Abstract
Prunus humilis Bunge (Cerasus humilis (Bunge) S. ya. Sokolov) (Rosaceae) is a small shrub native to China, highly valued for environmental protection and as a functional food. However, with the increasing scale of cultivation, crown gall disease has become a major concern. In this study, we aimed to isolate and identify the pathogenic strains causing crown gall disease in P. humilis, and investigate their physiological, phylogenetic, opine type, and biological control characteristics. We isolated 45 pathogenic strains from five regions of China. Using phenotypic and phylogenetic analysis of the 16S rRNA gene, we identified 12 strains as Rhizobium rhizogenes and 33 strains as members of the Agrobacterium tumefaciens species complex. Multi-locus sequence analysis of three housekeeping genes (rpoB, atpD, and recA) revealed that 11 strains belonged to A. fabacearum and 22 strains belonged to A. radiobacter within the A. tumefaciens species complex. Of the isolated strains, five displayed characteristics identical to the typical nopaline-type Ti plasmid, whereas the remaining 40 strains did not contain nopaline, octopine, or agropine plasmids. However, all strains possessed the trans-zeatin synthesizing gene and agrocinopine synthetase gene, with some also containing nopaline catabolism genes, suggesting the presence of a plasmid closely related to the nopaline type. Resistance to the agrocins produced by R. rhizogenes K1026 was observed in all strains. However, in a greenhouse experiment, K1026 demonstrated 100%, 100%, and 25% control efficiency against three P. humilis-derived pathogenic strains, respectively. These findings reveal unique characteristics of the pathogens responsible for crown gall disease in P. humilis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Crown gall disease is a bacterial disease affecting most dicotyledonous plants and some monocots. It is caused by organisms belonging to the Agrobacterium, Rhizobium, and Allorhizobium genera, with the key features of its pathogenesis including the possession of a tumor-inducing (Ti) plasmid (Van Larebeke et al., 1974). Plant cells infected by pathogens constantly produce auxin and cytokinin, stimulating cell proliferation, thus forming tumors (Gohlke & Deeken, 2014). As tumors grow, infected plants begin to grow poorly or die, causing severe economic losses.
In addition to producing hormones, cells infected by pathogens produce small molecular weight compounds called opines, which are not produced by normal plant cells and cannot be used by most microorganisms; however, Agrobacterium/Rhizobium can use opines. Different pathogenic strains induce different types of opine synthesis and the type of opine that a strain can induce depends on the type of Ti plasmid carried. Genes involved in the synthesis and metabolism of opines are often associated with other strain characteristics, such as conjugal transfer of the Ti-plasmid (Beck von Bodman et al., 1992) and resistance to agrocin 84 (Kim & Farrand, 1997). Agrocin 84, produced by the nonpathogenic R. rhizogenes K84/K1026, prevents crown gall disease caused by pathogens carrying nopaline-type or agrocinopine-type plasmids, but not octopine-type plasmids (Kerr, 2016). A careful analysis of the taxonomy of pathogens and their Ti plasmid characteristics is crucial for accurate risk assessment of the spread and damage caused by crown gall disease, and for the development of effective control measures.
The taxonomy of Agrobacterium spp. has undergone several revisions. Initially, Agrobacterium was classified into three species according to whether they were pathogenic: Agrobacterium tumefaciens, which produces tumors; Agrobacterium rhizogenes, which produces hairy roots, and Agrobacterium radiobacter, which is nonpathogenic. Later, it was found that the pathogenic genes were on plasmids rather than chromosomes and that these plasmids could be transferred under certain conditions (Teyssier-cuvelle et al., 2004). Therefore, the use of pathogenicity as a classification criterion was inappropriate. Therefore, according to the physiological and biochemical characteristics, Agrobacterium were divided into three biovars, each of which includes tumorigenic, rhizogenic, or nonpathogenic strains (Kersters et al., 1973). The biovars were then raised to the species level and were named A. tumefaciens, A. rhizogenes, and Agrobacterium vitis, respectively (Holmes & Roberts, 1981; Ophel & Kerr, 1990). Following this, the appellation ‘tumefaciens’ and ‘rhizogenes’ no longer indicated their pathogenic properties (Mafakheri et al., 2019). The results of 16S rRNA gene sequence analysis showed that A. rhizogenes and A. vitis are distantly related to A. tumefaciens (Sawada et al., 1993). Therefore, the first two were assigned to the genera Rhizobium and Allorhizobium and were referred to as Rhizobium rhizogenes and Allorhizobium vitis, respectively (Mousavi et al., 2015; Young et al., 2001). Analysis of whole-genome information of A. tumefaciens, using DNA-DNA hybridization, has revealed that this species is heterogeneous and constitutes a species complex (Costechareyre et al., 2010).
In the A. tumefaciens species complex, there are 15 genomospecies: G1–G9, G13, G14, G15, G19, G20, and G21 (Mafakheri et al., 2019; Singh et al., 2021). Only ten of these have species-level binomial names: A. fabacearum (G1; Delamuta et al., 2020), A. pusense (G2; Panday et al., 2011), A. tomkonis (G3; Singh et al., 2021), A. radiobacter (G4; Costechareyre et al., 2010), “A. deltaense” (G7; Yan et al., 2017a), “A. fabrum” (G8; Lassalle et al., 2011), A. salinitolerans (G9; Yan et al., 2017b), A. nepotum (G14; Puławska et al., 2012), A. burrii (G19) and A. shirazense (G20) (Mafakheri et al., 2022).
Prunus humilis Bunge (Cerasus humilis (Bunge) S. ya. Sokolov) (Rosaceae), native to China (Wang et al., 2018), is a small deciduous shrub with a well-developed root system and strong tolerance to drought and cold ( Yin et al., 2014, 2017). Its fruit contains very high levels of calcium, and is thus known as ‘calcium fruit’ in China (Mu et al., 2015). In addition, the fruits are rich in organic acids, amino acids, vitamins, flavonoids, anthocyanins, phenols, other nutrients, and bioactive substances (Mu et al., 2015; Wang et al., 2018). Therefore, P. humilis is very popular in environmental protection, such as in soil and sand erosion control, as well as a functional food for nutrition (Fu et al., 2020, 2021; Ran et al., 2022; Wang et al., 2017, 2018). With the increasing understanding of the potential functions of P. humilis, the planting area is increasing annually. In the process of large-scale cultivation, crown gall disease in P. humilis has developed gradually. According to our investigation, crown gall incidence in P. humilis orchards is approximately 30% and is spreading (Our unpublished data). Crown gall disease has become a bottleneck restricting the development of the P. humilis industry. There is only one published article on crown gall disease in P. humilis. In the study, four strains of A. radiobacter were identified in a nursery in the Ningxia Hui Autonomous Region, China (Bai et al., 2022). That paper is a good starting point, but the sample size is too small, and the plasmid characteristics of these pathogenic strains have not been reported. A larger sample size and more in-depth and systematic studies are needed to gain a better understanding of the taxonomy, diversity and plasmid characteristics of the pathogens involved.
In this study, we isolated and characterized tumorigenic bacteria from diseased P. humilis from five regions in China and determined their phylogenetic position through sequence analysis of 16S rRNA and three housekeeping genes (rpoB, atpD, and recA). Furthermore, we investigated the genes involved in the synthesis and metabolism of opines using PCR. We also evaluated the sensitivity of tumorigenic bacteria to the biocontrol strain R. rhizogenes K1026 and investigated the cause of resistance based on the agrocin 84 sensitivity gene. Finally, we evaluated the biocontrol efficacy of K1026 under greenhouse conditions. To the best of our knowledge, this is the first report on the diversity and plasmid characteristics of the pathogens that cause crown gall in P. humilis.
