Abstract
We analyzed the infection of the model plant Arabidopsis thaliana by the basidiomycete phytopathogen of cereals Sporisorium reilianum in order to use it as an experimental pathosystem model. Sterile plantlets of A. thaliana were grown on MS solid medium, and inoculated with haploid strains or mixtures of sexually compatible S. reilianum strains. Inoculated plants showed growth of filaments within their tissues, size reduction, a drastic increase in the formation of lateral roots, and a high production of plant pigments. Although symptoms were more severe in plants infected with sexually compatible strains, no spores were formed by the fungus. Among the pigments accumulated in the stunted plants we identified the anthocyanins cyanidin, cyanidin 3-glucoside, malvidin and pelargonidin. Congruently, the anthocyanin biosynthesis genes: chalcone synthase (CHS, AT5G13930) and dihydroflavonol reductase (DFR, AT5G42800) were over-expressed. The three genes encoding flavin monooxygenases: YUCCA7 (YUC7, AT2G33230), YUCCA8 (YUC8, AT4G28720), and YUCCA9 (YUC9, AT1G04180), and the transcription factor Jagged Lateral Organs (JLO, AT4G00220), all of them involved in the biosynthesis of auxins specific for root development, were also positively regulated. These data provide evidence that both haploid and the mixture of sexually compatible strains of S. reilianum can infect plants of A. thaliana; evidencing the usefulness of this pathosystem for the study of the genetic and metabolic reactions involved in S. reilianum virulence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Among the pathogenic microorganisms of plants, fungi are considered the most dangerous and devastating microbes. They cause severe damage in plants of economic and ecological importance throughout the world, affecting forests, horticulture, agriculture; and therefore also food security (van der Does and Rep 2017). Phytopathogenic fungi are mainly divided into necrotrophs and biotrophs according to their lifestyle. The first ones rapidly kill the plant to use it as nutritional material, whereas the second ones induce metabolic changes, and reprogram the fate of cell development in the host plant to allow their own growth without killing the host (Doehlemann et al. 2017).
Sporisorium reilianum is a biotropic dimorphic basidiomycete fungus with a saprobiotic yeast-like phase and a parasitic mycelial phase during its life cycle (Bhaskaran and Smith 1993). In nature, this fungus infects and causes the head smut disease in maize (Zea mays) and sorghum (Sorghum bicolor), two of the most economically important cereals in the world (Bhaskaran and Smith 1993; Martinez et al. 2002; Poloni and Schirawski 2016). The pathogenic phase of the fungus starts with the recognition and mating of two haploid yeast-like cells with compatible mating types through a pheromone receptor system, and formation of a dikaryotic filament that penetrates the plant roots (Martinez et al. 2002; Schirawski et al. 2005). Inside the host, the filaments colonize the plant, causing only chlorotic flecks and anthocyanin accumulation during the early stages of the infection process (Martinez et al. 2002); and it is only until the fungus invades the male and female inflorescences that occurs the formation of spores and the characteristics symptoms of the disease (Martinez et al. 2002; Schirawski et al. 2010). The spores are released to the environment and under favorable conditions germinate to form haploid yeast-like basidiospores that reinitiate the fungal life cycle (Martinez et al. 2002). Interestingly, S. reilianum exists in two forma specialis, which differ among them in the preference of their hosts; S. reilianum f. sp. reilianum (SRS) infects sorghum, and S. reilianum f. sp. zeae (SRZ) infects maize (Zuther et al. 2012). The preference of S. reilianum to infect maize or sorghum, is possibly due, as suggested by Poloni and Schirawski (2016), to the ability of each forma specialis of the fungus to avoid or respond to the defense mechanisms of their host plants.
An experimental strategy widely used to study the processes of pathogenesis from different microorganisms has been the use of non-natural or alternative hosts. In the literature there are many ancient and modern examples of the use of different and diverse alternative hosts to study the pathogenic processes developed by pathogenic fungi of animals or plants (some modern examples can be found in Perfect et al. 1986; Kontoyiannis and Lewis 2010; Martínez-Soto et al. 2013, 2016; Warpeha et al. 2013; Mazaheri-Naeini et al. 2015; Belhaj et al. 2017). Previously, our research group described the usefulness of the experimental pathosystem of A. thaliana with a fungal basidiomycete species closely related to S. reilianum, Ustilago maydis, also a maize pathogen (Méndez-Morán et al. 2005).
