Abstract
In this study, the Estonian population of Phytophthora infestans was characterized with mating type, sensitivity to metalaxyl, virulence on 11 potato R-gene differentials and 12 SSR markers to show the outcome of potential sexual reproduction in the population. During the three years 2010–2012, 141 P. infestans isolates, collected from 23 potato fields, showed quite a high and stable frequency of the A2 mating type, 48% of the total population. In 87% of all sampled potato fields, both mating types were recorded, suggesting continuous sexual reproduction of P. infestans and possible oospore production. Metalaxyl-sensitive isolates prevailed in all three years (68 out of 99 isolates). Amongst the 95 isolates tested, 51 virulence races were found. The race structure was diverse, and most pathotypes were unique, appearing only once; the two most common pathotypes, 1.2.3.4.6.7.10.11 and 1.2.3.4.7.10.11, comprised 35% of the population. The P. infestans population was genetically highly diverse and most of the multilocus genotypes (MLGs) appeared only once. Furthermore, all of the MLGs appeared in only one of the three sampling years. Our results confirm that the high diversity in the Estonian P. infestans population is most likely the result of frequent sexual reproduction, which benefits the survival, adaptability and diversity of the pathogen in the climate of North-Eastern Europe.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Late blight, caused by the oomycete pathogen Phytophthora infestans, which made its first appearance in Europe in the mid-1840s, remains a major threat to European potato crops. Despite recent active research and progress, late blight still requires vigilance and often numerous applications of fungicide for effective control (Cooke et al. 2011). The disease is a serious problem for Estonian potato production, particularly under favourable conditions, when it can destroy the whole potato haulm and can cause 20–25% or even more loss of yield in untreated fields (Runno-Paurson et al. 2010). Fungicides are used routinely in conventional potato production, but under favourable conditions for the disease, with heavy pressure from the pathogen, timing the first preventive spraying and later protection of large areas is complicated without using information from decision support systems (DSS) (Runno-Paurson et al. 2010).
The oomycete P. infestans is heterothallic with two mating types, A1 and A2, enabling the pathogen to reproduce both sexually and asexually (Fry et al. 1993). Before 1980, the population of the late blight pathogen, worldwide except in Mexico, appeared to be asexual and to consist of a single clonal lineage with only the A1 mating type. Nevertheless, moderately complex pathogen races dominated among the European populations (Shattock et al. 1977; Goodwin et al. 1994; Gisi and Cohen 1996). After the migration of new genotypes and the A2 mating type, apparently via potato imports from Mexico into Europe in the late 1970s, sexual reproduction of the pathogen became possible (Fry et al. 1993). In Europe, the A2 mating type was first reported in Switzerland in 1981 (Spielman et al. 1991) and six years later in 1987 in Estonia (Vorobyeva et al. 1991). New genotypes were more competitive, adaptable to the host and environment and therefore spread quickly across Europe displacing the old clonal lineage (Spielman et al. 1991; Gisi and Cohen 1996), which latterly has been found only rarely (Cooke et al. 2011). Sexual reproduction results in oospores which can survive in the soil for several years (Drenth et al. 1995); they may also survive cold temperatures which may even conserve their viability (Turkensteen et al. 2000; Lehtinen and Hannukkala 2004). The presence of sexual reproduction changes the epidemiology (Mayton et al. 2000) and increases the adaptability of the pathogen, and thus changes the way in which disease control must be approached (Cooke et al. 2011; Yuen and Andersson 2013).
