Abstract
Chromosome 7Hch of Hordeum chilense carries the Phytoene synthase 1 (Psy1) gene encoding the first step in the carotenoid biosynthetic pathway. As such it can be used in the improvement of seed carotenoid content in wheat. Four introgressions of chromosome 7Hch into wheat have been characterized by in situ hybridization of labeled DNA probes and by several sets of DNA markers. Chromosome-specific SSR were used for the identification of wheat chromosomes. Besides 113 conserved orthologous set (COS) markers were tested for homoeologous group 7, of which 97 amplified in H. chilense and 32 were polymorphic between H. chilense and wheat, and 28 expressed sequence tag (EST) barley markers previously allocated to chromosome 7. A total of 60 markers (32 COS and 28 EST) were allocated to chromosome 7Hch with 28 assigned to 7HchS and 22 to 7HchL. A combination of in situ probing and marker genotyping have shown that among the four introgressions there was a substitution line 7Hch (7D), a ditelosomic addition line for the long arm of 7Hch and two homozygous centric translocations 7HchS·2DS and 7HchS·5AL. The Psy1 gene was localized on the short arm of 7Hch. The positions of markers from the international barley consortium map (IBSC2012) were determined and the comparative arm location between H. chilense and H. vulgare is discussed. The genetic stocks characterized here include new wheat-H. chilense recombinations useful for genetic studies and with a potential for breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The reduction of genetic variability in crops is a serious threat to agriculture. The use of wild relatives in breeding programs of cultivated species provides plant breeders with a pool of useful genes to develop new varieties with an improved agronomic performance or quality characteristics. These resources are especially useful in the Triticeae. For instance, the development of synthetic wheat from the crosses between durum wheat and Aegilops tauschii Coss. represents a considerable effort for the use of the diversity available in wild relatives (van Ginkel and Ogbonnaya 2007) in wheat improvement. Similarly, alien resources have been widely used both in durum wheat (reviewed by Ceoloni et al. 2014) and bread wheat (reviewed by Khlestkina 2014) for the development of new varieties or for genetic studies.
Hordeum chilense Roem. et Schult is a diploid wild barley native to Chile and Argentina (Bothmer et al. 1995). It has a potential for breeding Triticeae species due to its high crossability with other members of this tribe, including both durum and common wheat (Martín et al. 1998). The development of introgression lines of H. chilense into wheat has potential for wheat breeding. Chromosome addition and substitution lines of H. chilense in common wheat were developed by Miller et al. (1982). These lines have been successfully used to determine the chromosome location of genes coding for important traits including resistance to greenbug (Schizaphis graminum) (Castro et al. 1996) and endosperm prolamins (Payne et al. 1987) located on chromosome 1Hch; resistance to Septoria tritici on chromosome 4Hch (Rubiales et al. 2000); tolerance to salt on chromosomes 1Hch, 4Hch and 5Hch (Forster et al. 1990), or restoration of fertility on chromosome 6Hch (Martín et al. 2008).
Wheat-H. chilense addition lines have also been used to transfer alien chromosome segments carrying genes of interest such as the resistance to cereal root-knot nematode (Meloidogyne naasi) from H. chilense to wheat (Person-Dedryver et al. 1990). Recently, Calderón et al. (2012) have reported the development of 4Hch introgression lines in durum wheat that may serve as a source of resistance to Septoria tritici blotch (STB) in wheat, although the characterization of these lines for STB resistance is still pending.
The yellow pigment content (YPC) is caused by carotenoids, mainly lutein (Atienza et al. 2007b; Mellado-Ortega and Hornero-Mendez 2012). The chromosome 7Hch has a potential for improving the seed carotenoid content in wheat since it carries the Phytoene synthase 1 (Psy1) which plays a major role in this trait (Ficco et al. 2014; Rodríguez-Suárez et al. 2014). The availability of molecular markers capable of distinguishing between H. chilense chromatin in wheat background constitutes an important tool for selection. Barley expressed sequence tag (EST) have been successfully transferred to H. chilense (Hagras et al. 2005a, b) and they have been used for physical mapping of chromosomes 4Hch (Said and Cabrera 2009), 1Hch (Cherif-Mouaki et al. 2011) and 3Hch (Said et al. 2012). In addition, the development of the conserved orthologous set (COS) (Quraishi et al. 2009) signifies an additional source of markers potentially useful for H. chilense. EST and COS markers allow comparative studies with barley and thus their transference to H. chilense is an important goal.
