Abstract
Soil salinity is major constraint for wheat production globally and breeding wheat cultivars for salt tolerance by conventional means is difficult. Therefore, understanding molecular components associated with salt tolerance is needed to facilitate breeding for salt tolerance in wheat. In this investigation, quantitative trait loci (QTL/s) associated with salt tolerance were identified using recombinant inbred lines (RILs) developed from a cross between Kharchia 65 (KH 65) and HD 2009 cultivars. Parents and RILs were evaluated under controlled and sodic stress conditions for 11 morpho-physiological and yield determining traits for two consecutive crop cycles. Simple sequence repeat (SSR) markers were employed for mapping studies. Using composite interval mapping approach, 11 QTLs on 6 chromosomal regions (1B, 2D, 5D, 6A, 6B and 7D) for 7 different traits were identified explaining proportion of the phenotypic variance (PVEs) (2.5–12.8%) under control condition. Three of the QTLs (QCph.iiwbr-2D.1, QCle.iiwbr-6A and QCle.iiwbr-6B) were most consistent in all the environments and explained PVEs (5.1–12.8%) under control condition. Twenty-five QTLs were detected on 7 chromosomal regions (1A, 1B, 2D, 4D, 5D, 6A and 7D) for 10 different traits explaining PVEs (2.6–15.1%) under salt stress. Six of the QTLs namely QSNa+.iiwbr-1B, QSK+.iiwbr-2D, QStn.iiwbr-4D, QSph.iiwbr-2D.1, QSph.iiwbr-6A and QSdth.iiwbr-2D were consistently reproducible in all the environments and explained PVEs ranging from 2.6 to 15.1%. SSR markers namely gwm 261, wmc 112, and cfd 84 were tightly linked with QTLs for K+ content; DTH and DTA; and TN and NE, respectively. Several QTLs contributing towards salt tolerance were present on 2D chromosome. Most of the QTLs linked with salt tolerant traits were inherited from KH 65 signifying the presence of several genes associated with salt tolerance in this cultivar. The information is very useful in marker assisted breeding to enhance salt tolerance in wheat.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Wheat is an important cereal crop which contributes significantly in food and nutritional security across the world. However, wheat production and productivity is suffered by several abiotic stresses i.e. drought, heat, cold and salt stress. Among these abiotic stresses, salt stress affects more than 800 million hectares of agricultural land globally including both saline and sodic soils (FAO 2014). In India, around 7 million hectares of land is salt affected and expected to increase over 16.2 million hectares by 2050 (CSSRI 2015). Salt affected sodic soils have excess of Na+ ion on exchange sites and high concentration of carbonate and bicarbonate anions associated with high pH (Sharma et al. 2004). High salt concentrations in soil affect plant growth and development, resulting in significant grain yield reduction (Munns and Tester 2008).
The mechanism of plant response to salt stress is a complex phenomenon occurring at cellular, tissue and whole plant level. The adverse effect of salt stress is considered mainly due to osmotic stress, ions toxicities and interference with the uptake of mineral nutrients in plants (Mba et al. 2007). Different physiological traits such as accumulation of compatible osmolytes, K+ selectivity and the exclusion of Na+ ion have been reported related to salt tolerance in wheat (Yeo et al. 1990; Munns et al. 2010; Rana et al. 2015). Among compatible osmolytes, proline has been reported to have role in imparting tolerance to salt stress by osmoregulation, maintaining a low NADPH: NADP+ ratio and in scavenging free radical during stress (Szabados and Savouré 2010; Rana et al. 2016). Several reports indicate that salt tolerance is polygenic in nature and significantly influenced by different environmental conditions (Foolad and Jones 1993). Since salt tolerance is complex and selection criteria are inadequate; breeding for salt tolerance by conventional means is difficult. Therefore, understanding molecular components associated with salt tolerance is needed to facilitate breeding for salt tolerance in wheat.
Although some single-gene effects for salt tolerance have been identified in higher plants like kna1 (Dubcovsky et al. 1996) for discrimination of K+ over Na+, (TaNHX1) Na+/H+ antiporter (Rana et al. 2015) and SOS1 (Cuin et al. 2008) in hexaploid wheat, Nax1 and Nax2 (Lindsay et al. 2004; James et al. 2006; Byrt et al. 2007) in dicoccoides, AtNHX1 and RAS1 in Arabidopsis (Apse et al. 1999; Ren et al. 2010) and OsNHX1 and SKC1 in rice (Fukuda et al. 1999; Ren et al. 2005), it is complex and polygenic in nature (Flowers 2004). Previous studies used QTL/marker trait association (MTA) analyses to explore genomic regions underlying salt tolerance related traits under field conditions in wheat (Dubcovsky et al. 1996; Díaz De León et al. 2011; Xu et al. 2012, 2013; Azadi et al. 2015; Oyiga et al. 2016; Shamaya et al. 2017; Asif et al. 2018; Liu et al. 2018). Though, field screening under natural and sodicity stress is important to identify tolerant genotype, soil heterogeneity and inherent spatial variability for salt concentration in field conditions often hamper the true expression of genotype. To avoid this constraint, specially designed microplots having controlled and sodic conditions were used for reliable screening under salt stress.
In this investigation, QTL mapping was done for salt tolerance using recombinant inbred lines (RILs) derived from a cross between the cultivars Kharchia 65 (KH 65) and HD 2009. KH 65 is known internationally for its salt tolerance and played a key role in the development of salt tolerant genotypes in India and elsewhere. However, little is known about the molecular mechanism of salt tolerance in KH 65 (Rana et al. 2016). Therefore, understanding molecular components associated with salt tolerance in KH 65 will help in breeding process for improving salt tolerance in wheat.
Materials and methods
Plant material, crop management and data collection
A set of 114 recombinant inbred lines (RILs) derived from a cross between salt tolerant Kharchia 65 (KH 65) and salt susceptible (HD 2009) wheat cultivars were used in this study. The cultivar KH 65 was developed in 1966 from the cross between Kharchia local and EG 953, while HD 2009 originated in 1975 from a cross between LR 64A and NAI 60. This study was carried out in the specially developed microplots (bins) of the size of 3 × 6 meters across and 1.5 meters deep and having rain cover of transparent sheet during 2015–2016 and 2016–2017 crop seasons at ICAR-Indian Institute of Wheat and Barley Research, Karnal, India. There are 16 microplots in total and 8 of them had controlled (pH 8.0) conditions and 8 with sodic (pH 9.2) conditions (Fig. 1a). Microplots with sodic conditions were developed by adding the required quantity of sodium bicarbonate (NaHCO3) and sodium carbonate (Na2CO3) in soil. Electrical conductivity (EC 1:2) of the soil solution (10 g soil mixed in 20.0 ml distilled water) was measured using electrical conductivity meter (Delux Make, 601) and presented in Fig. 1b. On an average, EC 1:2 was around 0.50 ds/m in microplots with controlled condition and 3.02 ds/m in sodic microplots. The RIL population along with their parents were planted in the third week of November using a randomized complete block design (RCBD) with two replications in both the conditions (control and sodic) in two rows of 75 cm each with row to row distance of 20 cm. The crop was irrigated normally, and fertilizer was applied as per recommended agronomic practices (120 kg N, 60 kg P2O5 and 40 kg K2O per ha) with full dosage of P2O5 and K2O at the time of sowing and N in three split doses. The minimum and maximum mean temperatures were 13.6 and 26.8 °C during 2015–2016 and 12.5 and 26.5 °C during 2016–2017, respectively. The crop was protected from rust and spot blotch diseases by spraying 625 g/ha propiconazole via two growth stages at GS 54 and GS 69 (Zadoks et al. 1974). Several morpho-physiological traits namely days to heading (DTH), days to anthesis (DTA), tiller number (TN), plant height (PH) in cm, number of earheads (NE), length of earhead (LE) in cm, 1000 grain weight (TGW) and grain yield (GY) in grams on row basis were recorded.
a Periodic mean pH maintained under control and stressed condition during 2015–2016 and 2016–2017 in microplot at ICAR-IIWBR, Karnal, India. b Periodic mean performance of soil electric conductivity under control and stressed condition during 2015–2016 and 2016–2017 in microplot at ICAR-IIWBR, Karnal, India
Measurement of ions and proline content (Pro)
Ten days old flag leaf was used for the evaluation of Na+, K+ and proline concentration. 100 mg flag leaf samples were dried for 48 h at 65 °C and digested with 0.5 ml of 0.5 N HNO3 for 2 h at 80 °C as per the method (Munns et al. 2010). Digested samples were centrifuged and diluted 100 times with distilled water. Concentrations of Na+ and K+ ions were measured by flame photometer using standards in the range of 0.25–20 ppm and expressed as milligram per gram dry weight (mg/g DW).
