Abstract
This study aimed to understand the prevalence, diversity, antibiotic resistance, β-lactamase gene types, and possibility of environmental survival of meropenem-resistant bacteria present in hospital effluents in Seoul, Korea. Water sampling was performed at five general hospitals in Seoul, Korea, in January 2017. Water samples were plated in triplicate on tryptic soy agar plates with 16 mg/L meropenem. Meropenem-resistant bacteria were selected and subjected to 16S rRNA analysis for species determination and PCR for identification of β-lactamase gene types. Resistant bacteria were cultured in sterilized surface water. Meropenem-resistant bacteria exhibited resistance to more than 12 antibiotics and possessed several β-lactamase genes, such as those encoding OXT-M, NDM-1, AmpC, and OXA. They were able to multiply and survive in sterilized surface water for up to 60 days. Multidrug-resistant bacteria represent an environmental health risk, as they can survive in the environment for an extended period of time. Therefore, these bacteria should be monitored before discharge.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The carbapenem class of antibiotics, including imipenem, doripenem, and meropenem, are used as a last resort against bacteria that are resistant to β-lactam antibiotics due to the presence of β-lactamase genes. Carbapenem antibiotics exhibit antimicrobial activity against members of Enterobacteria, including Escherichia coli, Klebsiella pneumoniae, Enterobacter cloacae, Citrobacter freundii, Proteus mirabilis, and Serratia marcescens. They are known to be especially effective against E. coli and K. pneumoniae isolates that are resistant to cephalosporins as well as Pseudomonas aeruginosa and Acinetobacter species. Among gram-positive bacteria, carbapenem antibiotics are effective against Staphylococcus species that are sensitive to methicillin but are ineffective against Staphylococcus aureus and enterococcus species that are resistant to methicillin (Papp-Wallace et al. 2011; Carbapenem 2016).
Once excreted by humans, antibiotics enter the biosphere via multiple routes (Fig. 1). In hospitals, a diverse array of pharmaceuticals, organic solvents, and radioactive materials are used for the purposes of disinfection, diagnosis, and research. In particular, a large amount of antibiotics used in patients as well as their metabolites are released without decomposition via patient feces and travel to the wastewater treatment plant of the hospital. Subsequently, antibiotics and their metabolites that are not decomposed at the wastewater treatment plant are discharged as hospital effluent into the public groundwater system. As with general sewage and wastewater, the discharge of hospital effluents into groundwater is legal as long as certain effluent standards are satisfied. Table 1 shows the effluent standards for water pollutants in Korea (Ministry of Environment 2015). The National Institute of Environmental Research analyzed the antibiotics found in 25 hospital effluent samples and detected the presence of penicillin G, ciprofloxacin, sulfamethoxazole, vancomycin, and erythromycin (National Institute Environmental Research 2015). Such improperly treated antibiotics likely interfere with the aquatic ecosystem and may eventually threaten human health (Berglund 2015). While most microorganisms found in hospital wastewater treatment plants are bacteria derived from the water used for washing devices or buildings, the infectious bacteria that are found in hospital wastewater treatment plants primarily come from patient feces, and these commonly harbor resistance to antibiotics. Thus, these bacteria in particular may interfere with the ecosystem upon release without proper management (Kümmerer and Henninger 2003; Nuñez and Moretton 2007; Diwan et al. 2010; Katouli et al. 2015).
This study aimed to assess the prevalence of bacteria resistant to carbapenem antibiotics in hospital effluents, as carbapenem antibiotics are used as the last resort against bacteria resistant to β-lactam antibiotics. Meropenem, a carbapenem antibiotic, was selected for use in isolating resistant bacteria. In this study, the frequency and species diversity of meropenem-resistant bacteria were examined as well as their resistance spectra against antimicrobial agents and their β-lactamase genes. In addition, the survivability of meropenem-resistant bacteria in the natural environment was investigated. Furthermore, the general water quality of the effluents in terms of pH, biological oxygen demand (BOD), chemical oxygen demand (COD), total nitrogen (TN), total phosphorus (TP), culturable heterotrophic bacteria, and total coliform bacteria was measured to determine the appropriateness of the water treatment.