In the present work, four wheat-H. chilense chromosome 7Hch introgression lines were cytologically characterized using FISH/GISH and chromosome-specific SSR markers. COS markers were assessed for transferability to H. chilense. The arm location within 7Hch was determined for COS and EST markers and compared to their location in barley 7H.
Materials and methods
Plant material
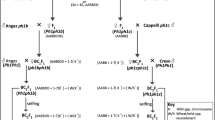
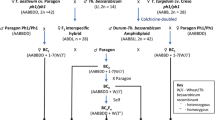
Hordeum chilense lines H1 and H7, common wheat ‘Chinese Spring’ (CS) and H. vulgare ‘Betzes’ were used as controls. Four wheat (CS)-H. chilense introgression lines involving chromosome 7Hch developed at the University of Córdoba were characterized. These lines were obtained by pollinating tritordeum line HT31 (amphiploid H. chilense and durum wheat, AABBHchHch, 2n = 6x = 42) with a wheat disomic addition line for gametocidal chromosome 2C from Aegilops cylindrica Host and F1 plants (AABBDHch + 2C) monosomic for the gametocidal 2C were backcrossed with CS followed by five generations of selfing (Cifuentes et al. 2005). The breeding procedure used to obtain these cytogenetic stocks was that described in (Said et al. 2012).
Fluorescence in situ hybridization (FISH)
Root tips were prepared as described by Said et al. (2012). Preparation of somatic metaphase chromosomes and FISH protocol were carried out as described by Cabrera et al. (2002). For chromosome identification, the pAs1 sequence (1 kb) isolated from A. tauschii Coss. by Rayburn and Gill (1986), the GAA-satellite sequence isolated from barley by Pedersen and Langridge (1997) and H. chilense genomic DNA were used as probes. The pAs1 probe hybridizes strongly to D-genome chromosomes of wheat (Mukai et al. 1993) and Hch-genome chromosomes from H. chilense (Cabrera et al. 1995). The GAA-satellite sequences probe allows identification of both A- and B-genome chromosomes (Pedersen and Langridge 1997). Using nick translation, the pAs1 and GAA-satellite sequences were labeled with biotin-16-dUTP (Roche Corporate) and total H. chilense DNA with digoxigenin-11-dUTP (Roche Corporate). Chromosome preparations were hybridized simultaneously with both pAs1 and H. chilense genomic DNA or GAA-satellite sequences and H. chilense genomic DNA as probes.
Biotin- and digoxigenin-labeled probes were detected with streptavidin-Cy3 conjugates (Sigma, St. Louis, MO, USA) and anti-digoxigenin-FITC (Roche Corporate) antibodies, respectively. Chromosomes were counterstained with 4′,6-diamidino-2-phenylindole (DAPI) and mounted in Vectashield (vector Laboratories, Inc). Signals were visualized using a Leica DMR epifluorescence microscope and images were captured with a SPOT CCD camera supported by the SPOT 2.1 software (Diagnostic Instruments, Inc., Sterling Heights, Michigan, USA). PhotoShop 7.0 software (Adobe Systems Inc., San José, California, USA) was used for processing the images.
Molecular markers analysis
A set of chromosome-specific single sequence repeats (SSR) for A- and D-genomes from wheat was used for the verification of the introgression lines (Supplementary file 1) (Sourdille et al. 2004; Röder et al. 1998). Amplifications were performed as described at GrainGenes, http://wheat.pw.usda.gov/cgi-bin/graingenes/browse.cgi?class=marker).
The presence of Psy1 gene from H. chilense was evaluated using the diagnostic cleaved amplified polymorphism (CAP) marker developed by Atienza et al. (2007a). A set of 113 COS markers (Quraishi et al. 2009) from wheat homoeologous group 7 was assessed for their utility in H. chilense (Supplementary file 2). In addition, twenty-eight barley EST markers previously transferred to H. chilense (Hagras et al. 2005a) were also screened (Supplementary file 3).
Total genomic DNA was isolated from young frozen leaf tissue using the CTAB method (Murray and Thompson 1980). Samples were stored at −20 °C until PCR amplification was carried out. The concentration of each sample was estimated using a NanoDrop 1000 Spectrophotometer (Thermo Scientific, Waltham, MA, USA). Amplifications were made using a TGradient thermocycler (Biometra, Göttingen, Germany) and performed with 40 ng of template DNA, in a 25 µl volume reaction including 2.5 µl of 10× PCR Buffer, 0.5 µM of each primer, 1.5–2.0 mM MgCl2, 2.4 mM dNTPs and 0.25 U of Taq DNA Polymerase (BIOTOOLS B&M Laboratories, Madrid, Spain). PCR conditions of COS markers were as follows: 3 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 30 s at optimal annealing temperature (Supplementary file 2), and 1 min at 72 °C. EST markers were amplified using the touchdown protocol previously reported by (Nasuda et al. 2005; Hagras et al. 2005a).