There are clear indications that changes have occurred in P. infestans populations worldwide in the last decade, which concur with the increasing severity of late blight outbreaks on potato and tomato crops (Montarry et al. 2010; Cooke et al. 2012; Fry et al. 2013; Li et al. 2013b; Chowdappa et al. 2015). In the Nordic region, there are indications of earlier outbreaks of late blight, requiring more frequent fungicide treatments per season to control the disease, probably partly because of oospore-derived infections (Hannukkala et al. 2007; Hannukkala 2012; Yuen and Andersson 2013). Long-term observation data from Finland and Estonia show that the first findings of blight now occur one month earlier than 20 years ago and blight outbreaks are more severe (Hannukkala et al. 2007; Runno-Paurson et al. 2013a). In 2014, late blight infection was registered earlier than ever before, already on 19 June in several large potato production fields in southern Estonia (monitored by E. Runno-Paurson).
In addition to oospore-caused infections, increasing severity of late blight outbreaks could also be associated with specific pathogen genotypes. To identify individual genotypes and to explore population diversity, molecular markers have to be used. Simple-sequence repeat (SSR) markers are neutral, co-dominant, polymorphic, single-locus molecular markers, and therefore are considered to be the most informative and effective in pathogen population studies (Cooke and Lees 2004; Lees et al. 2006). From the beginning of the twenty-first century SSR markers have been used to characterise P. infestans populations all over the world and at present this is the preferred genotyping method (Knapova and Gisi 2002; Lees et al. 2006; Montarry et al. 2010; Gisi et al. 2011; Cooke et al. 2012; Li et al. 2013a; Chowdappa et al. 2015). Genotyping results have revealed that most of the populations in Great Britain, France, Switzerland, Netherlands, Belgium as well as in North America, India and China are clonal with few dominating genotypes in the populations probably with rare sexual reproduction events (Montarry et al. 2010; Gisi et al. 2011; Cooke et al. 2012; Hu et al. 2012; Li et al. 2012, 2013b; Fry et al. 2013; Chowdappa et al. 2015). Conversely, recent studies in the Nordic countries (Finland, Sweden, Norway, Denmark), Russia and Poland have shown high genotypic diversity in P. infestans populations with many different genotypes present indicative of the relevance of sexual reproduction in these populations (Widmark et al. 2007; Brurberg et al. 2011; Sjöholm et al. 2013; Chmielarz et al. 2014; Statsyuk et al. 2014; Brylinska et al. 2016).
Estonian populations of P. infestans have previously been investigated with several phenotypic and molecular markers in 2001–2007. The P. infestans pathotype structure was highly diverse and complex and also a high number of fingerprint genotypes and considerable number of metalaxyl-resistant isolates were identified in Estonia (Runno-Paurson et al. 2009, 2010, 2012, 2013b, 2014). Previous studies have shown a steady almost equal ratio of A1 and A2 mating types in the population and furthermore, both mating types were found in most of the studied potato fields (more than 80%). All this indicates the high potential for sexual recombination in the pathogen population (Runno-Paurson et al. 2009, 2010, 2012, 2013b, 2014). Due to sexual reproduction, there is an ongoing change in genotypes and continuous diversification occurring in the Estonian P. infestans population, and this should be continuously monitored. Whereas late blight is still a serious threat to Estonian potato production, pathogen population studies are needed to efficiently control the pathogen population and manage late blight. Passing data on P. infestans genotypes through national and international networks to growers, advisors, breeders and agrochemical companies has already improved awareness and has had success in transfer of knowledge of population structure for adoption in practical late blight control (EuroBlight 2017; USAblight 2017).
In a previous study, nine polymorphic SSR markers were used to genotype Estonian P. infestans isolates from 2004 (Runno-Paurson et al. 2016), and a very high genotypic diversity was detected. In this more detailed study, more isolates from the main potato-growing areas in Estonia during a three-year period (2010–2012) were isolated and characterized phenotypically with mating type, resistance to metalaxyl, virulence assays and genotypically with a novel standardised SSR marker 12-plex approved and used by the EuroBlight network for monitoring P. infestans populations since 2013. Therefore, P. infestans genotypes and population structure in Estonia can be compared to the pathogen population situation in the whole of Europe. The main aim was to monitor recent changes in spatio-temporal variation in the population of P. infestans in Estonia. The following hypotheses were tested: 1) sexual reproduction is frequent in the population; 2) sensitivity to metalaxyl has increased; 3) a high number of virulence pathotypes are present; 4) no invasive clonal lineages are spreading or dominating in the population.