Proline content was extracted from 10 days old flag leaf as per the method (Bates et al. 1973). 50 mg of fresh leaf sample was homogenized in 3% sulphosalicylic acid (5 µl/mg FW), kept on ice for 5 min and centrifuged at 14,000 rpm for 10 min at room temperature and the supernatant was used for assaying proline content. The reaction mixture containing 200 µl glacial acetic acid, 200 µl ninhydrin reagent and 100 µl of supernatant was incubated for 20 min at 90 °C in water bath. The reaction was terminated by transferring the reaction mixture tubes on ice. 1 ml toluene was added in the reaction mixture and vortexed. The upper toluene phase was taken for measurement of proline using absorbance at 520 nm. Pro content was measured using proline as the calibration standard and expressed in microgram per gram fresh weight (μg/g FW).
Statistical analysis
Statistical analyses were accomplished for all the traits under both controlled and stressed conditions. Analysis of variance (ANOVA) was executed using the PROC GLM procedure inbuilt within the SAS v9.3 package (SAS Institute Inc.). Trait-wise broad-sense heritability was calculated for both the conditions separately using the formula \({\text{h}}^{2} = \frac{{\left( {\sigma_{\text{g}}^{2} } \right)}}{{\sigma_{g}^{2} + \frac{{\sigma_{\text{e}}^{2} }}{\text{r}}}}\) (Gitonga et al. 2014); where, \(\sigma_{\text{g}}^{2}\) = genetic variance, \(\sigma_{\text{e}}^{2}\) = environmental variance, r = number of replications. Pearson correlation coefficients were also analyzed using PROC CORR model included within @SAS package.
A multiple linear regression analysis was conducted for determining relative contribution of different parameters to the grain yield (Y) by applying the equation \(y = a + b_{1} x_{1} + b_{2} x_{2} + b_{3} x_{3} + \cdot \cdot \cdot + b_{i} x_{i}\) (Snedecor and Cochran 1980). Where, y is the dependent variable, the x’s are independent variables (measured traits) affecting dependent one, a is the intercept coefficient, and the b’s are the related coefficients of independent variables in predicting the dependent variable. Stepwise regression was used to identify most important variables contributing towards grain yield.
Genotyping, linkage mapping and QTL analysis
Genomic DNA from RILs and parents was extracted by the modified method (Benito et al. 1993). The DNA was quantified using a UV–vis spectrophotometer (NanoDrop, Thermo Scientific Products, USA) and final concentration was adjusted to 50 ng/μl. A total of 550 SSR markers (including barc, cfa, cfd, gwm, wmc) covering whole genome were used for parental polymorphism (Roder et al. 1998; Somers et al. 2004). Of these, 133 SSR markers exhibited polymorphisms between parents and were further employed for QTL analysis.
PCR reaction was performed using Biored Thermocycler (Bio-Rad, USA). The reaction volume was 10 µl, comprising 50 ng template genomic DNA, 1x PCR buffer, 1.25 mM of MgCl2, 200 µM of each deoxyribonucleotide (dNTP), 100 ng of each primer, 0.3 U of Taq DNA polymerase (Bangalore Genei, India). The standardized PCR program included an initial denaturation at 94 °C for 3 min, followed by 36 cycles of 94 °C for 45 s, annealing 50–65 °C (depending on the primer) for 30 s, and 72 °C for 45 s, and a final extension for 7 min at 72 °C before cooling to 4 °C. PCR products were separated by electrophoresis on 3.5% agarose at 95 V for 3 h, using TAE buffer. PCR amplified products were visualized and imaged using the gel documentation system (Bio-Rad, USA) for further scoring.
The bands generated by polymorphic SSR primers were scored by giving code `2’ for KH 65 type and `0’ for HD 2009 type band. Allelic segregation of the two genotypic classes (KH 65 and HD 2009) at each of the marker loci was analyzed using the chi square (χ2) test for segregation distortion from the expected 1:1 ratio in the RILs. The markers which exhibited 1:1 segregation pattern were only further used for linkage map construction. The linkage map was constructed from genotypic data of RILs using IciMapping v4.1 software with LOD threshold 3.0 between adjacent markers (Li et al. 2007). The QTL analysis was also performed using IciMapping v4.1 software by combined analysis of adjusted means of the phenotypic trait value and genotyping data via inclusive composite interval mapping (ICIM) algorithm for additive gene effect with function inbuilt in the software. For QTL analysis, LOD threshold was 2.5 and walking speed was set to 1 cM along the chromosomes for evaluation of significant QTL (Ribaut et al. 1997; Tuberosa et al. 2002; Srinivasa et al. 2014). Stability of QTL was acknowledged using LOD of QTL × environmental interaction (LODQEI) < 2.5. Linkage map was redrawn and QTL location was integrated within linkage map using map chart v2.32 (Voorrips 2002). Standard procedure of QTL nomenclature was followed using instructions (http://wheat.pw.usda.gov/ggpages/wgc/98/); of which, each QTL consisted of the letter Q for QTL, followed by an abbreviated trait name and regime, the institution designation (IIWBR) and the identity of the chromosome where QTL located.
Results
Phenotypic evaluation of RILs
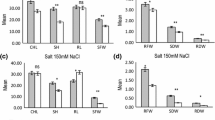
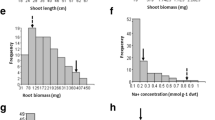
Though, both KH 65 (45.13 g) and HD 2009 (40.25 g) exhibited no significant difference in grain yield under controlled conditions (Supplementary Fig S1k), KH 65 showed significantly higher grain yield (40.25 g) as compared to HD 2009 (24.88 g) under sodic conditions. KH 65 also showed significantly lower reduction in TGW (6.09%) as compared to HD 2009 (20.78%) indicating its salt tolerant nature. Similarly, there was no significant difference in Na+ and K+ content in flag leaf of both KH 65 and HD 2009 under control condition, however, KH 65 exhibited a lower Na+ and higher K+ content as compared to HD 2009 under sodic condition (Fig. 2a–k). RILs exhibited a wider range of different traits as GY (10.25–44.50 g), Na+ content (2.10–10.10 mg/g DW), Pro (0.50–3.43 μg/g FW), K+ content (9.23–28.38 mg/g DW), TN (2.47–7.20), PH (75.50–112.19 cm), NE (2.05–6.13), LE (8.65–14.83 cm) and TGW (10.25–44.50 g) under sodic stress (Fig. 2a–k). The range in all traits exhibited the presence of transgressive segregants in both the directions in the RILs. Continuous distribution of traits under both control and sodic conditions showed polygenic nature of the traits (Supplementary Fig S1a-k and Fig. 2a–k). Significant variations (p < 0.001) were also observed for all the traits for year, genotype and year × genotype under both the conditions (Table 1).
