Abstract
Xanthomonas oryzae pv. oryzae (Xoo) poses a major risk to worldwide rice cultivation due to its ability to cause bacterial blight (BB). Identifying the Xoo race patterns, and using resistant genes specific to a particular race is a promising strategy to develop varieties with durable resistance. In the present research, 300 Xoo isolates were confirmed and purified from 40 rice-producing areas of Bangladesh to determine the existing races/pathotypes of Xoo. The sensitive rice varieties IR24, BRRI dhan49, and Purbachi showed susceptible reactions against the tested isolates. Fourteen monogenic differentials and 18 pyramid lines were challenged against 300 isolates of Xoo. Bacterial blight resistance genes Xa1, Xa2, Xa3, Xa4, xa5, Xa7, xa8, Xa10, Xa11, xa13, Xa14, Xa21, Xa23, and Xa27 were found in each monogenic differential. By analyzing patterns of the reaction of 300 Xoo isolates on monogenic differentials, 13 pathotypes/races were determined. The effectiveness of the host plant R genes Xa4, xa5, Xa7, xa8, Xa10, xa13, Xa14, Xa21, and Xa27 against bacterial blight has been determined by analyzing frequency resistances and the responses of near isogenic and pyramid lines. Races 1, 3, and 6 were dominantly widespread across the country and were regarded as important races since they had the greatest number of isolates (25%, 23.33%, and 9.67% respectively). Race 2 was the most ubiquitous among the pathotypes, whereas Race 3 was the most virulent, having circumvented every evaluated resistance gene. The bacterial-blight resistant R genes Xa21 and Xa27 have shown resistance against eight and ten races out of thirteen different races, respectively (i.e., 54.7 and 44.3% of the isolates tested). In the evaluation of 50 pyramid lines against the 5 most virulent races, the combinations of Xa4, Xa7, xa13, and Xa21 or the combinations of Xa4, xa5, Xa7, xa13, and Xa21 genes were effective. At present, the suitable and effective R genes i.e., xa5, Xa7, xa8, xa13, Xa21, Xa23, and Xa27 could be utilized for the development of a durable BB-resistant variety in Bangladesh.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The pathogen Xanthomonas oryzae pv. oryzae (Xoo) is responsible for bacterial blight (BB), a significant leaf disease that severely affects rice. Rice acts as an essential dietary component for more than 50% of the worldwide population (Khush, 2005; Phillips et al., 2024), the food security of Asian nations is contingent upon rice production. A significant annual reduction of 10–20% even up to 80% in rice production is due to the infection induced by bacterial blight disease (Xia et al., 2012). Typically, Xoo penetrates the rice leaf by the margin’s hydathodes and exists in the intercellular spaces of the surrounding epithelial tissues, and then migrates to the xylem arteries to create an infection caused by chronic bacterial blight (Niño-Liu et al., 2006). Distinguishing races of Xoo is possible according to their capacity to cause infection in various rice varieties. Currently, there are more than thirty recognized pathotypes of Xoo identified globally (Mishra et al., 2018; Reddy et al., 1979; Tekete et al., 2020). In favorable conditions, Xoo is capable of causing a maximum yield reduction of around 70% (Reddy et al., 1979). The degree of yield reduction depends on the stage of the crop, the level of sensitivity, and environmental conditions, even though BB impacts all stages of rice development (Mew, 1987; Ou, 1984).
The most economical and environmentally sustainable method of managing rice diseases is the use of resistant cultivars (Pinta et al., 2013). For the development of rice cultivars that are incompatible with a broad spectrum of Xoo races, experimenting with Xoo races is essential. The mutations of the pathogens are responsible for the breaking down of the host resistance resulting the severe emergence of bacterial blight disease. It has been observed that major rice-growing nations in Asia, including Bangladesh, India, Indonesia, China, Korea, Sri Lanka, Malaysia, Nepal, Japan, and the Philippines, exhibited a considerable degree of genetic variation in Xoo strains (Alam et al., 2016; Mishra et al., 2013; Nayak et al., 2008; Noer & Suryanto, 2018; Tekete et al., 2020). According to NIÑO-LIU et al. (2006), the pathogenic diversity of Xoo strains, which originated in the Chinese province of Yunnan, revealed that the strains are virulent and polymorphic along 12 Monogenic differentials. An evaluation of molecular and pathogenic variability of Xoo in India demonstrated that all isolates could overcome resistance genes except for xa5, Xa10, xa13, and Xa21. These four resistance genes were still effective in controlling the disease, suggesting their potential for developing durable bacterial blight resistant varieties (Rashid et al., 2021; L. J. Reddy et al., 2009).
In Bangladesh, 32 rice diseases have been documented (Haq et al., 2011; Khatun et al., 2021). Among those, BB is one of the most devastating diseases affecting rice production all over the world (Mew, 1987; Mew et al., 1993), including Bangladesh (Khan et al., 2009). Every year, the disease reappears in Bangladesh varying in severity (Jalaluddin & Kashem, 2013) and causes yield loss ranging from 5.8 to 30.4% may take place depending on the growth phases of crops and environmental factors (Ansari et al., 2019). The different pathotypes of Xoo that were found in Bangladesh’s main rice-growing areas and identified through investigating how those affected the Monogenic differentials of rice (Alam et al., 2016; Islam et al., 2016; Khan et al., 2009; Rashid et al., 2021). So far, a comprehensive study on analyzing the field evaluation of the Xa27 gene against different isolates of Xoo, has not yet been reported for the assessing of its resistancy and pathotypic variation in Bangladesh.
