Abstract
In total 58 Xanthomonas strains isolated from Araceae worldwide, together with 13 other phylogenetically-related Xanthomonas strains, were characterized using multilocus sequence analysis based on concatenated sequences of seven single copy orthologous genes, extracted from whole genome sequences. The analysis revealed a monophyletic clade of 48 strains, 44 isolated from Anthurium, identified as X. phaseoli pv. dieffenbachiae (Xpd) confirmed by nucleotide identity analysis. The other strains from aroids were identified as Xanthomonas euvesicatoria (2 strains), X citri (5 strains) and Xanthomonas sacchari (3 strains). Two TaqMan assays were designed for specific detection of Xpd, one targeting sequences of a hypothetical protein and one targeting a type I restriction endonuclease subunit S. The two assays showed similar reaction kinetics and were merged with an assay comprising an amplification and extraction control into a triplex assay. The assay was able to detect minimally 100 copies of a target sequence delivered as a gBlock, 100 fg of genomic DNA and 104 cells per mL in an Anthurium leaf extract.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Anthurium bacterial blight and leaf spot is caused by bacteria belonging to the genus Xanthomonas. In 1995 the pathogens from aroids were classified as Xanthomonas axonopodis pv. dieffenbachiae on the basis of DNA-DNA hybridization studies (Parkinson et al., 2009; Vauterin et al., 1995). The pathogen is not restricted to Anthurium, but can infect a number of aroids, amongst which Dieffenbachia, Philondendron, Aglaonema and Syngonium (Anonymous, 2009). The pathogen causes wet, chlorotic or necrotic leaf spots, blight and decay of plant tissues. Infections usually occur through hydathodes and therefore symptoms often start at the leaf margins and at the abaxial side of leaves. The pathogen is widely distributed and has been found on all continents (www.cabi.org/isc/datasheet/56931). X. axonopodis pv. dieffenbachiae is listed on EPPO alert list (A2).
Strains of xanthomonads pathogenic in ornamental aroids are highly heterogeneous as shown with various genetic techniques which comprised RAPD-PCR (Khoodoo & Jaufeerally-Fakim, 2004), rep-PCR (Rademaker et al., 2005), AFLP (Rademaker et al., 2005), rRNA restriction patterns (Berthier et al., 1993), single locus (gyrB) sequencing (Parkinson et al., 2009), multilocus sequence analysis (Constantin et al., 2016; Donahoo et al., 2013; Young et al., 2008), DNA-DNA-hybridization (Constantin et al., 2016; Donahoo et al., 2013) and Average Nucleotide Identity calculations (ANI) based on whole genome sequence analysis (Constantin et al., 2016).
Using rep-PCR techniques (BOX-, ERIC- and REP-PCR), X. axonopodis pv. dieffenbachiae strains were clustered in two subgroups of X. axonopodis, subgroup 9.4 comprising pathovars manihotis and phaseoli, and subgroup 9.6 comprising pathovars phaseoli fuscans, aurantifolii, vignaeradiatae, rhynchosia (cajani), bauhinae and sesbaniae (Rademaker et al., 2005). A polyphasic taxonomic approach, including multilocus sequence analysis, ANI values, DNA-DNA hybridization data and phenotypic analysis of strains resulted in a taxonomic revision of the X. axonopodis complex (Constantin et al., 2016). Strains isolated from Dieffenbachia, Philondendron and Anthurium were found to cluster within three different groups, each representing a different species. PG1 clustered with type strains of Xanthomonas citri and Xanthomonas fuscans, PGII with type strains of Xanthomonas euvesicatoria, X. perforans and Xanthomonas alfalfae, and PGIII with the type strain of X. phaseoli. No strains were found in aroids that clustered with the type strain of X. axonopodis. Taxonomic proposals were made to encompass strains of PGI as X. citri, PGII, comprising strains formerly designated X. perforans and X alfalfae as X. euvesicatoria, PGIII as X. phaseoli and PGIV as X. axonopodis.
X. phaseoli pv. dieffenbachiae (Xpd, PGIII) is the predominant causative agent of bacterial blight and leaf spot in Anthurium species (Constantin et al., 2017. Bioassays with some of these strains showed that they can be highly aggressive on Anthurium, although strains from X. citri and X. euvesicatoria could also cause symptomatic infections albeit with mild symptoms (Constantin et al., 2017)(unpublished results WUR, Wageningen, the Netherlands).
The primary aim of this study was to develop a specific triplex TaqMan assay for reliable detection of Xpd in various substrates. For this, studies on the taxonomic position and the genetic diversity of Xpd were undertaken. Xanthomonas strains isolated from various aroids (Anthurium, Aglaonema, Philodendron and Dieffenbachia) worldwide, but with an emphasis on Anthurium strains from the Netherlands, were included in the studies. Sequence information of the strains was generated or collected from public databases and analysed together with sequences of relevant type or pathotype strains using multilocus sequence analysis (MLSA). Specific primers and probes were selected for detection of Xpd using whole genome sequence information of target and non-target strains. A triplex TaqMan protocol was designed comprising assays against two target loci and including an extraction - amplification control. The specificity and sensitivity of the assay were determined and the assay was evaluated using Anthurium leaf material supplemented with various densities of Xpd.