Considering that S. reilianum is an important plant pathogen that may be used as a model organism to study the molecular bases of the interactions between pathogenic fungi and their hosts (Schirawski et al. 2010); and that A. thaliana is a model host to study the response of plants against pathogens (Davis 1993; Martínez-Soto et al. 2013, 2016; Belhaj et al. 2017), in the present study we have proceeded to analyze whether the fungus is able to infect A. thaliana. The main objective was the possible development of a useful pathosystem for the study of the virulence mechanisms of the fungus, and the resistance mechanisms of the plant. The results obtained suggest that indeed, this experimental pathosystem may be useful for these aims.
Materials and methods
Fungal strains and plant materials
The wild type and sexually compatible haploid strains of Sporisorium reilianum SRZ1 (a1b1) and SRZ2 (a2b2) used in this study were kindly made available by Dr. J. Schirawski (Aachen University, Germany). The sexual compatibility of the strains used was corroborated by means of the Fuz reaction devised for U. maydis (Banuett and Herskowitz 1989). Arabidopsis thaliana L. Landsberg erecta (Ler) plants were used as the host for S. reilianum infection.
Sporisorium reilianum growth and plant inoculation
The strains of S. reilianum were maintained in 50% glycerol at −70 °C, and recovered in liquid complete medium MC (Holliday 1974), with shaking at 25 °C for 20 h, and used as inocula for the different experiments described below. For plant inoculation, the cells were recovered by centrifugation at 2500×g for 10 min, washed three times with sterile distilled water (SDW) and suspended in approximately 5 ml of SDW. Finally, cell concentration was determined with a Neubauer cytometer (Hausser Scientific, Horsham PA, US), and the cell density was adjusted to 106 cells per mL. Mixtures containing the same cell number of sexually compatible strains (a1b1 X a2b2) of S. reilianum were obtained by combining the same volumes of each cell suspension containing 106 cells per mL.
A. thaliana seeds were freed of contaminants using chlorine, and were grown on solid MS synthetic medium (Murashige and Skoog 1962), under sterile conditions. Plantlets were then inoculated after 6 days of germination with 1–2 μl of the cell suspension containing (106 cells/mL; 1000–2000 cells) of haploid (a2b2) strains, or mixtures of the sexually compatible strains (a1b1 X a2b2) of S. reilianum, according to Méndez-Morán et al. (2005).
Determination of the symptoms in infected plants
Symptoms of infection and damage in plants were observed with a stereoscope (Leica MZ-8, Illinois, US) coupled to a Spot digital camera (Diagnostic Instruments, Houston TX, US), and a digital microscope Keyence VHX5000 (Keyence, Japan). The photographs of primary roots were taken with a DMCFX12 camera (Panasonic, Osaka, Japan). At different days post-infection (dpi), the plants were recovered, dried in an oven at 60 °C, and their dry weight was measured.
S. reilianum growth in A. thaliana plants
Growth of the fungus inside the plants was observed at different dpi. Plants and hyphae were stained with propidium iodide (P4864 Sigma-Aldrich) and solophenyl flavine respectively according to the method described by Hoch et al. (2005), and photographed by fluorescence microscopy using a LSM 510 META confocal scanning laser inverted microscope (Carls Zeiss, Oberkochen, Germany). Also the tissue was bleached with boiling 70% ethanol for 5 min, stained with calcofluor white (18,909 Sigma-Aldrich), observed by epifluorecent with a Leica DMRE microscope (Leica microsystems; Wetzlar, DE), and photographed with a Leica DFC450 C camera (Leica microsystems; Wetzlar, DE). The biomass of S. reilianum in inoculated plantlets was indirectly measured by ergosterol determination in fresh whole plants (250 mg fresh weight) as described by Martínez-Soto et al. (2013).