Mean performance based frequency distribution of different traits along with their mean values, standard deviation (std), coefficient of variation (cv) and heritability (h2) over two years (2015–2016 and 2016–2017) under sodic condition in the RIL population derived from a cross KH65 × HD2009. P1 Parent 1 (KH 65), P2 Parent 2 (HD 2009), Na+ Sodium content, K+ Potassium content, Pro Proline content, TN Tiller number per plant, PH Plant height, NE Number of earhead, LE Length of earhead, DTH Days to heading, DTA Days to anthesis, TGW Thousand grain weight, GY Grain yield
The coefficient of variation (CV) was higher for most of the traits except LE, DTH and TGW in sodic condition as compared to control condition. High heritability (h2) estimates were observed for GY (0.87), TGW (0.81), TN (0.91), PH (0.93), NE (0.87), LE (0.80), DTH (0.99), DTA (0.99) Na+ (0.92), K+ (0.91) and Pro (0.67) under control conditions (Supplementary Fig S1a-k) while comparatively lower estimates of h2 were exhibited for GY (0.66), TGW (0.68), TN (0.71), PH (0.85), NE (0.70), LE (0.70), DTH (0.97), DTA (0.99) Na+ (0.67), K+ (0.53) and Pro (0.69) under sodic condition (Fig. 2a–k).
There were wide ranges of phenotypic correlations among various parameters under control and sodic conditions. There were significant positive correlations between GY and TGW (p < 0.001), TN (p < 0.01), NE (p < 0.01), LE (p < 0.01), PH (p < 0.01) and Pro (p < 0.001) under control conditions. Similarly under sodic stress conditions, significant positive correlations were exhibited between GY and TGW (p < 0.001), TN (p < 0.01), NE (p < 0.001), LE (p < 0.01), PH (p < 0.01) and Pro (p < 0.001). Though there was no correlation between GY and Na+ and K+ content under control condition, significantly negative correlation was observed between GY and Na+ content (p < 0.01) and positive between GY and K+ content (p < 0.01) under sodic condition (Table 2).
Under control condition, significant and positive correlations were observed between Na+ content and DTH (0.27) and DTA (0.24); K+ content with proline content (0.30) and TGW (0.22); Pro with PH (0.39), LE (0.21), TGW (0.39) and GY (0.34); TN with NE (0.75), DTA (0.19), TGW (0.23) and GY (0.36); PH with LE (0.61), TGW (0.59) and GY (0.30); NE with TGW (0.59) and GY (0.26); LE with TGW (0.29) and GY (0.36); DTH with DTA (0.97); and TGW with GY (0.44). Whereas significant and negative correlations were observed between Na+ content and Pro (− 0.19), PH (− 0.20) and TGW (− 0.27); K+ content with DTH (− 0.19); PH with DTH (− 0.34) and DTA (− 0.31); DTA with TGW (− 0.39); and DTA with TGW (− 0.35) and presented in Table 2.
Under sodic condition, significant and positive correlations were observed between K+ content and TN (0.32), PH (0.43), NE (0.33), LE (0.32) and GY (0.49); between Pro and TN (0.41) and NE (0.43); between TN and PH (0.42) and LE (0.36); between PH and NE (0.39); and between NE and LE (0.35); whereas, significant and negative correlation were observed between Na+ content and K+ content (− 0.34), TN (− 0.26), NE (− 0.26), LE (− 0.38) and GY (− 0.34); between K+ content and DTA (− 0.21); between TN and DTH (− 0.24) and DTA (− 0.26); and between NE with DTH (− 0.23) and DTA (− 0.26) presented in Table 2.
Linkage map
Genetic map was constructed using 133 polymorphic SSR markers of which 49 SSRs mapped on D genome chromosomes, 44 SSRs on A genome chromosomes and 40 SSRs on B genome chromosomes. Chromosome 7A harbored highest markers coverage of 11 SSRs, and the chromosome 3A had lowest marker coverage with 2 SSRs. Whole linkage map covered a genetic length of 4725.16 cM with an average inter marker distance of 35.53 cM (Fig. 3).
Single marker analysis (SMA)
SMA was conducted using linear regression model for detection of significant associations between SSRs and each of the trait. A total of five SSR markers (wmc 601, gwm 437, gwm 261, wmc 112 and cfd 84) were identified associated with seven different traits under both control and sodic conditions. wmc 601 representing chromosome 2D showed tight linkage (0.4 cM) with the QTL (QCpro.iiwbr-2D) for Pro (p < 0.001) under control condition. The other marker gwm 437 on chromosome 7D was significantly associated (p < 0.001) to the QTL (QCgy.iiwbr-7D) for grain yield within 0.4 cM (Table 3).
Three SSRs (gwm 261, wmc 112 and cfd 84) were associated with five traits under sodic condition, of which three traits were associated with QTLs present on 2D indicating important role of the chromosome in salt tolerance. gwm 261 on chromosome 2D was linked significantly (p < 0.001) within 0.1 cM to K+ content (QSK+.iiwbr-2D) in both the years 2015–2016, 2016–2017 separately and also identified in pooled condition. The marker wmc 112 on linkage group 2D was significantly linked with DTH (QSdth.iiwbr-2D.1) within 0.2 cM in 2015–2016, and 0.8 cM for QTL (QSdth.iiwbr-2D.2) in 2016–2017 and pooled condition and with DTA within 0.2 cM in all three conditions. SSR marker cfd 84 on chromosome 4D was associated significantly (p < 0.001) to QTL (QStn.iiwbr-4D) for tiller number in all three environments separated to 4.5 cM, 0.5 cM and 4.5 cM respectively and for QTL (QSne.iiwbr-4D) NE in two environments 2016–2017 and pooled condition laid at a distance of 2.5 cM and 3.5 cM, respectively (Table 3).
Composite interval mapping (CIM)
QTLs for Na+ and K+ content
One QTL under control (Table 4) and seven QTLs under sodic condition (Table 5) were identified associated with Na+ content using CIM. Under control condition, one QTL (QCNa+.iiwbr-7D) was identified for Na+ content located on 7D chromosome in 2015–2016 and pooled mean flanked by SSR markers cfa 20140 and gdm 67 with LOD 3.1 and 2.9, respectively. The QTL explained phenotypic variation of 3.1% in 2015–2016 and 6.6% in pooled condition. The presence of HD 2009 alleles for QCNa+.iiwbr-7D resulted into increase in Na+ content under control condition (Supplementary Fig S2a and Table 4).
Seven QTLs (QSNa+.iiwbr-1B, QSNa+.iiwbr-2D, QSNa+.iiwbr-5D.1, QSNa+.iiwbr-5D.2, QSNa+.iiwbr-5D.3, QSNa+.iiwbr-6A and QSNa+.iiwbr-7D) representing chromosome 1B, 2D, 5D 6A and 7D were detected for Na+ content with PVEs ranging from 2.6 to 12.8% under sodic condition (Table 5). QSNa+.iiwbr-1B separated by SSR markers (barc 137 and wmc 144) was most consistent and stable across the environments. QSNa+.iiwbr-2D flanked by wmc 453 and wmc 601was identified on chromosome 2D in 2016–2017 and pooled condition with LOD scores 3.2 and 2.6 and explained phenotypic variation by 6.0% and 6.2%, respectively. QSNa+.iiwbr-5D.1 flanked by wmc 97-wmc 405 explained phenotypic variation 12.8% and 12.4% with LOD scores 3.5 and 3.3 during 2015–2016 and pooled conditions, respectively. QSNa+.iiwbr-5D.2 was identified only in 2016–2017 separated by cfd 18 and wmc 97 with LOD scores 2.8 and explained 7.0% phenotypic variation. QSNa+.iiwbr-5D.3 (flanked by wmc 405-cfa 2141) was identified in 2016–2017 with LOD scores 3.3 and PVE 6.8%. QSNa+.iiwbr-6A (flanked by wmc 256-gwm 169) and QSNa+.iiwbr-7D (flanked by gwm 111-gwm 130) were identified in 2016–2017 with LOD scores 2.7 and 3.0 and PVEs 8.9% and 6.0%, respectively. KH 65 alleles for the QTLs on chromosomes 1B, 5D and 6A and HD 2009 allele on chromosome 2D were associated with lower Na+ content while HD 2009 allele for the QTL on 7D chromosome was associated with higher Na+ content (Supplementary Fig S3a-e and Table 5).