At present, a total of 48 resistant R genes for BB have been identified on various chromosomes, and several of these genes have been characterized and identified in native wild rice. (Chen et al., 2020; Gu et al., 2004a; Kumar et al., 2012; Sinha et al., 2023; W.-Y. Song et al., 1995; Tan et al., 2004; Zhang et al., 2001) However, rapid alterations in Xoo’s pathogenicity and the emergence of novel Xoo races may decline the resistance acquired by R genes (Khan et al., 2014; Mew, 1987). Before trying to fix the issue of Xoo resistance breaking down, more researches need to be done on pathotypic variation, along with the effective resistant genes selection, and their different combinations to introduce broad-spectrum of resistance. It is critical to develop a thorough understanding of both the resistance mechanisms of hosts and the racial makeup of pathogen populations to develop an effective strategy for implementing resistance genotypes.
The monogenic differential having the Xa27 resistance gene was introduced for the first time in Bangladesh to identify the BB pathotypes. Present studies were undertaken to determine the pathotypic variation of Xoo in Bangladesh using 14 monogenic and 18 pyramid lines. The current investigation was also aimed to identify effective pyramid lines with different combinations of BB resistance genes for developing durable resistant rice cultivars in Bangladesh.
Materials and Methods
Bacterial blight-affected leaf collection
A total of 920 BB-infected leaf samples were collected from 40 widely rice-growing locations in Bangladesh (Fig. 1). Infected leaf samples were obtained from the following rice cultivars: BR3, BR11, BR22, BR23, BRRI dhan34, BRRI dhan46, and BRRI dhan49, Samba mashuri, Swarna, Kalizira, Paizom, and hybrid rice. Prior to Xoo isolation, the BB infected leaf samples were stored in air-dried paper envelopes at 4 °C.
Identification, purification, and isolation of Xoo isolates
A successful isolation of 350 Xoo isolates was obtained on peptone sucrose agar (PSA) medium by the methodology outlined by Rashid et al. (2021). Purified and identified by the methodology outlined by Jalaluddin and Kashem (2013), the isolated bacteria were thereafter briefly stored in a refrigerator set at 4 °C. The steps are shown in Fig. 2. During culture, few isolates were contaminated, or even fewer were not revived. Hence, for the final inoculation, 300 isolates out of 350 were effectively preserved in cryo-vials at −80 °C for further use.
Xoo isolates are confirmed via pathogenicity testing
The maintained bacterial isolates were subjected to three susceptible checks: IR24, BRRI dhan49, and Purbachi for Xoo confirmation by pathogenicity test. For Xoo inoculation, thirty-day-old seedlings were transferred from a net house to an earthen pot. PSA medium was used to culture the bacterial isolates to ensure optimal bacterial growth for two to three days at a temperature of 28 °C. A spectrophotometer was used to determine that the resuspended culture in distilled water had an optical density of OD600 = 1, which corresponds to a bacterial cell number of 3.3 × 108 CFU/ml method was followed to determine pathogenicity, as described by (Kauffman et al., 1973). Virulent isolates were identified 14 days after inoculation if they displayed bacterial blight lesions greater than 3 cm in diameter. The virulent isolates were kept at -80 °C in an NBY liquid medium containing 40% glycerol (Rashid et al., 2021). They were designated as BXoN (Bangladeshi Xanthomonas oryzae, where N denotes the isolate number).
Pathotypic variation of Xoo isolates determination
Materials for the plants and experimental site
The races were determined by analyzing the disease response of several Xoo isolates to monogenic differentials under artificial inoculation conditions. This research used a set of fourteen monogenic differentials i.e., IRBB1, IRBB2, IRBB3, IRBB4, IRBB5, IRBB7, IRBB8, IRBB10, IRBB11, IRBB13, IRBB14, IRBB21, IRBB23 and IRBB27; and 18 BB resistant pyramid lines IRBB50, IRBB51, IRBB52, IRBB53, IRBB54, IRBB55, IRBB56, IRBB57, IRBB58, IRBB59, IRBB60, IRBB61, IRBB62, IRBB63, IRBB64, IRBB65, IRBB66 and IRBB67 that were collected from the International Rice Research Institute (IRRI), Philippines. The identification of the race or races were accomplished by utilizing disease reactions to differential variations following the gene theory. At 14 days after inoculation, based on the cut leaf tip, the percentage of the 20 damaged leaves areas was measured. The percentage of the leaf regions impacted by the disease has been used to determine disease reactions. The classification of disease reactions was based on lesion length; lesions less than 3 cm were considered resistant (R), whereas lesions more than 3 cm were considered sensitive (S) (Li et al., 2009).