Materials and methods
Bacterial strains and growth conditions
Strains were collected from international culture collections and work collections at Naktuinbouw (Roelofarendsveen, the Netherlands), the Dutch National Plant Protection Organization (NVWA, Wageningen, NL) and Wageningen University & Research (WUR, Wageningen, NL) (Table 1). Strains from Araceae were isolated from symptomatic plants, but in most cases the pathogenicity of strains was not studied. The bacterial names were used as proposed by Constantin et al. (2016). Bacteria were grown for 48 h on TSA at 27 °C prior to use.
DNA extraction
Bacteria were collected from the agar surface of 9 cm diameter TSA plates in approximately 0.5 mL of water and stored at −20 °C until DNA extraction. The DNA was extracted using the Wizard Magnetic DNA purification System for Food (Promega, Leiden, The Netherlands). DNA yield was determined by using the Pico® Green I dye (Invitrogen, Bleiswijk, NL) and an Infinite® M200 pro microplate reader (Tecan, Männedorf, Switzerland) to measure fluorescence.
Illumina DNA sequencing, genome assembly and annotation
DNA concentration was brought to a concentration of 4 ng/μL for sequencing. Library preparation was performed on 50 μL of 4 ng/μL DNA using the protocol as recommended by Illumina and sequenced with the Novaseq technology at BaseClear B.V. (Leiden, NL). A paired-end DNA library with 2 X 150 bp reads was constructed for each strain. Subsequently, samples were demultiplexed and reads, containing adaptors were removed. FASTQ files were imported in CLC genomic workbench version 12.03 (Qiagen, Aarhus, Denmark). Reads were quality trimmed in CLC using a quality score of 0.05 and a maximum of two ambiguous nucleotides per read. Reads were filtered in length with a minimum number of nucleotides per read of 45.
De novo assembly was performed with the trimmed Paired-End data using automatic bubble size and a minimum contig length of 500 pb. A mapping mode of mapping the reads back to contigs was used with a length and similarity fraction of 0.8. Sequencing reads are available at NCBI under project number BioProject PRJNA716613. The accession numbers are provided in Table 1.
Genome assemblies were exported annotated using Prokka (version 1.14.6) (Seemann, 2014), with default settings for the bacteria.
Multilocus sequence analysis
Sequences of the seven housekeeping genes atpD (1407 bp), dnaK (1926 bp), efp (567 bp), glnA (1410 bp), gyrB (2445 bp), lrp (480 bp) and rpoD (1878 bp) were used. Sequences of these seven genes were used as previously described (Constantin et al., 2016), but the full length sequences were analyzed in our studies. Sequences were extracted from the annotated contigs using CLC. The genes were concatenated in alphabetic order into a total sequence of 10.113 bp. An alignment was made in CLC Work Bench and imported into MEGA 7.0 software for constructing a phylogenetic tree. A phylogenetic tree was inferred by using the Maximum Likelihood method based on the General Time Reversible model with 500 bootstrapping replications. Strains of X. citri pv. phaseoli LMG826 X. citri pv. citri LMG 9322 representing Phylogenetic Group I, X. euvesicatoria LMG27970 representing Phylogroup II, Xpd LMG 695 (in the collection catalogue listed as X. axonopodis pv. dieffenbachiae) representing Phylogenetic Group III, X. axonopodis pv. axonopodis LMG 982 representing Phylogenetic Group IV were used (Constantin et al., 2016) as a references. Stenotrophomonas maltophilia LMG 958 was used as an out-group.
Average nucleotide identity
Average Nucleotide Identity (ANI) scores between genomes of (patho)type strains and other Xanthomonas strains in the same clade or related clades were estimated using JSpeciesWS (online service offered by Ribocon, https://www.ribocon.com).
Designing of a TaqMan assay specific for Xpd
To design a TaqMan assay that only detects Xpd, the pathotype strain LMG 695 was used as a reference. Potential target sites were identified using CLC genomic workbench in a two-step filtering process: first identifying sequences that are conserved in all strains of the target group (Xpd) and removing sequences that are similar to non-target strains. In more detail, the genome sequence of pathotype strain LMG 695 (NZ_CP014347.1; 5.04 Mb) was dissected in 500 bp-long sequences (10074). These sequences were stringently mapped to all Xpd strains analyzed in the current study, and only those sequences (4694) were kept that mapped to all Xpd strains with 100% similarity. Subsequently this narrowed down set of sequences was mapped (CLC mapping settings: length fraction: 0.85, similarity fraction: 0.85, global alignment: no) to all other strains included in this study, removing sequences that mapped to any of the non-target strains. To assist the final selection and design step, these sequences were added as a track list to the genome of LMG 695. Before proceeding to the design step, the candidate regions were checked for sequence similarity with non-target organisms using BLAST in the nucleotide database of the NCBI Genbank.