Materials and methods
Samplings and bacterial isolation
From 2020–2021, we obtained 50 P. humilis plants with crown gall disease from 12 nurseries in five regions in China. These nurseries were in Jinzhong City, Shanxi Province; Kaiyuan City, Liaoning Province; Qinhuangdao City, Hebei Province; Changli County, Hebei Province; Tai'an City, Shandong Province. Tumor fragments were cut from plants, and washed with sterile distilled water. They were then patted dry with sterile filter paper, disinfected with 75% ethanol for 1 min, crushed in a sterile mortar, and ground with a small amount of normal saline solution. The mixture was transferred to a 2 mL sterile centrifuge tube and allowed to stand for 1 h. Then 100 μL of the supernatant was taken and applied to plates of selective media 1A and 2E, which could selectively separate A. tumefaciens and R. rhizogenes, respectively (Brisbane & Kerr, 1983). For rhizosphere soil, we took approximately 0.2 g using a sterile brush, placed it in a sterile 5 mL centrifuge tube, added 2 mL of sterile normal saline, before thoroughly shaking and mixing, standing for 1 h, and adding 100 μL of supernatant to selective media 1A and 2E. The plates, coated with supernatant, were placed in a 28 °C incubator and cultured in the dark for 2–7 days. Brick red colonies were selected from medium 1A and blue colonies were selected from medium 2E, and transferred onto fresh YEM (mannitol 10 g/L, yeast extract 3 g/L, NaCl 0.2 g/L, MgSO4·7H2O 0.2 g/L, K2HPO4·3H2O 0.5 g/L, agar 15 g/L, pH 7.2) ager plates, and pure single colonies were obtained by continuous streaking. Single colonies were cultured in YEM liquid medium at 28 °C for 1 day, then sterile glycerol was added to make a final volume of glycerol 20%, mixed, and stored at -80 ℃ for analyses.
Determination of pathogenicity and plasmid type
All isolated strains were evaluated for their tumorigenic ability. The test plants included carrot discs, P. humilis seedlings, sunflower seedlings, and tomato seedlings. Strains showing tumorigenicity on one of the materials were identified as pathogenic. Tumorigenicity on carrot discs was identified using the method described in Ryder et al. (1985). For P. humilis, sunflower, and tomato seedlings, the following steps were used. Tomatoes and sunflowers were grown from seeds. P. humilis was grown from cuttings. The stem of the seedling was punctured with a blunt needle and a drop of freshly cultured bacteria was placed on the wound site. Three to five stems were inoculated with each strain. After inoculation, the seedlings were cultured in a light incubator (14 h light at 25 °C, 10 h dark at 18 °C) and the bottom watering method was used to reduce the interference of spraying water on the inoculation sites. Thirty days later, the wounds were examined for tumor formation. The pots, substrates, and needles were sterilized by spraying 75% alcohol or autoclaving at 121 °C, 103 kPa for 20 min in advance. In addition, the virulence gene virD2 and cytokinin synthesis gene ipt of all isolates were detected using duplex PCR. These two genes are required for tumorigenesis and primers specifically amplified virD2 and ipt to generate 338 bp and 427 bp target bands, respectively (Haas et al., 1995). Fresh bacteria were cultured, and the plasmid was extracted as a PCR template, using the TIANprep Mini Plasmid Kit (TIANGEN, China), according to the manufacturer’s instructions. The PCR reaction mixture was as follows: 10 μL PCR mixture contained 5 μL of 2 × Taq Master Mix with loading dye (Vazyme, China), 3.2 μL of RNase-free water, 0.2 μL (10 pmol) of two primers, and 1 μL (100 ng) of template DNA. All PCR protocols conducted in this study used the same reaction mixture, except that 3.6 μL of RNase-free water was added when the reaction was not a duplex PCR. The PCR protocol was as follows: pre-denaturation at 98 ℃ for 30 s, followed by 35 cycles of 98 ℃ for 10 s, 40 s at their respective best annealing temperatures, 1 min extension at 72 ℃, with a final extension of 5 min at 72 ℃. Primer sequences and optimum annealing temperatures are listed in Supplementary Table S1. PCR products were separated using 1.5% agarose gel; positive amplification was detected on a gel imager, using D2000 DNA Ladder (Real-Times, China) as a Marker.
To determine the plasmid types, plasmid DNA PCR was performed with specific primers. The main target genes were nopaline synthetase gene (nos), octopine synthetase gene (ocs), agropine synthetase gene (ags), agrocinopine synthetase gene (acs), trans-zeatin synthesizing gene (tzs), and the right border of nopaline-type Ti plasmids (Hwang et al., 2013). Primer sequences and optimum annealing temperatures are listed in Table S1.
Phenotypic characterization
All strains identified as tumorigenic had their phenotypic characteristics analyzed. The main tests performed included analysis of: the production of 3-ketolactose; the ability to grow at 35 °C or in the presence of 2% (w/v) NaCl; alkaline or acid production from litmus milk, sodium malonate, erythritol, and ethanol; pigmentation in ferric ammonium citrate broth; and citrate utilization (Du & Lu, 2011).
Housekeeping genes sequencing and phylogenetic analysis
Fresh bacteria were cultured, and the DNA was extracted as PCR template using the TIANamp Bacteria DNA Kit (TIANGEN, China), according to the manufacturer’s instructions. Specific primers were used to amplify the 16S rRNA (Weisburg et al., 1991), rpoB (Aujoulat et al., 2011), atpD (Gaunt et al., 2001) and recA gene fragments. Primer sequences and optimum annealing temperatures are listed in Table S1. The amplified products were sent to the BGI TECH SOLUTIONS (BEIJING LIUHE) CO., LIMITED for sequencing. The amplified sequences were subjected to BLAST alignment in National Center for Biotechnology Information (NCBI) and uploaded to GenBank to obtain accession numbers. Gene sequences of related species were downloaded from the NCBI database and multiple sequence alignment was performed using ClustalW in MEGA 5 software (Tamura et al., 2011), followed by phylogenetic analysis using maximum likelihood. Phylogenetic analysis was performed based on each single gene as well as on the concatenated sequences of rpoB, atpD and recA genes, respectively. The best model for the phylogenetic tree was analyzed using MEGA MODELS. The stability of the phylogenetic trees was estimated with 1000 bootstrap replicates (Felsenstein, 1985).
Sensitivity to agrocin 84 and biological control with K1026
The sensitivity of the pathogenic strains to agrocin 84 produced by R. rhizogenes K1026 was tested using the method of Stonier (1960). Briefly, strain K1026 was inoculated on Stonier's agar plates and incubated at 28 °C for three days. The colony was removed with a sterile cotton swab and K1026 was killed with chloroform vapor. The plate was then covered with a thin layer of agar containing the test strains. After culturing at 28 °C for 1–2 days, the appearance of inhibition zones was observed, and the diameter was measured. “A. fabrum” strain C58T was used as a positive control. Strain K1026 was obtained from Professor Allen Kerr and Dr. Maarten Ryder at the University of Adelaide, South Australia.
Biocontrol of P. humilis crown gall disease by strain K1026 was performed in a greenhouse. The pathogenic strains and K1026 were cultured in Luria–Bertani (LB) and YEM medium for 1–2 days, respectively. The bacteria were collected by centrifugation and resuspended in 0.85% normal saline and the quantity was adjusted to 1.2 × 108 CFU/mL. The concentration of K1026 and pathogenic strains was adjusted by centrifugation and concentration (4 °C, 7,104 × g, 10 min). Appropriate volumes of the two bacterial solutions were mixed so that the number ratios of K1026 to pathogenic bacteria were 100:1, 10:1, 1:1, 1:10 or 1:100. The stems of 4-month-old P. humilis cuttings were stabbed with a blunt needle, and 3 μL of the bacterial mixture was inoculated on to the wound. Treatments inoculated with only K1026 were labeled ∞:0 and those inoculated with only the pathogenic strains were labeled 0:∞.
To investigate whether the inoculation order of pathogenic strains and K1026 affected the biological control effectiveness, we set up three treatments: K1026 inoculation before pathogenic strains, after pathogenic strains, and both bacteria inoculated at the same time. The gradients of advance inoculation time for K1026 was 12, 4, and 0.5 h, while the gradients of delayed inoculation time for K1026 was 10 and 30 min. The pathogenic strains were mixed with K1026 in equal proportions and inoculated at the wound site for simultaneous inoculation. The treatment inoculated with pathogenic bacteria was used as a positive control and the treatment inoculated with sterile water was used as a negative control. Five to ten cuttings were inoculated in each treatment and one spot was inoculated on each cutting. After inoculation, cuttings were placed in a greenhouse for normal management, and the bottom watering method was used to reduce potential interference from spraying water on the inoculation sites. After 50 days, the number of sites with galls was investigated. The ratio of the number of plants with galls to the number of plants inoculated was defined as the incidence rate.
Amplification of genes related to opine metabolism
The nopaline metabolism region contains four genes, ocd (coding for ornithine cyclodeaminase), arc (coding for arginase), noxA, and noxB, whose functions are closely related to the oxidative cleavage of nopaline (Lintig, 1991). There are eight genes in the loci related to agrocinopine A uptake and metabolism: accA–G and accR (Kim & Farrand, 1997). The related gene sequences of pTiC58 were downloaded from NCBI and used as templates to design primers using the NCBI primer BLAST. Primer sequences and optimum annealing temperatures are listed in Table S1. The amplified products were sent to the BGI TECH SOLUTIONS (BEIJING LIUHE) CO., LIMITED for sequencing. Multiple sequence alignment and phylogenetic analysis were performed with the gene sequences of related species according to previously described methods.