The use of pyramid lines and monogenic differentials to ascertain the pathotypes of Xoo isolates
In this investigation, fourteen monogenic differentials with known single resistance genes, i.e., IRBB1, IRBB2, IRBB3, IRBB4, IRBB5, IRBB7, IRBB8, IRBB10, IRBB11, IRBB13, IRBB14, IRBB21, IRBB23, and IRBB27 were utilized to determine the reaction pattern of 300 Xoo isolates collected from Bangladesh, while 18 pyramid lines containing multiple resistance genes were also assessed against same Xoo isolates. To differentiate among the Xoo pathotypes, pathogenicity assessment was done by inoculating on each group of genotypes during the maximum tillering stage.
Stringent evaluation of advanced materials against Xoo isolates
A total of 50 experimental materials, that were collected from the International Rice Research Institute (IRRI), Philippines, along with 2 resistant checks (IRBB60 and IRBB65) and 3 susceptible checks (BRRI dhan49, IR24, and purbachi) were used in both T. Aman and T. Aus season to assess the resistance against bacterial blight pathogen. The research was conducted in the Bangladesh Rice Research Institute’s (BRRI) experimental field, Plant Pathology Division, located in Gazipur, Bangladesh. Thirty-day-old seedlings were transplanted in the field maintaining 20 cm × 20 cm spacing. The experiment was conducted under field conditions by artificial inoculation. Over 48–72 hours at a temperature of 28 °C, a Petri dish containing PSA medium was utilized to cultivate 300 BB isolates. To prepare the inoculum for each isolate, the bacterial culture was diluted with distilled water. An adjustment was made to the inoculum concentration to roughly OD600 = 1 (3.3 × 108 CFU/ml). Plants were inoculated with 5 virulent isolates (BXo6, BXo13, BXo9, BXo22, BXo31) of Xanthomonas oryzae pv. oryzae (Xoo) during the Aus and T. Aman seasons at the highest tillering period by using the leaf clipping technique (Kauffman et al., 1973). Lesion length data from about 20 infected leaves were collected for disease grading fourteen days after inoculation. Click or tap here to enter text. Categorization of the entries was done by measuring the lesion length on the inoculated leaves as Highly Resistant (HR) <1 cm (Score 0), Resistant (R) 1–3 cm (Score 1), Moderately Resistant (MR) 3–5 cm (Score 3), Moderately Susceptible (MS) 5–10 cm (Score 5), Susceptible (S) 10–15 cm (Score 7), and Highly Susceptible (HS) >15 cm (Score 9) (Kim et al., 2015; Lore et al., 2011; Neelam et al., 2020). A scale was used to measure the length of the lesion, including the entire leaf-infected area of 20 leaves of each monogenic line (Table S1 and S2).
Results
Pathotype determination of Xoo isolates through the utilization of monogenic differentials and effective resistance genes
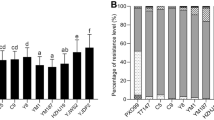
The 300 Xoo isolates exhibited characteristic symptoms of bacterial blight disease on 14 monogenic lines. Based on the response patterns exhibited by the Xoo isolates to Monogenic differentials, thirteen pathotypes/races were determined (Table 1). The most virulent race was 1 followed by 2, 6, and 12, while the least virulent race was 8 followed by 13, 9, 4, 7, etc. There were no resistance reactions observed in the genes Xa1, Xa2, xa3, and Xa11 against 300 Xoo isolates. In Table 1, Xa27, Xa23, Xa21, xa13, Xa7, Xa8, and xa5 genes were effective against BB races in Bangladesh. Here, the Xa21 gene exhibited the maximum frequency of resistance, i.e. the most effective (54.7%), to the greatest number of isolates, whereas the resistant frequencies of the remaining genes xa27, Xa13, xa7, Xa23, Xa8, xa13, and Xa5 were as follows: 44.3, 29.0, 15.00, 6.00, 5.33, and 5.0%, respectively (Table 1 and Fig. 3).
Isolates frequency and their location-wise distribution
The location-wise race distributions of 300 isolates originating from various regions in Bangladesh. The location-wise race distributions of 300 isolates show significant variation among the rice-growing regions (Table 3). Race 1(23.3%) and 2(7.33%) from Gazipur were found as the highest no of isolates, 8 and 4, respectively. This location also showed the maximum frequency of isolate distribution (6.33%). Race 3 (25%) was the predominant and most widely dispersed race, followed by Race 1 (23.33%), Race 6 (9.67%), and Race 7 (8.67%) also being identified as major races of the bacterial blight diseases in Bangladesh (Figs. 4 and 5).
The effect of Xoo isolates on resistant pyramid lines
In addition, the resistance and susceptibility of 18 pyramid lines to 300 tested BB isolates were evaluated to determine the prevalence of resistance for various gene combinations (Table 2). All sensitive checks demonstrated a compatible reaction to the tested isolates, confirming that Xoo isolates exhibited the virulence strain. In the current investigation, pyramid lines containing the combinations of Xa4, Xa7, xa13, and Xa21 or the combinations of Xa4, xa5, Xa7, xa13, and Xa21 genes displayed the greatest frequency of resistance (60%). Furthermore, pyramid line IRBB64 containing Xa4, Xa7, xa13, and Xa21 genes exhibited the second-highest frequency of resistance (59%), while IRBB60 (Xa4, xa5, xa13, and Xa21) and IRBB63 (xa5, Xa7, and xa13) genes exhibited 52% frequency of resistance. The frequency of resistance was greatest (52–60%) when the Xa21 gene was combined with the xa5, Xa7 or xa13 gene (Table 2 and Fig. 4).