One QTL QCK+.iiwbr-1B was identified for K+ content under control condition flanked by SSR markers wmc 619 and gwm 264 on 1B and explained phenotypic variation 2.5% and 3.4% during 2016–2017 and pooled condition with LOD 2.5 and 2.6, respectively. The allele for QCK+.iiwbr-1B with positive effect was inherited from tolerant parent KH 65 (Supplementary Fig S2b and Table 4). Similarly, one QTL (QSK+.iiwbr-2D) for K+ content detected under sodic condition flanked by SSR markers gwm 349-gwm 261was identified in all three conditions and explained 10.2%, 12.6% and 12.2% phenotypic variation with LOD scores 2.7, 3.2 and 3.1 in 2015–2016, 2016–2017 and pooled mean, respectively. SSR marker gwm 261 was the nearest marker tightly (0.1 cM distance) linked with the QTL and associated significantly (p < 0.001) with K+ content. The positive allele of the QTL was inherited from the tolerant parent KH 65 (Supplementary Fig S3f and Table 5).
QTLs for proline content (Pro)
One QTL (QCpro.iiwbr-2D) for proline content flanked by SSR markers wmc 601 and gwm 349 was identified only in 2015–2016. QCpro.iiwbr-2D was separated by 0.4 cM from wmc 601 with LOD score 2.6 and PVE 10.0% under control condition. Positive allele was transmitted by tolerant parent KH 65 associated with increased proline content (Supplementary Fig S2c and Table 4). However, under sodic condition, two QTLs QSpro.iiwbr-2D.1 (flanked by gwm 122-wmc 453) and QSpro.iiwbr-2D.2 (flanked by wmc 453-wmc 601) were identified for proline content only in 2016–2017 with LOD scores 2.5 and 3.1 and PVEs 5.5 and 6.2%, respectively. KH 65 alleles for both of the QTLs were associated with increased proline content (Supplementary Fig S3g and Table 5).
QTLs for tiller number (TN) and number of earheads (NE)
There was no QTL detected for TN and NE under controlled condition. However, three QTLs for TN and one QTL for NE associated with sodic stress conditions were identified. One QTL (QStn.iiwbr-2D) for TN on chromosome 2D was identified only in pooled mean flanked by wmc 453-wmc 601 and explained 5.6% phenotypic variation with LOD score 3.3.The presence of KH 65 alleles at the QTL was associated with higher TN on chromosomes 2D (Supplementary Fig S3h and Table 5).
QTLs for TN (QStn.iiwbr-4D) and NE (QSne.iiwbr-4D) were identified at approximately the same positions on chromosome 4D flanked by gwm 193-cfd 84 ad tightly linked to the SSR marker cfd 84 under sodic conditions (Supplementary Fig S3i and Table 5). QStn.iiwbr-4D were identified in 2015–2016, 2016–2017 and pooled mean with LOD scores 3.4, 4.0 and 4.1 and explaining 12.7, 15.1 and 14.9% phenotypic variations, respectively. QSne.iiwbr-4D was identified in 2016–2017 and pooled means with LOD scores 3.9 and 3.6, explaining 14.5 and 13.6% phenotypic variation respectively. The presence of KH 65 alleles at the QTL were associated with higher TN and NE on chromosomes 4D. Other QTL for TN (QStn.iiwbr-6A) on chromosome 6A was identified in 2015–2016 and pooled mean flanked by wmc 256-gwm 169 explaining 8.2% and 9.8% phenotypic variation with LOD scores 3.4 and 3.2 and respectively, significantly (p < 0.01) associated with SSR marker gwm 169 (Supplementary Fig S3j and Table 5). The presence of HD 2009 alleles at this locus was associated with decrease in TN on chromosome 6A.
QTLs for length of earhead (LE)
There was no QTL for LE under sodic condition. However, three QTL (QCle.iiwbr-5D, QCle.iiwbr-6A and QCle.iiwbr-6B) for LE were identified on 5D, 6A and 6B under control condition. QTL for LE (QCle.iiwbr-5D) identified in 2015–2016 and pooled mean flanked by cfa 2141-cfd 29 with LOD scores 4.1 and 3.4 explaining phenotypic variation 12.2% and 11.4% respectively (Supplementary Fig S2f and Table 4). QCle.iiwbr-6A identified in 2015–2016, 2016–2017 and pooled mean flanked by gwm 169-gwm 132 with LOD scores 3.2, 2.7 and 3.1 explained phenotypic variation 10.1%, 11.4% and 12.0% respectively (Supplementary Fig S2g and Table 4). QCle.iiwbr-6B identified in 2015–2016, 2016–2017 and pooled mean flanked by gwm 219-gwm 132 with LOD scores 3.0, 3.7 and 3.7 explaining phenotypic variation 10.7%, 12.2% and 12.8%, respectively (Supplementary Fig S2h and Table 4). The alleles from HD 2009 were associated with lower LE for the QTL on chromosomes 5D, 6A and 6B.
QTLs for plant height (PH)
QTLs for PH were identified at approximately the same positions under both control and sodic conditions on chromosome 2D (near to the SSR marker wmc 601) and 6A (near to the SSR marker gwm 169). QCph.iiwbr-2D.1 identified in 2015–2016, 2016–2017 and pooled mean flanked by wmc 601-gwm 349 with LOD score 2.9, 3.4, 3.3 and explaining PVEs 9.5%, 4.9% and 5.1% respectively and QCph.iiwbr-2D.2 identified in 2016–2017 and pooled mean flanked by wmc 453-wmc 601with LOD score 3.4, 3.3 and explaining phenotypic variation 5.3% and 6.5% respectively, under control condition (Supplementary Fig S2d and Table 4). QCph.iiwbr-6A identified in 2016–2017 and pooled mean flanked by gwm 169-gwm 132 with LOD score 3.3, 3.0 and explaining phenotypic variation 12.0% and 10.8% respectively; under control condition (Supplementary Fig S2e and Table 4).
QSph.iiwbr-2D.1 identified in 2015–2016, 2016–2017 and pooled mean flanked by wmc 601-gwm 349 with LOD score 3.8, 4.6, 4.7 and explaining phenotypic variation 11.4%, 4.8% and 5.4% respectively, likewise, QSph.iiwbr-2D.2 identified in 2016–2017 and pooled mean flanked by wmc 453-wmc 601 with LOD score 3.6, 4.2 and explaining phenotypic variation 4.9% and 5.8% respectively under sodic condition (Supplementary Fig S3k and Table 5). QSph.iiwbr-6A identified in 2015–2016, 2016–2017 and pooled mean flanked by gwm 169-gwm 132 with LOD score 3.8, 2.6, 3.9 and explaining phenotypic variation 13.1%, 9.8% and 11.9% respectively; under sodic condition (Supplementary Fig S3l and Table 5). Both the QTLs on 2D had alleles from KH 65 associated with higher PH while alleles for the QTLs on chromosomes 6A were derived from HD 2009 and associated lower PH under both control and sodic conditions.
QTLs for days to heading (DTH) and days to anthesis (DTA)
Two QTLs (QSdth.iiwbr-2D.1 and QSdth.iiwbr-2D.2) for DTH and one QTL (QSdta.iiwbr-2D) for DTA were associated with sodic stress. QTL for DTH (QSdth.iiwbr-2D.1) and DTA (QSdta.iiwbr-2D) on chromosome 2D are separated by 0.2 cM from SSR marker wmc 112. QTL for DTH (QSdth.iiwbr-2D.1) was identified in 2015–2016 with LOD (2.8) and PVE (10.5%). However, QSdth.iiwbr-2D.2 was identified in 2016–2017 and pooled mean with LOD 2.6 and 2.7 and PVEs 9.9% and 10.2%, respectively. Likewise, QTL for DTA (QSdta.iiwbr-2D) was identified in both the years as well as pooled mean with overall average LOD (2.6) and PVEs (10.1%, 9.9% and 10.1%), respectively (Supplementary Fig S3m and Table 5). For both traits, positive alleles on chromosome 2D were inherited from KH 65.