Materials and methods
Collection area and methods
Water sampling was performed at five randomly selected general hospitals in Korea in January 2017. In this study, individual hospitals are referred to by the designations J, K, P, S, and Y. For each hospital, 1 L of effluent was collected in a sterilized bottle, refrigerated, and transported to the laboratory. The pH of the collected effluent was measured using a pH meter (HM-30R, TOA, Japan), and the TN and TP contents were measured using a tool (Alliance, USA) for continuous measurement. The BOD and COD were analyzed according to the official testing methods for water quality (Ministry of Environment 2016).
Isolation of heterotrophic and antibiotic-resistant bacteria
To quantify the heterotrophic bacteria, samples were diluted with sterilized physiological saline and plated on plate count agar (Difco, USA) with 1 mL on each of three plates, which were then cultured for 48 h at 35 °C. To quantify the total coliforms, samples were diluted with sterilized physiological saline and plated on desoxycholate agar (Difco, USA) with 1 mL on each of three plates, which were then cultured for 24 h at 35 °C. To isolate meropenem-resistant bacteria, samples were plated on tryptic soy agar (TSA; Difco, USA) with medium containing 18 mg/L meropenem (Daewoong Pharmaceutical, Korea); 0.2 mL was plated on each of the five plates and cultured for 48 h at 35 °C.
Identification of meropenem-resistant bacteria and antibiotic resistance tests
Meropenem-resistant bacteria were subjected to 16S rRNA analysis for species determination. For identification, colony PCR was used to amplify the 16S rRNA genes of the resistant bacteria using pure colonies (Kim and Kim 2016). To determine the antibiotic sensitivities of the identified bacteria, disk diffusion assays were used. The following antibiotics were tested: tetracycline, clindamycin, erythromycin, lincomycin, gentamicin, ampicillin, quinupristin, dalfopristin, linezolid, mupirocin, ceftizoxime, chloramphenicol, ciprofloxacin, imipenem, meropenem, and colistin. The diameter of the inhibition zone was measured in mm and interpreted according to the Clinical Laboratory Standards Institute (CLSI) guidelines (CLSI 2013) as well as those of Sutter et al. (1973) and Finlay et al. (1997). The genes responsible for meropenem resistance were identified using a large-scale β-lactamase gene (bla) detection method (Lee et al. 2015).
Survivability of meropenem-resistant bacteria in the river
To investigate the survivability of the isolated meropenem-resistant bacteria in the environment, the surface water from an area near Han River aquatic facilities (boat yards) was collected and filtered through 5-A filter paper; then, 75 mL of the filtrate was poured into a 250-mL shaking conical flask for sterilization for 15 min at 121 °C. Each strain of the meropenem-resistant bacteria was cultured on a TSA agar plate for 48 h at 35 °C. Bacteria grown on the plate were dispersed using physiological saline (0.85% NaCl) before 0.1 mL of the solution was inoculated into the sterilized river water. After inoculation, the medium was cultured on a shaking incubator at 150 rpm and 25 °C, the average water temperature of the Han River in the summer, for approximately 60 days (Water Information System 2017). At each sampling time during the culture period, samples were taken and diluted in physiological saline for microbial counting using the plate colony count method. An equal amount of sterilized river water was added to the sample in order to maintain the nutritional supply and volume.
Results
The water qualities of the effluents from the five hospitals included in this study are described in Table 2. The pH ranged from 6.90 to 8.05, the BOD from 0.4 to 24.5 mg/L, the COD from 8.3 to 35.6 mg/L, the SS from 8.0 to 32.0 mg/L, the TP from 0.13 to 4.21 mg/L, and the TN from 4.27 to 24.2 mg/L.
As described in Table 3, heterotrophic bacteria were detected in samples from two hospitals, while meropenem-resistant bacteria were detected in the sample from one hospital. In the sample from K hospital, the number of heterotrophic bacteria was 1.7 × 103 CFU/mL, total coliform bacteria were not detected, and the number of meropenem-resistant bacteria was 8 CFU/mL. Meropenem-resistant bacteria were present at a rate of 0.47% among culturable heterotrophic bacteria in hospital effluents. In the sample from Y hospital, the number of heterotrophic bacteria was 1.2 × 103 CFU/mL, total coliform bacteria were not detected, and meropenem-resistant bacteria were not detected. In the samples from J, P, and S hospitals, where heterotrophic bacteria were not detected, a residual amount of chlorine was detected (data not shown), indicating that the effluent is undergoing chlorination at those hospitals.