Determination of anthocyanins
Visible changes in the amount of anthocyanins present in the inoculated plantlets were recorded by photography with a DMCFX12 camera (Panasonic, Osaka, Japan). To analyze quantitatively these pigments the method proposed by Abdel-Aal and Hucl (1999) was followed. Accordingly, fresh whole plants were ground with liquid N2 using a pestle and mortar. Five hundred mg of the resulting powder were then extracted under static conditions at 4 °C overnight with 15 mL of a mixture of 99% methanol:1 N HCl (90:10, by vol.). Anthocyanins were then analyzed by high performance liquid chromatography (HPLC) according to the method described by Kondo et al. (2002), and the conditions described by Garzón (2008), using for their identification the corresponding standard compounds of Sigma-Aldrich [cyanidin chloride (Cat. 79,457), cyanidin 3-O-glucoside chloride (Cat. 1,151,935), malvidin chloride (Cat. 68,120), pelargonidin chloride (Cat. P1659)] (Supplementary Fig. 1).
Gene expression analysis by quantitative real-time RT-PCR (qRT-PCR)
For qRT-PCR analyses, the body tissue or the roots of infected plants were collected, the total RNA was extracted using TRIzol (Invitrogen), and treated with DNAse I (Invitrogen). RNA concentration was measured by absorbance at 260 nm with a Nanodrop (Thermo Scientific, Waltham, MA, USA), and its integrity was determined by electrophoresis in agarose gels. Three biological replicates with the mixture of three technical replicas were done. qRT-PCR was performed using KAPA SYBR FAST qPCR Master Mix (2X) kit with ROX (Kapa Biosystems) according to the instructions of manufacturer in StepOne Real-Time PCR Systems (Applied Biosystems, Foster City, CA, USA). The genes corresponding to the analyzed transcripts, and the primers used appear listed in Supplementary Table 1, and the target gene expression levels were normalized using the gene UBQ10 (AT4G05320) (Weigel and Glazebrook 2002). The fold change in expression of the target genes in infected plants was calculated as described by Palomeros-Suárez et al. (2017).
Results
Symptoms of A. thaliana plants infected with S. reilianum
It was observed that both, haploid, and mixtures of sexually compatible strains of S. reilianum induced similar symptoms in inoculated plants of A. thaliana (see below). The results indicated that 95.0% (± 5.0%) and 98.4% (± 1.5%) of the plants infected with haploid strains or sexually compatible strains, respectively, showed symptoms of infection, although in none of them spores were formed.
Among the symptoms developed by the inoculated plants, stunting, darkening (later on identified as due to an overproduction of anthocyanins), and alterations in the root system were the most notorious ones, starting at 14 dpi, and being more clearly observed at 15 dpi (Fig. 1a–e). These symptoms became more severe with time. At 30 dpi, practically the whole plant was stained with red-purple pigments, and with a large number of lateral roots (Fig. 1g–k). Also, as part of the alterations in the plant root system, a decrease in the size of the primary root was observed (Fig. 2). In addition, S. reilianum reduced the size of inoculated Arabidopsis plants, their dry weight was approximately reduced 46.0% or 64.0% after 20 dpi in the plants infected with haploid strains or sexually compatible strains, respectively (Fig. 3).
Images of A. thaliana plants infected or not with S. reilianum strains. a and d, respectively, leaves and roots of plants infected with sexually compatible strains of S. relianum at 15 dpi; b and e, leaves and roots of plants infected with S. relianum haploid strains at 15 dpi. g and j, plant and roots of plant infected with sexually compatible strains of S. reilianum at 30 dpi; h and k, leaves and roots of plants infected with S. relianum haploid strains at 30 dpi. c and f, leaves and roots of a control plant at 15 days post-application of sterile distilled water (SDW); i and l, a whole plant and roots of a control plant at 30 days post-application of SDW. Notice in a, b, g and h; the presence of an intense red-purple colorations in the tissues of infected plants compared to tissues of control plants (c and i). Notice in d and e, initial alterations in the root system of inoculated plants at 15 dpi compared to roots of control plants (f). Notice in j and k, severe alteration in the root system of plants infected at 30 dpi compared to roots of not inoculated control plants (l)
Representative photographs of the length of the bodies and the main roots of A. thaliana plants infected with S. reilianum at 30 dpi. Left of plates, three control plants that received SDW only. Right of plates, three plants infected with mixture of sexually compatible (a) or haploid strains (b) of S. reilianum
Growth of A. thaliana plants infected or not with S. reilianum. Gray bar represents the dry weight of control plants that received SDW only. Black and white bars represent the dry weight of plants infected with sexually compatible or haploid (SRZ2) strains of A. thaliana respectively. Results from three independent experiments, with three replicas in each one using ten plants. Lines on each bar represent standard error values. Different letters denote significant differences (ANOVA, Tukey)
Growth of S. reilianum within Arabidopsis plantlets
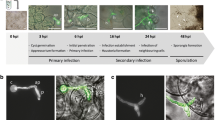
Using fluorescence microscopy, filaments of S. reilianum stained in green color was observed within the red fluorescent tissues of all infected plants (Fig. 4a–b). Also, the filaments of S. reilianum stained in blue color was observed inside the cells of infected plants tissue (Fig. 4c).