Materials and methods
Collection and isolation of P. infestans strains
Potato leaves infected by P. infestans were collected from 23 sites within the main potato-growing areas of Estonia (seven sites in 2010, eight sites in both 2011 and 2012) (Fig. 1, Table 1). The samples were collected from conventional production fields, organic fields and potato field trials, and at the beginning of late blight infection, in mid-outbreak (1–2 weeks later) and at the end of the growing season (>3 weeks later) each year. In the early stages of the outbreak, approximately 10–15% of the leaf area of the infected plants and less than 10% of plants were infected with late blight. In the later stages, about 20–30% of the leaf area and more than 50% of the plants were infected. Several fungicides with active ingredients such as metalaxyl + mancozeb, fluopicolide + propamocarb hydrochloride, cyazofamid, fenamidone + propamocarb hydrochloride, fluazinam, and mandipropamid were applied at different sites.
Two to fifteen isolates were cultured from each sampling site (Table 1). The plants from which samples were collected were located randomly across the field. From each plant, only single-lesion leaves were taken at random, excluding any with several or no lesions. Isolations were carried out and maintained using methods described by Runno-Paurson et al. (2009). All phenotypic tests were carried out every year of the study immediately after the isolations were finished (October to January). P. infestans isolates from this study are preserved at the Tartu Fungal Collection (TFC) in Estonia.
Phenotypic assays
Mating types were determined by the method described in Runno-Paurson et al. (2009). The tester isolates were 90,209 (A1) and 88,055 (A2) as described in Hermansen et al. (2000). Isolates forming oospores on plates with the A1 mating type were registered as A2; isolates that formed oospores with the A2 mating type were registered as A1.
Resistance to metalaxyl was tested using a modification of the floating leaflet method (Hermansen et al. 2000). Leaf disks (14 mm diameter) were cut with a cork borer from leaves of five-week-old greenhouse-grown plants. The susceptible cultivar ‘Berber’ was used. Six leaf disks were floated abaxial side up in Petri plates (50 mm diameter) each containing 7 mL of distilled water or metalaxyl in concentrations of 10.0 or 100.0 mg L−1 prepared from technical grade mefenoxam (Syngenta experimental compound (metalaxyl-M), CGA 329351A). The inoculation and trial incubation was done as described by Runno-Paurson et al. (2009). The isolates were rated resistant if they sporulated on leaf disks in 100 mg L−1 metalaxyl (Hermansen et al. 2000). Those sporulating on leaf disks in a metalaxyl concentration of 10 mg L−1, but not on leaves floating on 100 mg L−1 were rated intermediate, and those sporulating only in water were rated sensitive.
The virulence pathotype was determined in 2011 and 2012 with a detached-leaflet set of Black’s differentials of potato genotypes containing resistance genes R1-R11 from Solanum demissum (Malcolmson and Black 1966) (provided by the Scottish Agricultural Science Agency). Laboratory procedures were as described in Runno-Paurson et al. (2009). Pathotypes were not tested in 2010 for logistic reasons.
Genotyping
Pure-culture P. infestans isolates were grown on rye B agar plates for 2–4 weeks at 17 °C. Mycelium was harvested and frozen. DNA extractions were carried out with the DNeasy Plant Mini Kit (QIAGEN), according to the manufacturer’s instructions. Genotyping was performed at The James Hutton Institute (Scotland, UK), following the Li et al. (2013a) SSR marker 12-plex method with some modifications. One primer for each locus was labelled with different fluorescent labels (6FAM; NED; VIC; PET, Applied Biosystems) ensuring that no two markers with the same fluorescent dye had overlapping allele size ranges. PCR reactions were performed in a volume of 12.5 μl consisting of 1× Type-it Multiplex PCR Master Mix (QIAGEN Type-it Microsatellite PCR Kit, Cat. No. 206243), primers for each locus in optimal concentration (Online Resource 1) and 1 μl of template DNA (approximately 20 ng μl−1). Amplification reactions were carried out under the following conditions: 95 °C for 5 min followed by 28 cycles of 95 °C for 30 s, 58 °C for 90 s, and 72 °C for 20 s, and a final extension at 60 °C for 30 min.
