Abstract
Marinobacter is an ecologically important genus of Gammaproteobacteria found in diverse marine habitats, many species of which are capable of degrading hydrocarbons. In this study, we isolated a Marinobacter phage-host system from the surface waters of the Arabian Sea using enrichment culture methods, studied their growth characteristics and investigated the effect of salinity and nitrate concentrations on phage-host interactions. The bacterial isolate had maximum identity to Marinobacter salsuginis based on 16S rRNA similarities and was termed as Marinobacter sp., strain D1S9. It could tolerate up to 14% of NaCl with maximum growth at 11% NaCl. The host grew optimally between 35 and 40 °C and at pH 8. It had a generation time of 3.7 h with a mean growth rate of 0.27 h−1. The phage infected the host forming clear, round plaques of 1–2 mm diameter. It had a narrow host range restricted to the strain Marinobacter D1S9. The latent period and burst size of the phage were estimated to be 30 min and 106 phages per infected cell, respectively. The phage had an adsorption rate of 3.4 × 10−8 ml min−1 and retained 40.4% of its adsorption efficiency at 16% NaCl with a maximum at 4% NaCl (76.1%). Inorganic nitrate was found to have a direct role in controlling host growth and phage burst size.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Viruses are present in the marine environments in copious numbers and their distributions vary spatially and temporally (Bergh et al. 1989; Wommack et al. 1992; Yang et al. 2010; Parsons et al. 2012). Their activities have far-reaching influences in various marine ecosystems. Viral lytic infection results in host mortality and consequent recycling of organic nutrients in the oceans (Evans et al. 2003; Suttle 2007; Mojica et al. 2016). Majority of marine viruses are bacteriophages (Torrella and Morita 1979). Recent studies across the tropical and subtropical oceans revealed viral lysis as one of the major causes of prokaryotic mortality in the deep ocean when compared to the surface waters resulting in the release of dissolved organic carbon. It is estimated that viruses release 145 Gt of carbon per year in these oceans (Lara et al. 2017). Host specific infection and lysis aids in the suppression of prevalence of a single group and encourages the coexistence of diverse groups of prokaryotic organisms (Suttle 2007).
Phage-host interactions in the natural environment are influenced by physico-chemical factors such as temperature, salinity, nutrients, UV radiation, etc. (Finke et al. 2017). Availability of nutrients controls the physiological state of host which in turn determines the virus life strategies and lytic characteristics such as latent period and burst size (Mojica and Brussaard 2014). Several phage-host model systems were used to study the influence of nutrients on the interactions between the phage and its host (Moebus 1996a, b, 1997a, b; Wolf et al. 2004). However, salinity, temperature, and radiation have direct impact on viral stability and infectivity (Kukkaro and Bamford 2009).
Phage-host interactions in the natural environment are so complex that it is essential to monitor them under controlled laboratory conditions to understand how phages modify the functioning of their hosts and overall ecosystem. Marinobacter is a genus comprising numerous highly versatile marine bacteria. They have been isolated from different marine habitats such as oil producing wells and hydrothermal systems and from natural-gas wells. Some of them are known to degrade hydrocarbons, and others can influence the biogeochemical cycling of many inorganic elements (Huu et al. 1999; Handley and Lloyd 2013; Evans et al. 2018). In addition, there are reports that members of the genus Marinobacter could be used in bioreactors for drainage treatment from oil terminals. Phages infecting these bacteria have also been identified from these bioreactors that were supposed to improve the efficiency of the petroleum bioremediation process through phage-driven microbial loop (Rosenberg et al. 2010). The phages specific to hydrocarbon degrading Marinobacter could also be used for the screening of potential hosts in other waste water treatment plants as well. Information on the isolation of phages infecting Marinobacter is scarce with only one detailed study available to date (Zhu et al. 2018). In this study, we isolated and characterized a Marinobacter strain and the phage infecting it from surface water of the Arabian Sea. We hypothesize that nitrate concentration has a positive influence on host growth and subsequently on phage production.