Representative micrograph of an A. thaliana plant tissue infected with compatible strains of S. reilianum at 15 dpi. a and b, Plant tissue was stained with propidium iodide and appears in red color; fungal hyphae were stained with solophenyl flavine and appear in green color (white arrows). c, Fungal hyphae stained with calcofluor white (black arrow). Scale bar, 100 μm
To quantify the development of S. reilianum within the infected plants, the amount of ergosterol was measured. Ergosterol is a compound present in fungi, but absent in plants (Martin et al. 1990). It was found that the amount of this compound was slightly higher in the plants infected with sexually compatible strains than in plants infected with haploid strains (Fig. 5).
Determination of ergosterol in A. thaliana plants infected or not with S. reilianum. Empty squares joined with a solid line represent the ergosterol content in plantlets infected with sexually compatible strains of S. reilianum. Empty circles joined with a dotted line represent the ergosterol content of plantlets infected with haploid strains. Empty gray triangles joined with a gray solid line represent the ergosterol content of control plantlets that received SDW only. Results from three independent experiments with two replicas in each one, using 250 mg of fresh plant tissue. Bars represent standard error values. Different letters denote significant differences (ANOVA, Tukey)
Production of anthocyanins in infected Arabidopsis plants
The most apparent symptom of diseased plants was their intense purple color (Fig. 1), easily evident to the naked eye once extracted (Fig. 6a). Considering that this pigment might correspond to anthocyanins, we proceeded to extract it with HCl-methanol, and determine its concentration in infected and control plants by spectrophotometry (Fig. 6b). It was found that the infected plants contained high levels of anthocyanins, approximately twenty times more than the un-inoculated control plants (Fig. 6b).
Production of anthocyanins in A. thaliana plants infected or not with S. reilianum at 30 dpi. a, photograph of tubes containing the solution of anthocyanins extracted with HCl-methanol for 1 day at 4 °C in darkness, from A. thaliana plants infected or not with S. reilianum. b, Quantitative determination of total anthocyanins in A. thaliana plants infected or not with S. reilianum. Data were obtained from three different experiments, with three replicas in each one. Lines on each bar represent standard error values. Different letters denote significant differences (ANOVA, Tukey)
A quantitative analysis of the anthocyanins revealed that they were a mixture of the followings in order of abundance: cyanidin, pelargonidin, malvidin, and cyanidin 3-glucoside (Table 1), their amounts being slightly higher in plants infected with sexually compatible strains.
Expression of genes involved in the biosynthesis of plant pigments
Due to the high production of anthocyanins observed in the infected plants, the levels of expression of the genes encoding chalcone synthase (CHS) and dihydroflavonol reductase (DFR) were analyzed by qRT-PCR in infected plants at 23 dpi. The first gene is important in the biosynthesis pathway of flavonoids; and the second one is a gene involved in converting flavonoids to anthocyanins (Koes et al. 2005). Both genes were overexpressed in the infected plants, and especially higher levels of expression of CHS were observed in the plants infected with the sexually compatible strains (Fig. 7).
