Abstract
Diseases associated with tomato spotted wilt orthotospovirus (TSWV) are a serious threat to vegetable production worldwide. In 2019, leaf necrosis disease was observed on cucumber plants in Fukushima Prefecture, Japan. TSWV infection was identified in the diseased plants by immunostripe tests and RT-PCR analysis. Sequencing of the complete genome of this TSWV strain showed the highest identity with previously characterized TSWV isolates, and we therefore designated the cucumber-infecting variant as TSWV-Fukushima:cucumber 2019 (TS-FC19). Phylogenetic analyses based on the intergenic regions revealed that TS-FC19 and other TSWV isolates collected from the same area clustered in a distinct clade. Cucumber plants inoculated with TS-FC19 produced leaf necrosis symptoms, whereas other TSWV strains did not induce severe symptoms in this host. Together, our findings indicate that the cucumber leaf necrosis disease in Fukushima Prefecture is caused by a new strain of TSWV that appears to have evolved locally. This is the first detailed report on the biological characteristics of a cucumber-infecting TSWV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tomato spotted wilt orthotospovirus (TSWV) is a species of the genus Orthotospovirus in the family Tospoviridae, and is transmitted by several species of thrips (Family Thripidae) in a persistent-propagative manner (Rotenberg et al., 2015; Turina et al., 2016). All orthotospoviruses are transmitted by thrips (order Thysanoptera), and most of the orthotospovirus vectors are thrips in the family Thripidae, in particular, the genera Ceratothrioides, Dictyothrips, Frankliniella, Scirtothrips, Sericothrips and Thrips (Rotenberg et al., 2015). TSWV has an extremely wide range of host plants, with more than 900 species reported (Oliver & Whitfield, 2016).
The viral particles of orthotospoviruses are spherical and are enclosed in an outer coat composed of a lipid membrane of host origin. The virion is 80–120 nm in diameter and contains three minus-stranded/ambisense RNAs: the S-RNA (~2.9 kb), M-RNA (~4.8 kb), and L-RNA (~8.9 kb) (Turina et al., 2016). The S-RNA encodes a nonstructural (NSs) protein in the viral strand that inhibits the host RNA silencing machinery associated with viral infection (Takeda et al., 2002; Hedil & Kormelink, 2016) and this protein is also required for efficient thrips transmission (Margaria, Bosco, et al., 2014a). In addition, the complementary strand of the S-RNA encodes a nucleocapsid (N) protein that encapsidates the viral RNAs (de Haan et al., 1990). The M-RNA encodes a nonstructural (NSm) protein in the viral strand and the precursors of G1/G2 proteins in the complementary strand. NSm is involved in cell-to-cell movement of the virus (Kormelink et al., 1994; Soellick et al., 2000). The G1/G2 proteins bind to host-derived lipid membranes and are needed to form mature particles (Kormelink et al., 1992; Whitfield et al., 2004). The L-RNA encodes an RNA-dependent RNA polymerase on the complementary strand and is responsible for viral RNA replication (de Haan et al., 1991).
The S-RNA and M-RNA of orthotospoviruses have ambisense polarity with one open reading frame (ORF) each in the viral and the complementary strands. The two ORFs are separated by an intergenic region (IGR) that is adenine- and uracil-rich. The IGRs of the orthotospoviruses S-RNA and M-RNA are 500–600 and 200–250 bases in length respectively, with the exception of polygonum ringspot orthotospoviruses (PolRSV), which are shorter (van Knippenberg et al., 2005; Margaria, Miozzi, et al., 2014b). In the genus Orthotospovirus, there are extremely high levels of intraspecific homology in the amino acid sequence of the N protein, and all strains with more than 90% amino acid homology are considered to be one species (Oliver & Whitfield, 2016). On the other hand, the IGR region of the S-RNA is more diverse than the N protein within each species, and has therefore been useful for population analyses (Heinze et al., 2001).
Until the early 1990s, TSWV was the only recognized species in the genus Orthotospovirus, but with the development of new molecular genetic technologies, new species are being discovered, and there are now more than 25 recognized species in the genus. Thus far, 8 distinct orthotospovirus species have been reported in Japan (Fuji et al., 2022).