The results of the 16S rRNA analysis used to identify the resistant bacteria from K hospital grown on medium containing meropenem are presented. The effluent from K hospital produced two colonies of Bacillus licheniformis on the agar plate, one colony of Pseudomonas geniculata, and two colonies of Bacillus mageterium.
Table 4 reports the results of the antibiotic resistance tests of the meropenem-resistant bacteria. The B. licheniformis isolated from the effluent of K hospital exhibited resistance to all 16 antibiotics used in the study, including tetracycline, as did P. geniculata. B. megaterium showed resistance to 12 antibiotics, including clindamycin. Table 5 shows the water quality of the collected surface water, revealing that the effluent water in South Korea was properly treated according to quality standards.
The two colonies of B. licheniformis from the effluent of K hospital were shown to possess β-lactamase genes encoding CTX-M-3 and AmpC. P. geniculata carried genes encoding the CTX-M-3, NDM-1, and OXA-24 β-lactamases. Finally, B. megaterium was found to carry genes encoding the CTX-M-3 and OXA-10 β-lactamases.
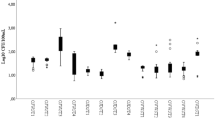
As depicted in Fig. 2, two of the isolated strains of meropenem-resistant bacteria, B. megaterium and P. geniculata, entered the exponential phase 1 day after inoculation into sterilized river water; this phase lasted for 4–6 days before the cultures settled into the stationary phase, where they remained for around 60 days. Among the isolated resistant bacteria, B. megaterium showed the fastest growth rate, while P. geniculata showed the slowest growth rate (data not shown).
Discussion
In this study, the prevalence, diversity, antibiotic resistance spectra, β-lactamase gene types, and environmental survivability of meropenem-resistant bacteria present in the effluents of selected hospitals in Seoul were investigated. Among the five hospitals included in the study, the effluent from K hospital contained 1.7 × 103 CFU/mL of heterotrophic bacteria, no total coliform bacteria, and 8 CFU/mL of meropenem-resistant bacteria.
B. licheniformis, a common soil bacterium that is also found in the feathers and chest hair of birds, was identified as meropenem-resistant in this study. B. licheniformis is a gram-positive, aerotropic, spore-forming, and mesophilic bacillus, whose optimum temperature for growth is around 50 °C, although it can tolerate higher temperatures. As the bacterium can produce alkaline serine protease, it has garnered much attention for its industrial potential (Bacillus licheniformis 2017). In the clinical setting, the bacterium is known to cause infection in patients with weakened immunity, leading to the development of symptoms such as bacteremia, peritonitis, food poisoning, eye infection, infectious endocarditis, and septicemia (Haydushka et al. 2012).
A previous study reported that B. licheniformis possesses a chromosomal ermD gene encoding the rRNA adenine N-6-methyltransferase, which confers resistance to the macrolid class of antibiotics such as erythromycin. Further in-depth study is required, however, to confirm whether the same ermD gene is responsible for the erythromycin resistance of the B. licheniformis identified in the present study. This bacterium has also been reported to be resistant to chloramphenicol, clindamycin, erythromycin, penicillin, tetracycline, and trimethoprim/sulfamethoxazole (Noor Uddin et al. 2015). In this study, B. licheniformis showed resistance to all 16 antibiotics tested, including tetracycline, clindamycin, erythromycin, lincomycin, gentamicin, vancomycin, ampicillin, quinupristin/dalfopristin, linezolid, mupirocin, ceftizoxime, chloramphenicol, ciprofloxacin, imipenem, meropenem, and colistin, implying a higher level of antibiotic resistance in B. licheniformis than that reported in previous studies. The β-lactamase produced by the blaCTX-M-3 gene of B. licheniformis identified in the present study is known to affect resistance to the oxyimino-β-lactam class of antibiotics, while AmpC β-lactamase, commonly found in gram-negative bacteria that are resistant to cephalosporin, is known to be involved in the hydrolysis of both the oxyimino-β-lactam and cephamycin antibiotics, thus explaining the resistance of the B. licheniformis isolate to ceftizoxime and ampicillin in the present study (Beta-lactamase 2017). Of particular interest is colistin, an antibiotic used in the treatment of NDM-1 metallo-β-lactamase multidrug-resistant Enterobacteriaceae such as P. aeruginosa, K. pneumoniae, and Acinetobacter, as it affects membrane stability by mediating the magnesium-calcium ion exchange at the lipopolysaccharides of the outer membranes of gram-negative bacteria (Guo et al. 2015). However, further research is required to verify whether the colistin resistance of B. licheniformis, a gram-positive bacterium without an outer membrane, is due to a barrier against the pharmaceutical permeability or to the acquisition of a resistance gene (Liu et al. 2016).