Two sets of primers/probe combinations were designed on 500 bp target-specific fragments using primer quest tool of Integrated DNA Technologies (IDT, Leuven, Belgium) with default settings (Table 2). The first set (Xpd866) was derived from a gene encoding for a hypothetical protein and consists of the forward primer FwXpd866 (5’-TACCTGCCTCGCCTCTT3′), reverse primer RvXpd866 (5’-GGATCGTCGGTCTTGTGTTT −3′) and the FAM labelled probe pXpd866 (5’CAACAGCGTGAGAAAGAAACTCGGCA-3′). The other set (Xpd4494) was derived from a gene encoding for a type I restriction endonuclease subunit S and consists of the forward primer fwXpd4494 (5’-GTATAGATGTACTGACGGCTCAC3′), reverse primer RvXpd4494 (5’-CGCGATCATTCCCGATACTT-3′) and an ATTO532 labelled probe pXpd4494 (5’-CGCTTGATTGCAGTTCCACTCAGGA-3′). Probes of TaqMan assays were double quenched with ZEN/Iowa Black FQ (IDT, Leuven, Belgium).
Specificity of the TaqMan assays
The specificity of the assays was tested using genomic DNA of 22 Xpd strains, 46 non-target Xanthomonas strains belonging to different species and a strain of S. maltophilia (Table 1). For most strains 0.2 ng of purified DNA was used, but for some strains a suspension of 109–1010 cells/mLwas boiled in 50 mM NaOH prior to testing. In Table 1 these results are marked with red.
For each TaqMan assay 5 μL of DNA or the boiled suspensions was mixed with 20 μL reaction mix containing 5 μL PerfeCTa multiplex qPCR ToughMix 5x (Quantabio, Beverly, USA), 100 nM probe and 300 nM of each forward and reverse primer. The reactions were performed in a Biorad CFX touch Real-Time PCR detection system (BioRad, Hercules, USA) using the following conditions: 95 °C for 2 min; 40 cycles of 95 °C for 15 s followed by 60 °C for 60 s. Analysis of the data was done by automatic threshold calculation within the Biorad system software. A Ct value ≤35 was considered positive.
Triplex TaqMan design
The two assays for Xpd were combined with an assay that quantify Acidovorax cattleya (Acat) (Bonants et al., 2019) into a triplex TaqMan (Table 2). The Acat assay consists of a forward primer Acat 2-F (5’-TGTAGCGATCCTTCACAAG- 3′), reverse primer Acat 2-R.
(5’-TGTCGATAGATGCTCACAAT - 3′) and a TexasRed labelled probe Acat 2-Pr (5’-CTTGCTCTGCTTCTCTATCACG - 3′). The triplex TaqMan was performed using the same PCR conditions and materials as the simplex tests. For the assays a primer concentrations of 300 nM was used and 100 nM probe in a total volume of 25 μL.
Sensitivity triplex TaqMan assay
The analytical sensitivity (detection threshold) of the triplex TaqMan assay was determined using gBlocks, genomic DNA and target bacteria administered to an Anthurium leaf extract. The gBlock, synthetic oligonucleotides (Integrated DNA Technologies, USA), contained concatenated sequences of the amplicons of the two assays for Xpd, an UTC Universal Template Control (UTC), as a control on the quality of the gBlock, interspaced with some extra flanking nucleotides (Table S1). A ten-fold serial dilution from 106 to 1 copies of the gBlocks DNA was used to determine the sensitivity of the triplex Taqman. Per reaction, 1 μL of 104 copies of a gBlock of Acat served as an internal control (Bonants et al., 2019).
For determining the detection threshold for genomic DNA, a ten-fold serial dilution of 1 ng-1 fg of genomic DNA from strain LMG 695 (IPO 1104) was used. Each sample was supplemented with 1 μL of 2 pg gDNA of Acat as an amplification control.