TSWV was first observed on dahlias showing necrosis symptoms in the Okayama Prefecture in Japan in 1970 (Inoue & Inoue, 1972). TSWV infecting tomato was then found in Nara Prefecture in 1972 (Kobatake et al., 1976). Control of the virus was initially effective because the infected dahlia plants, which acted as a virus reservoir, could be removed from the fields, and the main vector at the time, the Japanese flower thrip (Thrips setosus), was relatively easy to control by applying insecticides available at the time. However, a more persistent vector, the western flower thrip (Frankliniella occidentalis), invaded Japan in 1990, and TSWV was found in Chrysanthemum morifolium of Shizuoka Prefecture three years later (Kato & Hanada, 2000). The viral infection in C. morifolium plants can be latent, and F. occidentalis is difficult to control. For these reasons, TSWV has become widespread throughout Japan, and has damaged several solanaceous crops, such as tomato and pepper, and ornamental crops in almost all prefectures (Okuda, 2016).
TSWV has a wide host range, but infection of cucurbit plants has been observed only rarely. Massumi et al. (2007) detected TSWV in cultivated cucumber plants that were growing near thrip-susceptible tomatoes and weeds, resulting in a high population of viruliferous thrips. Abadkhah et al. (2018) also reported that TSWV could spread to squash and cucumber from infected tomatoes under experimental conditions. Cucurbits are more commonly infected with melon yellow spot orthotospovirus (MYSV) and watermelon silver mottle orthotospovirus (WSMoV) (Kato et al., 2000; Okuda et al., 2002). WSMoV (first reported in 1984 in Okinawa Prefecture by Hanada et al., 1993) causes chlorotic spots, mottle, and rolling of the leaf edge in watermelon. In addition, zucchini lethal chlorosis orthotospovirus (ZLCV) and melon severe mosaic virus (MSMV) have been reported in Brazil and Mexico, respectively (Bezerra et al., 1999; Ciuffo et al., 2009). An outbreak of WSMoV occurred in cucumber plants in Kanagawa Prefecture in 2016 (Shimada et al., 2019). Inoculation tests revealed that the virus causes leaf and fruit malformation and chlorotic spots in leaves of cucumber plants. MYSV was first identified in melons of Shizuoka Prefecture in 1992. It has recently been found in cucumbers, causing symptoms of mosaic, yellowing, necrotic spots, and vein necrosis in leaves. At present, this outbreak is gradually spreading to western Japan.
In June 2019, leaf necrosis was observed on cucumbers growing in a greenhouse and adjacent open fields in northern Nakadori, Fukushima Prefecture. The novel disease symptoms were observed only in these several adjacent fields. In the early stages of the disease, small spots appeared on the leaves, followed by widespread vesicular spotting. A simple immunostripe test confirmed that the plants were infected with TSWV. However, the pathogenicity and other biological characteristics of this cucumber-infecting TSWV were unknown.
In this study, we analyzed the biological and genetic characteristics of the TSWV strain found in Fukushima Prefecture, and reproduced the disease symptoms in cucumber. We determined the complete nucleotide (nt) sequence of the strain and investigated its relationships with previously reported isolates by phylogenetic analyses. We performed inoculation tests with several strains of TSWV on cucumber cultivars and other cucurbit plants. Furthermore, we investigated the occurrence and origin of this isolate by comparing the IGRs of the new TSWV isolate with other isolates from tomato and cucumber collected in the same area.
Materials and methods
Virus isolation
In June 2019, cucumber and tomato plants showing necrosis symptoms on leaves were collected from Nakadori, Fukushima Prefecture, Japan. An immuno-strip for TSWV (Agdia) was initially used to identify the virus according to the manufacturer’s instructions. One cucumber (FC19) and one tomato (FT19) sample with TSWV was cut into 5 mm square pieces and ground with 0.1 M phosphate buffer, pH 7.0, containing 0.1% of 2-mercaptoethanol. Each extract was used to mechanically inoculate Chenopodium quinoa plants using carborundum powder. Then, virus-containing extracts recovered after three passages through C. quinoa were used to mechanically inoculate cotyledons of cucumber plants cv. Courage 2. Upper leaves of cucumber plants showing necrotic symptoms were ground, and the extracts were used to mechanically inoculate Nicotiana benthamiana plants. Each inoculated leaves (FC19 and FT19) of N. benthamiana were collected at 9 days post inoculation and kept at −80 °C until further use.