Materials and methods
Isolation of bacterial host and phage
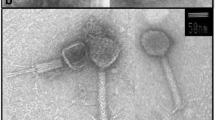
Both the bacterium and the phage were isolated from surface water of the Arabian Sea off the coast of Kochi (9.9586° N and 76.0825° E), southwest coast of India, during December 2014. Culturable bacteria were isolated by serial dilution and spread plating of the collected water sample on Zobell marine agar (ZMA). Pure cultures of morphologically different colonies were then established. Bacteriophages in the sample were screened against the bacterial isolates by soft agar overlay method after enriching 25 mL seawater with 10× Zobell marine broth (Middelboe et al. 2010). Plates were examined for the appearance of plaques after 48 h of incubation at ambient temperature. Clear plaques formed were scraped into phage buffer (450 mM NaCl, 50 mM MgSO4, 50 mM Tris, 0.01% gelatin, and pH 8) and incubated overnight at 4 °C. This was followed by centrifugation at 10,000×g for 10 min and subsequently, the supernatant was passed through a 0.22 μm pore size Acrodisc syringe filter (Sigma-Aldrich, India) to remove any bacterial cells. The filtrate was then plated to isolate a single plaque and this was repeated five times to get the purified phage stock. In order to check the chloroform sensitivity of the phage, phage stock having 1010 PFU mL−1 was treated with 20% final concentration of chloroform and incubated at 37 °C for 30 min. Plaque counts were determined by soft agar overlay method. The phage was characterized by whole genome sequencing (Aparna et al. 2019). Briefly, the viral particles were concentrated by PEG precipitation and ultrafiltration using Amicon Ultra-15 centrifugal filters. Viral DNA was extracted from the concentrated viral particles and sequenced using Illumina MiSeq platform. The detailed protocols and genome analyses are described by Aparna et al. (2019). The genome sequence of the isolated phage termed Marinobacter phage AS1 is available at GenBank under accession number MK088078. Transmission electron microscopy was used for the morphological characterization of the isolated phage (Aparna et al. 2019).
Phylogenetic, morphological, and biochemical characterization of the host
The bacterial host of the isolated phage was identified using 16S rDNA analysis. 16S rDNA region in the genome was amplified by colony PCR using primers: UF (GCACAAGCGGTGGAGCATGTGG) and UR (GCCCGGGAACGTATTCACCG) (Amann et al. 1995). A single isolated colony of the bacterium was mixed with 100 μl TE buffer and incubated at 100 °C for 10 min. The mixture was centrifuged at 1000×g for 10 min, and the supernatant was used as template in PCR with 16S rDNA universal primers. Amplified DNA was sequenced by Sanger dideoxy method and the sequence was compared against NCBI non-redundant database to analyze the phylogenetic lineage of the bacterium. The 16S rRNA partial sequence was submitted in GenBank with accession number MG948170.
Morphology of the bacterium was determined by Grams staining. Various biochemical tests for catalase, oxidase, citrate utilization, nitrate reduction, DNase, amylase, xylanase, lipase, laccase, protease, cellulase, asparaginase, urease, gelatinase, pectinase, phosphatase, indole production, methyl red, and Voges-Proscauer were carried out to identify metabolic potential of the isolated bacterium (Krieg and Holt 1984; Sánchez-Porro et al. 2003; Kasana et al. 2008).
Generation time and mean growth rate of the host
Generation time (G) and mean growth rate (μ) of the bacterium were estimated from its growth curve. For that, an overnight bacterial culture was added to fresh Zobell marine broth to obtain a final OD600 of 0.05 and incubated for 48 h at 37 °C. OD values were measured at regular intervals and plotted against time to generate the growth curve. Generation time and mean growth rate were calculated using the formulae:
Optimum temperature, pH, and salinity for the growth of host bacterium
The host bacterium was grown in Zobell marine broth at different temperatures (10, 20, 30, 35, 40, and 45 °C) and pH (6.5, 7.0, 7.5, 8.0, and 8.5), and in tryptone broth having different NaCl concentrations (0.5, 3, 5, 7, 9, 11, 14, and 16) for 24 h. Optical densities (OD600) were measured at the start and at the end of incubations.