Results
Isolation and identification
On 1A and 2E selective agar media, we isolated 231 bacterial colonies from tumors and soils collected from five different regions of China (Fig. 1). Of these isolates, 134 were identified as Agrobacterium spp. using 16S rRNA gene analysis. We selectively amplified the virD2 gene and the cytokinin synthesis gene (ipt). Both of these genes are essential for tumorigenesis, and they are found on the Ti plasmids of tumorigenic strains. Duplex PCR with ipt and virD2 primers differentiated the pathogenic isolates. A total of 45 strains yielded two specific amplification products, indicating the presence of both the virD2 and ipt genes in the T-DNA region. These results agree with those of the pathogenicity tests, which showed that all 45 strains were capable of causing crown gall symptoms in various plant species (Fig. 1). Phenotypic characteristics of the 45 tumorigenic isolates are shown in Table 1. Most of the tumorigenic isolates had physiological characteristics similar to those of previously described Agrobacterium/Rhizobium spp. strains (Zahradník et al., 2018).
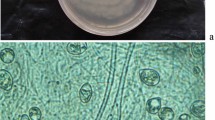
Culture characteristics and disease symptoms of pathogenic strains isolated from P. humilis. A: Results of double PCR amplification of virulence-related genes ipt and virD2. The first column is DL 2000 Marker. Columns 2–12 show amplification results for different strains. X-AA2 and X-AB3 were nontumorigenic strains. B: Colony morphology of strains on 1A selective plates. C: Colony morphology of strains on 2E selective plates. D–G: Disease symptoms on artificially inoculated plant (D: Carrot disc – upper discs were inoculated with tumorigenic strain X-AB2, lower were treated with nonpathogenic strain X-AA2; E: Tomato; F: Sunflower; G: P. humilis.) with A. tumefaciens strain X-AB2 isolated from P. humilis crown galls
Phylogenetic analysis of housekeeping gene
By phylogenetic analysis of the 16S rRNA, rpoB, atpD and recA genes, the isolates of the tumorigenic strains from P. humilis were identified. Genomic DNA of 45 pathogenic strains were extracted, and target genes were amplified by specific primers. All the sequences obtained were confirmed by BLAST and deposited in the NCBI database. Accession numbers (including the reference strains) are listed in Table S2. Amplification of the 16S rRNA gene produced 1465 bp DNA fragment. BLAST searches revealed that 16S rRNA of the 45 isolates had homology of more than 98% with A. tumefaciens species complex, as well as R. rhizogenes, A. rubi, A. rosae, and A. larrymoorei in the NCBI GenBank database. The phylogenetic tree based on 16S rRNA gene sequences confirmed the BLAST results (Fig. 2). The entire dataset was clearly divided into three groups: A. tumefaciens species complex (group A), A. vitis (group B) and R. rhizogenes (group C). The bootstrap values of the three groups were higher than 97%, confirming the reliability of the topological structure. P. humilis-derived pathogenic strains clustered within group A (33 strains) and group C (12 strains). However, several strains that did not belong to the A. tumefaciens species complex were clustered into group A, including A. rubi strain IFO 13261 T, A. skierniewicense strain Ch11T, and A. rosae strain NCPPB 1650 T. In addition, some strains in group A were not accurately identified at the species level. For example, strains X-AC1, X-AC2, X-AC3, X-AB1, X-AB2, X-A71, X-A73, X-A74, X-A75, X-A78, X-A82 and A. nepotum 39/7 T, A. fabacearum CNPSo 675 T, A. arsenijevicii KFB 330 T clustered on the same branch, indistinguishable from each other. This means that for the closely related A. tumefaciens species complex, the 16S rRNA gene alone could not be used to distinguish isolates at the species level.
Maximum likelihood phylogeny tree of the tumorigenic strains isolated from P. humilis based on 16S rRNA gene sequences. All strains beginning with X-A were isolated in this study. The symbols with different shapes in front of the isolates represent their geographic origins: green squares indicate Jinzhong City, Shanxi Province; black squares indicate Changli County, Hebei Province; blue triangles indicate Qinhuangdao City, Hebei Province; red triangles indicate Tai'an City, Shandong Province; red circles indicate Kaiyuan City, Liaoning Province. The Tamura 3-parameter model was used with gamma-distributed invariant sites (G + I). Bootstrap values > 70% are indicated at the nodes (1,000 replicates). Bradyrhizobium diazoefficiens strain USDA 110.T was used as an outgroup. All strains' accession numbers are listed in Table S2
To identify the species of the 33 strains in the A. tumefaciens species complex, we analyzed partial sequences of three housekeeping genes: rpoB, atpD and recA. We performed phylogenetic analysis using each single gene and the concatenated sequences of all three genes. In some cases, multiple strains had identical sequences for all three genes. To avoid redundancy, we selected only one strain from each group with identical sequences for the phylogenetic analysis. These strains were often isolated from the same location (Table S3).
The phylogenetic tree based on rpoB gene sequence using the maximum likelihood method is shown in Fig. 3. Available from the NCBI database, the dataset includes 14 taxa in the A. tumefaciens species complex (including 4 species without binomial names) and the related species A. arsenijevicii, which could not be distinguished by 16S rRNA. The "A. albertimagni" strain AOL15T was used as an outgroup. We observed that the rpoB gene is a more effective classifier of the A. tumefaciens species complex than 16S rRNA. Additionally, P. humilis-derived pathogenic strains are closely related to A. radiobacter and A. fabacearum. Strains represented by X-A70, X-AA1, and X-A94 clustered with A. radiobacter, strains represented by X-AB1 and X-A71 clustered with A. fabacearum, and strains represented by X-A234 and X-A282 clustered alone in another branch.
Maximum likelihood phylogeny tree of Agrobacterium spp. strains isolated from P. humilis based on rpoB gene sequences. All strains beginning with X-A were isolated in this study. The symbols with different shapes in front of the isolates represent their geographic origins: green squares indicate Jinzhong City, Shanxi Province; black squares indicate Changli County, Hebei Province; blue triangles indicate Qinhuangdao City, Hebei Province; red triangles indicate Tai'an City, Shandong Province; and red circles indicate Kaiyuan City, Liaoning Province. The Tamura 3-parameter model was used with gamma-distributed invariant sites (G + I). Bootstrap values > 70% are indicated at the nodes (1,000 replicates). "A. albertimagni" strain AOL15.T was used as an outgroup. All strains accession numbers are shown in Table S2
Phylogenetic trees based on atpD (Fig. 4) and recA (Fig. 5) gene sequences had very similar topologies. Similar to the tree established based on the rpoB gene, these trees also indicated that P. humilis-derived pathogenic strains were closely related to A. radiobacter and A. fabacearum. In addition, the strains represented by X-A234 and X-A282 did not cluster alone on another branch, but clustered together with A. radiobacter on a large branch. The recA-based tree had a higher bootstrap support of 99% on this node, while the atpD-based tree had a lower bootstrap support of 74% on this node.
Maximum likelihood phylogeny tree of Agrobacterium spp. strains isolated from P. humilis based on atpD gene sequences. All strains beginning with X-A were isolated in this study. The symbols with different shapes in front of the isolates represent their geographic origins: green squares indicate Jinzhong City, Shanxi Province; black squares indicate Changli County, Hebei Province; blue triangles indicate Qinhuangdao City, Hebei Province; red triangles indicate Tai'an City, Shandong Province; and red circles indicate Kaiyuan City, Liaoning Province. The Tamura 3-parameter model was used with gamma-distributed invariant sites (G + I). Bootstrap values > 70% are indicated at the nodes (1,000 replicates). "A. albertimagni" strain AOL15.T was used as an outgroup. All strains' accession numbers are shown in Table S2
Maximum likelihood phylogeny tree of Agrobacterium spp. strains isolated from P. humilis based on recA gene sequences. All strains beginning with X-A were isolated in this study. The symbols with different shapes in front of the isolates represent their geographic origins: green squares indicate Jinzhong City, Shanxi Province; black squares indicate Changli County, Hebei Province; blue triangles indicate Qinhuangdao City, Hebei Province; red triangles indicate Tai'an City, Shandong Province; and red circles indicate Kaiyuan City, Liaoning Province. The Tamura 3-parameter model was used with gamma-distributed (G). Bootstrap values > 70% are indicated at the nodes (1,000 replicates). "A. albertimagni" strain AOL15.T was used as an outgroup. All strains’ accession numbers are shown in Table S2
Compared with the tree established by each single gene, the phylogenies with the three concatenated genes (2121 bp) had the highest bootstrap support (Fig. 6). In the group of A. tumefaciens species complex, P. humilis-derived pathogenic strains were classified into two species: strains represented by X-A70, X-AA1, X-A94, X-A234 and X-A282 were A. radiobacter with a bootstrap value of 100%. The strains represented by X-AB1 and X-A71 were A. fabacearum with a bootstrap value of 100%.