Genome sequencing of the cucumber-infecting TSWV
Total RNA was extracted from leaf samples of N. benthamiana with TSWV infection using the RNeasy Plant Mini Kit (Qiagen) according to the manufacturer’s instructions. Reverse transcription (RT)-PCR was performed using the PrimeScript II High Fidelity One Step RT-PCR Kit (Takara Bio) with both degenerate and specific primers for TSWV (Supplementary Table S1). In general, each 25 μl reaction contained 12.5 μl One Step High Fidelity Buffer (1 ×), 0.5 μl PrimeScript II RT Enzyme Mix, 2 μl PrimeSTAR GXL polymerase, 0.5 μl of each primer (20 μM), 2.0 μl of total RNA (~1 μg), and RNase-free water. The TSWV cDNA was amplified under the following conditions: 1 cycle of 10 minutes at 45 °C and 2 minutes at 94 °C followed by 35 cycles of 10 sec at 98 °C, 15 sec at 55 or 60 °C (depending on primers used; see Supplementary Table S1), and 2 minutes at 68 °C. The PCR products were purified using the illustra GFX PCR DNA and Gel Band Purification Kit (Cytiva) and DNA sequence analysis was performed by the Biotechnology Center at Akita Prefectural University using the Big Dye™ Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems).
Phylogenetic analysis
The nucleotide and amino acid sequences were compared with those of previously characterized TSWV strains using GENETEX-Mac version 20.0.3 (Software Development). Phylogenetic trees were constructed using the neighbor-joining method of MEGA version 7.0 with 1000 bootstrap replications (Kumar et al., 2016).
Inoculation of experimental hosts
For these experiments, we inoculated cotyledons and the first true leaf of cucumber cv Courage 2 with the cucumber-infecting TSWV (TS-FC-19). We also inoculated cotyledons of commercial cultivars, including three cucumber varieties, one pumpkin, and one melon, with the same strain. In addition, we inoculated cotyledons of cucumber cv Courage 2 with a TSWV isolated from tomato (TS-FT19). We used the TSWV-pepper strain TS-P2 from the NARO Genebank (MAFF260146) as the representative strain for the symptomatology study. TS-P2 was grown in N. benthamiana plants before being used to inoculate the test plants, as described above. There were three independent biological replicates in the experiment.
Field surveys
A total of four diseased cucumber samples were collected between June 2019 and June 2021 from greenhouses and fields where cucumber and tomato are grown (FC19-F, FC20, FC21-1 and FC21-2; Supplementary Table S2). Total RNA was extracted from symptomatic plants and RT-PCR was carried out as described above. The IGRs of the M-RNA and S-RNA were amplified and analyzed.
Results
Molecular characterization of the cucumber-infecting TSWV
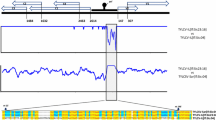
Immuno-strip assays of leaf samples from the cucumbers with necrosis symptoms revealed TSWV infection, which we designated TSWV-[Fukushima:cucumber:2019] (TS-FC19). Viral cDNA fragments were amplified from the cucumber samples by RT-PCR using both degenerate and specific TSWV primers. The complete nucleotide sequences consist of 8913 nt for the L-RNA (LC652640), 4834 nt for the M-RNA (LC652641), and 2966 nt for the S-RNA (LC652642). The genome organization is typical of previously characterized tospoviruses: the L-RNA carries a single ORF encoding RdRp on the complementary strand. The M-RNA encodes NSm on the virus strand and G1/G2 proteins on the complementary strand. The IGR sequence (between the stop codons of the NSm and G1/G2 ORFs) is 339 nt in length and contains a putative secondary structure with an AU rich region. The S-RNA encodes NSs on the virus strand and the N protein on the complementary strand, with an IGR of 539 nt.
BLAST searches with the complete nt sequences of the L-RNA, M-RNA, and S-RNA revealed that TS-FC19 has the highest identity with TSWV isolates LL-N.05 from Spain (97.5%), NC-3 from the USA (96.7%), and TSWV-12 from South Korea (96.9%). Amino acid sequence comparisons revealed high levels of identity (>97%) for the N, RdRp, NSs, NSm, and G1/G2 proteins with their homologous sequences in other TSWV isolates. Since there was >90% identity between the N protein of TS-FC19 and those of other TSWV isolates, TS-FC19 meets the criterion for classification as a variant of TSWV.