Host specificity and life cycle characterization of the phage
In order to investigate the host range of the phage, soft agar overlays were performed as described previously in the section with 10 Marinobacter strains and 4 Alteromonas strains obtained from microbial culture collection of the Marine Biotechnology Division, ICAR-Central Marine Fisheries Research Institute (CMFRI), Cochin, India.
Latent period and burst size, the two important parameters in phage characterization, were estimated by one step growth experiment as described previously (Middelboe et al. 2010). An overnight culture of host was added to 50 mL Zobell marine broth and incubated until the culture reached a cell density of ~108 CFU mL−1. One milliliter of this culture was added to phage stock of ~106 PFU mL−1 in triplicates, incubated for 10 min, and centrifuged at 6000×g for 10 min, and the pellet was resuspended in 1 mL of growth medium to remove any unadsorbed viruses. The centrifugation and washing processes were repeated thrice. Finally, 50 μL from the resuspended pellet was added to 50 mL of growth medium and incubated at 37 °C. Subsamples were taken at every 15 min interval to estimate the number of viruses as plaque forming units over a period of 2 h.
Determination of adsorption rate constant of the phage
A flask containing 9 mL of temperature equilibrated fresh Zobell marine broth at 37 °C was inoculated with 1 mL of overnight host bacterial culture. One milliliter of phage stock having 105 PFU mL−1 was added to the flask and mixed well. A control flask with 9 mL of the fresh medium and the same amount of phage particles was also maintained along with the experimental flask. Fifty microliter aliquots were removed from the flask and transferred to chilled tubes containing 950 μL growth medium at every 2 min interval until t = 16 min. A 50 μL subsample was also transferred to 950 μL medium to calculate the phage count at t = 0. All the tubes were chilled on ice and after completion of 16 min, the tubes were centrifuged at 10,000×g for 10 min at 4 °C. One-hundred microliter of the supernatant mixed with molten soft agar and the host culture was plated and the plaque forming units were calculated for each tube (Kropinski 2009). Bacterial counts were estimated by serial dilution and plating on Zobell marine agar plates. The adsorption rate constant was calculated as described by Kropinski (2009) using the formula:
Where, k is the adsorption rate constant; B is the bacterial cell concentration; t is the time taken for achieving 50% adsorption; P0 is the initial phage titer, and P is the phage titer at time t.
Influence of NaCl concentration on phage adsorption efficiency
A logarithmic phase culture of the host bacterium (~108 CFU mL−1) was centrifuged at 8000×g, and the pellet was resuspended in tubes with nutrient broth having different added NaCl concentrations (0.5–18%). Equal volume of phage stock having a final concentration of ~107 PFU mL−1 was added to each tube, and the tubes were incubated for 20 min. Titer of unbound viruses in the supernatant was determined for each NaCl concentration by soft agar overlay method after centrifugation at 10,000×g for 10 min (Kukkaro and Bamford 2009).
Phage infection under varying nitrate concentrations
Seawater samples with varying nitrate concentrations (5 μM and 0.3 μM) were used in the infection experiment after removing any bacteria by filtration. For each sample, three preparations were made: one flask with only virus and one with only host (V and B) and a third flask with both host and virus (B + V). To the water samples, equal number of host cells were added and incubated overnight to acclimatize with the growth conditions. Phages were added to a final multiplicity of infection (MOI) 0.1 and incubated at 37 °C. Plaque titer assay and plating on Zobell marine agar were performed at 0, 6, 24, and 48 h to estimate the number of phages and host, respectively. Latent period and burst size of the phage were also estimated under the two different conditions (Middelboe et al. 2010).
Results and discussion
Identification and biochemical characterization of the host
Based on 16S rDNA sequence similarities with the sequences in NCBI database, the host bacterium was closely related to Marinobacter salsuginis (100% similarity) and was designated as Marinobacter sp. strain D1S9. It is Gram negative, rod shaped bacterium forming raised, translucent, circular colonies of 2–3 mm diameter on Zobell marine agar after 32–36 h of incubation. It was oxidase, catalase, and protease positive and negative for all other tests.