Amplification products were resolved in 2 % agarose gels (ESTs and SSRs) or vertical PAGE gels at 8 % (w/v, C: 2.67 %) (COS and CAPs), stained with ethidium bromide or SafeView Nucleic Acid Stain (NBS Biologicals, Huntingdon, UK) incorporated in the gel. A 100 bp DNA Ladder (Solis BioDyne, Tartu, Estonia) was used as a standard molecular weight marker. Amplicon lengths were determined using Kodak Digital Science 1D software (version 2.0).
Comparative mapping
For COS markers (Quraishi et al. 2009), the rice locus (RAP) used to design each COS was considered. The RAP locus identifier was retrieved using the ID Converter tool (http://rapdb.dna.affrc.go.jp/tools/converter) (Supplementary file 2). COS markers were located in the Barley Genome Zipper (Mayer et al. 2011) using the RAP locus (Supplementary file 4). The barley full length cDNA corresponding to each rice locus was used for the determination of the barley Unigene corresponding to each COS marker. The Unigene sequences were aligned in Barleymap (Cantalapiedra et al. 2014, in press) to obtain their positions in the International Barley Sequencing Consortium map (IBSC2012) (Mayer et al. 2012) or the POPSEQ map (Mascher et al. 2013) (Supplementary file 5). For some COS, the RAP locus has no barley sequences in the Genome Zipper. In these cases, the positions were estimated considering the barley flanking sequences in the barley Genome Zipper.
The clone name used to design each EST marker (kindly provided by Prof. Tsujimoto, personal communication) was used to identify the corresponding barley Unigene at http://www.ncbi.nlm.nih.gov/unigene/. The Unigene sequence was used for alignment at Barleymap (http://floresta.eead.csic.es/barleymap) (Cantalapiedra et al. 2014, in press) as described for COS markers. Detailed information for BAWU markers is provided in Supplementary file 3.
Results
Cytogenetic and molecular characterization of wheat-H. chilense introgression lines involving chromosome 7Hch
FISH analysis with both pAs1 and H. chilense genomic DNA as probes allowed the identification of a pair of chromosomes 7Hch and the absence of wheat chromosomes 7D pair in one line with 42 chromosomes indicating that it was disomic for the substitution 7Hch(7D) (Fig. 1a, b). The absence of 7D was verified using Xcfd66-7DS (Fig. 2a) and Xbarc111-7DL (Fig. 2b) molecular markers. A pair of telocentric chromosomes for the long arm of chromosome 7Hch was identified in one line with 42 + 2t chromosomes, showing that this line was ditelosomic for the 7HchL (Fig. 1c, d). Two centromeric translocations involving chromosome 7HchS arm and wheat chromosomes were identified in the remaining two lines. One of these lines was homozygous for the 7HchS·2DS translocation and it was also ditelosomic for 2DL chromosome arm and nullisomic for chromosome 7D (Fig. 1e, f). Chromosome-specific markers showed the absence of 7D chromosome (Fig. 2a, b) and the presence of 2DS and 2DL (Fig. 2d, e). The other line was homozygous for the 7HchS·5AL translocation as demonstrated by double FISH with GAA-satellite sequence and genomic DNA from H. chilense as probes (Fig. 1g, h). Molecular markers specific for chromosome 5A confirmed the absence of 5AS and the presence of 5AL chromosome arms, respectively (Fig. 2c). All lines were fertile and vigorous. Table 1 shows the chromosome constitutions of wheat-H. chilense introgression lines involving chromosome 7Hch characterized in this study.