A proportion of P. geniculata isolated from clinical samples has been reported to show strong resistance to impenem and meropenem based on the presence of a β-lactamase gene similar to blaNDM-1 (Liu et al. 2016). New Delhi metallo-beta-lactamase-1 (NDM-1) is an enzyme sometimes referred to as a carbapenemase based on its ability to confer resistance to a wide array of β-lactam antibiotics. Bacteria that produce this enzyme are known as superbugs because they are difficult to treat, though they are known to be sensitive to polymyxin or tigecycline. The gene encoding NDM-1 is commonly found in gram-negative bacteria such as E. coli and K. pneumoniae and is known to be transferred to other neighboring bacteria via horizontal gene transfer (New Delhi metallo-beta-lactamase 2017). However, in the present study, the P. geniculata isolate exhibited resistance to colistin, a type of polymyxin, and possessed genes encoding the CTX-M-3 and OXA-24 β-lactamases, indicating the severity of antibiotic resistance.
B. megaterium is a gram-positive bacillus that forms spores and grows largely in soil. For industrial purposes, the bacterium is known to be useful for producing penicillin amidase in the manufacture of synthetic penicillin, amylase for use in the baking industry, and glucose dehydrogenase for use in blood sugar testing (Bacillus megaterium 2017). Although B. megaterium is rarely involved in infectious diseases, there have been occasional clinical reports of its involvement in keratitis, primary skin infection, and cerebral abscess (Korea Centers for Disease Control and Prevention 2017). In general, β-lactamase genes are more commonly found in gram-negative bacteria; however, the present study identified genes encoding CTX-M, AmpC, and OXA in the gram-positive bacteria B. licheniformis and B. megaterium, implying the widespread prevalence of antibiotic resistance among gram-positive bacteria.
According to data published by the Korea Center for Disease Control, 82.1% of the Acinetobacter species isolated from blood were found to be Acinetobacter baumannii, among which 94% were derived from hospital-acquired infections and 50% were derived from community-acquired infections. All of these infections were caused by bacteria resistant to two antibiotics belonging to the carbapenem class: meropenem and imipenem. In this study, meropenem-resistant bacteria were also found in hospital effluents, indicating that carbapenemase-producing bacteria have already begun to spread into the environment (Hrenovic et al. 2016).
As shown in Fig. 2, the antibiotic-resistant bacteria released from hospital effluents can survive and grow in river water with a low concentration of nutrients. In a similar experiment, A. baumannii was able to grow for more than 50 days in hospital effluents with a low concentration of nutrients. Thus, the resistant bacteria that are discharged may grow in the oligotrophic environment near recreational areas, thereby posing a potential threat to those who use the facilities in the area. Infection with resistant bacteria in recreational areas has been commonly reported in the USA, although it has not yet been reported in Korea (Leonard et al. 2015).
Bacteria that are resistant to the penicillin class of antibiotics often show resistance to non-β-lactam classes of antibiotics as well, indicating the common occurrence of multidrug-resistant bacteria in the environment. The present study also confirmed that all of the bacterial strains that were resistant to meropenem, part of the penicillin class of antibiotics, were in fact multidrug-resistant bacteria (Ash et al. 2002). Infection with these multidrug-resistant bacteria leads to not only expensive treatments and control measures to limit the prevalence or mortality rate but also the preservation of antibiotic resistance genes; thus, it is necessary to take precautions against these infections (Levy and Marshall 2004). Such multidrug-resistant bacteria may increase the health risks of hospital visits or consumption of contaminated food products and may render common antibiotic treatments marginally effective or ineffective. In the case of the USA, it is estimated that around 2 million people are infected by antibiotic-resistant bacteria, among which 23,000 die annually (Atlas World USA 2014). In Korea, infection with multidrug-resistant bacteria has also increased from 22,928 cases in 2011 to 38,074 cases in 2014, according to data from the Korea Center for Disease Control (Korea Joong Ang Daily 2015).