The effect of Xoo isolates on advanced lines
Out of 50 advanced lines, these 3 lines IR 129336:11–37 (Xa4-xa5-xa13(H)-Xa21(H)-Xa23), IR 127164:11–26 (Xa4-xa5(H)-Xa7(H)-xa13-Xa21(H)) and IR 129337:37–79 (Xa4-xa5-xa13(H)--Xa23) showed highly resistant to the 5 most virulent races, while 31 lines showed resistant reaction for both T. Aman and T. Aus seasons. Moreover, 14 lines showed moderately resistant reaction. In contrast, 3 lines showed the moderately susceptible reaction and 2 lines showed the susceptible reaction, while Swarna-Sub1 was highly susceptible (Table S1 and S2).
Discussion
Almost every rice-producing location on Earth has extensive documentation of the variations of virulence in Xoo isolates (Kaur et al., 2023; Song et al., 2023). The variability of the virulence of Xoo in Bangladesh is also the subject of considerable research. Thirteen pathotypes or races were identified from 300 Xoo isolates using 14 monogenic lines by their reaction patterns. Based on the reaction pattern of monogenic lines against BB isolates, previous studies also identified different pathotypes of Xoo in Bangladesh, India, Pakistan, and China (Gautam et al., 2015; Kaur et al., 2023; Rashid et al., 2021; Song et al., 2023). A monogenic line exhibited vertical resistance to a single isolate, while the same isolate exhibited diverse interactions with different Monogenic differentials; our finding corroborated with the previous studies as well (Rashid et al., 2021; C. Wang et al., 2005).
The distribution of significant pathotypic diversity was observed in 300 isolates collected from 40 different locations of Bangladesh. Gazipur and Cumilla had been noted to have the highest number of pathotypes, making those the most vulnerable locations to bacterial blight (Khan et al., 2009; Rashid et al., 2021). Additionally, the utilization of a greater number of isolates from these specific regions might contribute to maximum variation of Xoo races. Xoo, consisting of various pathotypes, exhibits diverse pathogenic characteristics in rice cultivars that possess varied resistance genes. Thus, it is imperative to comprehend the diversity of the Xoo population in order to effectively incorporate race-specific resistance genes in the program of variety development.
In our study, the most effective R-genes was Xa21 (54.7%), followed by Xa27 (44.3%), xa13 (29%), Xa7(15%), Xa23 (6%), and xa5(5%). Our study corroborated with the findings of previous studies, such as Khan et al. (2009) reported that the most commonly employed resistance genes for rice breeding to enhance resistance to bacterial blight is Xa21 across the world including Bangladesh. In addition, another study had also recorded a significant resistance of Xa23 to bacterial blight, which encodes an executor R protein to confer broad-spectrum of resistance to bacterial blight (Rashid et al., 2021; C. Wang et al., 2015). Xa27’s ectopic expression evaded the prerequisite for AvrXa27 and provided resistance to suitable strains, revealing that resistance is a result of Xa27 expression at the post-transcriptional level, with specificity being determined by the Xa27 promoter (Gu et al., 2005). Although the resistance spectrum displayed by Xa27 significantly overlapped with that of Xa21, the IRBB27 cultivars’ BB lesion length was less than the IRBB21’s ones, indicating that Xa27 provided greater resistance to the Xoo strains when evaluated compared to Xa21. (Gu et al., 2004b). So far, Xa27 has not been reported yet in Bangladesh. The present study reported first to differentiate the Bangladeshi Xoo isolates (BXoN) by utilizing the Xa27 gene. From our findings, it is Comprehensible that the effective single R genes, Xa21, Xa27, xa13, Xa7, Xa23, and xa5, in different combination could have great possibilities to develop a durable BB-resistant variety in Bangladesh.
Pathogenicity testing on monogenic differentials and pyramid lines, as well as the distribution of Xoo isolates and the number of isolates, were utilized to determine the extent of resistance among genotypes. The results of this study indicate that isolates from the same region had separate pathotypes, but isolates from other locations were found to have identical pathotypes. This discovery provided evidence that the 300 Xoo isolates that were examined are exceptionally dynamic, with distinct population structures observed in different localities. For instance, our previous study revealed the 10 and 08 races out of 12 from Comilla and Gazipur respectively (Rashid et al., 2021), whereas our present investigation found the 07 and 06 out of 13 races from the same locations. Region specific resistant variety could be developed with the conformity of result. Alam et al. (2016) documented variations in the virulence characteristics of Xoo isolates originating from the same regions. In addition, Ardales et al. (1996) concluded, an examination of various agro-ecosystems and cultivars in the Philippines revealed that the degree of variation among the hosts had a little impact on the diversity of pathogens.