Maximum likelihood phylogeny tree of Agrobacterium spp. strains isolated from P. humilis based on the concatenated gene sequences of rpoB, atpD and recA genes. All strains beginning with X-A were isolated in this study. The symbols with different shapes in front of the isolates represent their geographic origins: green squares indicate Jinzhong City, Shanxi Province; black squares indicate Changli County, Hebei Province; blue triangles indicate Qinhuangdao City, Hebei Province; red triangles indicate Tai'an City, Shandong Province; and red circles indicate Kaiyuan City, Liaoning Province. The General Time Reversible model was used with gamma-distributed invariant sites (G + I). Bootstrap values > 70% are indicated at the nodes (1,000 replicates). "A. albertimagni" strain AOL15.T was used as an outgroup. All strains' accession numbers are shown in Table S2
Based on the phylogenetic analysis of 16S rRNA, rpoB, atpD and recA genes, it can be concluded that the pathogenic strains isolated from P. humilis from five regions in China included three species: A. radiobacter, A. fabacearum and R. rhizogenes. Details of the strains included in each species were shown in Table S4.
From a geographic point of view, the strains causing P. humilis crown gall disease in different locations have own their characteristics. R. rhizogenes was found only in Tai'an, Shandong Province, while A. tumefaciens was isolated from the other four sites. A. radiobacter was the most widely distributed and was found in all five sites described here, while A. fabacearum was relatively less distributed and only found in Jinzhong City, Shanxi Province, and Kaiyuan City, Liaoning Province.
Determination of plasmid type
We investigated the opine synthesis genes of the 45 pathogenic strains using PCR amplification. PCR amplification for the octopine synthetase gene (ocs) and agropine synthetase gene (ags) resulted in negative amplification for all isolates. Only five isolates showed positive amplification of the nopaline synthetase gene (nos) (Table 2). Sequence alignment results showed that they were identical to the nos gene of the typical nopaline-type strain, “A. fabrum” C58T. This was surprising as previous studies have shown that the plasmids of crown gall pathogens isolated from the host of stone fruit trees are generally the nopaline type (Puławska et al., 2016; Wei et al., 2020). However, 89% of the isolates in this study could not be amplified with the nos gene primers. To address potential nonspecific binding between the template and primer due to DNA structure or sequence differences near the nos gene in our isolates, we selected NRB primer pairs for amplification. NRB primers were used to specifically amplify the right border of nopaline-type Ti plasmids. However, only “A. fabrum” C58 T and the five strains that showed positive nos amplification results mentioned above yielded the target band. To further clarify whether our isolates contained nopaline-type plasmids, we amplified nopaline metabolism genes. In general, plasmids that produce a particular opine also carry genes that degrade this opine. Of the four genes required for nopaline metabolism (noxA, noxB, arc, and ocd), we obtained positive amplification for only five strains, which were the same strains that previously showed positive nos amplification. “A. fabrum” C58 T, as a control, also showed positive amplification (Table 3). Twenty-five strains did not show any positive amplification. The other 15 strains showed partial amplification. Subsequently, we amplified the trans-zeatin synthesizing gene (tzs), which has been reported to be exclusive to nopaline-type plasmids (Regier & Morris, 1982). The amplification results showed that all 45 isolates had a specific PCR product of 998 bp for tzs. The sequence similarity was higher than 99.72% with the typical nopaline type strain, “A. fabrum” C58T.
We also amplified the agrocinopine synthetase (acs) gene. In recent studies, Tanaka classified acs genes into two groups based on phylogenetic analysis, with group 1 being associated with agrocinopine A and B synthesis and group 2 being associated with agrocinopine C and D synthesis. Group 1 contained two branches: acs1 and acs2. Some strains contained both acs1 and acs2 and some strains contained only acs1. The function of acs2 remains unknown (Tanaka et al., 2022). Agrocinopine C/D was specifically synthesized using agropine-type Ti-plasmids. Our isolates did not contain the agropine synthetase gene; therefore, we reasoned that they could not produce agrocinopine C/D. All isolates were positively amplified using specific primers for agrocinopine A/B synthetase. These results indicate that all crown gall strains isolated from P. humilis are likely to synthesize agrocinopine A/B. All isolates contained acs1 but only eight also contained acs2. The similarity between the two acs sequences of our isolates and the two acs of strain C58T was 100%.
Based on these data, we concluded that only five of the 45 pathogenic strains isolated from P. humilis were most likely nopaline plasmids. This is because they were most similar to the typical nopaline plasmid pTiC58 in terms of the PCR amplification profiles of the opine synthesis and metabolism genes. The other 40 strains did not contain nopaline plasmids. We also showed that they did not harbor octopine or agropine plasmids. Because these 40 strains all contained tzs and acs and some also contained noc-like genes, we suggest that they are closely related to nopaline plasmids.
Sensitivity to agrocin 84
When the 45 pathogenic strains were tested against agrocins produced by K1026 on Stonier's plates, none of the plates formed the expected inhibition zones, although the positive control strain “A. fabrum” C58T did. The experiment was repeated at least five times. All zones of C58 T were 50–65 mm in diameter with a sharp edge (Fig. 7). This result was significant; previous studies have shown that strains carrying nopaline/agrocinopine Ti plasmids are sensitive to agrocin 84 (Ellis, 2017). In this study, all strains contained tzs and acs genes, and five strains had positive nos amplification. However, all of them had resistance to antibiotics produced by K1026.
Bioassay of agrocin 84 sensitivity of selected Agrobacterium strains. Agrocin 84 production strain K1026 was inoculated in the center of Stonier’s medium plates. The upper layer was covered with a suspension of pathogenic strains in soft agars, with the P. humilis-derived strain X-AB2 on the left and the positive control strain C58 on the right
Characterization of acc genes
To explain the strains’ resistance to the antibiotics produced by K1026, we performed genetic analysis of the acc locus, which encodes uptake and catabolism of agrocinopines A and B and controls sensitivity to agrocin 84 produced by strain K1026. There are eight genes in this locus, namely accA–G and accR (Ellis, 2017; Kim & Farrand, 1997). We suspect that the gene sequences of P. humilis-derived strains at the acc locus are significantly different from that of the typical nopaline-type strain, and this difference leads to the resistance to agrocin 84. Therefore, in this study, eight primers were designed based on the acc genes of “A. fabrum” C58T to amplify accA-G and accR, respectively. Amplification results showed that nine strains could amplify all eight acc genes, and the other 36 strains could not amplify two or more. Based on the amplification results, the 45 strains were divided into seven types (Table 4). These types were independent of the geographic origin nor the species. Both R. rhizogenes and A. tumefaciens had unamplified acc genes. These results confirmed that the 45 pathogenic strains from P. humilis had significant differences with the typical nopaline-type strains at the acc locus. Among them, 36 strains were so different from C58T on accA-accE and accF that they did not have specific amplification on one or more genes, and these six genes have been proved to be indispensable in mediating the sensitivity of the nopaline-type strain to agrobacterin 84 (Hayman et al., 1993; Kim & Farrand, 1997). However, this difference does not fully explain the agrocin 84 resistance observed in this study. Nine strains were able to amplify all acc genes, and the PCR product sequencing results showed that their acc genes were 98.16% to 100% similar to the acc genes of C58T, however they were still resistant to agrocin 84 in vitro. So, we think that the acc locus is only one aspect of mediating agrocin 84 sensitivity, and there are other reasons at play.
Biological control with K1026 on P. humilis crown gall disease
Although the P. humilis-derived pathogenic strains were not sensitive to K1026 on the plate, we conducted greenhouse experiments to evaluate the biological control effect of K1026 on P. humilis crown gall disease. We selected strains belonging to three species (A. fabacearum, A. radiobacter, and R. rhizogenes) to make the evaluation more comprehensive. When only the pathogenic strains X-A282, X-AB1, and X-A271 were inoculated, the incidence of crown gall disease on P. humilis cuttings was 100% (Table 5). When the strain was inoculated with K1026 at a 1:1 ratio, tumor formation by X-A271 was completely inhibited and the incidence of galls with X-AB1 was reduced by 25%. Strain X-A282 was more difficult to control, and its incidence was still 100%. However, if the concentration of K1026 was increased to 100 times that of the pathogenic strains, the incidence of crown gall disease caused by X-A282 was reduced to 75% and tumor formation by X-AB1 was completely inhibited.