PCR products were diluted 0–100 times depending on the initial DNA concentration in the PCR reaction mix. The diluted PCR product (0.6 μl) was added to a mix containing 10.14 μl of deionized formamide (Hi-Di Formamide, Applied Biosystems) and 0.06 μl of GeneScan-500LIZ standard (Applied Biosystems). The fluorescent-labelled PCR products were analysed using an ABI3730 DNA Analyzer (Applied Biosystems) according to the manufacturer’s instructions. The size of the alleles was determined using GeneMapper v3.7 (Applied Biosystems) software.
Data analysis
Statistical analyses were performed with the SAS/STAT version 9.1 (SAS Institute Inc., Cary, NC, USA). Differences in the prevalence of the two mating types of P. infestans isolates between years and study sites were tested using a logistic analysis (GENMOD procedure in SAS) with a multinomial response variable (A1, A2, or both). Analogous logistic procedures were used to examine the differences in the resistance to metalaxyl (a multinomial response variable: resistant, intermediate or sensitive) between years, sites and also between different mating types. The dependence of specific virulence (percent of isolates that show virulence against particular R-genes) on years, sites and R-genes was analysed with type III ANOVA and Tukey HSD post-hoc tests (α = 0.05). In all analyses, “year” and “site” were treated as categorical variables. Race diversity was calculated with the normalized Shannon’s diversity index (Sheldon 1969). The dependence of race complexity on isolation time was analysed with one-way ANOVA and Tukey HSD test, as were the differences in the Shannon’s index values between collecting years and sites.
For genetic analysis, P. infestans subpopulations were defined according to sampling years and collecting sites. Data analysis was performed using Microsoft Office Excel 2013 Add-in GenAlEx version 6.501 (Peakall and Smouse 2006, 2012) and package poppr (Kamvar et al. 2014) within R version 3.1.0. All of the isolates were included in the calculations of single allele and allele pair (genotype) frequencies at all studied loci. Gene diversity (H) was calculated for all the loci according to the formula of H = 1-∑xj2, where xj is the frequency of the jth allele at the locus (Nei 1978). An H value equal to 0 implies no diversity at that locus and the limit of H is 1, which corresponds to the highest possible diversity. Inbreeding coefficient (FIS) and fixation index (FST) were calculated over all collecting sites for 12 SSR loci.
Multilocus genotypes (MLGs) were compiled for all the isolates using all 12 SSR loci. A single difference in an allele size was considered enough to discriminate a unique MLG. If an isolate had one missing locus it was regarded as identical to another MLG if the other eleven loci were identical. This could possibly lead to a lower estimation of genotypic diversity but not substantially because high diversity is expected in the population. Genotypic diversity was calculated by a normalized Shannon’s diversity index (Hs) (Sheldon 1969). Values for Hs may range from 0 (only single MLG present) to 1 (each isolate with a different MLG). The differences in the Shannon’s index values between collecting years and sites were analysed with one-way ANOVA and Tukey HSD test.
A clone-corrected data set was constructed by including only one isolate of each MLG. Deviations from the Hardy-Weinberg equilibrium at loci were tested with a chi-square test using this data set. Random recombination between pairs of SSR loci was assessed with the standardized index of association (r̄d) which was calculated with 999 simulations (Agapow and Burt 2001). It is less biased than the index of association (IA) as it accounts for the number of loci used for analysis.
