, 124.1 with Zuch being the progeny of Vusch × Szakal. Moreover Chwat with the pedigree Dukat × (Flamingsnova × Swan) and Krezus resulting from crossing the Góral × ((Flamingsnova × Swan mut.) × Dukat)) were also resistant to the same isolates 11.1, 12.2, 15.1, 23.1, 107.2 and 132.1 (IP_9). In addition, the same infection profiles were observed for Cacko and Deresz (IP_6), Haker and Skrzat (IP_11), as well as Arden, Koneser, Bohun and Grajcar (IP_12). However, the infection patterns of these genotypes did not correspond to any of the reference line infection patterns. This resistance might be polygenic or conditioned by allelic forms of known genes or genes not yet described. Only Celer presented an infection profile (IP_16) corresponding to the pattern of line Pc39 suggesting the presence of Pc39 in this cultivar. An allelism test conducted to verify this postulation indicates that the resistance of the Pc39 line and Celer is conditioned by either the same, allelic or tightly linked gene with less than 1% recombination. This result also confirms the reliability of the detached leaf assay used.
The above data has been supplemented with field experiments conducted in Czesławice. This is the location hosting field trials of a number of Avena genotypes with different Pc genes, leading to an expectation of high selection pressure and diversity within the occurring crown rust populations. Each year based on the level of rust infection of control cultivar Marvelous severe infection was observed. The majority of genotypes presented fully susceptible phenotypes in both growing stages. High levels of resistance frequently incur yield penalties (Brown 2002; Brown and Rant 2013; Ning et al. 2017) so it is possible that selection for yield as a priority may have led to elimination of some resistances during breeding. In a breeding programme resistance rarely is the most important target. Polish breeding stations conduct resistance selection mainly in the field. This strategy favours selection of resistance conditioned by genes of small effects determining less and slower pathogen growth (Niks et al. 2015). This type of resistance may be present in some of the newest cultivars Elegant Komfort and Harnaś, which were completely susceptible at the seedling stage, however showing relatively high level of resistance in each years field trials. Further research will be required to evaluate the source, durability and suitability of this APR.
This study is a part of the continuing effort to support and refine strategies for oat breeding programs (Sowa et al. 2016; Sowa and Paczos-Grzeda 2014; Paczos-Grzęda et al. 2018a; b; Paczos-Grzęda and Sowa 2019). The highly variable nature of P. coronata requires periodic monitoring of the occurrence and impact of pathogen populations as well as evaluating the effectiveness of resistance in breeding materials. The study demonstrates that screened historic cultivars are not a good source of new genes providing resistance to crown rust, however analyzed materials represented only a small part of old European oat genepool. Old oat cultivars developed in distinct breeding programs worldwide may exhibit different genetic diversity due to the local breeding strategies as well as historical and regional trends (van de Wouw et al. 2010; He and Bjørnstad 2012). This genepool may be potentially wider, as a significant loss of diversity occured at the transition from landraces and old cultivars to modern varieties (Nersting et al. 2006; He and Bjørnstad 2012). Novel and broad spectrum crown rust resistance were identified within Spanish (Sánchez-Martín et al. 2012; Montilla-Bascon et al. 2013, 2015) and Egyptian (Sánchez-Martín et al. 2017) oat landraces. Thus the possibility of uncovering the effective resistances harbored in obsolete, unexploited resources is not excluded.
The infection analysis gave also a new insight into the resistance of Polish modern cultivars in the adult plant stage. Resistance of these cultivars being the result of many years of selection in breeding programs can be valuable and promising for future work.