Implementing host resistance is the most sustainable, environment friendly, and cost-effective method of combating the bacterial blight disease (Gautam et al., 2015). Additionally, every resistant Xa genes associated with bacterial blight disease has already been catalogued along with its country of origin and source (Khan et al., 2014). In this experiment, the Xa7, xa13, Xa21, and Xa27 genes were shown to be resistant to the majority of the bacterial isolates, while the xa5, xa8 and Xa23 genes exhibited only modest resistance to the isolates. Although, the majority of the isolates of our study exhibited resistance to the Xa21 gene, which corroborates other findings, while a considerable proportion of those from Bangladesh, Korea, Sri Lanka, Pakistan, and Nepal demonstrated virulence towards Xa21 (Alam et al., 2016; Khan et al., 2012; Mazzola et al., 1994). Alam et al. (2016) determined, in accordance with the virulence profiles of 96 Xoo isolates, that Monogenic differentials carrying R-genes xa5 exhibited the highest resistance performance (66.67%) in Bangladesh, followed by Xa2 and Xa21 (65.63%). Furthermore, Adhikari et al. (1995) and Yang et al. (2013) reported identical findings, namely that the xa5 gene provided resistance to the majority of Xoo isolates. While the Xa21 gene has been identified as the most efficacious resistant gene against bacterial blight disease, additional Xa genes are essential for enhancing the action of the Xa21 gene and attaining long-lasting resistance (Jeung et al., 2006). In Pakistan, sixteen pyramid lines having two to five R genes were employed in an experiment against sixteen Xoo isolates; the results indicated that the xa13 and xa21 genes provided resistance to the majority of the isolates (Khan et al., 2014). In Bangladesh, based on the reaction pattern of 300 Xoo isolates on 18 pyramid lines, among those lines, IRBB65 (Xa4, Xa7, xa13, and Xa21), IRBB66 (Xa4, xa5, Xa7, xa13, and Xa21), IRBB64 (Xa4, Xa7, xa13, and Xa21), IRBB60 (Xa4, xa5, xa13, and Xa21) and IRBB63 (xa5, Xa7, and xa13) may be the best sustainable donor parent for the development of bacterial blight-resistant varieties. Furthermore, from screening assessment of 50 advanced lines during T. Aman (July to November) and T. Aus (mid-March to August) seasons, the efficient single R genes’ combinations, i.e., (Xa4, xa5, Xa7, xa13, Xa21), (Xa4, xa5, xa13, xa21, Xa23), and (Xa4, xa5, xa13, Xa23) offered highly resistant against the 5 most virulent races of Xoo. In agreement with the resuts of the present study, several authors showed similar findings (Sukhwinder-Singh et al., 2003; Sundaram et al., 2008; Wang et al., 2020). The different findings of monogenic, pyramid and advanced lines showed integrity in terms of R gene resistance and suggested that the combination of Xa4 or/and xa5 or/and Xa7 or/and xa13 or/and Xa21 or/and Xa23 or/and Xa27 could make the great pave to develop the most durable BB-resistant variety. Planning and designing are necessary to execute comprehensive research on population structures and their genetic compositions with regard to host-plant resistance in every rice-growing country to develop BB-resistant varieties. However, pathotype research with differential cultivars can fail to reveal the true extent of genetic variability present within a given pathogen population (Yashitola et al., 1997).
The bacterial blight-resistant variety can be developed for a country or within a country or in a particular location by a resistance breeding program utilizing the effective Xa genes identified in this work. Furthermore, Plant Pathologists and Plant Breeders will find the results of this research beneficial in gaining a deeper comprehension of the attributes of bacterial blight races in Bangladesh. This understanding will facilitate the development of BB-resistant varieties and the formulation of an effective and sustainable management approach to control the bacterial blight disease.
Conclusion
Our investigation on the pathogenicity test of 300 Xoo isolates collected from 40 districts in Bangladesh revealed 13 distinct pathotypes. A proportion of Xoo bacterial isolates comprising 48.33% belonged to pathotype/race 1 and 3. The resistant genes xa5, Xa7, xa8, xa13, Xa21, and Xa27 were effective against the 300 Xoo isolates. The gene combinations of advanced lines including IR 129336:11–37 (Xa4-xa5-xa13(H)-Xa21(H)-Xa23), IR 127164:11–26 (Xa4-xa5(H)-Xa7(H)-xa13-Xa21(H)), and IR 129337:37–79 (Xa4-xa5-xa13(H)--Xa23) were also highly resistant to BB disease in rice growing zones of Bangladesh. These genes may therefore be useful for the development of a sustainable bacterial blight resistant cultivars. Additionally, gene pyramiding could be an efficient approach for developing durable resistant varieties to bacterial blight disease in Bangladesh.
Data availability
The datasets created and/or analyzed during the current investigation are available upon reasonable request from the respective authors.
References
Adhikari, T. B., Cruz, C., Zhang, Q., Nelson, R. J., Skinner, D. Z., Mew, T. W., & Leach, J. E. (1995). Genetic diversity of Xanthomonas oryzae pv. Oryzae in Asia. Applied and Environmental Microbiology, 61(3), 966–971.
Alam, M. S., Islam, M. R., Hossain, I., Bhuiyan, M. R., & Khan, M. A. I. (2016). Pathotypic variation of Xanthomonas oryzae pv. Oryzae in Bangladesh. Archives of Phytopathology and Plant Protection, 49(1–4), 31–42.
Ansari, T. H., Ahmed, M., Akter, S., Mian, M. S., Latif, M. A., & Tomita, M. (2019). Estimation of rice yield loss using a simple linear regression model for bacterial blight disease. Bangladesh Rice Journal, 23(1), 73–79.