In addition the sensitivity was determined with Xpd supplemented to an Anthurium leaf extract. Six whole leaves (12 g) of cultivar ‘White Champignon’ in a BioReba bag with a synthetic intermediate layer (BioReba, Kanton Reinach, Swiss) were crushed using a sample crusher (AAA lab equipment B.V. Roelofarendsveen). Subsequently, 24 mL of 0.05 M PBS (2.5 g of NaH2PO4, 2.52 g Na2HPO4 and 8.2 g NaCl in 1 Liter water, pH 7.2) was added and after homogenization a serial dilution of bacterial cells of strain IPO1104 was supplemented in a ten-fold serial density of 107–101 cells/mL. From each dilution 1 mL was pipetted in a strip of Qiagen tubes (Qiagen, Aarhus, Denmark) in replicates. The tubes were centrifuged for 10 min at 6000 g and the supernatant discarded. The pellet was supplemented with 50 μL of a 1000-fold diluted suspension of Acat cells with an optical density at 600 nm of 0.8 (ca. 8. 108 cells/ml). DNA extraction was performed with the AGOWA maxi kit (Nucleics, Woollahra, Australia) according to the manufacturer’s instructions.
Data analysis
Ct-values were imported from the QuantStudio software (ThermoFisher Scientific, Waltham, USA). Ct-values were plotted against the 10log number of gBlock or gDNA copies, or the density of bacterial cells. The log of the densities in the leaf extract, and (linear) trend lines were determined and the coefficient of determination (R2) were calculated using MS Excel. The amplicon efficiency was calculated from the slope of the calibration curves, as 101/slope − 1 (Kubista et al., 2006). To compare the two assays in the triplex TaqMan, the Ct-values were plotted against each other, to calculate the coefficient of determination (R2).
Effect of the concentration of the leaf extract on the sensitivity of the TaqMan.
To determine the maximum weight of material that can be used in a composite sample, DNA was extracted from increasing amounts of the central parts of the leaves including part of the petiole in fixed volume of buffer, i.e. 6.2–27.1 g in 100 mL of 0.01 M PBS which were homogenized in, representing different pooling strategies. A suspension of Acat (NBC 430) was supplemented in a density of approximately 105 cells per mL before the DNA extraction as extraction and amplification control, to obtain a Ct-value between 27 and 30 after 40 cycles. DNA extraction was carried out using the mag maxi DNA extraction kit (LGC Genomics, Berlin, Germany) on a KingFisher KF96 system following the manufacturer’s instructions.
Results
Strain characterization
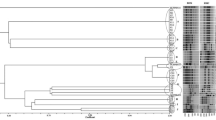
In the multilocus sequence analysis (MLSA) based on concatenated sequences of seven housekeeping genes (atpD, dnaK, efp, glnA, gyrB, lrp and rpoD), strains isolated from Anthurium in different parts of the world (the Netherlands, Poland, Spain, Italy, Dominican Republic, Ghana and the USA), together with four strains from other aroids (Aglaonema, Philodendron and two of Dieffenbachia), clustered in a monophyletic group with the pathotype LMG 695 of X. phaseoli pv. dieffenbachiae (Xpd) (Fig. 1). Seven SNP’s were found to be unique and specific for the set of 48 strains in the 10.113 nt sequence analysed. In addition, a 6 bp long deletion in rpoD was found to be specific for six strains (PD 2179, PD 2170, LMG 695, LMG 12752, NBC 785 and A2108).
Maximum-likelihood phylogenetic tree of concatenated nucleotide sequences of the atpD, dnaK, efp, glnA, gyrB, lrp and rpoD genes from 73 Xanthomonas strains. Bootstrap values greater than 50% are shown for 500 replicates. Concatenated sequences of Stenotrophomonas maltophilia were used as an out-group. T = type strain; PT = pathotype strain
Not surprisingly, the cluster most related to Xpd comprised two strains of X. phaseoli pv. phaseoli isolated from bean (CFBP 2534 and NCPPB 1811) and five strains of X. phaseoli pv. manihotis (CFBP 7153, NBC 1192, NBC 1194, NBC 1264 and NBC 1265). X. axonopodis pv. begoniae (CFBP 2524 and NBC 96) was the closest related species to X. phaseoli.
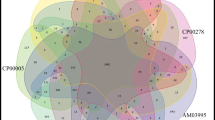
Using the ANI score, strains isolated from aroids were not only identified as X. phaseoli, but also as X. citri, X. euvesicatoria and Xanthomonas sacchari (Table S5–8). All strains of Anthurium, however, were identified as Xpd, with the exception of one strain from Hawaii, identified as X. sacchari.
Specificity
The specificity of the two assays (Xpd866 and Xpd4494) in the triplex TaqMan was determined on the basis of an in silico analysis using the blastn function in NCBI database. Both assays were highly specific and full matches of the amplicon sequences, 74 nt for Xpd866 and 100 nt for Xpd4494, were found only with target strains LMG 25940 and LM G695. In X. arboricola and Xanthomonas dyei the closest non-target sequences were found for Xpd866 and Xpd4494, respectively, but only with a low level of homology.