Generation time and mean growth rate
A growth curve of D1S9 was plotted as OD600 vs time after 48 h incubation in Zobell marine broth at 37 °C (Fig. 1). The bacterium had a generation time of 3.7 h with a mean growth rate of 0.27 h−1.
Optimum temperature, pH, and salinity for Marinobacter D1S9
Optimum growth of the host occurred between 35 and 40 °C, at pH 8.0, and at 11% NaCl concentration (Fig. 2). The host was moderately halophilic with good growth in the range 3–14% NaCl. Optimum pH of the bacterium is 8, closer to the pH of seawater. Factors such as salinity, temperature, and pH influence the growth of bacteria in marine systems. Temperature tolerance of bacteria is governed by the temperature sensitivity of key enzymes, membrane permeability, etc. (MacLeod 1965). Salt dependence, on the other hand, is a fundamental difference between marine and freshwater bacteria in terms of their physiology and ecological distribution. Na+ ion pumps are required for membrane transport of metabolites from the environment. Moreover, the osmotic effect of the ions helps in the retention of solutes within the cell (MacLeod 1986). Many strains of the genus Marinobacter have been reported to be moderately halophilic with specific enzyme activities and may play crucial roles in different marine environments (Yoon et al. 2003; Martin et al. 2003; Montes et al. 2008; Kumar and Khare 2012). They efficiently degrade aliphatic and polycyclic aromatic hydrocarbons and acyclic compounds and hence play important roles in mitigation of oil pollution (Duran 2010). They have frequently been isolated from hydrocarbon polluted sites (Lal et al. 2015). Marinobacter sp. has been used for saline industrial wastewater treatment due to its denitrifying efficiency, salt tolerance, pH tolerance, and broad carbon use range (Li et al. 2013).
Plaque characteristics and host specificity of the phage
The phage formed clear, round plaques of 1–2 mm diameter on bacterial lawn indicating complete lysis of the cells. Transmission electron microscopic analysis revealed the virus to be a Podovirus having a dsDNA genome with an icosahedral head and a short non-contractile tail. The detailed genome analysis of this phage is already described by Aparna et al. (2019). Phage AS1 lost 99.6% of its activity after treatment with 20% chloroform. In spite of the absence of lipids, one third of the tailed viruses are reported to be chloroform sensitive (Ackermann 2006). For example, Filamentous, dsDNA viruses of family Inoviridae are chloroform sensitive, despite the fact that they contain no lipid component (Cann 2016).
The isolated phage did not infect any of the Marinobacter or Alteromonas strains tested in this study, indicating a narrow host range restricted to the strain Marinabacter D1S9. Phage infection mechanisms and host resistance mechanisms are two possible factors determining host range of a particular phage. Adaptation to specialize for a single host bestows the phage with advantages over selecting for a broad host range (Koskella and Meaden 2013).
One-step growth curve of the phage
The latent period as well as burst size of infection of phage AS1were estimated from one step growth curve (Fig. 3). The latent period lasted for about 30 min and followed by a steady increase in phage counts. This rise in counts of about 2 h resulted in a burst size of ~ 106 plaque forming units per infected cell. Compared to Marinobacter phage B23 (Zhu et al. 2018), which has a longer latent period and smaller burst size, AS1 has shorter latent period and a larger burst size. The two parameters vary largely among phages. A short latent period at high host densities helps the phage to establish multiple infections (Abedon 1989).
Adsorption rate constant and adsorption efficiency of the phage at varying NaCl concentrations
Phage AS1 had a fairly fast adsorption rate of 3.4 × 10−8 mL min−1 when used to infect the host in Zobell marine broth at 37 °C. The phage showed maximum adsorption efficiency at 4% NaCl and had considerable stability up to 16% NaCl (Fig. 4). Fourty percent of adsorption efficiency was retained up to 16% NaCl and dropped to 29.2% at 18% NaCl. Study by Kukkaro and Bamford (2009) on virus-host interactions in a wide range of saline environments reported drastic variations in phage adsorption rates ranging from 2.9 × 10−13 to 1.2 × 10−9. Membrane bound viruses were more sensitive to variations in salt concentration than non-lipid containing ones. The efficiency of phage AS1 to infect the host under a broad salinity range indicates its ability to thrive under varying natural salinity conditions.