FISH with the pAs1 (red) probe to mitotic metaphase chromosomes: a disomic substitution 7Hch (7D) in CS; c ditelosomic addition of 7HchL chromosome arm in CS and; e homozygous translocation 7HchL·2DS in CS; In (b, d, f) double FISH signals with the pAs1 (red) and H. chilense genomic DNA (green) as probes hybridized to the same metaphase as in (a, c, e), respectively; g metaphase chromosomes of homozygous 7HchS·5AL translocation line probed with GAA-satellite sequence (red); h double FISH signals with GAA-satellite sequence (red) and H. chilense genomic DNA (green) as probes hybridized to the same metaphase as in g. H. chilense chromatin is visualised by green FITC fluorescence whereas wheat chromosomes are counterstained with blue DAPI. Scale bar is 10 µm. (Color figure online)
A set of 113 COS markers corresponding to homoeologous group 7 of wheat (Quraishi et al. 2009) was used for characterizing the introgression lines (Supplementary file 2). Ninety-seven (85.8 %) COS markers were successfully amplified in H. chilense. Out of the remaining 16, six only amplified in wheat and ten failed to amplify in both wheat and H. chilense. Thirty-two COS (32.9 %) were polymorphic between H. chilense and wheat, which permitted the characterization of the cytogenetic stocks described above (Table 2). All the polymorphic COS were located on chromosome 7Hch as demonstrated by their presence in 7Hch (7D) substitution line. The translocation lines T7HchS·2DS and T7HchS·5AL and the ditelosomic line Dt 7HchL were used for the assignation of markers to a specific arm of chromosome 7Hch. Twenty-one COS were assigned to 7HchS and nine to 7HchL. The remaining two COS were simultaneously amplified in both 7HchS and 7HchL arms. A set of 28 barley ESTs markers (coded BAWU) located in 7Hch (Hagras et al. 2005a) was also used (Supplementary file 3). All these markers were successfully amplified in the substitution line 7Hch (7D). Seven markers were located in 7HchS, thirteen in 7HchL arm, and eight markers were found in both arms (Table 2). Examples of amplification of EST and COS markers are given in Fig. 3. A total of sixty markers allowed the distinction of chromosome 7Hch in wheat background and they are therefore useful for the selection of introgressions of this chromosome. The four wheat-H. chilense introgressions were genotyped for the presence of Psy1 gene (Fig. 4). The presence of Psy1 gene on the wheat substitution 7Hch (7D) line and both T7HchS·2DS and T7HchS·5AL translocation lines as well as their absence in the ditelosomic line for the 7HchL arm confirmed the location of this gene on the short arm of chromosome 7Hch (Atienza et al. 2007a).
The Align tool implemented in BarleyMap (http://floresta.eead.csic.es/barleymap) (Cantalapiedra et al. 2014, in press) was used to determine the positions of COS and EST in barley maps (Table 2, Supplementary files 2 and 4). The majority of the markers unambiguously assigned to 7HchS and 7HchL were located in the same barley 7H arm (Table 2). The most notable exceptions were Psy1, which is located in the distal part of 7HL, and BAWU21 and BAWU763, which are located in the distal part of 7HS in barley. Besides, two BAWU markers (BAWU302 and BAWU857) were not found in Barleymap and three (BAWU381, BAWU559 and BAWU752) were located on different barley chromosomes, as previously reported (Hagras et al. 2005b). Four of the COS markers (COS557, COS561, COS562 and COS571) were not found in the Barley Genome Zipper (Mayer et al. 2011) so that their position in barley could not be investigated.
Discussion
The cytogenetic stocks analyzed in this work were obtained using the chromosome 2Cc from Ae. cylindrica which induces chromosome breaks in wheat (Endo 1988). This approach was subsequently applied to barley (Shi and Endo 1999) and rye (Friebe et al. 2000). The chromosome 2Cc induces frequent breakage in the centromeric region of barley chromosomes, with high rate of telocentric chromosomes and whole-arm translocations (Endo 2009). Accordingly, this strategy has allowed the development of structural changes in H. chilense for chromosomes 4Hch (Said and Cabrera 2009), 1Hch (Cherif-Mouaki et al. 2011) and 3Hch (Said et al. 2012) of H. chilense.
Alien introgression is an important strategy in both common (Khlestkina 2014) and durum wheats (Ceoloni et al. 2014). In particular, alien introgressions for homoeologous group 7 have received significant attention for the use of Lr19 and Sr25 from Thynopirum ponticum (Popd.) Barkworth & D.R., Dewey (2n = 10x = 70) or a fusarium head blight resistance locus from Thynopirum elongatum (Ceoloni et al. 2014). Similarly, the development of wheat-Thynopirum recombinants for chromosome 7 has also resulted in improved seed carotenoid content in durum (Ceoloni et al. 2014) and common wheat (Marais and Marais 1990; Ahmad et al. 2013) presumably due to the presence of alien phytoene synthase genes. To our knowledge, no previous translocation lines involving the chromosome 7Hch into wheat have been described and, thus, these cytogenetic stocks constitute an important resource for studying the effect of this chromosome on wheat grain color. Besides, these cytogenetic stocks have allowed the localization of a set of DNA-based markers to specific arms in H. chilense. These markers would facilitate the selection of new genotypes carrying shorter regions of chromosome 7Hch. Some COS and EST markers were found in both arms which suggest that they are located in the pericentromeric region as was concluded in barley when analyzing chromosome arms derived from flow-sorting (Muñoz-Amatriaín et al. 2011).