A limitation of the present study is that only a small number of hospitals were investigated, and therefore, effects due to the size of the hospital or the characteristics of the respective wastewater treatment plants could not be fully assessed. In addition, as the environmental survivability of the bacteria was estimated under specific experimental conditions, the influence of seasonal variations could not be tested. Thus, future research should include an in-depth study on the effects of hospital size, the method of wastewater treatment, and the environmental survivability of resistant bacteria across different seasons. Although the microbiological monitoring of hospital effluents includes total coliform bacteria, these were not detected in hospital effluents, whereas carbapenem-resistant bacteria were detected. This implies that it may be necessary to update the types of bacteria monitored based on the effluent characteristics (Hrenovic et al. 2017).
Conclusions
This study investigated the species and prevalence of meropenem-resistant bacteria found in hospital effluents as well as their antibiotic resistance spectra, β-lactamase gene types, and environmental survivability. The multidrug-resistant bacteria released through hospital effluents can survive and grow in the water near recreational areas used by large numbers of people, suggesting the need for continuous monitoring of not only total coliforms or E. coli but also multidrug-resistant bacteria, which pose significant health risks.
References
Ash, R. J., Mauck, B., & Morgan, M. (2002). Antibiotic resistance of gram-negative bacteria in rivers, United States. Emerg Infect Dis, 8(7), 713–716.
Atlas World USA (2014). Antibiotics in animals tied to risk of human infection. https://www.atlasworldusa.com/blog/antibiotics-in-animals-tied-to-risk-of-human-infection. Accessed 8 October 2017.
Bacillus licheniformis (2017). https://en.wikipedia.org/wiki/Bacillus_licheniformis. Accessed 27 June 2017.
Bacillus megaterium (2017). https://en.wikipedia.org/wiki/Bacillus_megaterium. Accessed October 2017.
Berglund, B. (2015). Environmental dissemination of antibiotic resistance genes and correlation to anthropogenic contamination with antibiotics. Infection, Ecology & Epidemiology, 5, 28564.
Beta-lactamase (2017). https://en.wikipedia.org/wiki/Beta-lactamase. Accessed 29 September 2017.
Carbapenem (2016). https://en.wikipedia.org/wiki/carbapenem. Accessed 26 July 2016.
Clinical Laboratory Standards Institute. (2013). Performance standards for antimicrobial susceptibility testing; Twenty-third informational supplement. M100-S23. Wayne: Clinical Laboratory Standards Institute.
Diwan, V., Tamhankar, A. J., Khandal, R. K., Sen, S., Aggarwal, M., Marothi, Y., Iyer, R. V., Sundblad-Tonderski, K., & Stålsby-Lundborg, C. (2010). Antibiotics and antibiotic-resistant bacteria in waters associated with a hospital in Ujjain, India. BMC Public Health, 10, 414.
Finlay, J. E., Miller, L. A., & Poupard, J. A. (1997). Interpretive criteria for testing susceptibility of staphylococci to mupirocin. Antimicrobial Agents and Chemotherapy, 41(5), 1137–1139.
Guo, F. P., Fan, H. W., Liu, Z. Y., Yang, Q. W., Li, Y. J., & Li, T. S. (2015). Brain abscess caused by Bacillus megaterium in an adult patient. Chinese Medical Journal, 128(11), 1552–1554.
Haydushka, I. A., Markova, N., Kirina, V., & Atanassava, M. (2012). Reccurrent sepsis due to Bacillus lichenformis. Journal of Global Infectious Diseases, 4(1), 82–83.
Hrenovic, J., Goic-Brasic, I., Kazazic, S., Kovacic, A., Ganjto, M., & Tonkic, M. (2016). Carbapenem-resistant isolates of Acinetobacter baummanni in a municipal wastewater treatment plant, Croatia, 2014. Euro Surveillance, 21(15), 30195.
Hrenovic, J., Ganjto, M., & Goic-Barisic, I. (2017). Carbapenem-resistant bacteria in a secondary wastewater treatment plant. Water SA, 43(2), 186–191.
Katouli, M., Thompson, J. M., Gündoğdu, A., & Stratton, H. M. (2015). Antibiotic resistant bacteria in hospital wastewaters and sewage treatment plants. Science Forum and Stakeholder Engagement: Building Linkages, Collaboration and Science Quality. http://www98.Griffith.edu.au/dspace/bitstream/handle/10072/51288/86245_1.pdf?sequence=1. Accessed 3 March 2015.