Influence of nitrate on phage life cycle characteristics
The latent period of infection was estimated to be three hours for both 5 μM and 0.3 μM nitrate incubations (Fig. 5). The viral burst size varied greatly between the two nitrate levels. In the presence of 5 μM nitrate, the burst size was 325, while in the presence of 0.3 μM nitrate only 28 viral progenies released per bacterial cell. At higher nitrate concentration, a gradual increase in viral counts was observed reaching a maximum (2.9 × 108 PFU mL−1) in 6 h, followed by a steady decline. Bacterial counts did not vary during first six hours, but decreased gradually thereafter. Also at lower nitrate concentration of 0.3 μM, the viral counts showed a rapid increase for the first 6 h (3.2 × 107 PFU mL−1) that subsequently decreased till 24 h, stabilized and increased to reach 3.2 × 107 PFU mL−1 at 48 h. Initial bacterial count at lower nitrate level was half of that at 5 μM at 0 h. No considerable variation in growth was observed until 6 h of incubation (6.7 × 106 CFU mL−1), decreased substantially (4.8 × 105 CFU mL−1) by 24 h and increased thereafter till 48 h (9.6 × 105 CFU mL−1). This increase in bacterial counts after 24 h (Fig. 5b) may have contributed by the increased availability of recycled nutrients as a result of the high viral lysis rates after 6 h. In the bacterial control (BC) flask without viruses, a continuous decrease was observed up to 48 h (3.4 × 105 CFU mL−1). This might be due to the depletion of nutrients in the flask. In both low and high nitrate concentrations, bacterial counts at the end of the experiment were higher than the counts in control incubations indicating the replenishment of nutrients due to viral lysis in the experimental flasks (B + V). Viral lysis of host cells in (B + V) flasks release dissolved nutrients from dead cells that can be sources of nutrients for surviving bacteria. In viral control (VC) flasks, counts remained stable till the end of the experiment.
Influence of varying nitrate concentrations on the Marinobacter phage-host system (a) counts of phage and host (represented as plaque forming units and colony forming units, respectively) at different time intervals under high nitrate concentration (5 μM) and (b) counts under low nitrate concentration (0.3 μM). BC, bacterial count in the control flask; VC, viral count in the control flask; B, bacterial count in the experimental flask with both bacteria and virus; V, viral count in the experimental flask with both bacteria and virus
Several studies have reported the influence of nutrient addition on virus-host interactions (Wolf et al. 2004; Motegi et al. 2009; Motegi and Nagata 2007; Williamson and Paul 2004). Experimental evidences show that burst size is linked to the supply of nutrients to the host cells, which increases with the increase in bacterial growth rate (Middelboe 2000; Nabergoj et al. 2018). In a study, investigating the influence of physiological status of the marine haptophyte Phaeocystis pouchetii on its interactions with virus PpV01, host growth conditions had no significant effect on phage latent period, but burst size varied substantially (Bratbak et al. 1998). Studies have shown that viral production was positively influenced by nitrate addition to seawater (Motegi and Nagata 2007). Viral lytic production is strongly influenced by the trophic status of the system (Payet and Suttle 2013). Nitrogen is an essential element for protein synthesis, the depletion of which indirectly affects phage production by reducing the host growth. In our experiment, the decline in host cell number was faster in high nitrate than in low nitrate concentration, implying the importance of this nutrient in host growth and phage production.
Conclusions
Studies on phage-host interactions are key to understanding the specific interplay between the two in the marine ecosystem. Cultivation experiments combined with genomics provide insights into the role of phages in the functioning of hosts in the environment. In this paper, we describe the isolation and characterization of a Marinobacter phage-host system and discuss the importance of nitrate and salinity status on phage life cycle and adsorption efficiency, respectively. Both the host and the phage exhibited tolerance to a wide range of salinities with the host growing at salinities up to 11% NaCl and the virus retaining 40% of its adsorption efficiency at 16% NaCl. Further, nitrate concentrations have significant positive influence on host growth and phage burst size. Further mesocosm studies are necessary to understand the effects of natural nutrient conditions on phage-host interactions.