In addition, genomic DNA was tested of 22 strains of Xpd, 46 other Xanthomonas strains from which eight belonging to other X. phaseoli pathovars and one strain of S. maltophilia (Table 1). A positive reaction with low Ct values (21.2–24.5) was found for all 22 Xpd strains in both assays. All non-target strains were negative in both assays. A weak reaction above the detection threshold (Ct values of 38.9 and 39) was only found with two strains (PD 3347 and PD 2170) of the closely-related pathogen X. phaseoli pv. manihotis in assay Xpd4494.
Analytical sensitivity and linearity of the TaqMan assays
Both assays (Xpd866 and Xpd4494) were able to detect up to 100 copies of the gBlock, 100 fg of genomic DNA and 104 cells per mL in Anthurium leaf extract (Fig. 2, Table S2–4). If the Ct-values were plotted against the logarithm of the copy number, a regression coefficient higher than 0.98 was found within the dynamic range of the assay (Fig. 2). This was not influenced by the target or the matrix, gBlock, gDNA or cells in Anthurium leaf extract. The efficiency of the Xpd866 for gBlocks, gDNA and cells in leaf extract were 103.5, 88.9 and 93.8%, respectively. The estimated efficiency of the Xpd4494 for gBlocks, gDNA and cells in leaf extract were 92.7, 102.2 and 103.1%, respectively. The two assays for Xpd showed highly similar Ct values in experiments with the serially diluted targets. If the Ct-values of the serial dilutions of gBlocks, gDNA or cell-suspensions in leaf extracts for the two assays were plotted against each other, the regression coefficient (R2) was always higher than 0.99 (Fig. S1). This was despite the difference in the delta Rn values for the assays, caused by the difference in fluorophore (FAM versus Atto532) used in the two assays (Fig. 3). The Ct-values of the Acat TaqMan assay ranged in all samples with the gBlock and gDNA between 27.7 and 29.1 (Table S3 and S4). For the cells in leaf extract (0.5 g/mL), the Ct values for Acat ranged between 31.7 and 34.0 (Table S3).
Calibration curves of TaqMan assays for detection of Xanthomonas phaseoli pv. dieffenbachiae (Xpd). In A, two TaqMan assays (Xpd866 and Xpd4494) were individually calibrated using a gBlock comprising the amplicon sequences of both assays. The Ct values and the logarithm of the number of gBlock copies were plotted on the x and y axis, respectively. In B, the TaqMan was calibrated in a multiplex format with both assays using genomic DNA (gDNA). The Ct values and the logarithm of the weight of the gDNA of Xpd strain NCPPB 1833 were plotted on the x and y axis, respectively. In C, the TaqMan was also calibrated in a multiplex format with both assays using cell suspensions of Xpd strain NCPPB 1833 in an extract of Anthurium leaves. The Ct values and the logarithm of the density of Xpd cells were plotted on the x and y axis, respectively. The error bars represent standard deviations
Amplification plot of the two TaqMan assays, Xpd866 labelled with FAM and Xpd4494 labelled with Atto532 and designed for detection of Xanthomonas phaseoli pv. dieffenbachiae (Xpd). For the plots a gBlock comprising the amplicon sequences of both assays was used. An assay for Acidovorax cattleya (Acat), labelled with Texas Red, was used as amplification control. RFU = relative fluorescence units
Effect of the leaf extract concentration
The analytical sensitivity of the triplex TaqMan assay was determined for Anthurium leaf concentrations ranging between 0.062 and 0.271 g per mL (Table 3). Reactions was considered positive if the Ct values of both assays (Xpd866 and Xpd4494) were ≤ 35. Detection thresholds were always determined with 5 μl of sample as template. The detection threshold was 104 cells per mL for all leaf extract concentrations except for a concentration of 0,171 g/mL in which the threshold was 103 cells/mL. Ct values of the Acat TaqMan assay were relatively constant (27.6–31.4), but at the highest density of Xpd (108 cells per mL), the reaction of Acat TaqMan was inhibited.
Discussion
A set of 58 Xanthomonas strains of aroids, were characterized using MLSA on concatenated sequences of seven housekeeping genes as described by Constantin et al. (2016), as a first step in the development of a TaqMan assay for specific detection of Xpd. All strains isolated from Anthurium plus strains of Aglaonema, Philodendron and two of Dieffenbachia grouped into the clade with the type strain of the pathotype of Xpd (LMG 695), previously named X. axonopodis pv. dieffenbachiae (Constantin et al., 2016). In accordance with the work of Constantin et al. (2016), also strains isolated from aroids were identified as X. euvesicatoria and X. citri. The use of pathovar names for these two species is premature as no pathogenicity data are available.
For the first time, X. sacchari is described in association with Anthurium, but the pathogenicity has still to be determined. In our studies, three strains from Anthurium had an ANI score near to 95% with the type strain of X. sacchari, one of which (strain D109 from Hawaii, USA) exceeded the threshold of 95% for species delineation. A Xanthomonas strain closely-related to X. sacchari is known to be able to cause a grain rot of rice (Mirghasempour et al., 2020). The data suggest a genomic species complex for X. sacchari and a broad host range as strains were initially found in association with sugar cane, but later also with banana plants (Studholme et al., 2011) and with rice seeds (Bansal et al., 2021; Cottyn et al., 2001; Fang et al., 2015).