The high transference rate of COS markers to H. chilense (85.8 %) was expected since these markers were intended for comparative studies among grasses (Quraishi et al. 2009). All the EST markers used in this work were previously assigned to chromosome 7Hch (Hagras et al. 2005a). Barley ESTs show a lower transference rate to H. chilense than COS. Indeed, from a set of 1,165 barley markers, a total of 510 (43.8 %) successfully amplified in H. chilense (Hagras et al. 2005a). Nevertheless, these markers are very useful for the identification of H. chilense chromatin into wheat background (Atienza et al. 2007c; Castillo et al. 2014; Said and Cabrera 2009; Cherif-Mouaki et al. 2011; Said et al. 2012). Previous studies have yielded a limited set of markers useful for H. chilense including RAPDs, SCARs, SSRs and STS (reviewed by Hernandez 2005) with barley and wheat SSRs providing a transferability rate of 53.5 % (Hernandez et al. 2002). The transferability of COS markers reported in this work clearly outperforms both ESTs and SSRs. Besides, these markers show a good collinearity among different grasses and they are therefore expected to be highly informative for comparative mapping between H. chilense and barley in future studies.
Until now, comparative mapping of carotenoid-related genes (Rodríguez-Suárez and Atienza 2012) and polyphenol oxidase genes (Rodríguez-Suárez and Atienza 2014) has shown good collinearity between H. chilense and Triticeae species. Nevertheless, breaks in the collinearity between H. chilense and barley have been previously reported (Hagras et al. 2005b; Said and Cabrera 2009). In this work, the arm location of COS and EST markers in H. chilense showed a good correspondence with barley. The most notorious change refers to the distal parts of barley 7H since the markers located in these regions are found in the opposite arms in H. chilense. This is especially important in the case of Psy1 since this gene influences grain carotenoid content in wheat and tritordeum (Ficcó et al. Ficco et al. 2014; Rodríguez-Suárez et al. 2014). This gene was located in 7HchS using H. chilense-wheat chromosome addition lines (Atienza et al. 2007a; Rodríguez-Suárez et al. 2011), whereas it is found in the long arm of the homoeologous group 7 of wheat and barley (Ficco et al. 2014; Rodríguez-Suárez et al. 2011). On the contrary, the COS318 marker is mapped in the distal part of 7HchL (Rodríguez-Suárez et al. 2012; Rodríguez-Suárez and Atienza 2012) in agreement with the position in barley (Mascher et al. 2013). COS318 is located close to Psy1 in barley whereas Psy1 and COS318 are located in different arms in H. chilense. Furthermore, the EST markers at the distal positions of 7HS are located in 7HchL. These results might indicate an inversion between the distal parts of 7HchS and 7HchL. Nevertheless, further studies would be required to confirm this hypothesis. In any case, inter-specific hybrids between H. vulgare and H. chilense are possible, but are always sterile, and chromosome doubling to produce amphiploids has been unsuccessful (Thomas and Pickering 1985). Inversion affecting homoeologous chromosomes could be responsible for failures in chromosome pairing leading to the lack of fertility in interspecific hybrids between H. vulgare and H. chilense.
In conclusion, the genetic stocks characterized here include new wheat-H. chilense recombinations useful for genetic studies and with a potential for breeding. Furthermore, these lines allowed the identification of new molecular markers useful for selecting new introgressions involving 7Hch. Finally, the distal parts of 7H and 7Hch do not maintain the collinearity, which includes the position of Psy1.