Phylogenetic analyses
We performed phylogenetic analyses of TS-FC19 and other TSWV isolates, using the amino acid sequences of the N, NSm, NSs, RdRp, and G1/G2 proteins. The phylogenetic trees generated from these analyses revealed that TS-FC19 is closely related to previously characterized isolates of TSWV (Supplementary Figs. S1, S3, S5, S7 and S9). In the tree based on the N protein, TS-FC19 was placed in a cluster of 7 TSWV isolates [accession numbers (amino acid identity): APG79442 (99.2%), QNJ34562 (98.4%), AAU95399 (98.8%), APG79492 (98.8%), APG79494 (99.2%), ACQ85224 (99.2%) and ACQ85225(98.8%)], most of which came from Solanaceae hosts including tomato, tobacco, potato, and capsicum (Supplementary Fig. S1). However, the unique amino acid position of TS-FC19 was not found when the sequence was compared with the above 7 isolates (Supplementary Fig. S2). In the NSm tree, TS-FC19 was placed in a cluster of 14 TSWV isolates [accession numbers (amino acid identity): ANE20450 (98.0%), ANE20449 (98.0%), AGM53717 (98.6%), AAV28055 (98.6%), AAF80978 (98.3%), QWT83577 (98.6%), AYW35339 (98.6%), BAK57498 (98.6%), ADR66465 (99.0%), AKC94842 (98.6%), AVP72413 (98.3%), CAA63801(98.0%), AMH38316 (99.3%) and AAP13904 (97.3%)] with Solanaceae and Asteraceae hosts (Supplementary Fig. S3). In comparison with the above 14 isolates, TS-FC19 unique sequences were found in amino acid positions A172, E173 and N191 (Supplementary Fig. S4). In the tree generated using the NSs sequences of the most closely related TSWV isolates, TS-FC19 was placed on a distinct branch (Supplementary Fig. S5). Therefore, the amino acid comparison was performed with the top 10 isolates [accession numbers (amino acid identity): ABD92813 (98.2%), CBX23910 (98.0%), ASW33985 (98.0%), CBX23908 (97.8%), AGM53725 CBX23922 (97.8%), CBX23913 (98.0%), AEK21289 (97.7%), CBX23918 (97.8%) and CBX23911 (97.8%)] selected on BLAST analysis, and 7 unique positions (N88, S150, V157, V172, S288, N310 and I352) were found in NSs (Supplementary Fig. S6). In the RdRp tree, TS-FC19 mainly clustered with 7 TSWV isolates [accession numbers (amino acid identity): AAL55403 (98.0%), QDD67728 (97.7%), QAU55721 (98.2%), QAU55720 (98.2%), AKC94841 (98.6%), AMH38315 (98.1%) and ATL64764 (97.8%)] from Solanaceae hosts (Supplementary Fig. S7). Unique positions were found in 31 amino acids (L48, A88, P141, R314, V343, E417, I572, I717, I778, A839, E864, T925, E1048, D1137, K1247, L1385, I1511, T1516, K1679, M1708, D2354, N2455, K2496, N2578, N2595, V2692, T2715, D2746, I2848, K2863, D2871) (Supplementary Fig. S8). In the G1/G2 tree, TS-FC19 was placed with 17 TSWVs isolated from Nicotiana sp., Capsicum sp., Chrysanthemum, tomato, lettuce, common chickweed and dahlia hosts (Supplementary Fig. S9). The amino acid comparison was performed with the top 10 isolates [accession numbers (amino acid identity): ADR66466 (97.8%), ADR66478 (97.3%), QYA72379 (98.3%), ADR66486 (97.1%), QEL52525 (97.8%), QWT83576 (97.5%), AVP72414 (97.5%), AEK06237 (97.9%), AAV28056 (97.8%), ADR66474 (97.5%)] selected on BLAST analysis, and 7 unique positions (G89, V169, I276, K298, E422, L550, G603, C779, M896, N956, R991, G1116) were found in G1/G2 (Supplementary Fig. S10).