Genotypic differentiation was assessed by calculating the Bruvo’s genetic distance between the MLGs (Bruvo et al. 2004). This difference was used in a principal coordinate analysis (PCoA) to visualize any clustering based on isolate collecting year. Pairwise population Nei’s genetic distance (Nei 1978) and Wright’s fixation index (FST) values were calculated to estimate the genetic differentiation among the three years. The significance of pairwise FST values was tested by 999 permutations.
Results
P. infestans isolates collected from 23 sites in 2010–2012 were analysed for mating type and SSR marker genotype (141 isolates), a subset of 99 isolates for metalaxyl response and 95 for virulence (isolates only from 2011 and 2012).
Both A1 and A2 mating types were found in all three study years. Among 141 isolates, 73 were A1 mating type and 68 were A2 mating type (Table 1). Both mating types were present at 20 of 23 sites (87%). Although the number of A1 and A2 mating types varied with year, there were no statistically significant differences between years (Chi-square = 1.99, df = 2, p = 0.37). Sample size was too small to detect any statistically significant differences in the proportion of A1 and A2 between sampling sites (Chi-square = 26.39, df = 22, p = 0.24). No statistically significant association between response to metalaxyl and mating type was found (Chi-square = 0.08, df = 2, p = 0.96).
Of the 99 isolates analysed for metalaxyl response, 12 were resistant, 19 were intermediate and 68 were sensitive (Table 1). No statistically significant differences were found between sampling years (Chi-square = 3.23, df = 4, p = 0.52) and sampling sites (Chi-square = 41.22, df = 40, p = 0.42).
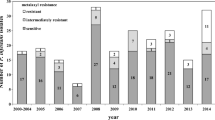
All 11 known virulence factors were found among the 95 tested isolates (Fig. 2). A statistically significant difference in the prevalence of virulence factors (R1–R11) was observed in the two sampling years (F(10, 154) = 69.81, p < 0.001). Virulence factors 9 (2011: 10.3 ± 4.7%; 2012: 4.2 ± 2.7%), 5 (2011: 13.4 ± 5.0%; 2012: 4.2 ± 4.2%) and 8 (2011: 17.0 ± 7.2%; 2012: 1.8 ± 1.8%) were relatively rare (Fig. 2). No statistically significant differences in virulence factors were found between years (F(1,150) = 2.64, p = 0.11). However, there were statistically significant differences between study sites (F(14,139) = 3.60, p < 0.001).
Estimated mean frequency of virulence to potato R-genes in the Estonian population of Phytophthora infestans over all study sites during 2011–2012. Data are presented as means ± SE of the values for each location. Different letters upon the bars indicate significant differences at α = 0.05 (ANOVA, Tukey HSD test), whereas uppercase and lowercase letters are for samples from 2011 and 2012, respectively
There was a high level of race diversity with 52 pathotypes identified among the 95 isolates tested (Table 2). The average number of virulence factors per isolate was 6.3 and ranged among sites from 4.3 to 8.5 and among years from 6.0 in 2012 to 6.6 in 2011. The most common races were 1.2.3.4.7.10.11 and 1.2.3.4.6.7.10.11, represented by 33 (35%) isolates. In 2011, 24 (60%) of the pathotypes were found only once, and in 2012, 18 (45%) of the detected pathotypes were unique (Table 2). The overall normalized Shannon’s diversity index was very high, 0.74, but with no statistically significant differences between years (F(1,10) = 0.05, p = 0.83).