Expression of A. thaliana genes involved in the biosynthesis of anthocyanins. Bars represent the expression of Chalcone synthase (CHS) and Dihydroflavonol reductase (DFR) genes. Data were obtained from three different experiments, with two replicas in each one and are related to the expression s of the constitutive gene UBQ10. Lines on each bar represent standard error values
Expression of genes involved in the biosynthesis of auxins
As described above, a large number of lateral roots were observed in the infected plants (Fig. 1), accordingly, we analyzed by qRT-PCR the levels of expression of YUC7, YUC8, and YUC9 genes encoding enzymes of the flavin monooxygenase family; and the one encoding the transcription factor Jagged Lateral Organs (JLO), all of them involved in the biosynthesis of auxins specific for root development (Zhao 2012; Rast-Somssich et al. 2017). Interestingly, all the genes analyzed were overexpressed in the infected plants, and particularly was the expression of the transcription factor JLO (Fig. 8).
Expression of A. thaliana genes involved in the biosynthesis of auxins. Expression of three Flavin monooxygenase genes: YUC7, YUC8 and YUC9; and the transcription factor Jagged Lateral Organs (JLO); all of them involved in the biosynthesis of auxins related to root development. Data were obtained from three different experiments, with two replicas in each one, and are related to the expression of the constitutive gene UBQ10. Lines on each bar represent standard error values
Discussion
In this work, the capacity of haploid strains or mixtures of sexually compatible strains of S. reilianum SRZ to invade A. thaliana, was demonstrated. This result is noticeable considering that A. thaliana is a plant phylogenetically distant from maize, the natural host of the fungus. This result is similar to our previous demonstration that also Ustilago maydis, a close relative of S. reilianum (Schirawski et al. 2010), is able to invade A. thaliana under axenic conditions (Méndez-Morán et al. 2005; Martínez-Soto et al. 2013, 2016). In addition, it was recently described that U. maydis is also able to infect Medicago truncatula (Mazaheri-Naeini et al. 2015). All these experimental pathosystems offer the possibility to analyze the mechanisms involved in the virulence of these important cereal pathogens, and the plant resistance under controlled conditions.
Our results show that similarly to the U. maydis-A. thaliana pathosystem (Méndez-Morán et al. 2005; Martínez-Soto et al. 2013, 2016), there was no production of teliospores in the virulent process developed by sexually compatible strains of S. reilianum in A. thaliana; which are formed only in maize plants infected by the dikaryotic filament of sexually compatible strains (Banuett and Herskowitz 1989; Banuett 1995; Martinez et al. 2002). However, contrary to the infection of A. thaliana by U. maydis, where the fungus grows inside and on the surface of the plant tissues, and the symptoms of infection appear at early stages [1 or 2 dpi (Méndez-Morán et al. 2005; Martínez-Soto et al. 2013, 2016)], S. reilianum only colonizes the tissues of the Arabidopsis plant, and the symptoms of infection appear at later time points (14 dpi), becoming very severe at about 23–30 dpi. It is interesting to notice that of the main symptoms developed in Arabidopsis infected by S. reilianum, stunting and over-production and accumulation of anthocyanins, are also present in common and susceptible maize lines infected with S. reilianum SRZ (Martinez et al. 2002); although in the pathosystem here presented, these symptoms are noticeably exaggerated.
The development of these symptoms in the infected Arabidopsis plants by S. reilianum can be a combination of the defense mechanisms developed by the plant against the fungus, and the ability of the pathogen to manipulate the plant metabolism, similarly as it does in its natural host (Poloni and Schirawski 2016). We found that the high production and accumulation of anthocyanins in the infected plants, was accompanied by an over-expression of the genes encoding chalcone synthase (CHS) and dihydroflavonol reductase (DFR). These genes are involved in the synthesis pathway of anthocyanins and phytoalexins (Koes et al. 2005; Zuther et al. 2012). In the literature, it has been described that sorghum plants infected with S. reilianum SRZ show an over-expression of DFR, and the formation of red spots in the tissues of the host plants, corresponding to an overproduction of the phytoalexin Luteolinidin, as a defense mechanism against the growth and development of the fungus (Zuther et al. 2012). Likewise, Ibraheem et al. (2015) demonstrated that 3-deoxyanthocyanidins are involved in the defense and resistance of maize transgenic plants against the fungus Colletotrichum graminicola. It is known that anthocyanins inhibit the growth of pathogenic microorganisms of plants, mainly bacteria, inducing cellular malformations, destruction of the structural integrity of their cell wall and membrane, and affecting their general metabolism (reviewed by Khoo et al. 2017). Also, recently it was suggested that an increase in the synthesis and accumulation of anthocyanins in plants induces a significant reduction in their size and in the length of their primary roots (Li et al. 2018). Although the cause and effect of this relationship was not clarified, this suggestion agrees with our observations on the phenotype of Arabidopsis plants infected with S. reilianum.