Ardales, E. Y., Leung, H., Vera Cruz, C. M., Mew, T. W., Leach, J. E., & Nelson, R. J. (1996). Hierarchical analysis of spatial variation of the rice bacterial blight pathogen across diverse agroecosystems in the Philippines. Phytopathology, 86(3), 241–252.
Chen, S., Wang, C., Yang, J., Chen, B., Wang, W., Su, J., Feng, A., Zeng, L., & Zhu, X. (2020). Identification of the novel bacterial blight resistance gene Xa46 (t) by mapping and expression analysis of the rice mutant H120. Scientific Reports, 10(1), 12642.
Gautam, R. K., Singh, P. K., Sakthivel, K., Srikumar, M., Kumar, N., Kumar, K., Singh, A. K., & Roy, S. D. (2015). Analysis of pathogenic diversity of the Rice bacterial blight pathogen (Xanthomonas oryzae pv. Oryzae) in the Andaman Islands and identification of effective resistance genes. Journal of Phytopathology, 163(6), 423–432. https://doi.org/10.1111/jph.12338
Gu, K., Tian, D., Yang, F., Wu, L., Sreekala, C., Wang, D., Wang, G.-L., & Yin, Z. (2004a). High-resolution genetic mapping of Xa27 (t), a new bacterial blight resistance gene in rice, Oryza sativa L. Theoretical and Applied Genetics, 108, 800–807.
Gu, K., Tian, D., Yang, F., Wu, L., Sreekala, C., Wang, D., Wang, G.-L., & Yin, Z. (2004b). High-resolution genetic mapping of Xa27(t), a new bacterial blight resistance gene in rice, Oryza sativa L. TAG. Theoretical and Applied Genetics. Theoretische Und Angewandte Genetik, 108(5), 800–807. https://doi.org/10.1007/s00122-003-1491-x
Gu, K., Yang, B., Tian, D., Wu, L., Wang, D., Sreekala, C., Yang, F., Chu, Z., Wang, G.-L., White, F. F., & Yin, Z. (2005). R gene expression induced by a type-III effector triggers disease resistance in rice. Nature, 435(7045), 1122–1125. https://doi.org/10.1038/nature03630
Haq, M., Mia, M. A. T., Rabbi, M. F., & Ali, M. A. (2011). Incidence and severity of rice diseases and insect pests in relation to climate change. Climate Change and Food Security in South Asia (pp 445–457). Springer.
Islam, M. R., Alam, M. S., Khan, A. I., Hossain, I., Adam, L. R., & Daayf, F. (2016). Analyses of genetic diversity of bacterial blight pathogen, Xanthomonas oryzae pv. Oryzae using IS1112 in Bangladesh. Comptes Rendus Biologies, 339(9–10), 399–407.
Jalaluddin, M., & Kashem, M. A. (2013). Pathogenic variability in Xanthomonas oryzae pv oryzae in Bangladesh.
Jeung, J. U., Heu, S. G., Shin, M. S., Vera Cruz, C. M., & Jena, K. K. (2006). Dynamics of Xanthomonas oryzae pv. Oryzae populations in Korea and their relationship to known bacterial blight resistance genes. Phytopathology, 96(8), 867–875.
Kauffman, H. E., Reddy, A. P. K., Hsieh, S. P. Y., & Merca, S. D. (1973). An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Disease Report, 57(6), 537–541 https://eurekamag.com/research/000/019/000019488.php
Kaur, A., Rana, R., Bansal, K., Patel, H. K., Sonti, R. V., & Patil, P. B. (2023). Insights into the diversity of transcription activator-like effectors in Indian Pathotype strains of Xanthomonas oryzae pv. Oryzae. Phytopathology®, 113(6), 953–959. https://doi.org/10.1094/PHYTO-08-22-0304-SC
Khan, J. A., Arshad, H. M. I., Saleem, K., Sandhu, A. F., Hasnain, S., & Babar, M. M. (2012). Evaluation of resistance genes in rice against local isolates of Xanthomonas oryzae pv. Oryzae in Punjab Province of Pakistan. Archives of Phytopathology and Plant Protection, 45(15), 1826–1839.
Khan, M. A., Naeem, M., & Iqbal, M. (2014). Breeding approaches for bacterial leaf blight resistance in rice (Oryza sativa L.), current status and future directions. European Journal of Plant Pathology, 139, 27–37.
Khan, M. A. I., Kabir, M. S., Monsur, M. A., Ali, M. A., & Mia, M. A. T. (2009). Pathogenic diversity of Xanthomonas oryzae pv. Oryzae in Bangladesh. Bangladesh Journal of Plant Pathology, 25(1/2), 1–6.
Khatun, M., Nessa, B., Salam, M., & Kabir, M. (2021). Strategy for Rice disease Management in Bangladesh. Bangladesh Rice Journal, 25(1), 23–36. https://doi.org/10.3329/brj.v25i1.55177
Khush, G. S. (2005). What it will take to feed 5.0 billion rice consumers in 2030. Plant Molecular Biology, 59, 1–6.
Kim, S.-M., Suh, J.-P., Qin, Y., Noh, T.-H., Reinke, R. F., & Jena, K. K. (2015). Identification and fine-mapping of a new resistance gene, Xa40, conferring resistance to bacterial blight races in rice (Oryza sativa L.). Theoretical and Applied Genetics, 128, 1933–1943.