References
Boczkowska M, Onyśk A (2016) Unused genetic resources: a case study of Polish common oat germplasm. Ann Appl Biol 169:155–165. https://doi.org/10.1111/aab.12289
Boczkowska M, Harasimiuk M, Onyśk A (2015) Studies on genetic variation within old Polish cultivars of common oat. Cereal Res Commun 43:12–21. https://doi.org/10.1556/crc.2014.0025
Brown JKM (2002) Yield penalties of disease resistance in crops. Curr Opin Plant Biol 5:339–344. https://doi.org/10.1016/S1369-5266(02)00270-4
Brown JKM, Rant JC (2013) Fitness costs and trade-offs of disease resistance and their consequences for breeding arable crops. Plant Pathol 62:83–95. https://doi.org/10.1111/ppa.12163
Cabral AL, Gnanesh BN, Fetch JM et al (2014) Oat fungal diseases and the application of molecular marker technology for their control. In: Goyal A, Manoharachary C (eds) Future challenges in crop protection against fungal pathogens. Springer, New York, pp 343–358
Carson ML (2008) Virulence frequencies in oat crown rust in the United States from 2001 through 2005. Plant Dis 92:379–384. https://doi.org/10.1094/PDIS-92-3-0379
Carson ML (2011) Virulence in oat crown rust (Puccinia coronata f. sp. avenae) in the United States from 2006 through 2009. Plant Dis 95:1528–1534. https://doi.org/10.1094/PDIS-09-10-0639
CDL (2018) Cereal Disease Laboratory. Resistance genes. Oat. Pc (crown rust). http://www.ars.usda.gov/midwest-area/stpaul/cereal-disease-lab/docs/resistance-genes/resistance-genes/. Accessed 4 Oct 2018
Chaves MS, Martinelli JA, Wesp CdeL, Graichen FAS (2008) The cereal rusts: an overview. Pest Technol 2:38–55
Chong J, Zegeye T (2004) Physiologic specialization of Puccinia coronata f. sp. avenae, the cause of oat crown rust, in Canada from 1999 to 2001. Can J Plant Pathol 26:97–108. https://doi.org/10.1080/07060660409507119
Chong J, Gruenke J, Dueck R et al (2008) Virulence of oat crown rust Puccinia coronata f. sp. avenae in Canada during 2002–2006. Can J Plant Pathol 30:115–123. https://doi.org/10.1080/07060660809507502
Chong J, Gruenke J, Dueck R et al (2011) Virulence of Puccinia coronata f. sp. avenae in the eastern prairie region of Canada during 2007-2009. Can J Plant Pathol 33:77–87. https://doi.org/10.1080/07060661.2010.546957
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302. https://doi.org/10.2307/1932409
Fetch T, McCallum B, Menzies J et al (2011) Rust diseases in Canada. PS&C 4:86–96
Gnanesh BN, Fetch JM, Zegeye T, McCartney CA, Fetch T (2014) Oat. In: Pratap A, Kumar J (eds) Alien gene transfer in crop plants. Springer, New York, pp 51–73
He X, Bjørnstad Å (2012) Diversity of North European oat analyzed by SSR, AFLP and DArT markers. Theor Appl Genet 125(1):57–70. https://doi.org/10.1007/s00122-012-1816-8
Hsam SLK, Peters N, Paderina EV et al (1997) Genetic studies of powdery mildew resistance in common oat (Avena sativa L.) I. Cultivars and breeding lines grown in Western Europe and North America. Euphytica 96:421–427
http://www.coboru.pl Centralny Ośrodek Badania Odmian Roślin Uprawnych [Research Centre for Cultivar Testing]
Król M, Mucha S (1977) Charakterystyka i technologia uprawy odmian owsa. IUNG, Puławy
McCallum BD, Fetch T, Chong J (2007) Cereal rust control in Canada. Aust J Agric Res 58(6):639–647
McNeal FM, Konzak CF, Smith EP, Tate WS, Russell TS (1971) A uniform system for recording and processing cereal research data. USDA-ARS Bull 34:121
Menzies JG, Xue A, Dueck R, Greunke J (2015) Virulence of Puccinia coronata f. sp. avenae in Canada; 2010 to 2014. In: 14th International Cereal Rust and Powdery Mildew Conference 5–8 July 2015. Denmark, Copenhagen, p 95
Miedaner T (2016) Breeding strategies for improving plant resistance to diseases. In: Al-Khayri JM, Jain SM, Johnson DV (eds) Advances in plant breeding strategies: agronomic, abiotic and biotic stress traits. Springer, Cham, pp 1–707
Miedaner T, Korzun V (2012) Marker-assisted selection for disease resistance in wheat and barley breeding. Phytopathology 102:560–566. https://doi.org/10.1094/PHYTO-05-11-0157
Montilla-Bascón G, Sánchez-Martín J, Rispail N, Rubiales D, Mur LAJ, Langdon T, Griffiths I, Howarth C, Prats E (2013) Genetic diversity and population structure among oat cultivars and landraces. Plant Mol Biol Rep 31:1305–1314. https://doi.org/10.1007/s11105-013-0598-8
Montilla-Bascón G, Rispail N, Sánchez-Martín J, Rubiales D, Mur LAJ, Langdon T, Howarth CJ, Prats E (2015) Genome-wide association study for crown rust and powdery mildew resistance in an oat (Avena sativa L.) collection of commercial varieties and landraces. Front Plant Sci 6:103. https://doi.org/10.3389/fpls.2015.00103