Heinze et al. (2001) reported that the S-RNA IGR sequences of TSWV isolates varied according to their geographic origin. To determine the phylogenetic and geographic relationships among 7 TSWV isolates collected from a greenhouse and fields in Fukushima, Japan, we constructed phylogenetic trees using their S-RNA and M-RNA IGR sequences. In both trees, TS-FC19 clustered with the 6 other Fukushima isolates (TS-FC19-F, TS-FT19, TS-FC20, TS-FC21-A, TS-FC21–1, and TS-FC21–2) and these 7 isolates were in a different clade from other TSWV isolates (Supplementary Figs. S11 and S12).
Infectivity and symptomatology of TSWV-FC19
The infectivity and pathogenicity of TS-FC19 was initially assessed in cucumber plants. All cucumber cultivars inoculated with TS-FC19 developed necrotic lesions on inoculated cotyledons, and the virus moved to upper leaves, causing necrosis and chlorosis at 7 days post inoculation (Fig. 1 and Supplementary Fig. S13). In cotyledons of cucumber cv. Courage 2, the average numbers of lesions formed at 7 days post inoculation were 57.9 for TS-FC19 and 249.7 for TS-P2 (Fig. 2a–e). However, the average numbers of lesions on the first leaf at 7 days post inoculation were 591.9 for TS-FC19 and 27.6 for TS-P2 (Fig. 2d–g).
Symptoms on cucumber leaves inoculated with TSWV. The cotyledons and first leaves were inoculated with the two isolates. a Cotyledon inoculated with buffer as healthy control (a). b, c Cotyledons infected with TS-FC19 (b) and TS-P2 (c). d, e, f First leaves infected with Healthy (d), TS-FC19 (e) and TS-P2 (f). g Average number of lesions obtained in cotyledons and first true leaves of cucumber plants infected with TS-FC19 and TS-P2. The error bars show standard error; n = 3
Necrosis symptoms also developed when either TS-FC19 or TS-P2 was used to inoculate the first true leaves of cucumber plants (Fig. 3a, e). However, the two strains showed different patterns of pathogenicity in the cucumber plants. TS-FC19 moved systemically and caused necrosis in the upper leaves (Fig. 3b to d). On the other hand, although some TS-P2 spread to the second and third leaves, the numbers of necrotic lesions in upper leaves were much lower than in plants infected with TS-FC19 (Fig. 3f to h). The average numbers of lesions on the first and second leaves at 7 days post inoculation were actually higher in the TS-P2 infected plants (161.8 and 103.75 respectively) than in the TS-FC19 infected plants (127.4 and 20.2 respectively) (Fig. 3i). However, the numbers of lesions on the third and fourth leaves of TS-P2 infected plants were only 0.75 and 0, respectively, while those on the third and fourth leaves of TS-FC19 infected plants were much higher: 1268.6 and 129, respectively (Fig. 3c and i).
Symptoms on cucumber leaves after inoculation of first leaves with TSWV. a–h First (a, e), second (b, f) third (c, g) and fourth (d, h) leaves of plants infected with TS-FC19 (a to d) and TS-P2 (e to h). i Average number of lesions obtained in the first to fourth leaves of cucumber plants infected with TS-FC19 and TS-P2. The error bars show standard error; n = 3
The TSWV isolate from tomato, TS-FT19, was able to systemically infect cucumber plants cv. Courage 2 and induce necrosis symptoms on upper leaves, indicating that this isolate can also cause cucumber necrosis disease (Supplementary Fig. S14).
When TS-FC19 was tested on pumpkin and melon, the plants developed necrotic lesions on the inoculated cotyledons at 7 days post inoculation, but no symptoms appeared on the upper leaves. Furthermore, TS-FC19 was not detected in newly emerged leaves (Supplementary Fig. S15).
Discussion
Diseases caused by thrip-borne orthotospoviruses have emerged as a major threat to vegetable and flower production (Oliver & Whitfield, 2016). Identifying the viruses involved, and understanding their biology are the first steps toward developing disease management strategies. RT-PCR analyses and immuno-strip assays revealed that TS-FC19 isolates were present in samples collected from cucumber growing areas in Fukushima Prefecture in the years 2019 to 2021. The presence of TS-FC19 in diseased plants suggested that the leaf necrosis disease of cucumber in Fukushima Prefecture was caused by a strain of TSWV.