Among all the 141 isolates characterised with SSR markers, 51 alleles were detected at 12 loci, which were all polymorphic (Online Resource 2). The number of alleles differed among the loci, ranging from two at loci SSR2 and Pi70 to nine at locus G11. In addition, at locus D13 a null allele was observed meaning that this locus was not amplified for as many as 48% of the isolates. Variability was detected in the allele frequencies between collecting years for nearly all the determined loci. 18 alleles out of all 51 had an overall frequency lower than 0.05 and were considered as rare alleles, which mainly appeared in the isolates collected in the same year. Two loci, D13 and G11, stood out with a high proportion of rare alleles in contrast to most of the loci where only one rare allele was detected. Gene diversities (H) for each SSR locus were calculated by further processing allele frequencies (Online Resource 2). The H value ranged between 0.08 (locus Pi70) to 0.8 (locus G11) with the mean of 0.53 over all loci. As the H value estimates the mean expected heterozygosity at the locus, the lowest value H = 0.08 at locus Pi70 makes it the least informative locus, because of one frequent dominant homozygous genotype 192/192 (f = 0.92) (Online Resource 3). By contrast, at locus G11 with the highest diversity nine alleles were observed and 13 heterozygous and only six homozygous genotypes were detected. However, the most frequent genotypes at locus G11 were homozygous 156/156 (f = 0.13), 154/154 (f = 0.1) and 162/162 (f = 0.1) (Online Resource 3). At the second-highest-diversity locus SSR4 (H = 0.74), 11 out of 14 genotypes were heterozygous and even five genotypes had a frequency f ≥ 0.1. The lowest number of genotypes were observed at loci Pi70 (genotypes 192/192, 192/195) and SSR2 (173/173, 173/175, 175/175). Overall, 90 genotypes were detected and genotype frequencies varied between collecting years for all the determined loci (Online Resource 3). The rarest genotypes at each locus were detected mainly in one collecting year, for example genotypes 142/156, 152/208, 154/208, 164/164 at locus G11. At locus D13 as many as 8 out of 12 genotypes were rare and observed in one collecting year (Online Resource 3).
Four loci (D13, G11, Pi04 and Pi4B) were not in Hardy-Weinberg equilibrium. The inbreeding coefficient (FIS) was negative for all but two loci indicating an excess of heterozygosity at most of the loci and asexual reproduction within sites (Table 3). The fixation index (FST) at all SSR loci varied between 0.101 and 0.394, with a mean of 0.220, which shows some degree of genetic differentiation between subpopulations collected from different sites (Table 3). However, the standardized index of association (r̄d) between loci did not differ significantly from zero (r̄d = 0.003) and also the null hypothesis of no linkage among loci failed to be rejected (p = 0.26), which could be explained by recombination of alleles into new genotypes during sexual recombination.
Multilocus genotypes (MLGs) were the result of the combination of alleles from all 12 SSR loci. In total, 96 MLGs were identified among the studied isolates, including 69 (72%) MLGs which appeared only once so were unique. Most of the MLGs that were identified more than once were associated with isolates collected from the same field and only seven (26%) repeated MLGs were discovered from separate sites. Moreover, all of the MLGs appeared in only one sampling year and not in other years. Genotypic diversity calculated by the normalized Shannon’s diversity index was altogether very high (Hs = 0.89), being highest in 2010 (Hs = 0.90) and somewhat lower in 2012 (Hs = 0.86) and 2011 (Hs = 0.84), but with no statistically significant differences between years (F(2,20) = 2.78, p = 0.09). Among sites, the highest genotypic diversity (Hs = 1) was identified in six fields out of 23 as all the isolates collected from these sites had unique MLGs.
Principal coordinate analysis (PCoA) did not reveal any clustering based on isolate collecting year (Fig. 3). These results were in concordance with the low values of pairwise FST (0.007–0.020) and Nei’s genetic distance (0.021–0.035) calculations, which additionally showed no differentiation between subpopulations collected in different years (Table 4).
Discussion
Whereas late blight is still a severe threat to Estonian potato production, continuous monitoring of local highly evolving P. infestans population is needed for effective late blight management. Therefore, recent changes in spatio-temporal variation were monitored with analysis of mating type, sensitivity to metalaxyl, virulence on 11 potato R-gene differentials and 12 SSR markers to show the outcome of possible sexual reproduction in the population. Increase in sensitivity to metalaxyl was presumed so metalaxyl-based fungicides could be efficiently used for late blight control. Frequent sexual reproduction was expected in the pathogen population as well as a high number of virulence pathotypes and high genetic diversity with no spread of clonal lineages.