QTLs for thousand grain weight (TGW)
QTLs for TGW were identified at approximately the same positions under control (QCtgw.iiwbr-2D) and sodic conditions (QStgw.iiwbr-2D.2) on chromosome 2D flanked by wmc 453-wmc 601 (close to the SSR marker wmc 601). QCtgw.iiwbr-2D was found in 2016–2017 (LOD score 2.9, phenotypic variation 10.6%) and pooled mean (LOD score 2.6, phenotypic variation 9.6%) under control condition (Supplementary Fig S2i and Table 4). Likewise, two QTLs for TGW identified under sodic condition, QStgw.iiwbr-2D.1 was detected in 2015–2016 and pooled mean flanked by wmc 601-gwm 349 having LOD score 2.8, 3.3 and explaining PVEs 12.3%, and 11.8%, respectively. While QStgw.iiwbr-2D.2 identified only in 2016–2017 having LOD score 2.6 and explained phenotypic variation 8.9% (Supplementary Fig S3n and Table 5). KH 65 allele was associated with higher TGW under sodic condition.
QTLs for grain yield (GY)
One QTL (QCgy.iiwbr-7D) on chromosome 7D flanked by gwm 437-wmc 405 was associated with GY with LOD score 2.6, explaining 11.8% phenotypic variation under control condition. SSR marker gwm 437 was the nearest marker with the distance 0.4 cM to the QTL. The presence of HD 2009 allele for QCgy.iiwbr-7D was associated with higher GY (Supplementary Fig S2j and Table 4). A total of two QTLs for GY were detected under sodic condition, QTL (QSgy.iiwbr-1A) located at 1A was identified for GY in 2016–2017 (LOD = 2.6, PVE = 5.9%), and pooled mean (LOD = 3.3, PVE = 6.2%) and flanked by wmc 24-barc 17. SSR marker barc 17 was the nearest marker to the QTL at a distance of 28.3 cM during 2016–2017 and 27.3 cm in pooled condition (Supplementary Fig S3o and Table 5). Another QTL for GY located at 2D (QSgy.iiwbr-2D) was identified only in pooled mean and flanked by wmc 453-wmc 601 with LOD scores 3.6 explaining 4.2% phenotypic variation (Supplementary Fig S3p and Table 5). wmc 601 was the nearest marker associated to the QTL. For both the QTLs, QSgy.iiwbr-1A and QSgy.iiwbr-2D alleles were derived from KH 65 associated with higher GY. QTL for GY located at 6A (QSgy.iiwbr-6A) in 2016–2017 (LOD = 2.7, phenotypic variation = 5.3%) and pooled mean (LOD = 3.7, phenotypic variation = 5.5%) was flanked by wmc 256-gwm 169 marker was derived from HD 2009 allele and associated with lower yield (Supplementary Fig. S3q and Table 5).
Discussion
The analysis of variance indicated large diversity in parental lines KH 65 and HD 2009 in terms of traits taken under the study and hence the RILs developed using these lines are suitable for identification of QTLs associated with salt tolerance. There were significant genotype x environmental interactions for each trait and thus making it difficult to select desirable traits under salt stress condition. Similar observations have been made by several other studies under abiotic stresses (Blum 1988; Reynolds et al. 1994; Paliwal et al. 2012; Tiwari et al. 2013). There was continuous distribution of the traits in the RIL population indicating that the traits were quantitatively inherited (Supplementary Fig S1a-k and Fig. 2a–k) and also exhibited transgressive segregation towards both directions. This suggested that both parental lines carried few different alleles contributing significantly under sodic condition. This also been reported in different populations by several workers (Amin et al. 2011; Oyiga et al. 2016; Hussain et al. 2017; Asif et al. 2018). The heritability estimates was lower under sodic condition for all the traits as compared to control condition because of higher G x E interaction under more stressful environment. Moderate to high heritability was also reported for salt conditions (Akbarpour et al. 2015; Oyiga et al. 2016). This exhibited that selection may be effective for the improvement of salt tolerance in wheat (Farshadfar et al. 2008).
Since salt affected sodic soils have excess of Na+ ion on exchange sites, plants growing under sodic conditions accumulate more of ions like Na+ resulting in ionic imbalance as well as toxic effects of the accumulated ions (Tavakkoli et al. 2011). Na+ content was negatively correlated with all traits except DTH and DTA. (Khan et al. 2009; Genc et al. 2010) also reported negative correlation of Na+ content with GY under salinity condition. Accumulation of compatible osmolytes and K+ ions over Na+ ion can have adaptive mechanism under salt stress in wheat (Yeo et al. 1990; Munns et al. 2010; Rana et al. 2015). In this investigation significant and positive correlations were observed between GY and K+ content, proline, TN, PH, NE, LE and TGW under sodic stress. The existence of positive correlation between GY and its associated traits have also been previously reported (del Pozo et al. 2016; Ayed-Slama et al. 2018). Therefore, combinations of some of the traits can be used in identifying salt tolerant genotypes in wheat.
Since low Na+ concentration and high K+ content in leaf have been recognized as an important cellular mechanism that supports plant adaptation in salt stress (Munns and James 2003; El-Hendawy et al. 2017; Siddiqui et al. 2017; Mbarki et al. 2018), QTLs analysis was conducted to identify loci associated with Na+ and K+ contents in flag leaf under sodic stress conditions. Although, some single-gene effects for salt tolerance have been identified in higher plants like kna1 (Dubcovsky et al. 1996) and (TaNHX1) Na+/H+ antiporter (Rana et al. 2015) in wheat, AtNHX1 and RAS1 in Arabidopsis (Apse et al. 1999; Ren et al. 2010) and OsNHX1, SKC1 in rice (Fukuda et al. 1999; Ren et al. 2005), the study showed it is polygenic in nature. In this investigation, Six of the loci (QSNa+.iiwbr-1B, QSNa+.iiwbr-5D.1, QSNa+.iiwbr-5D.2, QSNa+.iiwbr-5D.3 and QSNa+.iiwbr-6A) derived from KH 65 and one locus (QSNa+.iiwbr-2D) derived from HD 2009 were associated significantly with lower Na+ content contributing PVEs ranging from 2.6 to 12.8%. The QTL (QSNa+.iiwbr-7D) derived from HD 2009 was associated with higher accumulation of Na+ content in flag leaf. Several other reports also indicated the presence of QTL on chromosome 2A (Oyiga et al. 2016, 2018), 2B (Xu et al. 2012), 5D (Spielmeyer et al. 2007), 6A (Asif et al. 2018), 7A (Xu et al. 2012; Masoudi et al. 2015; Asif et al. 2018) and 7D (Masoudi et al. 2015) associated with salt tolerance. The QSNa+.iiwbr-5D.1 exhibited phenotypic variation (12.4 to 12.8%) for lower Na+ content under salt stress and showed strong association (p < 0.001) with SSR marker wmc 97 inherited from tolerant parent KH 65.
A major QTL QSK+.iiwbr-2D for accumulation of K+ content was identified on 2D chromosome contributing 10.2–12.6% phenotypic variation under salt stress conditions. SSR marker gwm 261 showed tight linkage (0.1 cM) with the QTL and inherited from tolerant parent KH 65. The QTL was identified in all environments under sodic condition. SSR marker gwm 261 was present at 23 cM position in consensus map (Somers et al. 2004). Earlier, (Amin and Diab 2013) also reported QTL for K+ content on chromosome 2D. Some of the previous studies concluded that maintenance of cellular K+ concentration above a certain threshold value in the presence of excess external Na+ is critical for growth and salt tolerance (Zhu et al. 1998) and a positive relationship between K+ and salt tolerance was reported (Bağci et al. 2007; Chen et al. 2007; Cuin et al. 2008). Since, The QTL QSK+.iiwbr-2D is effective under all conditions, it has utility for using as selection criteria in salt tolerance breeding programmes.