The clade of Xpd comprises strains that can differ in their aggressiveness on their host of origin (Constantin et al., 2017), but several were found highly aggressive on Anthurium including IPO1838 and 1839 (unpublished results). It has been evidenced that strains isolated from other aroids, including Dieffenbachia and Syngonium were aggressive on Anthurium, but the identity of these strains is not known (Chase et al., 1992). More recently this was also reported by Cottyn et al. (2018).
For detection of Xpd, various methods have been described, based on the use of semi-selective media (Laurent et al., 2009; Norman & Alvarez, 1989), serology (Lipp et al., 1992), DNA-based amplification (Lu et al., 2012, Robene-Soustrade et al., 2006, Chabirand et al., 2014,) and methods combining assays of different principles (Khoodoo et al., 2005; Norman & Alvarez, 1994). Immunocapture PCR using Xanthomonas-specific monoclonal antibodies and primers designed from sequence characterized amplified regions (SCARS) allowed detection of strains from Aroids at a level of 102–103 cfu/mL (Khoodoo et al., 2005). This assay successfully detected Xpd strain D15–3 from Hawaii (USA) which were also included in our study. However, it also detected strain D93, designated in our studies as X. sacchari, indicating that the assay is not entirely Xpd specific. For on-site detection of Xpd, a loop-mediated isothermal amplification (LAMP) assay has been developed allowing detection of 104 cfu/mL and 1–10 fg of pure genome DNA (Jun-Hai et al., 2015). We presume that the assay has been developed for Xpd, but only Chinese strains were included and a (pathovar) type strain was lacking in their studies.
In 2019, a duplex quantitative (TaqMan) real-time PCR assay for detection of Xpd was described, targeting sequences of a gene coding for an ABC transporter (Jouen et al., 2019). The authors used an assay based on sequences encoding for chalcone synthase in Anthurium andreanum as an extraction and amplification control. The assay was able to detect as low as 18 bacterial cells per reaction. The specificity of the assay was evaluated both in situ and in silico with a representative panel of 50 Xpd strains from different regions worldwide and with closely related Xanthomonas strains. Only a strain of X. euvesicatoria, not pathogenic on Anthurium, was found positive.
In this paper, a second TaqMan assay for detection of Xpd has been described. TaqMan assays are the method of choice in many inspection laboratories for routine detection of plant pathogens, due to its specificity, sensitivity, robustness and the possibilities for multiplexing and quantification (Martinelli et al., 2015; Mirmajlessi et al., 2015). We developed a triplex TaqMan, including assays detecting Xpd specific sequences of two loci and an assay for Acidovorax cattleya to check for extraction and amplification efficiency, similarly as described by (Bonants et al., 2019). In comparison with the assay described by Jouen et al. (2019), the added value of the triplex assay describes in this paper lies primarily in its double check for the presence of Xpd, by targeting Xpd specific sequences of two different genes. In addition, the use of Acidovorax cattleya as in internal control has the advantage that it is useful for detection of Xpd in any matrix, not only in plant material. The specificity was high according to the in silico analysis and this was confirmed using DNA from 22 target and 47 non target strains. The analytical sensitivity of the assay for gBlocks (100 copies) and genomic DNA (100 fg) was high but for detection of the target in Anthurium leaf extracts a minimum of 104 cells per mL was required to exceed the detection threshold (Ct 35). In case of a low bacterial density in symptomless plant material, an enrichment in or on a selective growth medium may be required before TaqMan analysis similarly as used for other plant pathogenic bacteria (Schaad et al., 1995; Schaad et al., 1999; Song et al., 2004). The assays for the two target specific loci resulted in very similar reaction values. The diagnostic sensitivity and specificity were 100%.
In practice, the triplex TaqMan assay was highly robust. Only minor effects were found of the concentration leaf material between 0.6 and 0.27 g per mL on the analytical sensitivity of the triplex TaqMan assay. In addition, the cultivar type had no noticeable influence on the TaqMan values. Results were independent for cut or potting plants and independent of the color of leaves or flowers (data not shown).
References
Anonymous. (2009). PM 7/23 (2): Xanthomonas axonopodis pv. dieffenbachiae. EPPO Bulletin,39, 393–402.
Bansal, K., Kaur, A., Midha, S., Kumar, S., Korpole, S., & Patil, P. B. (2021). Xanthomonas sontii sp. nov., a non-pathogenic bacterium isolated from healthy basmati rice (Oryza sativa) seeds from India. Antonie van Leeuwenhoek,114, 1935–1947.