In this study, both A1 and A2 mating types were found in Estonian P. infestans populations in all study years. The frequency of the A2 mating type remained quite stable over the three years, varying from 41% (20 out of 49) in 2012 to 55% (28 out of 51) in 2011, with an average of 48% over all isolates. These findings of mating types in Estonia are generally comparable to those found previously in Estonia (Runno-Paurson et al. 2014) and recently in Finland (Hannukkala 2012), other Nordic countries (Denmark, Norway, Sweden) (Lehtinen et al. 2008), Latvia (Aav et al. 2015), Lithuania (Runno-Paurson et al. 2015), Poland (Chmielarz et al. 2014), the Russian Federation (Moscow region) (Statsyuk et al. 2013) and the Czech Republic (Mazakova et al. 2006).
There are several P. infestans populations in Europe, in the UK (2006–07), France (2006–07), Switzerland (2006–07) and Belgium (2007), where A2 mating type frequency has increased to high levels compared to previous studies (Gisi et al. 2011). These findings are related to the significant changes that have occurred in P. infestans European populations, including the UK. From 2004 to 2008, the extent of the A2 mating type increased from very low (<5%) up to 80% among the UK population mainly due to one specific invasive P. infestans genotype 13_A2 (also known as “Blue_13”) (Cooke et al. 2012). The same genotype dominated the Dutch population from its first appearance in 2004 until 2009 (Li et al. 2012). It has also been found in other European countries including Poland (Chmielarz et al. 2014), which indicates the possibility of 13_A2 spreading to nearby populations, although as yet we found none in Estonia. Furthermore, no other clonal lineage appeared either. It is suggested that it is more difficult for a single genotype to invade a population with high diversity, compared to one where only a few genotypes dominate (Chmielarz et al. 2014).
Behind the high level of diversity in the Estonian P. infestans population is the presence of both mating types in the same fields enabling potential sexual reproduction and diverse MLGs together with a limited spread of these genotypes through the asexual cycle. In addition to the overall high genetic diversity, in six sampling fields all of the isolates had unique MLGs. Furthermore, all of the MLGs were detected in only one sampling year and none of them dominated in the whole population. These results are in accordance with the previous P. infestans population genotyping studies from Estonia in 2004 (Runno-Paurson et al. 2016), the Nordic countries (Brurberg et al. 2011; Sjöholm et al. 2013), Poland (Chmielarz et al. 2014; Brylinska et al. 2016) and Russia (Statsyuk et al. 2014). Variation in genotypes among Estonian populations is so vast it totally differentiates from that in Western and Central European countries like the UK, France, the Netherlands and Switzerland (Montarry et al. 2010; Gisi et al. 2011; Cooke et al. 2012; Li et al. 2012), where genotype structure is clonal and only a few genotypes dominate.
Isolates with the same MLG collected from the same field at the same time referred to clonal (asexual) reproduction occurred within 74% of sampled sites. Also negative FIS values in SSR loci (Table 3) are expected in asexually reproducing populations (Balloux et al. 2003). In contrast to the clonal reproduction within fields, only minimal clonal spread was observed between fields nearby (seven MLGs out of 96 were found in two separate fields). Additionally, FST values (0.101–0.394) support some differentiation of local field populations. Besides, none of the invasive clonal lineages common to Western Europe were detected. Similarly, studies in the Nordic countries showed a high level of clonal reproduction within fields, but no clonal spread between fields and the majority of genotypic variation within collecting sites (Sjöholm et al. 2013; Montes et al. 2016).