One novel QTL QCpro.iiwbr-2D for proline content contributing 10.0% PVE was identified on 2D chromosome under controlled condition in a single environment 2015–2016. SSR marker wmc 601 was found tightly linked (0.4 cM) with the QTL using both SMA and CIM procedures. Two novel QTLs namely QSpro.iiwbr-2D.1 and QSpro.iiwbr-2D.2 were identified under stressed condition inherited from tolerant parent KH 65. Moreover, better accumulation of proline reduces the deleterious effects on plant growth, development and stomatal conduction under stress condition (Sairam et al. 2005; Munns and Tester 2008; Rana et al. 2015).
Three QTLs for TN (QStn.iiwbr-2D, QStn.iiwbr-4D and QStn.iiwbr-6A) and one QTL (QSne.iiwbr-4D) for NE were identified in salt stress conditions. The QTL QStn.iiwbr-2D on chromosome 2D was associated with SSR marker wmc 453. QTL for TN (QStn.iiwbr-4D) and NE (QSne.iiwbr-4D) were co-located on chromosome 4D at 184 cM and 186 cM position, respectively in the linkage map and found tightly linked to the SSR marker cfd 84 (0.5 to 4.5 cM for TN and 2.5 to 3.5 cM for NE under different conditions). The presence of KH 65 alleles on chromosomes 2D and 4D were associated with increased TN and NE under salt tolerance. QTLs for TN and NE on chromosome 4D were robust and highly reproducible. These two QTLs explained 12.7–15.1% phenotypic variation. QTL for TN on 2D and 4D and NE on 4D chromosomes also identified (Díaz De León et al. 2011). QTL for TN on chromosome 4B that was linked to the SSR marker gwm 6 was previously identified (Genc et al. 2010). While, QTL for NE on chromosome 1A, 2A, 2D and 5B were reported earlier (Heidari et al. 2011; Amin and Diab 2013). QStn.iiwbr-6A allele from HD 2009 was associated with decrease in TN under salt stress condition. There was no report available of QTL for TN on 6A. Although, kna1 gene located on 4D chromosome was identified for exclusion of Na+ content and discrimination of K+ content in root under salt stress (Dubcovsky et al. 1996), in this investigation QTLs for TN and NE were present on 4D.
Three QTLs (QCle.iiwbr-5D, QCle.iiwbr-6A and QCle.iiwbr-6B) for LE were identified on 5D, 6A and 6B chromosomes under control condition. None of the locus for LE was detected under salt stress condition. QTLs for LE harbored alleles from HD 2009 genotype associated with increase in LE under control conditions were novel. QTL for LE on chromosome 1A that was linked to the RFLP marker Xabc156 was reported (Kumar et al. 2007).
QTLs for PH were identified approximately at the same positions under both control and sodic conditions on chromosome 2D and 6A. The KH 65 alleles associated with the QTL for PH on chromosomes 2D increased PH while HD 2009 alleles associated with the QTL for PH on chromosomes 6A were associated with decrease in PH under both control and sodic condition. In earlier reports, QTLs were identified for PH located on 3B chromosome under salt stress (Zhang et al. 2011) and 5B chromosome under control condition (Ghaedrahmati et al. 2014). In this investigation, QTLs for PH (QSph.iiwbr-2D.1 and QSph.iiwbr-2D.2), TGW (QStgw.iiwbr-2D) and GY (QSgy.iiwbr-2D) were identified on 2D chromosome under sodic condition and SSR marker wmc 601 was found linked with all the QTLs (Table 6). Further, KH 65 type alleles of these QTLs for PH, TGW and GY were effective in increasing their value under stress condition. Some other reports also indicated the presence of QTLs for these traits located at 2D Chromosome (Huang et al. 2004, 2006; Cuthbert et al. 2008).
Two QTLs (QSdth.iiwbr-2D.1 and QSdth.iiwbr-2D.2) for DTH and one QTL (QSdta.iiwbr-2D) for DTA, were found tightly linked to the SSR marker wmc 112. KH 65 alleles for both DTH and DTA were found associated with decrease in DTH and DTA under salt stress condition. Earlier reports showed the presence of QTLs for DTH and DTA on 2D chromosome under control condition (Börner et al. 2002; Huang et al. 2003; Kumar et al. 2007) and under salt stress condition (Díaz De León et al. 2011). Photoperiodic gene (Ppd1) which has strong intra-allelic interaction, photoperiodic response or pleiotropic effect on number of moropho-physiological and yield related traits also present on 2D chromosome (Mclntosh 1998; Börner et al. 2002; Kumar et al. 2007).
Two QTLs (QStgw.iiwbr-2D.1 and QStgw.iiwbr-2D.2) explaining PVEs 12.3 and 8.9% respectively under salt stress condition and one QTL (QCtgw.iiwbr-2D) explaining PVE 10.6% under control condition were detected for TGW and found associated with SSR marker wmc 601. (Heidari et al. 2011; Díaz De León et al. 2011; Masoudi et al. 2015) also reported QTLs on 2D chromosome associated with TGW under salt stress condition. However, (Huang et al. 2004, 2006; Cuthbert et al. 2008) reported the association between TGW and SSR marker wmc 601 under control condition only.
One QTL (QCgy.iiwbr-7D) under control condition and 3 QTLs (QSgy.iiwbr-1A, QSgy.iiwbr-2D and QSgy.iiwbr-6A) under salt stress condition were identified for GY. QCgy.iiwbr-7D was identified on 7D chromosome contributing 11.8% PVE under control condition only in one environment (2016–2017) and inherited from HD 2009 was novel. While, QSgy.iiwbr-1A, QSgy.iiwbr-2D inherited from the KH 65 were found associated with increased yield. QSgy.iiwbr-6A harbored alleles from HD 2009 contributing PVEs ranging 4.2–6.2% was associated with lower yield under salt stress condition. Earlier reports indicated the presence of QTLs for salt tolerance on chromosome 1A (Huang et al. 2004; Cuthbert et al. 2008; Azadi et al. 2015), 2D (Kumar et al. 2007; Kuchel et al. 2007; Azadi et al. 2015) and 6A (Heidari et al. 2011). Most of the QTLs identified in this investigation under sodic stress represented chromosomes having QTLs identified by many workers under salinity stress mentioned in the review paper by Mujeeb-Kazi et al. 2019). This is possible because of some common features between the two conditions as reviewed by Rengasamy 2010.
Conclusions
In this present investigation, 25 QTLs were detected on 7 chromosomal regions (1A, 1B, 2D, 4D, 5D, 6A and 7D) for 10 different traits explaining PVEs (2.6–15.1%) under salt stress. Among these QTLs, 6 QTLS as QSNa+.iiwbr-1B, QSK+.iiwbr-2D, QStn.iiwbr-4D, QSph.iiwbr-2D.1, QSph.iiwbr-6A and QSdth.iiwbr-2D were consistently reproducible in all environments and explained PVEs 2.6–2.8%, 10.2–12.6%, 12.7–15.1%, 4.8–11.4%, 9.8–13.1% and 9.9–10.1%, respectively. SSR markers namely gwm 261, wmc 112, and cfd 84 were tightly linked with QTLs for K content; DTH and DTA; and TN and NE, respectively. Many of the QTLs linked with salt tolerant traits were identified on 2D chromosome indicating its role in imparting salt tolerance. The information is very useful in marker assisted breeding to enhance salt tolerance in wheat.