Nazareno ES, Li F, Smith M et al (2018) Puccinia coronata f. sp. avenae: a threat to global oat production. Mol Plant Pathol 19:1047–1060. https://doi.org/10.1111/mpp.12608
Nersting LG, Andersen SB, von Bothmer R, Gullord M, Jorgensen RB (2006) Morphological and molecular diversity of Nordic oat through one hundred years of breeding. Euphytica 150:327–337. https://doi.org/10.1007/s10681-006-9116-5
Niks RE, Qi X, Marcel TC (2015) Quantitative resistance to biotrophic filamentous plant pathogens: concepts, misconceptions, and mechanisms. Annu Rev Phytopathol 53:445–470. https://doi.org/10.1146/annurev-phyto-080614-115928
Ning Y, Liu W, Wang GL (2017) Balancing immunity and yield in crop plants. Trends Plant Sci 22:1069–1079. https://doi.org/10.1016/j.tplants.2017.09.010
Paczos-Grzęda E (2004) Pedigree, RAPD and simplified AFLP-based assessment of genetic relationships among Avena sativa L. cultivars. Euphytica 138:13–22. https://doi.org/10.1023/B:EUPH.0000047055.99322.7a
Paczos-Grzęda E (2007) Wykorzystanie metod ISSR i RAPD oraz analizy rodowodów do oceny podobiemstwa międzyodmianowego Avena sativa (Avena sativa cultivars similarity estimation based on RAPD and ISSR methods and pedigree analysis). Zesz Probl Post Nauk Rol 517:547–558
Paczos-Grzęda E, Sowa S (2019) Virulence structure and diversity of Puccinia coronata f. sp. avenae P. Syd. & Syd. in Poland during 2013–2015. Plant Dis PDIS-10-18-1820-RE. https://doi.org/10.1094/pdis-10-18-1820-re
Paczos-Grzęda E, Sowa S, Koroluk A, Langdon T (2018a) Characteristics of resistance to Puccinia coronata f. sp. avenae in Avena fatua. Plant Dis 102:1–9. https://doi.org/10.1094/pdis-03-18-0528-re
Paczos-Grzęda E, Sowa S, Boczkowska M, Langdon T (2018b) Detached leaf assays for resistance to crown rust reveal diversity within populations of Avena sterilis L. Plant Dis. https://doi.org/10.1094/pdis-06-18-1045-re
Periyannan S, Milne RJ, Figueroa M et al (2017) An overview of genetic rust resistance: from broad to specific mechanisms. PLoS Pathog 13:1–6. https://doi.org/10.1371/journal.ppat.1006380
Pradesh A, Ragimekula N, Varadarajula NN et al (2013) Marker assisted selection in disease resistance breeding. J Plant Breed Genet 1:90–109
Sánchez-Martín J, Rubiales D, Sillero JC, Prats E (2012) Identification and characterization of sources of resistance in Avena sativa, A. byzantina and A. strigosa germplasm against a pathotype of Puccinia coronata f.sp. avenae with virulence against the Pc94 resistance gene. Plant Pathol 61:315–322. https://doi.org/10.1111/j.1365-3059.2011.02514.x
Sánchez-Martín J, Rispail N, Flores F, Emeran AA, Sillero JC, Rubiales D, Prats E (2017) Higher rust resistance and similar yield of oat landraces versus cultivars under high temperature and drought. Agron Sustain Dev 37:3. https://doi.org/10.1007/s13593-016-0407-5
Šebesta J, Harder DE (1983) Occurrence and distribution of virulence in Puccinia coronata var. avenae in Europe, 1977-1980. Plant Dis 67:56–59
Simons MD (1985) Crown rust. In: Roelfs AP, Bushnel WR (eds) The cereal rust, vol 2. Diseases, distribution, epidemiology, and control. Academic Press, Orlando, pp 131–172
Simons MD, Martens JW, McKenzie RIH et al (1978) Oats: a standardized system of nomenclature for genes and chromosomes and catalog of genes governing characters. US Department of Agricultural Handbook No. 509. US Department of Agriculture, Madison, WI
Sowa S, Paczos-Grzęda E (2014) Puccinia coronata f. sp. avenae virulence in south-eastern Poland in 2014. Folia Pomer Univ Technol Stetin Agric Aliment Pisc Zootech 336(43):157–166. https://doi.org/10.21005/oe.2017.86.1.06
Sowa S, Paczos-Grzęda E, Koroluk A et al (2016) Resistance to Puccinia coronata f. sp. avenae in Avena magna, A. murphyi, and A. insularis. Plant Dis 100:1184–1191. https://doi.org/10.1094/PDIS-06-15-0671-RE
Spiss L (2003) History of oat breeding in Poland. Biul Inst Hod i Aklim Roślin 7–11
Świerczewski A, Mazaraki M (1993) Hodowla owsa (Oat breeding). In: Mazurek J, Wojcieska U, Mazurek J, Król M (eds) Biologia i agrotechnika owsa (Biology and agrotechnology of oats). IUNG, Pulawy, pp 129–161
Tibco Software Inc. (2017) Statistica 13.1 (data analysis software system) Palo Alto, CA
van de Wouw M, Kik C, van Hintum T, van Treuren R, Visser B (2010) Genetic erosion in crops: concept, research results and challenges. Plant Genet Resour 8:1–15. https://doi.org/10.1017/S1479262109990062
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sowa, S., Paczos-Grzęda, E. A study of crown rust resistance in historical and modern oat cultivars representing 120 years of Polish oat breeding. Euphytica 216, 12 (2020). https://doi.org/10.1007/s10681-019-2545-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-019-2545-8