References
Ahmad FT, Asenstorfer RE, Soriano IR, Mares DJ (2013) Effect of temperature on lutein esterification and lutein stability in wheat grain. J Cereal Sci 58:408–413
Atienza SG, Avila CM, Martin A (2007a) The development of a PCR-based marker for Psy1 from Hordeum chilense, a candidate gene for carotenoid content accumulation in tritordeum seeds. Aust J Agric Res 58:767–773
Atienza SG, Ballesteros J, Martin A, Hornero-Mendez D (2007b) Genetic variability of carotenoid concentration and degree of esterification among tritordeum (×Tritordeum Ascherson et Graebner) and durum wheat accessions. J Agric Food Chem 55:4244–4251
Atienza SG, Martín AC, Martín A (2007c) Introgression of wheat chromosome 2D or 5D into tritordeum leads to free-threshing habit. Genome 50:994–1000
Bothmer R, Jacobsen N, Baden C, Jorgensen RB, Linde-Laursen I (1995) An ecogeographical study of the genus Hordeum. Systematic and ecogeographics studies on crop genepools, International Plant Genetic Resources Institute, Rome
Cabrera A, Friebe B, Jiang J, Gill BS (1995) Characterization of Hordeum chilense chromosomes by C-banding and in situ hybridization using highly repeated DNA probes. Genome 38:435–442
Cabrera A, Martin A, Barro F (2002) In-situ comparative mapping (ISCM) of Glu-1 loci in Triticum and Hordeum. Chromosome Res 10:49–54
Calderón MC, Ramírez MC, Martín A, Prieto P (2012) Development of Hordeum chilense 4Hch introgression lines in durum wheat: a tool for breeders and complex trait analysis. Plant Breed 131:733–738. doi:10.1111/j.1439-0523.2012.02010.x
Cantalapiedra CP, Boudiar R, Casas AM, Igartua E, Contreras-Moreira B (2014) BARLEYMAP: physical and genetic mapping of nucleotide sequences and annotation of surrounding loci in barley. Mol Breed (in press)
Castillo A, Atienza SG, Martín AC (2014) Fertility of CMS wheat is restored by two Rf loci located on a recombined acrocentric chromosome. J Exp Bot. doi: 10.1093/jxb/eru388
Castro AM, Martin A, Martin LM (1996) Location of genes controlling resistance to greenbug (Schizaphis Graminum Rond.) in Hordeum chilense. Plant Breed 115:335–338
Ceoloni C, Kuzmanović L, Forte P, Gennaro A, Bitti A (2014) Targeted exploitation of gene pools of alien Triticeae species for sustainable and multi-faceted improvement of the durum wheat crop. Crop Pasture Sci 65:96–111
Cherif-Mouaki S, Said M, Alvarez JB, Cabrera A (2011) Sub-arm location of prolamin and EST-SSR loci on chromosome 1Hch from Hordeum chilense. Euphytica 178:63–69
Cifuentes Z, Said M, Cabrera A (2005) Terminal deletions in Hordeum chilense induced by gametocidal activity of chromosome 2C from Aegilops cylindrica. Chromosome Res 13(Suppl 1):146
Endo TR (1988) Induction of chromosomal structural changes by a chromosome of Aegilops cylindrica L. in common wheat. Hereditas 79:366–370
Endo TR (2009) Cytological dissection of barley genome by the gametocidal system. Breed Sci 59:481–486
Ficco DBM, Mastrangelo AM, Trono D, Borrelli GM, De Vita P, Fares C, Beleggia R, Platani C, Papa R (2014) The colours of durum wheat: a review. Crop Pasture Sci 65:1–15. doi:http://dx.doi.org/10.1071/CP13293
Friebe B, Kynast RG, Gill BS (2000) Gametocidal factor-induced structural rearrangements in rye chromosomes added to common wheat. Chromosome Res 8:501–511
Forster BP, Philips MS, Miller TE, Baird E, Powell W (1990) Chromosome location of genes ontrolling tolerance to salt (NaCl) and vigour in Hordeum vulgare and H. chilense. Heredity 65:99–107
Hagras AA, Kishii M, Sato K, Tanaka H, Tsujimoto H (2005a) Extended application of barley EST markers for the analysis of alien chromosomes added to wheat genetic background. Breed Sci 55:335–341
Hagras AA, Kishii M, Tanaka H, Sato K, Tsujimoto H (2005b) Genomic differentiation of Hordeum chilense from H. vulgare as revealed by repetitive and EST sequences. Genes Genet Syst 80:147–159
Hernandez P (2005) Comparison among available marker systems for cereal introgression breeding: a practical perspective. Euphytica 146:95–100
Hernandez P, Laurie DA, Martin A, Snape JW (2002) Utility of barley and wheat simple sequence repeat (SSR) markers for genetic analysis of Hordeum chilense and tritordeum. Theor Appl Genet 104:735–739