References
Abedon, S. T. (1989). Selection for bacteriophage latent period length by bacterial density: a theoretical examination. Microbial Ecology, 18, 79–88.
Ackermann, H. W. (2006). Classification of bacteriophages. In R. Calendar (Ed.), The bacteriophages (pp. 8–16). New York: Oxford University Press.
Amann, R. I., Ludwig, W., & Schleifer, K. H. (1995). Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiological Reviews, 59, 143–169.
Aparna, S., Parvathi, A., Ram, A. P., & Sime-Ngando, T. (2019). Genome analysis of Marinobacter phage AS1 suggests its close interactions with host Marinobacter sp. Aquatic Microbial Ecology, 83, 119–129.
Bergh, O., Borsheim, K. Y., Bratbak, G., & Heldal, M. (1989). High abundance of viruses found in aquatic environments. Nature, 340, 467–468.
Bratbak, G., Jacobsen, A., Heldal, M., Nagasaki, K., & Thingstad, F. (1998). Virus production in Phaeocystis pouchetii and its relation to host cell growth and nutrition. Aquatic Microbial Ecology, 16, 1–9.
Cann, A. J. (2016). Particles. In A. J. Cann (Ed.), Principles of molecular virology (pp. 27–57). Cambridge: Academic.
Duran, R. (2010). Marinobacter. In K. N. Timmis (Ed.), Handbook of hydrocarbon and lipid microbiology (p. 1289). Berlin, Heidelberg: Springer.
Evans, C., Archer, S. D., Jacquet, S., & Wilson, W. H. (2003). Direct estimates of the contribution of viral lysis and microzooplankton grazing to the decline of a Micromonas spp. population. Aquatic Microbial Ecology, 30, 207–219.
Evans, M. V., Panescu, J., Hanson, A. J., Welch, S. A., Sheets, J. M., Nastasi, N., et al. (2018). Members of Marinobacter and Arcobacter influence system biogeochemistry during early production of hydraulically fractured natural gas wells in the Appalachian basin. Frontiers in Microbiology, 9, 2646.
Finke, J., Hunt, B., Winter, C., Carmack, E., & Suttle, C. (2017). Nutrients and other environmental factors influence virus abundances across oxic and hypoxic marine environments. Viruses, 9, 152.
Handley, K. M., & Lloyd, J. R. (2013). Biogeochemical implications of the ubiquitous colonization of marine habitats and redox gradients by Marinobacter species. Frontiers in Microbiology, 4, 136. https://doi.org/10.3389/fmicb.2013.00136.
Huu, N. B., Denner, E. B., Ha, D. T., Wanner, G., & Stan-Lotter, H. (1999). Marinobacter aquaeolei sp. nov., a halophilic bacterium isolated from a Vietnamese oil-producing well. International Journal of Systematic and Evolutionary Microbiology, 49, 367–375.
Kasana, R. C., Salwan, R., Dhar, H., Dutt, S., & Gulati, A. (2008). A rapid and easy method for the detection of microbial cellulases on agar plates using Gram’s iodine. Current Microbiology, 57, 503–507.
Koskella, B., & Meaden, S. (2013). Understanding bacteriophage specificity in natural microbial communities. Viruses, 5, 806–823.
Krieg, N. R., & Holt, J. G. (1984). Bergey's manual of systemic bacteriology. Baltimore: Wiliams & Wilkins.
Kropinski, A. M. (2009). Measurement of the rate of attachment of bacteriophage to cells. In M. R. Clokie & A. M. Kropinski (Eds.), Methods in molecular biology (pp. 151–155). New York: Humana Press.
Kukkaro, P., & Bamford, D. H. (2009). Virus–host interactions in environments with a wide range of ionic strengths. Environmental Microbiology Reports, 1, 71–77.