It is known that anthocyanins regulate all cellular processes involved in plant growth and development, such as cellular division and growth, tissue development, organ development, development and architecture of the root system, including lateral root development (reviewed by Olatunji et al. 2017). It is also known that phytohormones regulate the expression of genes involved in the synthesis of auxins that play a role in the formation and development of the root system of plants (Cai et al. 2014). In this sense it has been described that jasmonic acid regulates the expression of the genes YUC8 and YUC9 encoding the flavin monooxygenase enzymes which inhibit the elongation of the primary root, and induce the formation of lateral roots (reviewed by Olatunji et al. 2017), just as occurred in Arabidopsis plants infected with S. reilianum. Also, it has been described that these two genes, together with YUC3, YUC5, and YUC7; are responsible for the production of auxins in the roots of A. thaliana (Zhao 2012); and additionally that the expression of the Jagged Lateral Organs (JLO) transcription factor is required to coordinate the development of the entire root system of the plant (Rast-Somssich et al. 2017). In line with all these observations, we found that all these genes were over-expressed in the Arabidopsis plants that were infected with S. reilianum, being particularly noticeable the high levels of expression of the JLO gene, which as described above, is the master transcription factor that regulates all the genetic machinery of the plant root formation.
It is important to recall that S. reilianum SRZ has a biotrophic behavior during the infection of its natural host, maize (Schirawski et al. 2010). Similarly, during the early stages of the pathogenic process developed in Arabidopsis plants, from 1 to ca. 13 dpi, S. reilianum showed a biotrophic behavior without causing apparent symptoms of infection; but at later stages of the infection (ca. 14 to 30 dpi under our conditions), there occurred accumulation of anthocyanins, stunting, and the formation of lateral roots. Possibly at this late stage of the infective process, there occurs induction in the production of jasmonic acid, as a defense mechanism of the plant against S. reilianum, and in turn, this induces the expression of YUC genes as described above; causing the formation of a large number of lateral or secondary roots, and inhibiting the growth of the primary root. Interestingly, we observed that in A. thaliana plants infected with U. maydis there occurred increase in jasmonic acid formation (Martínez-Soto et al. 2016).
In summary, the results here presented constitute a strong evidence of the usefulness of this experimental pathosystem for the study of the mechanisms of virulence of the fungus, and those of the resistance of the plant. Considering the experimental advantages of A. thaliana as an alternative host because of its facility for genetic manipulation, its availability of mutants, small size, and the short time that elapses before the appearance of symptoms, contrasting with that of its natural host, its usefulness becomes evident.
References
Abdel-Aal, E. S. M., & Hucl, P. (1999). A rapid method for quantifying total anthocyanins in blue aleurone and purple pericarp wheats. Cereal Chemistry, 76, 350–354.
Banuett, F. (1995). Genetics of Ustilago maydis, a fungal pathogen that induces tumors in maize. Annual Review of Genetics, 29, 179–208.
Banuett, F., & Herskowitz, I. (1989). Different a alleles of Ustilago maydis are necessary for maintenance of filamentous growth but not for meiosis. Proceedings of the National Academy of Sciences of the United States of America, 86, 5878–5882.
Belhaj, K., Cano, L. M., Prince, D. C., Kemen, A., Yoshida, K., Dagdas, Y. F., Etherington, G. J., Schoonbeek, H. J., van Esse, H. P., Jones, J. D. G., Kamoun, S., & Schornack, S. (2017). Arabidopsis late blight: infection of a nonhost plant by Albugo laibachii enables full colonizationby Phytophthora infestans. Cellular Microbiology, 19, e12628. https://doi.org/10.1111/cmi.12628.
Bhaskaran, S., & Smith, R. H. (1993). Carbohydrates, inertase activity, growth and dimorphism in Sporisorium reilianum. Mycopathologia, 122, 35–41.
Cai, X. T., Xu, P., Zhao, P. X., Liu, R., Yu, L. H., & Xiang, C. B. (2014). Arabidopsis ERF109 mediates cross-talk between jasmonic acid and auxin biosynthesis during lateral root formation. Nature Communications, 5, 5833. https://doi.org/10.1038/ncomms6833.