Kumar, P. N., Sujatha, K., Laha, G. S., Rao, K. S., Mishra, B., Viraktamath, B. C., Hari, Y., Reddy, C. S., Balachandran, S. M., & Ram, T. (2012). Identification and fine-mapping of Xa33, a novel gene for resistance to Xanthomonas oryzae pv. Oryzae. Phytopathology, 102(2), 222–228.
Li, G., Song, C., Pang, X., Yang, Y., & Wang, J. (2009). Analysis of pathotypic and genotypic diversity of Xanthomonas oryzae pv. Oryzae in China. Journal of Phytopathology, 157(4), 208–218.
Lore, J. S., Vikal, Y., Hunjan, M. S., Goel, R. K., Bharaj, T. S., & Raina, G. L. (2011). Genotypic and pathotypic diversity of Xanthomonas oryzae pv. Oryzae, the cause of bacterial blight of rice in Punjab state of India. Journal of Phytopathology, 159(7–8), 479–487.
Mazzola, M., Leach, J. E., Nelson, R., & White, F. F. (1994). Analysis of the interaction between Xanthomonas oryzae pv. Oryzae and the rice cultivars IR24 and IRBB21. Phytopathology (USA), 84(4).
Mew, T. W. (1987). Current status and future prospects of research on bacterial blight of rice. Annual Review of Phytopathology, 25(1), 359–382.
Mew, T. W., Alvarez, A. M., Leach, J. E., & Swings, J. (1993). Focus on bacterial blight of rice. Plant Disease, 77(1), 5–12.
Mishra, D., Vishnupriya, M. R., Anil, M. G., Konda, K., Raj, Y., & Sonti, R. V. (2013). Pathotype and genetic diversity amongst Indian isolates of Xanthomonas oryzae pv. Oryzae. PLoS One, 8(11), e81996.
Mishra, R., Joshi, R. K., & Zhao, K. (2018). Genome editing in rice: Recent advances, challenges, and future implications. Frontiers in Plant Science, 9(September). https://doi.org/10.3389/fpls.2018.01361
Nayak, D., Bose, L., Reddy, P., & Nayak, P. (2008). Host-pathogen interaction in rice-bacterial blight pathosystem. Journal of Plant Protection Research, 48, 371–384.
Neelam, K., Mahajan, R., Gupta, V., Bhatia, D., Gill, B. K., Komal, R., Lore, J. S., Mangat, G. S., & Singh, K. (2020). High-resolution genetic mapping of a novel bacterial blight resistance gene xa-45 (t) identified from Oryza glaberrima and transferred to Oryza sativa. Theoretical and Applied Genetics, 133, 689–705.
Niño-Liu, D. O., Ronald, P. C., & Bogdanove, A. J. (2006). Xanthomonas oryzae pathovars: Model pathogens of a model crop. Molecular Plant Pathology, 7(5), 303–324.
Noer, Z., & Suryanto, D. (2018). Pathotype profile of Xanthomonas oryzae pv. Oryzae isolates from north Sumatera. IOP Conference Series: Earth and Environmental Science, 122(1), 12142.
Ou, S. H. (1984). Exploring tropical rice diseases: A reminiscence. Annual Review of Phytopathology, 22(1), 1–11.
Phillips, J., Durand-Morat, A., Nalley, L. L., Graterol, E., Bonatti, M., de la Pava, K. L., Urioste, S., & Yang, W. (2024). Understanding demand for broken rice and its potential food security implications in Colombia. Journal of Agriculture and Food Research, 15, 100884.
Pinta, W., Toojinda, T., Thummabenjapone, P., & Sanitchon, J. (2013). Pyramiding of blast and bacterial leaf blight resistance genes into rice cultivar RD6 using marker assisted selection. African Journal of Biotechnology, 12(28), 4432–4438.
Rashid, M. M., Nihad, S. A. I., Khan, M. A. I., Haque, A., Ara, A., Ferdous, T., Hasan, M. A., & Latif, M. A. (2021). Pathotype profiling, distribution and virulence analysis of Xanthomonas oryzae pv. Oryzae causing bacterial blight disease of rice in Bangladesh. Journal of Phytopathology, 169(7–8), 438–446.
Reddy, A. P. K., Mackenzie, D. R., Rouse, D. I., & Rao, A. V. (1979). Relationship of bacterial leaf blight severity to grain yield of rice. Phytopathology, 69(9), 967–969.
Reddy, L. J., Sirisha, C., & Rao, K. (2009). Assessment of genetic and pathogenic diversity of Xanthomonas oryzae pv. Oryzae on high yielding local variety, tella hamsa, from farmer fields in Gagillapur and Kompally, Andhra Pradesh. Taiwania, 54(3), 241–247.
Sinha, P., Kumar, D., Sk, H., Solanki, M., Gokulan, C. G., Das, A., Miriyala, A., Gonuguntala, R., Elumalai, P., & Kousik, M. B. V. (2023). Fine mapping and sequence analysis reveal a promising candidate gene encoding a novel NB-ARC domain derived from wild rice (Oryza officinalis) that confers bacterial blight resistance. Frontiers in Plant Science, 14, 1173063.