We also examined the effect of the inoculation sequence of K1026 and the pathogenic strains on the incidence (Table 6). When the two were inoculated at the wound site on P. humilis stems at a ratio of 1:1, pre-inoculated K1026 helped reduce the incidence and improve the effect of control. As long as the biocontrol strain K1026 was inoculated 0.5 h before pathogenic strain inoculation, the incidence of X-A282, the most pathogenic strain, could be reduced from 100 to 71.4% and that of strain X-AB1 from 75 to 62.5%. In contrast, when K1026 was inoculated later than the pathogenic strains, the crown gall incidence significantly increased, with even a 10 min delay increasing the incidence from 0 to 44.4% (X-A271) and 75 to 100% (X-AB1). When the delayed inoculation time of K1026 was extended to 0.5 h, the incidence of the three strains increased to 100% and the biocontrol effect was significantly reduced.
These results confirmed that although the pathogenic strains from P. humilis were not sensitive to agrocin produced by K1026 on plates, K1026 still had a certain control effect against P. humilis crown gall disease. Increasing the concentration of K1026 and inoculating K1026 before exposure to pathogenic strains can effectively reduce the incidence and improve the control effect. It should also be noted that K1026 had different control effects on different strains. For example, for R. rhizogenes X-A271, it can achieve 100% control efficiency. However, for A. radiobacter X-A282, only a maximum effectiveness of 25% was achieved.
Discussion
There are three crown gall causal species of P. humilis in China, and the A. tumefaciens species complex is the primary
To the best of our knowledge, there is only one published study on P. humilis crown gall disease, in which four strains of A. radiobacter were isolated and identified from a nursery in Ningxia Hui Autonomous Region, China (Bai et al., 2022). However, the study’s sample size is too small to represent the causal species classification and diversity of P. humilis crown gall disease.
Many studies on closely related species of P. humilis (Wang et al., 2015), such as plums, apricots, sweet cherries, and peaches, have shown that in addition to A. tumefaciens, R. rhizogenes is also isolated from the grown gall or rhizosphere soils of these trees. According to available data, the proportions of the two groups isolated from different hosts in different countries vary greatly. In New Zealand and Australia, the A. tumefaciens species complex dominates peaches, almonds, and plums (Keane et al., 1970). In Poland, Puławska obtained 33 pathogenic strains from cherry, peach, and plum in five different locations, of which 31 were R. rhizogenes (Joanna Puławska & Kałużna, 2012). In Algeria, 96 tumorigenic bacteria have been obtained from peach, of which R. rhizogenes accounts for 80%. From apricots and almonds, 25 strains were obtained, all belonging to the A. tumefaciens species complex. For cherry and plum, the A. tumefaciens species complex accounted for 59 and 50%, respectively (Bouzar et al., 1991). In China, Hao investigated pathogenic strains in peach galls from six different regions and found that 86% of them were R. rhizogenes (Hao et al., 2019). More than 100 strains were isolated from gall tissue and rhizosphere soil of susceptible cherries in Tai'an, Shandong province, and identified more than 96% of the strains as R. rhizogenes (Wei et al., 2020).
Therefore, it is difficult to determine whether A. tumefaciens or R. rhizogenes prefers these species. In our study, diseased P. humilis were collected from five regions in China, all of which are major cultivation areas and representative of P. humilis production. From these materials, we isolated 45 tumorigenic strains. Among these, 33 strains (73%) belonged to the A. tumefaciens species complex. All R. rhizogenes strains were isolated only from Tai'an City, Shandong Province. In general, the A. tumefaciens species complex is the main species causing crown gall in P. humilis in China.
Based on phylogenetic analysis of the 16S rRNA, rpoB, atpD and recA gene, we identified three species in the gall or rhizosphere soil of P. humilis in China namely R. rhizogenes, A. fabacearum and A. radiobacter. A. fabacearum and A. radiobacter are often found in crown galls or rhizosphere soils of stone fruits (Costechareyre et al., 2010; Puławska et al., 2016). They are closely related and phenotypic characteristics cannot distinguish between them. Previous studies have shown that the 16S rRNA gene is highly conserved in the Agrobacterium spp., making it difficult to identify them to a species level (Delamuta et al., 2020; Singh et al., 2021). In our study, the results were consistent with those previous works. R. rhizogenes could be well identified by 16S rRNA gene. But within the A. tumefaciens species complex, the 16S rRNA gene could not distinguish the strains at the species level, nor related species A. rubi, A. skierniewicense and A. rosae. Phylogenetic analysis based on other housekeeping genes helps to identify them. In this study, three housekeeping gene (rpoB, atpD and recA) provided a much better phylogenetic relationship among Agrobacterium spp. The rpoB-based phylogenetic tree indicates that the strains represented by X-A234 and X-A282 cluster in a single branch, suggesting they may have separate taxonomic status. However, the phylogenetic trees based on atpD and recA showed that these strains clustered together with A. radiobacter. The MLSA tree proved this again with a bootstrap value of 100%, indicating that MLSA is an efficient way to identify Agrobacterium species.
We also observed differences according to geographic regions. R. rhizogenes was only found in samples from Tai'an, A. fabacearum was only found in samples from Kaiyuan and Jinzhong, and A. radiobacter could be isolated from samples collected in all the five regions. Moreover, within the same species, there were differences between different regions, which could be clearly seen from the phylogenetic tree based on the concatenated genes sequences. These results indicate the high diversity and complexity of the species causing crown gall of P. humilis in China.
The opine synthesis and metabolism genes of the P. humilis-derived pathogenic strains possess unique characteristics
Previous studies have shown that the plasmids of crown gall pathogens isolated from the host of stone fruit trees are generally nopaline type ( Puławska et al., 2016; Wei et al., 2020). We used several pairs of widely used primers to amplify the genes related to opine synthesis, including nopaline synthetase gene (nos), octopine synthetase gene (ocs), agrocinopine synthetase gene (acs) and agropine synthetase gene (ags). The results showed that ocs and ags could not be amplified in all 45 pathogenic strains from P. humilis, while nos could be amplified in five strains (X-A70, X-A81, X-A234, X-A252, X-A254) and acs could be amplified in all strains. We also amplified the trans-zeatin synthesizing gene (tzs), and all 45 pathogenic strains from P. humilis obtained target products. The presence of the tzs gene is a typical feature of nopaline-type plasmids. Those strains that carry octopine or agropine plasmids did not contain it (Beaty et al., 1986; Regier & Morris, 1982). Thus, PCR amplification of opine synthetase and tzs gene indicated that five of the 45 pathogenic strains isolated from P. humilis were most likely nopaline plasmids, and the other 40 strains are closely related to nopaline plasmids.
The primers used in this paper to amplify nos and the T-DNA right border of nopaline-type Ti plasmids were designed based on the conserved region of the typical nopaline-type strain C58T. Only those strains with high similarity to the corresponding gene sequences of C58T could obtain specific amplification. In previous studies, it has been reported that nos genes in nopaline-type strains do not seem to be conserved. Shao pointed out that the nos gene of nopaline-type plasmid pTikerr108 had only 85% nucleotide similarity with pTiC58 through plasmid sequencing results (Shao et al., 2019). Kuzmanović isolated two R. rhizogenes tumorigenic strains C5.7 and C6.5 from the same crown gall on the "Colt "cherry rootstock, which carried the nopaline-type Ti plasmid pTiC5.7/pTiC6.5. Comparative genomic analysis showed that the genes on the right side of T-DNA of pTiC5.7/pTiC6.5 (iaaH-to-nos) had less than 94% nucleotide similarity with the genes carried by pTiC58. The nos gene of pTiC5.7/pTiC6.5 shared only 88% of its nucleotides with that of pTiC58 (Kuzmanović & Puławska, 2019). When Hwang used the nos primers in this paper to identify the opine type of five tumorigenic A. tumefaciens strains, only three strains had positive amplification, and the other two strains (1D1108 and 1D132) could not be amplified. He later confirmed that these five strains were nopaline-type plasmids (Hwang et al., 2013). In our study, PCR product sequencing of the nos gene in the X-A70 strains showed 100% similarity to the corresponding sequences of C58T, suggesting that these strains are most similar to the typical nopaline-type strain. However, the other 40 strains could not obtain positive amplification of nos, nor could they obtain positive amplification of T-DNA right border, indicating that the nos genes of these 40 strains were less similar to those of C58T.