Berthier, Y., Verdier, V., Guesdon, J. L., et al. (1993). Characterization of Xanthomonas campestris pathovars by ribosomal RNA gene restriction patterns. Applied and Environmental Microbiology,59, 851–859.
Bonants, P., Griekspoor, Y., Houwers, I., et al. (2019). Development and evaluation of a triplex TaqMan assay and next-generation sequence analysis for improved detection of Xylella in plant material. Plant Disease,103, 645–655.
Chabirand, A., Jouen, E., Pruvost, O., et al. (2014). Comparative and collaborative studies for the validation of a nested PCR for the detection of X. axonopodis pv. dieffenbachiae from Anthurium samples. Plant Pathology,63, 20–30.
Chase, A. R., Stall, R. E., Hodge, N. C., & Jones, J. B. (1992). Characterization of X. campestris strains from aroids using physiological, pathological, and fatty-acid analysis. Phytopathology,82, 754–759.
Constantin, E. C., Cleenwerck, I., Maes, M., et al. (2016). Genetic characterization of strains named as X. axonopodis pv. dieffenbachiae leads to a taxonomic revision of the X. axonopodis species complex. Plant Pathology,65, 792–806.
Constantin, E. C., Haegeman, A., Van Vaerenbergh, J., et al. (2017). Pathogenicity and virulence gene content of Xanthomonas strains infecting Araceae, formerly known as X. axonopodis pv. dieffenbachiae. Plant Pathology,66, 1539–1554.
Cottyn, B., Regalado, E., Lanoot, B., De Cleene, M., Mew, T., & Swings, J. (2001). Bacterial populations associated with rice seed in the tropical environment. Phytopathology,91, 282–292.
Cottyn, B., Constantin, E., & Maes, M. (2018). Consequences of the taxonomic revision of X. axonopodis pv. dieffenbachiae and of the analysis of its associated pathogenicity for the recommendation of regulation (EPPO A2 List). EPPO Bulletin,48, 242–244.
Donahoo, R. S., Jones, J. B., Lacy, G. H., Stromberg, V. K., & Norman, D. J. (2013). Genetic analyses of X. axonopodis pv. dieffenbachiae strains reveal distinct phylogenetic groups. Phytopathology,103, 237–244.
Fang, Y., Lin, H., Wu, L., et al. (2015). Genome sequence of Xanthomonas sacchari R1, a biocontrol bacterium isolated from the rice seed. Journal of Biotechnology,206, 77–78.
Jouen, E., Chiroleu, F., Maillot-Lebon, V., et al. (2019). A duplex quantitative real-time PCR assay for the detection and quantification of X. phaseoli pv. dieffenbachiae from diseased and latently infected anthurium tissue. Journal of Microbiological Methods,161, 74–83.
Jun-Hai, N., Yue-Rong, G., Jun-Mei, Y., Qing-Yun, L., Guang-Sui, Y., Cun, W., & Yu, R. (2015). Development and evaluation of a loop-mediated isothermal amplification assay for rapid detection of bacterial blight pathogen (Xanthomonas axonopodis pv. dieffenbachiae) in anthurium. European Journal of Plant Pathology,142, 801–813.
Khoodoo, M. H. R., & Jaufeerally-Fakim, Y. (2004). RAPD-PCR fingerprinting and southern analysis of X. axonopodis pv. dieffenbachiae strains isolated from different aroid hosts and locations. Plant Disease,88, 980–988.
Khoodoo, M. H. R., Sahin, F., & Jaufeerally-Fakim, Y. (2005). Sensitive detection of X. axonopodis pv. dieffenbachiae on Anthurium andreanum by immunocapture-PCR (IC-PCR) using primers designed from sequence characterized amplified regions (SCAR) of the blight pathogen. European Journal of Plant Pathology,112, 379–390.
Kubista, M., Andrade, J. M., Bengtsson, M., et al. (2006). The real-time polymerase chain reaction. Molecular Aspects of Medicine,27, 95–125.
Laurent, P., Chabirand, A., Jouen, E., et al. (2009). A new semi-selective medium for the isolation of X. axonopodis pv. dieffenbachiae, the etiological agent of anthurium bacterial blight. Letters in Applied Microbiology,49, 210–216.
Lipp, R. L., Alvarez, A. M., Benedict, A. A., & Berestecky, J. (1992). Use of monoclonal antibodies and pathogenicity tests to characterize strains of X. campestris pv. dieffenbachiae from aroids. Phytopathology,82, 677–682.
Lu, M., Zhou, X., Zeng, W., Chen, Y., & Liu, Q. (2012). Specific PCR Detection of X. axonopodis pv. dieffenbachiae. Acta Phytopathologica Sinica,42, 1–9.
Martinelli, F., Scalenghe, R., Davino, S., et al. (2015). Advanced methods of plant disease detection. A review. Agronomy for Sustainable Development,35, 1–25.