Song, W.-Y., Wang, G.-L., Chen, L.-L., Kim, H.-S., Pi, L.-Y., Holsten, T., Gardner, J., Wang, B., Zhai, W.-X., & Zhu, L.-H. (1995). A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science, 270(5243), 1804–1806.
Song, Z., Zheng, J., Zhao, Y., Yin, J., Zheng, D., Hu, H., Liu, H., Sun, M., Ruan, L., & Liu, F. (2023). Population genomics and pathotypic evaluation of the bacterial leaf blight pathogen of rice reveals rapid evolutionary dynamics of a plant pathogen. Frontiers in Cellular and Infection Microbiology, 13. https://doi.org/10.3389/fcimb.2023.1183416
Sukhwinder-Singh, Sodhi, M., Vikal, Y., George, M. L. C., Bala, G. S., Mangat, G. S., Garg, M., Sidhu, J. S., & Dhaliwal, H. S. (2003). DNA fingerprinting and virulence analysis of Xanthomonas oryzae pv. oryzae isolates from Punjab, northern India. Euphytica, 130, 107–115. https://doi.org/10.1023/A:1022329024651
Sundaram, R. M., Vishnupriya, M. R., Biradar, S. K., Laha, G. S., Reddy, G. A., Rani, N. S., Sarma, N. P., & Sonti, R. V. (2008). Marker assisted introgression of bacterial blight resistance in Samba Mahsuri, an elite indica rice variety. Euphytica, 160, 411–422.
Tan, G.-X., Ren, X., Weng, Q.-M., Shi, Z.-Y., Zhu, L.-L., & He, G.-C. (2004). Mapping of a new resistance gene to bacterial blight in rice line introgressed from Oryza officinalis. Yi Chuan Xue Bao= Acta Genetica Sinica, 31(7), 724–729.
Tekete, C., Cunnac, S., Doucouré, H., Dembele, M., Keita, I., Sarra, S., Dagno, K., Koita, O., & Verdier, V. (2020). Characterization of new races of Xanthomonas oryzae pv. Oryzae in Mali informs resistance gene deployment. Phytopathology, 110(2), 267–277.
Wang, C., Su, C., Zhai, H., & Wan, J. (2005). Identification of QTLs underlying resistance to a virulent strain of Xanthomonas oryzae pv. Oryzae in rice cultivar DV85. Field Crops Research, 91(2–3), 337–343.
Wang, C., Zhang, X., Fan, Y., Gao, Y., Zhu, Q., Zheng, C., Qin, T., Li, Y., Che, J., Zhang, M., Yang, B., Liu, Y., & Zhao, K. (2015). XA23 is an executor R protein and confers broad-Spectrum disease resistance in Rice. Molecular Plant, 8(2), 290–302. https://doi.org/10.1016/j.molp.2014.10.010
Wang, S., Liu, W., Lu, D., Lu, Z., Wang, X., Xue, J., & He, X. (2020). Distribution of bacterial blight resistance genes in the main cultivars and application of Xa23 in rice breeding. Frontiers in Plant Science, 11, 555228.
Xia, C., Chen, H., & Zhu, X. (2012). Identification, mapping, isolation of the genes resisting to bacterial blight and application in rice. Molecular Plant Breeding. https://doi.org/10.5376/mpb.2012.03.0012
Yang, S.-Q., Liu, S.-Y., Zhao, S., Yu, Y.-H., Li, R.-B., Duan, C.-J., Tang, J.-L., & Feng, J.-X. (2013). Molecular and pathogenic characterization of new Xanthomonas oryzae pv. Oryzae strains from the coastline region of Fangchenggang city in China. World Journal of Microbiology and Biotechnology, 29, 713–720.
Yashitola, J., Krishnaveni, D., Reddy, A. P. K., & Sonti, R. V. (1997). Genetic diversity within the population of Xanthomonas oryzae pv. Oryzae in India. Phytopathology, 87(7), 760–765.
Zhang, Q., Wang, C. L., Zhao, K. J., Zhao, Y. L., Caslana, V. C., Zhu, X. D., Li, D. Y., & Jiang, Q. X. (2001). The effectiveness of advanced rice lines with new resistance gene Xa23 to rice bacterial blight. Rice Genetics Newsletter, 18, 71–72.
Acknowledgements
The authors wish to express their gratitude to the International Rice Research Institute (IRRI) for supplying the pyramid lines and near isogenic lines utilized in the process of race identification.
Funding
The National Agricultural Technology Program-Phase II (NATP-2; Project ID: 091) provided funding to the authors for this experiment.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors reported no possible conflicts of interest.
Supplymentary information
ESM 1
Table S1: Assessing rice genotypes for bacterial blight (BB) during T. Aus season. Table S2: Assessing rice genotypes for bacterial blight (BB) during T. Aman season (DOCX 47 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Latif, M.A., Rahman, L., Islam, N. et al. Pathotypic diversity of Xanthomonas oryzae pv. oryzae, and stringent evaluation of resistance lines of Rice in Bangladesh. Eur J Plant Pathol (2024). https://doi.org/10.1007/s10658-024-02900-6
Accepted:
Published:
DOI: https://doi.org/10.1007/s10658-024-02900-6