Similar phenomena were observed in the amplification of genes related to nopaline metabolism. The four pairs of noc-related gene amplification primers used in this paper were also designed with C58T as the target. Only the five strains represented by X-A70 amplified all four noc-related genes; 25 of the other 40 strains did not get any amplification, and some genes in 15 strains were specifically amplified. Although some studies have indicated that the noc region of the nopaline-type plasmid is strongly conserved (Shao et al., 2019), some differences were observed in the noc region of the 45 strains in this study compared with the typical nopaline-type strain C58T.
There are some unique characteristics in the sensitivity to agrocin 84 of the P. humilis-derived pathogenic strains
All P. humilis-derived strains contained acs1, and some strains also contained acs2. Phylogenetic analysis (Fig. S1) revealed that the sequences of these strains were closely related to typical nopaline-type strains and formed a cluster with a group described by Tanaka et al. (2022). This group of strains is responsible for the production of agrocinopines A and B in tumors. Strains that produce agrocinopines A and B are typically sensitive to agrocin 84 (Kim & Farrand, 1997). However, none of the strains obtained from P. humilis showed sensitivity in vitro. The functions most closely related to agrocin 84 sensitivity are the agrocinopine A/B uptake and metabolism operon (Kim & Farrand, 1997). It is located on the Ti plasmid and is called the acc locus, which is composed of eight genes (accA-accF, accG, and accR) (Hayman & Farrand, 1988). accA-accE are required for the transport of agrocin 84 and agrocinopine A/B, and is responsible for the susceptibility to agrocin 84 (Hayman et al., 1993). Mutations in accF and accG, which are responsible for metabolic opines, lead to the loss of the ability to grow on agrocinopine A/B as the sole carbon source. The accG mutation had no effect on agrocin 84 sensitivity. Although accF mutants can still take up agrocin 84, they are resistant to it, which means that agrocin 84 must be activated by its accF-encoded function to exert its toxic effect (Kim & Farrand, 1997). accR encodes a transcriptional repressor that acts as a regulator of the acc locus and the conjugal transfer of the Ti plasmid. accR mutations lead to the constitutive expression of acc and tra (Beck von Bodman et al., 1992). Insertional mutations in accR result in hypersensitivity to agrocin 84 (Kim & Farrand, 1997). In conclusion, these results suggest that accA-accE and accF are required for sensitivity to agrocin 84(Kim & Farrand, 1997).
In this study, eight primers were designed based on the acc genes of the typical nopaline-type strain “A. fabrum” C58T. When using these primers to amplify 45 pathogenic strains of P. humilis, 36 strains were found to have one or more negative amplification of accA-accE and accF, which could explain the resistance of these 36 strains to agrocin 84 in vitro. It also showed that the acc genes of the pathogenic strains of P. humilis have some differences from that of the typical nopaline-type strain “A. fabrum” C58T. The other nine strains were able to obtain positive amplification of all acc genes, and these genes had 98.16% to 100% similarity to the acc genes of C58T, but they were still resistant to agrocin 84 in vitro. Therefore, we believe that acc locus is only one aspect of mediating agrocin 84 sensitivity, and there are other reasons that play an important role.
The observed characteristics captured our attention. Both the synthesis and metabolism of opine and sensitivity to agrocin 84 are directly linked to the pathogen’s virulence and biocontrol. Given the rising incidences of P. humilis crown gall disease in large-scale cultivation, it is crucial to establish a connection between these characteristics and crown gall control measures during production.
Although we did not detect the desired inhibition zone under pure culture conditions, K1026 showed 100% control efficacy against two P. humilis-derived pathogenic strains in a greenhouse experiment. This was not surprising as past studies have reported that strain K1026 can control pathogenic strains that are resistant to agrocin 84 (López et al., 1989). The biological control of K1026 has a complex mechanism, with the production of agrocin being only one part (Kerr, 2016). The mechanism by which strain K1026 controls agrocin 84 resistant strains has not been fully defined (Penyalver et al., 2000). Regardless, 45 pathogenic strains isolated from P. humilis rhizosphere soils and crown galls collected in five regions of China were resistant to agrocin 84. Based on these results and the fact that K1026 has not been approved for sale in China, new biocontrol strains are urgently needed to prevent the outbreak of P. humilis crown gall disease.
In conclusion, we isolated and identified 45 pathogenic strains of P. humilis crown gall disease from five regions in China, including R. rhizogenes, A. fabacearum, and A. radiobacter. The A. tumefaciens species complex accounted for the majority of species. The pathogens that cause crown gall in P. humlilis have some unique characteristics in the synthesis and metabolism of opines. We speculate that during the long period of coevolution between Agrobacterium/Rhizobium and their specific host P. humilis, the strains developed mechanisms different from those of its ancestors to coordinate the synthesis and metabolism of opines, among other unknown aspects. Since our study was primarily based on PCR amplification of specific genes, further investigations remain warranted to fully assess the Ti plasmid of P. humilis pathogenic strains. In future studies, we intend to sequence the Ti plasmids to investigate the similarities, differences, and evolutionary relationships between these and other Ti plasmids, which should provide novel insights into the diversity of Ti plasmids of Agrobacterium/Rhizobium and ultimately help to develop more effective measures to prevent and manage crown gall disease.
References
Aujoulat, F., Jumas-Bilak, E., Masnou, A., Salle, F., Faure, D., Segonds, C., et al. (2011). Multilocus sequence-based analysis delineates a clonal population of Agrobacterium (Rhizobium) radiobacter (Agrobacterium tumefaciens) of human origin. Journal of Bacteriology, 193(10), 2608–2618. https://doi.org/10.1128/JB.00107-11
Bai, J., Yao, T., Lan, X., Yang, Y., Wang, Z., & Wang, X. (2022). Isolation of Agrobacterium tumefaciens/ biovar 1 from the crown gall of Cerasus humilis in China. Journal of Plant Pathology, 104(1), 141–148. https://doi.org/10.1007/s42161-021-00947-6
Beaty, J. S., Powell, G. K., Lica, L., Regier, D. A., MacDonald, E. M. S., Hommes, N. G., & Morris, R. O. (1986). Tzs, a nopaline Ti plasmid gene from Agrobacterium tumefaciens associated with trans-zeatin biosynthesis. Molecular and General Genetics MGG, 203(2), 274–280. https://doi.org/10.1007/BF00333966
Beck von Bodman, S., Hayman, G. T., & Farrand, S. K. (1992). Opine catabolism and conjugal transfer of the nopaline Ti plasmid pTiC58 are coordinately regulated by a single repressor. Proceedings of the National Academy of Sciences, 89(2), 643–647. https://doi.org/10.1073/pnas.89.2.643
Bouzar, H., Daouzli, N., Krimi, Z., Alim, A., & Khemici, E. (1991). Crown gall incidence in plant nurseries of Algeria, characteristics of Agrobacterium tumefaciens strains, and biological-control of strains sensitive and resistant to agrocin 84. Agronomie, 11(10), 901–908. https://doi.org/10.1051/agro:19911007
Brisbane, P., & Kerr, A. (1983). Selective media for three biovars of Agrobacterium. Journal of Applied Bacteriology, 54(3), 425–431. https://doi.org/10.1111/j.1365-2672.1983.tb02638.x
Costechareyre, D., Rhouma, A., Lavire, C., Portier, P., Chapulliot, D., Bertolla, F., et al. (2010). Rapid and efficient identification of Agrobacterium species by recA allele analysis. Microbial Ecology, 60(4), 862–872. https://doi.org/10.1007/s00248-010-9685-7
Delamuta, J. R. M., Scherer, A. J., Ribeiro, R. A., & Hungria, M. (2020). Genetic diversity of Agrobacterium species isolated from nodules of common bean and soybean in Brazil, Mexico, Ecuador and Mozambique, and description of the new species Agrobacterium fabacearum sp. nov. International Journal of Systematic and Evolutionary Microbiology, 70(7), 4233–4244. https://doi.org/10.1099/ijsem.0.004278
Du, L., & Lu, F. (2011). Experimental technique in microbiology (1st ed.). China Light Industry Press.