Khlestkina E (2014) Current applications of wheat and wheat–alien precise genetic stocks. Mol Breed 34:273–281
Marais GF, Marais AS (1990) The assignment of a Thinopyrum distichum (Thunb.) Löve-derived translocation to the long arm of wheat chromosome 7D using endopeptidase polymorphisms. Theor Appl Genet 79:182–186
Martín A, Martín LM, Cabrera A, Ramírez MC, Giménez MJ, Rubiales P, Hernández P, Ballesteros J (1998) The potential of Hordeum chilense in breeding Triticeae species. In: Jaradat AA (ed) Triticeae III. Science Publications, Enfield, pp 377–386
Martín AC, Atienza SG, Ramirez MC, Barro F, Martin A (2008) Male fertility restoration of wheat in Hordeum chilense cytoplasm is associated with 6HchS chromosome addition. Aust J Agric Res 59:206–213
Mascher M, Muehlbauer GJ, Rokhsar DS, Chapman J, Schmutz J, Barry K, Muñoz-Amatriaín M, Close TJ, Wise RP, Schulman AH, Himmelbach A, Mayer KFX, Scholz U, Poland JA, Stein N, Waugh R (2013) Anchoring and ordering NGS contig assemblies by population sequencing (POPSEQ). Plant J 76:718–727. doi:10.1111/tpj.12319
Mayer KFX, Martis M, Hedley PE, Šimková H, Liu H, Morris JA, Steuernagel B, Taudien S, Roessner S, Gundlach H, Kubaláková M, Suchánková P, Murat F, Felder M, Nussbaumer T, Graner A, Salse J, Endo T, Sakai H, Tanaka T, Itoh T, Sato K, Platzer M, Matsumoto T, Scholz U, Doležel J, Waugh R, Stein N (2011) Unlocking the barley genome by chromosomal and comparative genomics. Plant Cell 23:1249–1263. doi:10.1105/tpc.110.082537
Mayer KFX, Waugh R, Langridge P, Close TJ, Wise RP, Graner A, Matsumoto T, Sato K, Schulman A, Ariyadasa R, Schulte D, Poursarebani N, Zhou R, Steuernagel B, Mascher M, Scholz U, Shi B, Madishetty K, Svensson JT, Bhat P, Moscou M, Resnik J, Muehlbauer GJ, Hedley P, Liu H, Morris J, Frenkel Z, Korol A, Bergès H, Taudien S, Felder M, Groth M, Platzer M, Himmelbach A, Lonardi S, Duma D, Alpert M, Cordero F, Beccuti M, Ciardo G, Ma Y, Wanamaker S, Cattonaro F, Vendramin V, Scalabrin S, Radovic S, Wing R, Morgante M, Nussbaumer T, Gundlach H, Martis M, Poland J, Pfeifer M, Moisy C, Tanskanen J, Zuccolo A, Spannagl M, Russell J, Druka A, Marshall D, Bayer M, Swarbreck D, Sampath D, Ayling S, Febrer M, Caccamo M, Tanaka T, Wannamaker S, Schmutzer T, Brown JWS, Fincher GB, Stein N (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–716
Mellado-Ortega E, Hornero-Mendez D (2012) Isolation and identification of lutein esters, including their regioisomers, in tritordeum (×Tritordeum Ascherson et Graebner) grains: evidence for a preferential xanthophyll acyltransferase activity. Food Chem 135:1344–1352
Miller TE, Reader SM, Chapman V (1982) The addition of Hordeum chilense chromosomes to wheat. In: Broertjes C (ed) Proceedings of international symposium on Eucarpia on induced variability in plant breeding, Wageningen, Pudoc, pp 79–81
Mukai Y, Nakahara Y, Yamamoto M (1993) Simultaneous discrimination of the three genomes in hexaploid wheat by multicolour fluorescence in situ hybridization using total genomic and highly repeat DNA probes. Genome 36:489–494
Muñoz-Amatriaín M, Moscou MJ, Bhat PR, Svensson JT, Bartoš J, Suchánková P, Šimková H, Endo TR, Fenton RD, Lonardi S, Castillo AM, Chao S, Cistué L, Cuesta-Marcos A, Forrest KL, Hayden MJ, Hayes PM, Horsley RD, Makoto K, Moody D, Sato K, Vallés MP, Wulff BBH, Muehlbauer GJ, Doležel J, Close TJ (2011) An improved consensus linkage map of barley based on flow-sorted chromosomes and single nucleotide polymorphism markers. Plant Genome 4:238–249. doi:10.3835/plantgenome2011.08.0023
Murray YHG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Nasuda S, Kikkawa Y, Ashida T, Rafiqul Islam AKM, Sato K, Endo TR (2005) Chromosomal assignment and deletion mapping of barley EST markers. Genes Genet Syst 80:357–366
Payne PI, Holt LM, Reader SM, Miller TE (1987) Chromosomal location of genes coding for endosperm proteins of Hordeum chilense, determined by two-dimensional electrophoresis of wheat-H. chilense chromosome addition lines. Biochem Genet 25:53–65