References
Akbarpour OA, Dehghani H, Rousta MJ (2015) Evaluation of salt stress of Iranian wheat germplasm under field conditions. Crop Pasture Sci 66:770–781
Amin AY, Diab AA (2013) QTL mapping of wheat (Triticum aestivum L.) in responce to salt stress. Int J Bio-Technol Res 3:47–60
Amin A, Eslam MH, Seyed AM et al (2011) Screening of recombinant inbred lines for salinity tolerance in bread wheat (Triticum aestivum L.). Afr J Biotechnol 10:12875–12881. https://doi.org/10.5897/AJB11.1729
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285:1256–1258
Asif MA, Schilling RK, Tilbrook J et al (2018) Mapping of novel salt tolerance QTL in an Excalibur × Kukri doubled haploid wheat population. Theor Appl Genet 131:2179–2196. https://doi.org/10.1007/s00122-018-3146-y
Ayed-Slama O, Bouhaouel I, Chamekh Z et al (2018) Genetic variation of salt-stressed durum wheat (Triticum turgidum subsp. durum Desf.) genotypes under field conditions and gynogenetic capacity. J Genet Eng Biotechnol 16:161–167. https://doi.org/10.1016/j.jgeb.2017.11.004
Azadi A, Mardi M, Hervan EM et al (2015) QTL mapping of yield and yield components under normal and salt-stress conditions in bread wheat (Triticum aestivum L.). Plant Mol Biol Rep 33:102–120. https://doi.org/10.1007/s11105-014-0726-0
Bağci SA, Ekiz H, Yilmaz A (2007) Salt tolerance of sixteen wheat genotypes during seedling growth. Turk J Agric For 31:363–372
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207
Benito C, Figueiras AM, Zaragoza C et al (1993) Rapid identification of Triticeae genotypes from single seeds using the polymerase chain reaction. Plant Mol Biol 21:181–183
Blum A (1988) Plant breeding for stress environments. CRC Press, Boca Raton
Börner A, Schumann E, Fürste A et al (2002) Mapping of quantitative trait loci determining agronomic important characters in hexaploid wheat (Triticum aestivum L.). Theor Appl Genet 105:921–936
Byrt CS, Platten JD, Spielmeyer W et al (2007) HKT1;5-like cation transporters linked to Na+ exclusion loci in wheat, Nax2 and Kna1 [OA]. Plant Physiol 143:1918–1928. https://doi.org/10.1104/pp.106.093476
Chen Z, Pottosin II, Cuin TA et al (2007) Root plasma membrane transporters controlling K+/Na+ homeostasis in salt-stressed barley. Plant Physiol 145:1714–1725
CSSRI (2015) CSSRI vision 2050. Central soil salinity research institute, Karnal, Haryana, India. http://www.cssri.org. Accessed 28 Oct 2018
Cuin TA, Betts SA, Chalmandrier R, Shabala S (2008) A root’s ability to retain K+ correlates with salt tolerance in wheat. J Exp Bot 59:2697–2706
Cuthbert JL, Somers DJ, Brûlé-Babel AL et al (2008) Molecular mapping of quantitative trait loci for yield and yield components in spring wheat (Triticum aestivum L.). Theor Appl Genet 117:595–608. https://doi.org/10.1007/s00122-008-0804-5
del Pozo A, Yáñez A, Matus IA et al (2016) Physiological traits associated with wheat yield potential and performance under water-stress in a mediterranean environment. Front Plant Sci 7:987
Díaz De León JL, Escoppinichi R, Geraldo N et al (2011) Quantitative trait loci associated with salinity tolerance in field grown bread wheat. Euphytica 181:371–383. https://doi.org/10.1007/s10681-011-0463-5
Dubcovsky J, Maria GS, Epstein E (1996) Mapping of the K+/Na+ discrimination locus Kna1 in wheat. Theor Appl Genet 92:448–454
El-Hendawy SE, Hassan WM, Al-Suhaibani NA et al (2017) Comparative performance of multivariable agro-physiological parameters for detecting salt tolerance of wheat cultivars under simulated saline field growing conditions. Front Plant Sci 8:435
FAO (2014) Land and plant nutrition management service. FAO soils portal. Food and agriculture organization of the united nations. http://www.fao.org. Accessed 28 Oct 2018
Farshadfar E, Mahjouri S, Aghaee M (2008) Detection of epistasis and estimation of additive and dominance components of genetic variation for drought tolerance in durum wheat. J Biol Sci 8:598–603
Flowers TJ (2004) Improving crop salt tolerance. J Exp Bot 55:307–319. https://doi.org/10.1093/jxb/erh003
Foolad MR, Jones RA (1993) Mapping salt-tolerance genes in tomato (Lycopersicon esculentum) using trait-based marker analysis. Theor Appl Genet 87:184–192
Fukuda A, Nakamura A, Tanaka Y (1999) Molecular cloning and expression of the Na+/H+ exchanger gene in Oryza sativa 1. Biochim Biophys Acta 1446:149–155
Genc Y, Oldach K, Verbyla AP et al (2010) Sodium exclusion QTL associated with improved seedling growth in bread wheat under salinity stress. Theor Appl Genet 121:877–894. https://doi.org/10.1007/s00122-010-1357-y
Ghaedrahmati M, Mardi M, Naghavi MR et al (2014) Mapping QTLs associated with salt tolerance related traits in seedling stage of wheat (Triticum aestivum L.). J Agric Sci Technol 16:1413–1428
Gitonga VW, Koning-Boucoiran CF, Verlinden K et al (2014) Genetic variation, heritability and genotype by environment interaction of morphological traits in a tetraploid rose population. BMC Genet 15:146
Heidari B, Sayed-Tabatabaei BE, Saeidi G et al (2011) Mapping QTL for grain yield, yield components, and spike features in a doubled haploid population of bread wheat. Genome 54:517–527. https://doi.org/10.1139/g11-017
Huang XQ, Cöster H, Ganal MW, Röder MS (2003) Advanced backcross QTL analysis for the identification of quantitative trait loci alleles from wild relatives of wheat (Triticum aestivum L.). Theor Appl Genet 106:1379–1389
Huang XQ, Kempf H, Ganal MW, Röder MS (2004) Advanced backcross QTL analysis in progenies derived from a cross between a German elite winter wheat variety and a synthetic wheat (Triticum aestivum L.). Theor Appl Genet 109:933–943
Huang XQ, Cloutier S, Lycar L et al (2006) Molecular detection of QTLs for agronomic and quality traits in a doubled haploid population derived from two Canadian wheats (Triticum aestivum L.). Theor Appl Genet 113:753–766
Hussain B, Lucas SJ, Ozturk L, Budak H (2017) Mapping QTLs conferring salt tolerance and micronutrient concentrations at seedling stage in wheat. Sci Rep 7:15662. https://doi.org/10.1038/s41598-017-15726-6
James RA, Davenport RJ, Munns R (2006) Physiological characterization of two genes for Na+ exclusion in durum wheat, Nax1 and Nax2 1. Plant Physiol 142:1537–1547. https://doi.org/10.1104/pp.106.086538
Khan MA, Shirazi MU, Khan MA et al (2009) Role of proline, K/Na ratio and chlorophyll content in salt tolerance of wheat (Triticum aestivum L.). Pak J Bot 41:633–638
Kuchel H, Williams KJ, Langridge P et al (2007) Genetic dissection of grain yield in bread wheat. I. QTL analysis. Theor Appl Genet 115:1029–1041. https://doi.org/10.1007/s00122-007-0629-7
Kumar N, Kulwal PL, Balyan HS, Gupta PK (2007) QTL mapping for yield and yield contributing traits in two mapping populations of bread wheat. Mol Breed 19:163–177
Li H, Ye G, Wang J (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175:361–374. https://doi.org/10.1534/genetics.106.066811
Lindsay MP, Lagudah ES, Hare RA, Munns R (2004) A locus for sodium exclusion (Nax1), a trait for salt tolerance, mapped in durum wheat. Funct Plant Biol 31:1105–1114. https://doi.org/10.1071/FP04111
Liu Y, Liu Y, Zhang Q et al (2018) Genome-wide association analysis of quantitative trait loci for salinity-tolerance related morphological indices in bread wheat. Euphytica 214:176. https://doi.org/10.1007/s10681-018-2265-5
Masoudi B, Mardi M, Hervan EM et al (2015) QTL mapping of salt tolerance traits with different effects at the seedling stage of bread wheat. Plant Mol Biol Rep 33:1790–1803. https://doi.org/10.1007/s11105-015-0874-x