Ellis, J. G. (2017). Can plant microbiome studies lead to effective biocontrol of plant diseases? Molecular Plant-Microbe Interactions, 30(3), 190–193. https://doi.org/10.1094/MPMI-12-16-0252-CR
Felsenstein, J. (1985). Confidence-limits on phylogenies - an approach using the bootstrap. Evolution, 39(4), 783–791. https://doi.org/10.2307/2408678
Fu, H., Mu, X., Wang, P., Zhang, J., Fu, B., & Du, J. (2020). Fruit quality and antioxidant potential of Prunus humilis Bunge accessions. Plos One, 15(12), e0244445. https://doi.org/10.1371/journal.pone.0244445
Fu, H., Qiao, Y., Wang, P., Mu, X., Zhang, J., Fu, B., & Du, J. (2021). Changes of bioactive components and antioxidant potential during fruit development of Prunus humilis Bunge. Plos One, 16(5), e0251300. https://doi.org/10.1371/journal.pone.0251300
Gaunt, M. W., Turner, S. L., Rigottier-Gois, L., Lloyd-Macgilp, S. A., & Young, J. P. (2001). Phylogenies of atpD and recA support the small subunit rRNA-based classification of rhizobia. International Journal of Systematic and Evolutionary Microbiology, 51(6), 2037–2048. https://doi.org/10.1099/00207713-51-6-2037
Gohlke, J., & Deeken, R. (2014). Plant responses to Agrobacterium tumefaciens and crown gall development. Frontiers in Plant Science, 5. https://doi.org/10.3389/fpls.2014.00155
Haas, J., Moore, L., Ream, W., & Manulis, S. (1995). Universal PCR primers for detection of phytopathogenic Agrobacterium strains. Applied and Environmental Microbiology, 61(8), 2879–2884. https://doi.org/10.1128/AEM.61.8.2879-2884.1995
Hao, F., Li, G., Wang, B., Cai, Z., You, Y., & Niu, S. (2019). Identification of pathogens and their virulence of crown gall disease of peach from different peach production regions. Guangdong Agricultural Sciences, 46(7), 86–91. https://doi.org/10.16768/j.issn.1004-874X.2019.07.013
Hayman, G. T., & Farrand, S. K. (1988). Characterization and mapping of the agrocinopine-agrocin 84 locus on the nopaline Ti plasmid pTiC58. Journal of Bacteriology, 170(4), 1759–1767. https://doi.org/10.1128/jb.170.4.1759-1767.1988
Hayman, G. T., Beck von Bodman, S., Kim, H., Jiang, P., & Farrand, S. K. (1993). Genetic analysis of the agrocinopine catabolic region of Agrobacterium tumefaciens Ti plasmid pTiC58, which encodes genes required for opine and agrocin 84 transport. Journal of Bacteriology, 175(17), 5575–5584. https://doi.org/10.1128/jb.175.17.5575-5584.1993
Holmes, B., & Roberts, P. (1981). The Classification, identification and nomenclature of agrobacteria. Journal of Applied Bacteriology, 50(3), 443–467. https://doi.org/10.1111/j.1365-2672.1981.tb04248.x
Hwang, H.-H., Wu, E. T., Liu, S.-Y., Chang, S.-C., Tzeng, K.-C., & Kado, C. I. (2013). Characterization and host range of five tumorigenic Agrobacterium tumefaciens strains and possible application in plant transient transformation assays. Plant Pathology, 62(6), 1384–1397. https://doi.org/10.1111/ppa.12046
Keane, P., Kerr, A., & New, P. (1970). Crown gall of stone fruit. 2. Identification and nomenclature of Agrobacterium isolates. Australian Journal of Biological Sciences, 23(3), 585-. https://doi.org/10.1071/BI9700585
Kerr, A. (2016). Biological control of crown gall. Australasian Plant Pathology, 45(1), 15–18. https://doi.org/10.1007/s13313-015-0389-9
Kersters, K., Deley, J., Sneath, P., & Sackin, M. (1973). Numerical taxonomic analysis of Agrobacterium. Journal of General Microbiology, 78(OCT), 227–239. https://doi.org/10.1099/00221287-78-2-227
Kim, H., & Farrand, S. K. (1997). Characterization of the acc operon from the nopaline-type Ti plasmid pTiC58, which encodes utilization of agrocinopines A and B and susceptibility to agrocin 84. Journal of Bacteriology, 179(23), 7559–7572. https://doi.org/10.1128/jb.179.23.7559-7572.1997
Kuzmanović, N., & Puławska, J. (2019). Evolutionary relatedness and classification of tumor-inducing and opine-catabolic plasmids in three Rhizobium rhizogenes strains isolated from the same crown gall tumor. Genome Biology and Evolution, 11(6), 1525–1540. https://doi.org/10.1093/gbe/evz091
Lassalle, F., Campillo, T., Vial, L., Baude, J., Costechareyre, D., Chapulliot, D., et al. (2011). Genomic species are ecological species as revealed by comparative genomics in Agrobacterium tumefaciens. Genome Biology and Evolution, 3, 762–781. https://doi.org/10.1093/gbe/evr070
López, M. M., Gorris, M. T., Salcedo, C. I., Montojo, A. M., & Miró, M. (1989). Evidence of biological control of Agrobacterium tumefaciens strains sensitive and resistant to agrocin 84 by different Agrobacterium radiobacter strains on stone fruit trees. Applied and Environmental Microbiology, 55(3), 741–746. https://doi.org/10.1128/aem.55.3.741-746.1989
Mafakheri, H., Taghavi, S. M., Puławska, J., de Lajudie, P., Lassalle, F., & Osdaghi, E. (2019). Two novel genomospecies in the Agrobacterium tumefaciens species complex associated with rose crown gall. Phytopathology, 109(11), 1859–1868. https://doi.org/10.1094/PHYTO-05-19-0178-R
Mafakheri, H., Taghavi, S. M., Khezerpour, K., Kuzmanović, N., & Osdaghi, E. (2022). Genomic analyses of rose crown gall-associated bacteria revealed two new Agrobacterium species: Agrobacterium burrii sp. nov. and Agrobacterium shirazense sp. nov. Phytopathology, 112(6), 1208–1213. https://doi.org/10.1094/PHYTO-11-21-0463-SC
Mousavi, S. A., Willems, A., Nesme, X., de lajudie, P., & Lindstrom, K. (2015). Revised phylogeny of Rhizobiaceae: Proposal of the delineation of Pararhizobium gen. nov., and 13 new species combinations. Systematic and Applied Microbiology, 38(2), 84–90. https://doi.org/10.1016/j.syapm.2014.12.003
Mu, X. P., Aryal, N., Du, J. M., Du, J. J., Brisbane, P., & Kerr, A. (2015). Oil content and fatty acid composition of the kernels of 31 genotypes of Chinese dwarf cherry [Cerasus humilis (Bge.) Sok.]. Journal of Horticultural Science & Biotechnology, 90(5), 525–529. https://doi.org/10.1080/14620316.2015.11668709TY
Ophel, K., & Kerr, A. (1990). Agrobacterium vitis sp. nov for strains of Agrobacterium biovar 3 from grapevines. International Journal of Systematic Bacteriology, 40(3), 236–241. https://doi.org/10.1099/00207713-40-3-236
Panday, D., Schumann, P., & Das, S. K. (2011). Rhizobium pusense sp nov., isolated from the rhizosphere of chickpea (Cicer arietinum L.). International Journal of Systematic and Evolutionary Microbiology, 61, 2632–2639. https://doi.org/10.1099/ijs.0.028407-0
Penyalver, R., Vicedo, B., & Lopez, M. (2000). Use of the genetically engineered Agrobacterium strain K1026 for biological control of crown gall. European Journal of Plant Pathology, 106(9), 801–810. https://doi.org/10.1023/A:1008785813757
Puławska, Joanna, & Kałużna, M. (2012). Phylogenetic relationship and genetic diversity of Agrobacterium spp. isolated in Poland based on gyrB gene sequence analysis and RAPD. European Journal of Plant Pathology, 133(2), 379–390. https://doi.org/10.1007/s10658-011-9911-2
Puławska, Joanna, Willems, A., De Meyer, S. E., & Süle, S. (2012). Rhizobium nepotum sp. nov. isolated from tumors on different plant species. Systematic and Applied Microbiology, 35(4), 215–220. https://doi.org/10.1016/j.syapm.2012.03.001
Puławska, J., Warabieda, W., & Ismail, E. (2016). Identification and characterization of bacteria isolated from crown galls on stone fruits in Poland. Plant Pathology, 65(6), 1034–1043. https://doi.org/10.1111/ppa.12482
Ran, B., Guo, C.-E., Zhang, Y., Han, C., Cao, T., Huang, H., et al. (2022). Preventive effect of Chinese dwarf cherry [Cerasus humilis (Bge.) Sok.] fermentation juice on dextran sulfate sodium-induced ulcerative colitis rats through the regulation of IgA and the intestinal immune barrier. Food & Function, 13(10), 5766–5781. https://doi.org/10.1039/D1FO04218A