Kim, Y. J., & Kim, Y. G. (2016). Study on antibiotic resistant bacteria in surface water receiving pharmaceutical complex effluent. Journal of Environmental Health Science, 42(6), 34–40.
Korea Centers for Disease Control and Prevention (2017). http://cdc.go.kr/CDC/intr/CdcKrIntro0201.jsp?menuIds=HOME001-MNU1154-MNU0005-MNU0011&cid=75353. Accessed 10 October 2017.
Korea Joong Ang Daily (2015). http://news.joins.com/article/19094395. Accessed 20 November 2015.
Kümmerer, K., & Henninger, A. (2003). Promoting resistance by the emission of antibiotics from hospitals and households into effluent. Clinical Microbiology and Infection, 9(12), 1203–1214.
Lee, J. J., Lee, J. H., Kwon, D. B., Jeon, J. H., Park, K. S., Lee, C. R., & Lee, S. H. (2015). Fast and accurate large-scale detection of β-lactamase genes conferring antibiotic resistance. Antimicrobial Agents and Chemotherapy, 59, 5967–5975.
Leonard, A. F., Zhang, L., Balfour, A. J., Garside, R., & Gaze, W. H. (2015). Human recreational exposure to antibiotic resistant bacteria in coastal bathing waters. Environment International, 82, 92–100.
Levy, S. B., & Marshall, B. (2004). Antibacterial resistance worldwide: Causes, challenges and responses. Nature Medicine, 10(12), 122–129.
Liu, Y. Y., Wang, Y., Walsh, T. R., Yi, L. X., Zhang, R., Spencer, J., Doi, Y., Tian, G., Dong, B., Huang, X., Yu, L. F., Gu, D., Ren, H., Chen, X., Lv, L., He, D., Zhou, H., Liang, Z., Liu, J. H., & Shen, J. (2016). Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infectious Diseases, 16(2), 161–168.
Ministry of Environment (2015). http://www.law.go.kr/DRF/lawService.do?OC=jaa806&target=law&MST=166074&type=HTML&mobileYn=&efYd=20150101. Accessed 3 March 2015.
Ministry of Environment (2016). Water pollution test standards. http://www.me.go.kr/gg/web/board/read.do?menuId=2246&boardId=788130&boardMasterId=228&condition.hideCate=1. Accessed 20 March 2017.
National Institute Environmental Research (2015). http://img.kisti.re.kr/originalView/originalView.jsp. Accessed 3 March 2015.
New Delhi metallo-beta-lactamase 1 (2017). https://en.wikipedia.org/wiki/New_Delhi_metallo-beta-lactamase_1. Accessed 8 October 2017.
Noor Uddin, G. M., Larsen, M. H., Christensen, H., Aarestrup, F. M., Phu, T. M., & Dalsgaard, A. (2015). Identification and antimicrobial resistance of bacteria isolated from probiotic products used in shrimp culture. PLoS One, 10(7), e0132338.
Nuñez, L., & Moretton, J. (2007). Disinfectant-resistant bacteria in Buenos Aires city hospital wastewater. Brazilian Journal of Microbiology, 38, 644–648.
Papp-Wallace, K. M., Endimiani, A., Taracila, M. A., & Bonomo, R. A. (2011). Carbapenems: Past, present, and future. Antimicrobial Agents and Chemotherapy, 55(11), 4943–4960.
Sutter, V. L., Kwock, Y. Y., & Finegold, S. M. (1973). Susceptibility of Bacteroides fragilis to six antibiotics determined by standardized antimicrobial disc susceptibility testing. Antimicrobial Agents and Chemotherapy, 3(2), 188–193.
Water Information System (2017). http://water.nier.go.kr/waterMeasurement/selectWater.do. Accessed 20 February 2017.
Funding
This research was supported by Basic Science Research Program through the National Research Foundation of Korean (NRF) funded by the Ministry of Science, ICT, and Future Planning (2018R1C1A1A02037363).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Hwang, S.H., Kim, Y.J. Meropenem-resistant bacteria in hospital effluents in Seoul, Korea. Environ Monit Assess 190, 673 (2018). https://doi.org/10.1007/s10661-018-7071-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-018-7071-y