Kumar, S., & Khare, S. K. (2012). Purification and characterization of maltooligosaccharide-forming α-amylase from moderately halophilic Marinobacter sp. EMB8. Bioresource Technology, 116, 247–251.
Lal, D., Jindal, S., Kumari, H., Jit, S., Nigam, A., Sharma, P., Kumari, K., & Lal, R. (2015). Bacterial diversity and real-time PCR based assessment of linA and linB gene distribution at hexachlorocyclohexane contaminated sites. Journal of Basic Microbiology, 55, 363–373.
Lara, E., Vaqué, D., Sà, E. L., Boras, J. A., Gomes, A., Borrull, E., et al. (2017). Unveiling the role and life strategies of viruses from the surface to the dark ocean. Science Advances, 3, e1602565.
Li, R., Zi, X., Wang, X., Zhang, X., Gao, H., & Hu, N. (2013). Marinobacter hydrocarbonoclasticus NY-4, a novel denitrifying, moderately halophilic marine bacterium. SpringerPlus, 2, 1–9.
MacLeod, R. A. (1965). The question of the existence of specific marine bacteria. Bacteriological Reviews, 29, 9–23.
MacLeod, R. A. (1986). Salt requirements for membrane transport and solute retention in some moderate halophiles. FEMS Microbiology Reviews, 2, 109–113.
Martin, S., Márquez, M. C., Sánchez-Porro, C., Mellado, E., Arahal, D. R., & Ventosa, A. (2003). Marinobacter lipolyticus sp. nov., a novel moderate halophile with lipolytic activity. International Journal of Systematic and Evolutionary Microbiology, 53, 1383–1387.
Middelboe, M. (2000). Bacterial growth rate and marine virus–host dynamics. Microbial Ecology, 40, 114–124.
Middelboe, M., Chan, A., & Bertelsen, S. K. (2010). Isolation and life cycle characterization of lytic viruses infecting heterotrophic bacteria and cyanobacteria. In S. Wilhelm, M. Weinbauer, & C. Suttle (Eds.), Manual of aquatic viral ecology (pp. 118–133). Waco: American Society of Limnology and Oceanography, Inc.
Moebus, K. (1996a). Marine bacteriophage reproduction under nutrient-limited growth of host bacteria. I. Investigations with six phage-host systems. Marine Ecology Progress Series, 144, 1–2.
Moebus, K. (1996b). Marine bacteriophage reproduction under nutrient-limited growth of host bacteria. II. Investigations with phage-host system [H3: H3/1]. Marine Ecology Progress Series, 144, 13–22.
Moebus, K. (1997a). Investigations of the marine lysogenic bacterium H24. II. Development of pseudolysogeny in nutrient-rich broth culture. Marine Ecology Progress Series, 148, 229–240.
Moebus, K. (1997b). Investigations of the marine lysogenic bacterium H24. III. Growth of bacteria and production of phage under nutrient-limited conditions. Marine Ecology Progress Series, 148, 241–250.
Mojica, K. D., & Brussaard, C. P. (2014). Factors affecting virus dynamics and microbial host–virus interactions in marine environments. FEMS Microbiology Ecology, 89, 495–515.
Mojica, K. D., Huisman, J., Wilhelm, S. W., & Brussaard, C. P. (2016). Latitudinal variation in virus-induced mortality of phytoplankton across the North Atlantic Ocean. The ISME Journal, 10, 500.
Montes, M. J., Bozal, N., & Mercade, E. (2008). Marinobacter guineae sp. nov., a novel moderately halophilic bacterium from an Antarctic environment. International Journal of Systematic and Evolutionary Microbiology, 58, 1346–1349.
Motegi, C., & Nagata, T. (2007). Enhancement of viral production by addition of nitrogen or nitrogen plus carbon in subtropical surface waters of the South Pacific. Aquatic Microbial Ecology, 48, 27–34.
Motegi, C., Nagata, T., Miki, T., Weinbauer, M. G., Legendre, L., & Rassoulzadegand, F. (2009). Viral control of bacterial growth efficiency in marine pelagic environments. Limnology and Oceanography, 54, 1901–1910.