Regier, D., & Morris, R. (1982). Secretion of trans-zeatin by Agrobacterium tumefaciens - a function determined by the nopaline Ti plasmid. Biochemical and Biophysical Research Communications, 104(4), 1560–1566. https://doi.org/10.1016/0006-291X(82)91429-2
Ryder, M., Tate, M., & Kerr, A. (1985). Virulence properties of strains of Agrobacterium on the apical and basal surfaces of carrot root disks. Plant Physiology, 77(1), 215–221. https://doi.org/10.1104/pp.77.1.215
Sawada, H., Ieki, H., Oyaizu, H., & Matsumoto, S. (1993). Proposal for rejection of Agrobacterium-tumefaciens and revised descriptions for the genus Agrobacterium and for Agrobacterium radiobacter and Agrobacterium rhizogenes. International Journal of Systematic Bacteriology, 43(4), 694–702. https://doi.org/10.1099/00207713-43-4-694
Shao, S., van Heusden, G. P. H., & Hooykaas, P. J. J. (2019). Complete sequence of succinamopine Ti-plasmid pTiEU6 reveals its evolutionary relatedness with nopaline-type Ti-plasmids. Genome Biology and Evolution, 11(9), 2480–2491. https://doi.org/10.1093/gbe/evz173
Singh, N. K., Lavire, C., Nesme, J., Vial, L., Nesme, X., Mason, C. E., et al. (2021). Comparative genomics of novel Agrobacterium G3 strains isolated from the international space station and description of Agrobacterium tomkonis sp. nov. Frontiers in Microbiology, 12, 765943. https://doi.org/10.3389/fmicb.2021.765943
Stonier, T. (1960). Agrobacterium tumefaciens conn ii: Production of an antibiotic substance. Journal of Bacteriology, 79(6), 889–898. https://doi.org/10.1128/jb.79.6.889-898.1960
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., & Kumar, S. (2011). Mega5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 28(10), 2731–2739. https://doi.org/10.1093/molbev/msr121
Tanaka, A., Ryder, M. H., Suzuki, T., Uesaka, K., Yamaguchi, N., Amimoto, T., et al. (2022). Production of agrocinopine a by Ipomoea Batatas agrocinopine synthase in transgenic tobacco and its effect on the rhizosphere microbial community. Molecular Plant-Microbe Interactions, 35(1), 73–84. https://doi.org/10.1094/MPMI-05-21-0114-R
Teyssier-cuvelle, S., Oger, P., Mougel, C., Groud, K., Farrand, S., & Nesme, X. (2004). A highly selectable and highly transferable Ti plasmid to study conjugal host range and Ti plasmid dissemination in complex ecosystems. Microbial Ecology, 48(1), 10–18. https://doi.org/10.1007/s00248-003-2023-6
Van Larebeke, N., Engler, G., Holsters, M., Van Den Elsacker, S., Zaenen, I., Schilperoort, R., & Schell, J. (1974). Large plasmid in Agrobacterium tumefaciens essential for crown gall inducing ability. Nature, 252(5479), 169–170. https://doi.org/10.1038/252169a0
von Lintig, J. (1991). Positive regulators of opine-inducible promoters in the nopaline and octopine catabolism regions of Ti plasmids. Molecular Plant-Microbe Interactions, 4(4), 370. https://doi.org/10.1094/MPMI-4-370
Wang, P., Li, L., Du, J., Zhang, J., Ji, W., Mu, X., & Cao, Q. (2015). Genetic diversity among wild Chinese dwarf cherry from different regions and its closely related plant species based on RAPD analysis. Journal of Plant Genetic Resource, 16(1), 119–126. https://doi.org/10.13430/j.cnki.jpgr.2015.01.019
Wang, P., Jia, L., Du, J., Zhang, J., Mu, X., & Ding, W. (2017). Improvement of soil quality by Chinese dwarf cherry cultivation in the Loess Plateau steep hill region. Acta Prataculturae Sinica, 26(03), 65–74.
Wang, P., Mu, X., Du, J., Gao, Y. G., Bai, D., Jia, L., et al. (2018). Flavonoid content and radical scavenging activity in fruits of Chinese dwarf cherry (Cerasus humilis) genotypes. Journal of Forestry Research, 29(1), 55–63. https://doi.org/10.1007/s11676-017-0418-3
Wei, Y., Maarten, R., Li, H., Li, J., Hu, J., & Yang, H. (2020). Identification and characterrization of pathogenic bacteria isolated from crown galls on cherry trees in Tai’an. Plant Protection, 46(6), 22–29. https://doi.org/10.16688/j.zwbh.2020124
Weisburg, W., Barns, S., Pelletier, D., & Lane, D. (1991). 16S ribosomal DNA amplification for phylogenetic study. Journal of Bacteriology, 173(2), 697–703. https://doi.org/10.1128/JB.173.2.697-703.1991
Yan, J., Li, Y., Han, X. Z., Chen, W. F., Zou, W. X., Xie, Z., & Li, M. (2017a). Agrobacterium deltaense sp nov., an endophytic bacteria isolated from nodule of Sesbania cannabina. Archives of Microbiology, 199(7), 1003–1009. https://doi.org/10.1007/s00203-017-1367-0
Yan, J., Li, Y., Yan, H., Chen, W. F., Zhang, X., Wang, E. T., et al. (2017b). Agrobacterium salinitolerans sp nov., a saline-alkaline-tolerant bacterium isolated from root nodule of Sesbania cannabina. International Journal of Systematic and Evolutionary Microbiology, 67(6), 1906–1911. https://doi.org/10.1099/ijsem.0.001885
Yin, ZePeng, Li, S., Ren, J., & Song, X. S. (2014). Role of spermidine and spermine in alleviation of drought-induced oxidative stress and photosynthetic inhibition in Chinese dwarf cherry (Cerasus humilis) seedlings. Plant Growth Regulation, 74(3), 209–218. https://doi.org/10.1007/s10725-014-9912-1
Yin, Zepeng, Ren, J., Zhou, L., Sun, L., Wang, J., Liu, Y., & Song, X. (2017). Water deficit mechanisms in perennial shrubs Cerasus humilis leaves revealed by physiological and proteomic analyses. Proteome Science, 15. https://doi.org/10.1186/s12953-017-0117-1
Young, J., Kuykendall, L., Martinez-Romero, E., Kerr, A., & Sawada, H. (2001). A revision ofRhizobium Frank 1889, with an emended description of the genus, and the inclusion of all species of Agrobacterium Conn 1942 and Allorhizobium undicola de Lajudie et al. 1998 as new combinations: Rhizobium radiobacter, R. rhizogenes, R. rubi, R. undicola and R. vitis. International Journal of Systematic and Evolutionary Microbiology, 51, 89–103. https://doi.org/10.1099/00207713-51-1-89
Zahradník, J., Nunvar, J., Pařízková, H., Kolářová, L., Palyzová, A., Marešová, H., et al. (2018). Agrobacterium bohemicum sp. nov. isolated from poppy seed wastes in central Bohemia. Systematic and Applied Microbiology, 41(3), 184–190. https://doi.org/10.1016/j.syapm.2018.01.003
Acknowledgements
We are grateful to Professor Allen Kerr and Dr Maarten Ryder at the University of Adelaide, South Australian for providing the Rhizobium rhizogenes strain K1026. We are indebted to their continued inspiration for crown gall disease research.
Author information
Authors and Affiliations
Contributions
Material preparation, data collection, verification and analysis were performed by Rong Xiao, Xiao-Peng Mu, Jian-Cheng Zhang, Shuai Zhang and Chun-Fen Zhang. Data analysis and software usage were performed by Rong Xiao and Shu Deng. Methodology, germplasm management, data management, project management were performed by Peng-Fei Wang. Methodology, project management and funding were performed by Jun-Jie Du. Manuscript writing was conducted by Rong Xiao with input from the remaining authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of financial or non-financial interest. This study is original, has not been published previously, and is not under consideration for publication elsewhere.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiao, R., Mu, XP., Zhang, JC. et al. Isolation and characterization of tumorigenic bacteria associated with crown gall disease of Prunus humilis Bunge in China. Eur J Plant Pathol 166, 463–483 (2023). https://doi.org/10.1007/s10658-023-02675-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-023-02675-2