Pedersen C, Langridge P (1997) Identification of the entire chromosome complement of bread wheat by two-colour FISH. Genome 40:589–593
Person-Dedryver F, Jahier J, Miller TE (1990) Assessing the resistance to cereal root-knot nematode, Meloidogyne naasi, in a wheat line with the added chromosome arm 1HchS of Hordeum chilense. J Genet Breed 44:291–295
Quraishi UM, Abrouk M, Bolot S, Pont C, Throude M, Guilhot N, Confolent C, Bortolini F, Praud S, Murigneux A, Charmet G, Salse J (2009) Genomics in cereals: from genome-wide conserved orthologous set (COS) sequences to candidate genes for trait dissection. Func Integr Genomics 9:473–484
Rayburn AL, Gill BS (1986) Isolation of a D-genome specific repeated DNA sequence from Aegilops squarrosa. Plant Mol Biol Rep 4:102–109
Röder M, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Rodríguez-Suárez C, Atienza SG (2012) Hordeum chilense genome, a useful tool to investigate the endosperm yellow pigment content in the Triticeae. BMC Plant Biol 12:200
Rodríguez-Suárez C, Atienza SG (2014) Polyphenol oxidadese genes in Hordeum chilense and implications in tritordeum breeding. Mol Breed. doi 10.1007/s11032-014-0145-9
Rodríguez-Suárez C, Atienza SG, Piston F (2011) Allelic variation, alternative splicing and expression analysis of Psy1 gene in Hordeum chilense Roem. Et Schult. PLoS One 6:e19885
Rodríguez-Suárez C, Giménez M, Gutiérrez N, Ávila C, Machado A, Huttner E, Ramírez M, Martín A, Castillo A, Kilian A, Atienza SG (2012) Development of wild barley (Hordeum chilense)-derived DArT markers and their use into genetic and physical mapping. Theor Appl Genet 124:713–722. doi:10.1007/s00122-011-1741-2
Rodríguez-Suárez C, Mellado-Ortega E, Hornero-Méndez D, Atienza SG (2014) Increase in transcript accumulation of Psy1 and e-Lcy genes in grain development is associated with differences in seed carotenoid content between durum wheat and tritordeum. Plant Mol Biol 84:659–673
Rubiales D, Reader SM, Martin A (2000) Chromosomal location of resistance to Septoria tritici in Hordeum chilense determined by the study of chromosomal addition and substitution lines in ‘Chinese Spring’ wheat. Euphytica 115:221–224
Said M, Cabrera A (2009) A physical map of chromosome 4Hch from H. chilense containing SSR, STS and EST-SSR molecular markers. Euphytica 167:253–259. doi:10.1007/s10681-009-9895-6
Said M, Recio R, Cabrera A (2012) Development and characterisation of structural changes in chromosome 3Hch from Hordeum chilense in common wheat and their use in physical mapping. Euphytica 188:429–440. doi:10.1007/s10681-012-0712-2
Shi F, Endo TR (1999) Genetic induction of structural changes in barley chromosomes added to common wheat by a gametocidal chromosome derived from Aegilops cylindrica. Genes Genet Syst 74:49–54
Sourdille P, Singh S, Cadalen T, Brown-Guedira GL, Gay G, Qi L, Gill BS, Dufour P, Murigneux A, Bernard M (2004) Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Func Integr Genomics 4:12–25
Thomas HM, Pickering RA (1985) Comparison of the hybrid Hordeum chilense × H. vulgare, H. chilense × H. bulbosum, H. chilense × Secale cereale and the amphiploid of H. chilense × H. vulgare. Theor Appl Genet 69:519–522
van Ginkel M, Ogbonnaya F (2007) Novel genetic diversity from synthetic wheats in breeding cultivars for changing production conditions. Field Crops Res 104:86–94
Acknowledgments
M. G. Mattera was recipient of a fellowship from Ministerio de Economía y Competitividad (BES-2012-055961). The authors thank Prof. Tsujimoto (Tottori University, Japan), for providing barley EST primer sequences. This research was funded by Grant AGL2011-24399, from Ministerio de Economía y Competitividad including FEDER funding. We are grateful to Ana Pozo for her technical assistance.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mattera, M.G., Ávila, C.M., Atienza, S.G. et al. Cytological and molecular characterization of wheat-Hordeum chilense chromosome 7Hch introgression lines. Euphytica 203, 165–176 (2015). https://doi.org/10.1007/s10681-014-1292-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-014-1292-0