TSWV causes significant crop yield losses in many parts of the world. In this study, we describe the molecular characterization of a TSWV strain isolated from cucumber in Fukushima Prefecture, Japan. Phylogenetic analysis showed that the cucumber TSWV isolates clustered separately from other TSWV isolates, but the genomic RNA sequences and the amino acid sequence of the N protein showed more than 90% sequence identity with other TSWV strains, and thus the cucumber strain was designated as TSWV-FC19. Phylogenetic analyses based in the IGRs of the TSWV isolates collected in 2019–2021 suggested that TS-FC19 may have evolved locally. It is possible that TSWV infecting indigenous hosts (i.e., tomato) may have developed the ability to cause leaf necrosis symptoms in cucumber plants. This hypothesis is supported by the fact that a TSWV isolate from tomato (TS-FT19) collected near the area where TS-FC19 was found could also infect cucumber and cause necrosis symptoms. The IGR shows a high level of diversity among TSWV isolates and may confer a selective advantage by promoting replication and suppressing host defense (Hedil & Kormelink, 2016). Additionally, the amino acid changes in NSs, especially the 7 unique sequence positions, may have contributed to the ability to infect cucumbers. Although orthotospoviruses have been detected in cucurbit plants, including field-grown cucumber, the pathogenicity of TSWV in cucumber has not been studied previously (Karavina & Gubba, 2017). Therefore, we conducted inoculation tests of TS-P2 and TS-FC19 using cucumber plants, and found differences in pathogenicity between these strains. Although TS-P2 produced larger numbers of lesions on inoculated cotyledons and first leaves, it did not spread systemically. TS-FC19 spread systemically and induced many necrotic lesions in upper leaves. In addition, some TS-FC19-inoculated plants developed wilt symptoms on the stem, leading to plant death. These results indicate that TS-FC19 is compatible and pathogenic with cucumber.
Cucumber mosaic virus (CMV), zucchini yellow mosaic virus (ZYMV), and watermelon mosaic virus (WMV) infect field-grown cucurbit plants, including cucumber, in Japan. Immuno-stripe tests showed that some field grown cucumbers in Fukushima Prefecture were infected with a mixture of TS-FC19 and CMV, but ZYMV and WMV were not detected (data not shown). However, neither CMV nor ZYMV was detected in the plants from which TS-FC19 was originally isolated. These observations, along with the results of the inoculation tests, indicate that the cucumber leaf necrosis disease was caused by TSWV.
All the cucumber cultivars used in our study were susceptible to TS-FC19 and showed disease symptoms. Also, TS-FC19 had stronger infectivity in cucumbers than the TS-P2 strain. A report by Margaria et al. (2015) shows that a Tsw resistance-breaking isolate (TSWV-p331) emerged via reassortment from two evolutionarily distinct strains. Therefore, TS-FC19 may have emerged via a reassortment event. Further experiments are needed to clarify the roles of TSWV-encoded virulence-related genes (Feng et al., 2020) in this difference in pathogenicity. TSWV is commonly found in vegetables and flowers, mainly in the families Solanaceae, Brassicaceae, Asteraceae, and Gentianaceae, but there have been no previous reports of TSWV infecting cucurbits in Japan. Further experiments with insect vectors are needed to investigate the possibility that TSWV may have multiplied inside the thrips and evolved into a mutant strain that causes pathogenicity in cucurbits. For future investigations of the geographic distribution and host range of TS-FC19, we will design primers that can specifically amplify the IGR sequences by RT-PCR.
Thus far, TS-FC19 has been detected only in a few greenhouses and adjacent fields in Fukushima Prefecture, and no regional spread has been confirmed. However, it will be important to monitor cucumber and other crops in the future, so that any emergence or spread of new TSWV variants can be detected as early as possible. It has been suggested that mixed infections of closely related tospoviruses can result in chimeric strains that may emerge as new pathogens (Butković et al., 2021). The occurrence of WSMoV and MYSV is increasing in cultivated cucumbers in Japan, and it will be important to evaluate the risk of new viruses arising from multiple infections with TSWV in advance.
References
Abadkhah, M., Koolivand, D., & Eini, O. (2018). A new distinct clade for Iranian tomato spotted wilt virus isolates based on the polymerase, nucleocapsid, and non-structural genes. Plant Pathology Journal, 34, 514–531.