Davis, K. R. (1993). Arabidopsis thaliana as a model for plant-pathogen interactions. In R. Hammershmidt (Ed.), The American phytopathological society (pp. 1–134). APS Press: St. Paul.
Doehlemann G., Ökmen B., Zhu W., & Sharon A. (2017). Plant pathogenic fungi. Microbiology Spectrum, 5. https://doi.org/10.1128/microbiolspec.FUNK-0023-2016.
Garzón, G. A. (2008). Anthocyanins as natural colorants and bioactive compounds. A review. Acta Biológica Colombiana, 13, 27–36.
Hoch, H. C., Galvani, C. D., Szarowski, D. H., & Turner, J. N. (2005). Two new fluorescent dyes applicable for visualization of fungal cell walls. Mycologia, 97, 580–588.
Holliday, R. (1974). Ustilago maydis. In R. C. King (Ed.), The handbook of genetics (pp. 575–595). New York: Plenum Press.
Ibraheem, F., Gaffoo, I., Tan, Q., Shyu, C. R., & Chopra, S. (2015). A sorghum MYB transcription factor induces 3-deoxyanthocyanidins and enhances resistance against leaf blights in maize. Molecules, 20, 2388–2404.
Khoo, H. E., Azlan, A., Tang, S. T., & Lim, S. M. (2017). Anthocyanidins and anthocyanins: colored pigments as food, pharmaceutical ingredients, and the potential health benefits. Food & Nutrition Research, 61, 1361779.
Koes, R., Verweij, W., & Quattrocchio, F. (2005). Flavonoids: a colorful model for the regulation and evolution of biotechemical pathways. Trends in Plant Science, 10, 236–242.
Kondo, S., Hiraoka, K., Kobayashi, S., Honda, C., & Terahara, N. (2002). Changes in the expression of anthocyanin biosynthetic genes during apple development. Journal of the American Society for Horticultural Science, 127, 971–976.
Kontoyiannis, D. P., & Lewis, R. E. (2010). Studying fungal pathogenesis in Drosophila. Microbe, 5, 291–295.
Li, N., Wu, H., Ding, Q., Li, H., Li, Z., Ding, J., & Li, Y. (2018). The heterologous expression of Arabidopsis PAP2 induces anthocyanin accumulation and inhibits plant growth in tomato. Functional & Integrative Genomics, 18, 341–353. https://doi.org/10.1007/s10142-018-0590-3.
Martin, F., Delareulle, C., & Hilbert, J. L. (1990). An improved ergosterol assay to estimate fungal biomass in ectomycorrhizas. Mycological Research, 94, 1059–1064.
Martinez, C., Roux, C., Jauneau, A., & Dargent, R. (2002). The biological cycle of Sporisorium reilianum f. sp. zeae: an overview using microscopy. Mycologia, 94, 505–514.
Martínez-Soto, D., Robledo-Briones, A. M., Estrada-Luna, A. A., & Ruiz-Herrera, J. (2013). Transcriptomic analysis of Ustilago maydis infecting Arabidopsis reveals important aspects of the fungus pathogenic mechanisms. Plant Signaling & Behavior. https://doi.org/10.4161/psb.25059.
Martínez-Soto, D., Pérez-Garcia, F. E., & Ruiz-Herrera, J. (2016). Arabidopsis infection by haploid or diploid strains of Ustilago maydis reveals its capacity as a necrophic or biotrophic phytopathogen. Fungal Genomics and Biology. https://doi.org/10.4172/2165-8056.1000133.
Mazaheri-Naeini, M., Sabbagh, S. K., Martinez, Y., Séjalon-Delmas, N., & Roux, C. (2015). Assessment of Ustilago maydis as a fungal model for root infection studies. Fungal Biology, 119, 145–153.
Méndez-Morán, L., Reynaga-Peña, C. G., Springer, P. S., & Ruiz-Herrera, J. (2005). Ustilago maydis infection of the nonnatural host Arabidopsis thaliana. Phytopathology, 95, 480–488.
Murashige, T., & Skoog, F. (1962). A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiologia Plantarum, 15, 473–497.