Mirghasempour, S. A., Huang, S., Studholme, D. J., & Brady, C. L. (2020). A grain rot of rice in Iran caused by a Xanthomonas strain closely related to X. sacchari. Plant Disease,104, 1581–1583.
Mirmajlessi, S. M., Loit, E., Maend, M., & Mansouripour, S. M. (2015). Real-time PCR applied to study on plant pathogens: potential applications in diagnosis-a review. Plant Protection Science,51, 177–190.
Norman, D., & Alvarez, A. (1989). A rapid method for presumptive identification of X. campestris pv. dieffenbachiae and other Xanthomonads. Plant Disease,73, 654–658.
Norman, D. J., & Alvarez, A. M. (1994). Rapid detection of X. campestris pv. dieffenbachiae in Anthurium plants with a miniplate enrichment ELISA system. Plant Disease,78, 954–958.
Parkinson, N., Cowie, C., Heeney, J., & Stead, D. (2009). Phylogenetic structure of Xanthomonas determined by comparison of gyrB sequences. International Journal of Systematic and Evolutionary Microbiology,59, 264–274.
Rademaker, J. L. W., Louws, F. J., Schultz, M. H., et al. (2005). A comprehensive species to strain taxonomic framework for Xanthomonas. Phytopathology,95, 1098–1111.
Robene-Soustrade, I., Laurent, P., Gagnevin, L., Jouen, E., & Pruvost, O. (2006). Specific detection of X. axonopodis pv. dieffenbachiae in anthurium (Anthurium andreanum) tissues by nested PCR. Applied and Environmental Microbiology,72, 1072–1078.
Schaad, N. W., Cheong, S., Tamaki, S., Hatziloukas, E., & Panopoulos, N. J. (1995). A combined biological and enzymatic amplification (BIO-PCR) technique to detect Pseudomonas syringae pv. phaseolicola in bean seed extracts. Phytopathology,85, 243–246.
Schaad, N., Berthier-Schaad, Y., Sechler, A., & Knorr, D. (1999). Detection of Clavibacter michiganensis subsp. sepedonicus in potato tubers by BIO-PCR and an automated real-time fluorescence detection system. Plant Disease,83, 1095–1100.
Seemann, T. (2014). Prokka: rapid prokaryotic genome annotation. Bioinformatics,30, 2068–2069.
Song, W., Kim, H., Hwang, C., & Schaad, N. (2004). Detection of Acidovorax avenae ssp. avenae in Rice Seeds Using BIO-PCR. Journal of Phytopathology,152, 667–676.
Studholme, D. J., Wasukira, A., Paszkiewicz, K., et al. (2011). Draft genome sequences of X. sacchari and two banana-associated xanthomonads reveal insights into the Xanthomonas group 1 clade. Genes,2, 1050–1065.
Vauterin, L., Hoste, B., Kersters, K., & Swings, J. (1995). Reclassification of Xanthomonas. International Journal of Systemic Bacteriology,45, 472–489.
Young, J. M., Park, D. C., Shearman, H. M., & Fargier, E. (2008). A multilocus sequence analysis of the genus Xanthomonas. Systematic and Applied Microbiology,31, 366–377.
Acknowledgements
This research was funded by the Dutch Ministry of Agriculture, nature and Food Safety (LNV) in the Topsector Program “Horticulture and Starting Materials” under the theme “Phytosanitary Important for NL” (project number: TU-18095) and its partners (Naktuinbouw, Glastuinbouw NL, Plantum and VBN). The work is supported by COST (European Cooperation in Science and Technology) via grant CA16107 EuroXanth (https://euroxanth.eu).
Data availability statement
Data available on request from the authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors herewith declare that they have no conflict of interest. This study does not involve studies with human participants or animals performed by any of the authors.
Supplementary Information
ESM 1
Figure S1 Correlation in Ct values between TaqMan assay Xpd866 and Xpd4494 for detection of Xanthomonas phaseoli pv. dieffenbachiae (Xpd). The Ct values of Xpd866 and Xpd4494 were plotted on the x and y axis, respectively. In A, results for a gBlock, comprising the amplicon sequences of both assays, is shown in two individual TaqMan assays. In B, for genomic DNA and in C for a bacterial suspension in an Anthurium leaf extract. (PPTX 50 kb)
Table S1
(DOCX 13 kb)
Table S2–S4
(DOCX 15 kb)
Table S5–S8
(XLSX 17 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
van der Wolf, J., Krijger, M., Mendes, O. et al. Molecular characterization of Xanthomonas species isolated from Araceae and the development of a triplex TaqMan assay for detection of Xanthomonas phaseoli pv. dieffenbachiae. Eur J Plant Pathol 163, 167–179 (2022). https://doi.org/10.1007/s10658-022-02466-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-022-02466-1