Bezerra, I. C., Resende, R. D., Pozzer, L., Nagata, T., Kormelink, R., & de Avila, A. C. (1999). Increase of tospoviral diversity in Brazil with the identification of two new tospovirus species, one from chrysanthemum and one from zucchini. Phytopathology, 89, 823–830.
Butković, A., González, R., & Elena, S. F. (2021). Revisiting Orthotospovirus phylogeny using full-genome data and testing the contribution of selection, recombination and segment reassortment in the origin of members of new species. Archives of Virology, 166, 491–499.
Ciuffo, M., Kurowski, C., Vivoda, E., Copes, B., Masenga, V., Falk, B. W., & Turia, M. (2009). A new tospovirus sp in cucurbit crops in Mexico. Plant Disease, 93, 467–474.
de Haan, P., Wagemakers, L., Peters, D., & Goldbach, R. (1990). The S RNA segment of tomato spotted wilt virus has an ambisense character. Journal of General Virology, 71, 1001–1008.
de Haan, P., Kormelink, R., Resende, D., van Poelwijk, F., Peters, D., & Goldbach, R. (1991). Tomato spotted wilt virus L RNA encodes a putative RNA polymerase. Journal of General Virology, 72, 2207–2216.
Feng, M., Cheng, R., Chen, M., Guo, R., Li, L., Feng, Z., Wu, J., Xie, L., Hong, J., Zhang, Z., Kormelink, R., & Tao, X. (2020). Rescue of tomato spotted wilt virus entirely from complementary DNA clones. Proceedings of the National Academy of Sciences of the United States of America, 117, 1181–1190.
Fuji, S., Mochizuki, T., Okuda, M., Tsuda, S., Kagiwada, S., Sekine, K.-T., Ugaki, M., Natsuaki, K. T., Isogai, M., Maoka, T., Takeshita, M., Yoshikawa, N., Mise, K., Sasaya, T., Kondo, H., Kubota, K., Yamaji, Y., Iwanami, T., Ohshima, K., et al. (2022). Plant viruses and viroids in Japan. Journal of General Plant Pathology, 88, 105–127.
Hanada, K., Tsuda, S., Kameya-Iwaki, M., & Tochhara, H. (1993). Distinct properties of nucleocapsid of a watermelon isolate of tomato spotted wilt virus. Annals of the Phytopathological Society of Japan, 59, 500–506.
Hedil, M., & Kormelink, R. (2016). Viral RNA silencing suppression: The enigma of bunyavirus NSs proteins. Viruses, 8, 208.
Heinze, C., Letschert, B., Hristova, D., Yankulova, M., Willingmann, P., Karadjova, O., Atanassov, A., & Adam, G. (2001). Variability of the N-protein and the intergenic region of the S-RNA of tomato spotted wilt tospovirus (TSWV). New Microbiologica, 24, 175–187.
Inoue, T., & Inoue, N. (1972). Tomato spotted wilt virus occurred in Dahlia plant. Nogaku-kennkyu, 54, 74–90.
Karavina, C. & Gubba, A. (2017). Detection and characterization of Tomato spotted wilt virus infecting field and greenhouse-grown crops in Zimbabwe. European Journal of Plant Pathology, 49, 933–944.
Kato, K., & Hanada, K. (2000). A necrotic disease of chrysanthemum (Chrysanthemum morifolium Ramat) caused by tomato spotted wilt virus (TSWV) in Japan. Kyushu Plant Protection Research, 46, 61–65.
Kato, K., Hanada, K., & Kameya-Iwaki, M. (2000). Melon yellow spot virus: A distinct species of the genus Tospovirus isolated from melon. Phytopathology, 90, 422–426.
Kobatake, H., Osaki, T., Yoshioka, A., & Inoue, T. (1976). Spotted wilt disease of tomatoes in Japan. Annals of the Phytopathological Society of Japan, 42, 287–294.
Kormelink, R., de Haan, P., Meurs, C., Peters, D., & Goldbach, R. (1992). The nucleotide sequence of the M RNA segment of tomato spotted wilt virus, a bunyavirus with two ambisense RNA segments. Journal of General Virology, 73, 2795–2804.
Kormelink, R., Storms, M., van Lent, J., Peters, D., & Goldbach, R. (1994). Expression and subcellular location of the NSm protein of tomato spotted wilt virus (TSWV), a putative viral movement protein. Virology, 200, 56–65.