Olatunji, D., Geelen, D., & Verstraeten, I. (2017). Control of endogenous auxin levels in plant root development. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms18122587.
Palomeros-Suárez, P. A., Massange-Sánchez, J. A., Sánchez-Segura, L., et al. (2017). AhDGR2, an amaranth abiotic stress-induced DUF642 protein gene, modifies cell wall structure and composition and causes salt and ABA hyper-sensibility in transgenic Arabidopsis. Planta, 245, 623–640.
Perfect, J. R., Savani, D. V., & Durack, D. T. (1986). Comparison of itraconazole and fluconazole in treatment of cryptococcal meningitis and candida pyelonephritis in rabbits. Antimicrobial Agents and Chemotherapy, 29, 579–583.
Poloni, A., & Schirawski, J. (2016). Host specificity in Sporisorium reilianum is determined by distinct mechanisms in maize and sorghum. Molecular Plant Pathology, 17, 741–754.
Rast-Somssich, M. I., Zádníkova, P., Schmid, S., Kieffer, M., Kepinski, S., & Simon, R. (2017). The Arabidopsis Jagged Lateral Organs (JLO) gene sensitizes plants to auxin. Journal of Experimental Botany, 68, 2741–2755.
Schirawski, J., Heinze, B., Wagenknecht, M., & Kahmann, R. (2005). Mating type loci of Sporisorium reilianum: novel pattern with three a and multiple b specificities. Eukaryotic Cell, 4, 1317–1327.
Schirawski, J., Mannhaupt, G., Münch, K., et al. (2010). Pathogenicity determinants in smut fungi revealed by genome comparison. Science, 330, 1546–1548.
van der Does, H. C., & Rep, M. (2017). Adaptation to the host environment by plant-pathogenic fungi. Annual Review of Phytopathology, 55, 427–450.
Warpeha, K. M., Park, Y. D., & Williamson, P. R. (2013). Susceptibility of intact germinating Arabidopsis thaliana to human fungal pathogens Cryptococcus neoformans and C. gattii. Applied and Environmental Microbiology, 79, 2979–2988.
Weigel, D., & Glazebrook, J. (2002). Arabidopsis: A laboratory manual. New Yok: Cold Spring Harbor Laboratory Press.
Zhao, Y. (2012). Auxin biosynthesis: a simple two-step pathway converts tryptophan to indole-3-acetic acid in plants. Molecular Plant, 5, 334–338.
Zuther, K., Kahnt, J., Utermark, J., Imkampe, J., Uhse, S., & Schirawski, J. (2012). Host specificity of Sporisorium reilianum is tightly linked to generation of the phytoalexin luteolinidin by Sorghum bicolor. Molecular Plant-Microbe Interactions, 25, 1230–1237.
Acknowledgments
This work was partially supported by Consejo Nacional de Ciencia y Tecnología (Conacyt), México. Thanks are given to Prof. Jan Schirawski, Institute of Applied Microbiology, RWTH Aachen University, Germany, for making available the S. reilianum strains SRZ2 and SRZ1. We also thank Mayela F. Salazar-Chávez from the Irapuato Unit of Centro de Investigación y de Estudios Avanzados del Instituto Politécnico Nacional (Cinvestav), Leandro A. Núñez-Muñoz and Brenda J. Vargas-Hernández from Zacatenco Unit of Cinvestav; Renata Cruz-Calderon from Instituto Tecnológico Superior de Los Reyes, TSLR; and Drs. José I. Reyes-Olalde, and Stefan de Folter from the Unit of Advance Genomics, Cinvestav, for their assistance respectively in strain maintenance, preparation of some reagents, statistical analyses, and microscopic analyses.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they are aware of the ethical responsibilities required by the European Journal of Plant Pathology for the submission of manuscripts. Also, they declare no conflict of interests and that they were informed of the submission of the manuscript.
Rights and permissions
About this article
Cite this article
Martínez-Soto, D., Velez-Haro, J.M., León-Ramírez, C.G. et al. The cereal phytopathogen Sporisorium reilianum is able to infect the non-natural host Arabidopsis thaliana. Eur J Plant Pathol 153, 417–427 (2019). https://doi.org/10.1007/s10658-018-1567-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-018-1567-8