Nabergoj, D., Modic, P., & Podgornik, A. (2018). Effect of bacterial growth rate on bacteriophage population growth rate. MicrobiologyOpen, 7, e00558.
Parsons, R. J., Breitbart, M., Lomas, M. W., & Carlson, C. A. (2012). Ocean time-series reveals recurring seasonal patterns of virioplankton dynamics in the northwestern Sargasso Sea. The ISME Journal, 6, 273.
Payet, J. P., & Suttle, C. A. (2013). To kill or not to kill: the balance between lytic and lysogenic viral infection is driven by trophic status. Limnology and Oceanography, 58, 465–474.
Rosenberg, E., Bittan-Banin, G., Sharon, G., Shon, A., Hershko, G., Levy, I., & Ron, E. Z. (2010). The phage-driven microbial loop in petroleum bioremediation. Microbial Biotechnology, 3, 467–472.
Sánchez-Porro, C., Martin, S., Mellado, E., & Ventosa, A. (2003). Diversity of moderately halophilic bacteria producing extracellular hydrolytic enzymes. Journal of Applied Microbiology, 94, 295–300.
Suttle, C. A. (2007). Marine viruses—major players in the global ecosystem. Nature Reviews Microbiology, 5, 801–812.
Torrella, F., & Morita, R. Y. (1979). Evidence by electron micrographs for a high incidence of bacteriophage particles in the waters of Yaquina Bay, Oregon: ecological and taxonomical implications. Applied and Environmental Microbiology, 37, 774–778.
Williamson, S. J., & Paul, J. H. (2004). Nutrient stimulation of lytic phage production in bacterial populations of the Gulf of Mexico. Aquatic Microbial Ecology, 36, 9–17.
Wolf, A., Zheng, T., Witzel, K. P., & Jost, G. (2004). Impact of initial phage/host ratio and nutrient addition on coexistence in a phage-host system. Aquatic Microbial Ecology, 35, 131–139.
Wommack, K. E., Hill, R. T., Kessel, M., Russek-Cohen, E., & Colwell, R. R. (1992). Distribution of viruses in the Chesapeake Bay. Applied and Environmental Microbiology, 58, 2965–2970.
Yang, Y., Motegi, C., Yokokawa, T., & Nagata, T. (2010). Large-scale distribution patterns of virioplankton in the upper ocean. Aquatic Microbial Ecology, 60, 233–246.
Yoon, J. H., Shin, D. Y., Kim, I. G., Kang, K. H., & Park, Y. H. (2003). Marinobacter litoralis sp. nov., a moderately halophilic bacterium isolated from sea water from the East Sea in Korea. International Journal of Systematic and Evolutionary Microbiology, 53, 563–568.
Zhu, M., Wang, M., Jiang, Y., You, S., Zhao, G., Liu, Y., Yang, Q., Liu, Q., Liu, Z., Gong, Z., & Shao, H. (2018). Isolation and complete genome sequence of a novel Marinobacter phage B23. Current Microbiology, 75, 1619–1625.
Acknowledgments
The authors are grateful to Dr. Sunil Kumar Singh, the Director, CSIR-NIO, Goa and Dr. T. Pankajakshan, the Scientist-in-Charge, NIO (RC), Cochin for their support and advice. Authors are grateful to the Director, Indian National Centre for Ocean Information Services, Hyderabad and Dr. Jyothibabu R, Principal Scientist, for the funding through grant-in aid project, GAP 2807. AS is grateful to University Grants Commission (UGC), New Delhi, India for financial support for senior research fellowship grant. The authors are grateful to Dr. Sandhya S. V., ICAR-Central Marine Fisheries Research Institute, Cochin for providing Marinobacter and Alteromonas cultures.
Funding
This work was supported by the Indian National Centre for Ocean Information Services through grant-in-aid project [NIO GAP 2807].
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Aparna, S., Parvathi, A. & Kaniyassery, A. Isolation and characterization of a moderately halophilic Marinobacter phage-host system from the Arabian Sea. Environ Monit Assess 192, 199 (2020). https://doi.org/10.1007/s10661-020-8166-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-020-8166-9