Kumar, S., Stecher, G., & Tamura, K. (2016). MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Molecular Biology and Evolution, 33, 1870–1874.
Margaria, P., Bosco, L., Vallino, M., Ciuffo, M., Mautino, G. C., Tavella, L., & Turina, M. (2014a). The NSs protein of tomato spotted wilt virus is required for persistent infection and transmission by Franklinella occidentalis. Journal of Virology, 88, 5788–5802.
Margaria, P., Miozzi, L., Ciuffo, M., Pappu, H., & Turina, M. (2014b). The complete genome sequence of polygonum ring spot virus. Archives of Virology, 159, 3149–3152.
Margaria, P., Ciuffo, M., Rosa, C., & Turina, M. (2015). Evidence of a tomato spotted wilt virus resistance-breaking strain originated through natural reassortment between two evolutionary-distinct isolates. Virus Research, 196, 157–161.
Massumi, H., Samei, A., Hosseini Pour, A., Shaabanian, M., & Rahimian, H. (2007). Occurrence, distribution, and relative incidence of seven viruses infecting greenhouse-grown cucurbits in Iran. Plant Disease, 91, 159–163.
Okuda, M. (2016). Tospoviruses occurring in and outside Japan. Japanese Journal of Phytopathology, 82, 169–184.
Okuda, M., Takeuchi, S., Taba, S., Kato, K., & Hanada, K. (2002). Melon yellow spot virus and watermelon silver mottle virus: Outbreak of cucurbit infecting tospovirus in Japan. Acta Horticulturae, 588, 143–148.
Oliver, J. E., & Whitfield, A. E. (2016). The genus Tospovirus: Emerging bunyaviruses that threaten food security. Annual Review of Virology, 3, 101–124.
Rotenberg, D., Jacobson, A. L., Schneweis, D. J., & Whitfield, A. E. (2015). Thrips transmission of tospoviruses. Current Opinion in Virology, 15, 80–89.
Shimada, R., Okuda, M., & Uekusa, H. (2019). Characteristics of watermelon silver mottle virus (WSMoV) found in Kanagawa prefecture and reproduction of symptoms on cucumber caused by WSMoV. Japanese Journal of Phytopathology, 85, 108–111.
Soellick, T. R., Uhrig, J. F., Bucher, G. L., Kellmann, J. W., & Schreier, P. H. (2000). The movement protein NSm of tomato spotted wilt tospovirus (TSWV): RNA binding, interaction with the TSWV N protein, and identification of interacting plant proteins. Proceedings of the National Academy of Sciences of the United States of America, 97, 2373–2378.
Takeda, A., Sugiyama, K., Nagano, H., Mori, M., Kaido, M., Mise, K., Tsuda, S., & Okuno, T. (2002). Identification of a novel RNA silencing suppressor, NSs protein of tomato spotted wilt virus. FEBS Letters, 532, 75–79.
Turina, M., Kormelink, R., & Resende, R. O. (2016). Resistance to tospovirusus in vegetable crops: Epidemiological and molecular aspects. Annual Review of Phytopathology, 54, 347–371.
van Knippenberg, I., Goldbach, R., & Kormelink, R. (2005). Tomato spotted wilt virus S-segment mRNAs have overlapping 3′-ends containing a predicted stem-loop structure and conserved sequence motif. Virus Research, 110, 125–131.
Whitfield, A. E., Ullman, D. E., & German, T. L. (2004). Expression and characterization of a soluble form of tomato spotted wilt virus glycoprotein GN. Journal of Virology, 78, 13197–13206.
Acknowledgments
We thank the NARO Genebank for providing the TSWV-P2 strain.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Human and animal rights
This article does not contain any studies with human participants or animals.
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
The GenBank/EMBL/DDBJ accession number for the sequence reported in this paper is LC652640, LC652641, and LC652642 (TS-FC19).
Rights and permissions
About this article
Cite this article
Kon, T., Watanabe, N., Ootake, H. et al. Occurrence and characterization of tomato spotted wilt orthotospovirus isolated from cucumber. Eur J Plant Pathol 163, 789–797 (2022). https://doi.org/10.1007/s10658-022-02515-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-022-02515-9