Mba C, Afza R, Jain SM et al (2007) Induced mutations for enhancing salinity tolerance in rice. In: Jenks MA, Hasegawa PM, Jain SM (eds) Advances in molecular breeding toward drought and salt tolerant crops. Springer, Berlin, pp 413–454
Mbarki S, Sytar O, Zivcak M et al (2018) Anthocyanins of coloured wheat genotypes in specific response to salt stress. Molecules 23:1518
Mclntosh RA (1998) Catalogue of gene symbols for wheat. In: 9th international wheat genetics symposium. Saskatoon, Canada, pp 123–142
Mujeeb-Kazi A, Munns R, Rasheed A et al (2019) Breeding strategies for structuring salinity tolerance in wheat. In: Sparks DL (ed) Advances in agronomy. Elsevier, Amsterdam, pp 121–187
Munns R, James RA (2003) Screening methods for salinity tolerance: a case study with tetraploid wheat. Plant Soil 253:201–218
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Munns R, Wallace PA, Teakle NL, Colmer TD (2010) Measuring soluble ion concentrations (Na+, K+, Cl-) in salt-treated plants. In: Sunkar R (ed) Plant stress tolerance. Springer, Berlin, pp 371–382
Oyiga BC, Sharma RC, Shen J et al (2016) Identification and characterization of salt tolerance of wheat germplasm using a multivariable screening approach. J Agron Crop Sci 202:472–485. https://doi.org/10.1111/jac.12178
Oyiga BC, Sharma RC, Baum M et al (2018) Allelic variations and differential expressions detected at quantitative trait loci for salt stress tolerance in wheat: GWAS reveals gene loci for salt tolerance in wheat. Plant, Cell Environ 41:919–935. https://doi.org/10.1111/pce.12898
Paliwal R, Röder MS, Kumar U et al (2012) QTL mapping of terminal heat tolerance in hexaploid wheat (T. aestivum L.). Theor Appl Genet 125:561–575. https://doi.org/10.1007/s00122-012-1853-3
Poustini K, Siosemardeh A (2004) Ion distribution in wheat cultivars in response to salinity stress. Field Crops Res 85:125–133
Rana V, Ram S, Sedhil R et al (2015) Physiological, biochemical and morphological study in wheat (Triticum aestivum L.) RILs population for salinity tolerance. J Agric Sci 7:119–128
Rana V, Ram S, Nehra K, Sharma I (2016) Expression of genes related to Na+ exclusion and proline accumulation in tolerant and susceptible wheat genotypes under salt stress. Cereal Res Commun 44:404–413. https://doi.org/10.1556/0806.44.2016.009
Ren Z-H, Gao J-P, Li L-G et al (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146
Ren Z, Zheng Z, Chinnusamy V et al (2010) RAS1, a quantitative trait locus for salt tolerance and ABA sensitivity in Arabidopsis. Proc Natl Acad Sci 107:1–6
Rengasamy P (2010) Soil processes affecting crop production in salt-affected soils. Funct Plant Biol 37:613. https://doi.org/10.1071/FP09249Ren
Reynolds MP, Balota M, Delgado MIB et al (1994) Physiological and morphological traits associated with spring wheat yield under hot, irrigated conditions. Funct Plant Biol 21:717–730
Ribaut J-M, Jiang C, Gonzalez-de-Leon D et al (1997) Identification of quantitative trait loci under drought conditions in tropical maize. 2. Yield components and marker-assisted selection strategies. Theor Appl Genet 94:887–896
Roder MS, Korzun V, Wendehake K et al (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Sairam RK, Srivastava GC, Agarwal S, Meena RC (2005) Differences in antioxidant activity in response to salinity stress in tolerant and susceptible wheat genotypes. Biol Plant 49:85–91
Shamaya NJ, Shavrukov Y, Langridge P et al (2017) Genetics of Na+ exclusion and salinity tolerance in Afghani durum wheat landraces. BMC Plant Biol 17:209. https://doi.org/10.1186/s12870-017-1164-6
Sharma R, Rao B, Saxena R (2004) Salt affected soils in India—current assessment. In: Advances in sodic land reclamation. 9–14 February, Lucknow, India, pp 1–26
Siddiqui MN, Mostofa MG, Akter MM et al (2017) Impact of salt-induced toxicity on growth and yield-potential of local wheat cultivars: oxidative stress and ion toxicity are among the major determinants of salt-tolerant capacity. Chemosphere 187:385–394
Snedecor GW, Cochran WG (1980) Statistical methods, 7th edn. Iowa State University Press, Ames, Iowa, USA
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114. https://doi.org/10.1007/s00122-004-1740-7
Spielmeyer W, Hyles J, Joaquim P et al (2007) A QTL on chromosome 6A in bread wheat (Triticum aestivum) is associated with longer coleoptiles, greater seedling vigour and final plant height. Theor Appl Genet 115:59–66. https://doi.org/10.1007/s00122-007-0540-2
Srinivasa J, Arun B, Mishra VK et al (2014) Zinc and iron concentration QTL mapped in a Triticum spelta × T. aestivum cross. Theor Appl Genet 127:1643–1651. https://doi.org/10.1007/s00122-014-2327-6
Szabados L, Savouré A (2010) Proline: a multifunctional amino acid. Trends Plant Sci 15:89–97. https://doi.org/10.1016/j.tplants.2009.11.009
Tavakkoli E, Fatehi F, Coventry S et al (2011) Additive effects of Na+ and Cl– ions on barley growth under salinity stress. J Exp Bot 62:2189–2203
Tiwari C, Wallwork H, Kumar U et al (2013) Molecular mapping of high temperature tolerance in bread wheat adapted to the Eastern Gangetic Plain region of India. Field Crops Res 154:201–210. https://doi.org/10.1016/j.fcr.2013.08.004
Tuberosa R, Sanguineti MC, Landi P et al (2002) Identification of QTLs for root characteristics in maize grown in hydroponics and analysis of their overlap with QTLs for grain yield in the field at two water regimes. Plant Mol Biol 48:697–712
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78. https://doi.org/10.1093/jhered/93.1.77
Xu YF, An DG, Liu DC et al (2012) Mapping QTLs with epistatic effects and QTL × treatment interactions for salt tolerance at seedling stage of wheat. Euphytica 186:233–245. https://doi.org/10.1007/s10681-012-0647-7
Xu Y, Li S, Li L et al (2013) Mapping QTLs for salt tolerance with additive, epistatic and QTL × treatment interaction effects at seedling stage in wheat. Plant Breed 132:276–283. https://doi.org/10.1111/pbr.12048
Yeo AR, Yeo ME, Flowers SA, Flowers TJ (1990) Screening of rice (Oryza sativa L.) genotypes for physiological characters contributing to salinity resistance, and their relationship to overall performance. Theor Appl Genet 79:377–384
Zadoks JC, Chang TT, Konzak CF (1974) A decimal code for the growth stages of cereals. Weed Res 14:415–421. https://doi.org/10.1111/j.1365-3180.1974.tb01084.x
Zhang LP, Xu XQ, Zhao CP et al (2011) QTL analysis of plant height in photoperiod-thermo sensitive male sterile wheat. Mol Plant Breed 2:92–97
Zhu J-K, Liu J, Xiong L (1998) Genetic analysis of salt tolerance in Arabidopsis: evidence for a critical role of potassium nutrition. Plant Cell 10:1181–1191
Acknowledgements
The authors would like to extend their sincere appreciation to the support and facilities provided by ICAR-Indian Institute of Wheat and Barley Research (IIWBR), Karnal, India under the network project for salt tolerance funded by Indian Council of Agricultural Research (ICAR), New Delhi, India. We are grateful for the support of RD for conducting experiment, data collection, analysis and manuscript writing, SR for overall guidance in conducting experiments, analysis of data and writing manuscript, VR for generating data on SSR, VKM for data analysis and manuscript writing, VP and GPS for reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Devi, R., Ram, S., Rana, V. et al. QTL mapping for salt tolerance associated traits in wheat (Triticum aestivum L.). Euphytica 215, 210 (2019). https://doi.org/10.1007/s10681-019-2533-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-019-2533-z