Comparing Estonian P. infestans population SSR marker results (Online Resource 2) with those from other Northern (Finland, Sweden, Norway, Denmark) and Eastern European (Poland) countries revealed both similarities and dissimilarities in allele frequencies and occurrence, although each study had slightly different sample collecting strategy. Dominating alleles were the same at all loci in Estonian and Polish populations, but allele frequencies were variable and the number of different alleles was higher in every studied locus within the Polish population probably due to extensive sampling from more collecting sites. But there were also alleles that were detected in the Estonian population that were missing in Poland, for example, allele 215 at locus Pi4B, 208 at G11, 138 at D13 (Chmielarz et al. 2014; Brylinska et al. 2016). Although the Nordic populations have been characterized with a different SSR marker set, some of the loci (Pi04, Pi4B, D13, G11) were the same (Brurberg et al. 2011; Sjöholm et al. 2013). Allele frequencies of the most common alleles at these loci were similar between Estonian and the Nordic populations, besides some of the alleles appeared in the Estonian population, but not in the Nordic countries, for example, allele 215 at locus Pi4B, 206 at G11, 142 at D13 (Brurberg et al. 2011; Sjöholm et al. 2013). Altogether, P. infestans isolates from these countries are likely to be genetically different, but all referred populations share the most common alleles at SSR loci although reported with variable frequencies.
The present study also showed that metalaxyl-sensitive isolates prevailed (69%) in the population. The frequency of metalaxyl sensitivity was quite homogeneous between different study sites and years, except for a small decrease in 2012, when the frequency of metalaxyl-sensitive isolates decreased from 78.9% in 2010 and 73.5% in 2011 to 60.9% in 2012. The product Ridomil Gold MZ 68WG, which contains metalaxyl-M and mancozeb is still in common use as an effective fungicide at the beginning of a late blight control strategy by potato growers in Estonia. Moreover, in Estonia, sales of Ridomil Gold increased steadily from 2011 to 2015 (producer’s information, 2015). The increase in metalaxyl resistance coincides with increased use of metalaxyl-based fungicides in potato fields in 2012 when late blight occurred earlier and disease pressure was extra high. In contrast, weather conditions in 2010 and 2011 were unfavourable for late blight and disease appeared in the fields quite late in both seasons (Runno-Paurson et al. 2013a). These results contrast with those of previous studies carried out in 2001–2007 in Estonia, where the majority of the tested isolates were resistant or tolerant to metalaxyl (Runno-Paurson et al. 2009, 2010, 2012, 2013b, 2014). However, the increase in the frequency of metalaxyl-sensitive isolates was noticed in 2006 and 2007 (Runno-Paurson et al. 2012, 2014). This increasing trend has continued, and perhaps can be explained by more moderate use of metalaxyl fungicide than in the early 2000s. Our research findings about the dominance of metalaxyl-sensitive isolates in the Estonian population of P. infestans corroborate results from recent studies from other Northern and Eastern European countries, such as Finland, Sweden, Denmark and Norway (Lehtinen et al. 2008; Runno-Paurson et al. 2014; Montes et al. 2016), Latvia (Aav et al. 2015), Lithuania (Runno-Paurson et al. 2015), Russia (Moscow region, Statsyuk et al. 2013), Belarus (Pobedinskaya et al. 2011) and Poland (Chmielarz et al. 2014; Brylinska et al. 2016). Indeed, they differ completely from the findings in France in 2006–07 and the UK in 2006–07 (Gisi et al. 2011), where the frequency of metalaxyl-resistant isolates was very high probably due to high incidence of genotype 13_A2.
The Estonian P. infestans virulence race structure found in this study was diverse and complex, affected by a high diversity of potato genotypes from which the isolates were obtained. On average more than half of the races were unique, and the two most common pathotypes 1.2.3.4.6.7.10.11 and 1.2.3.4.7.10.11 comprised only 35% of the population. These findings are similar to those of previous studies in Estonia (Runno-Paurson et al. 2009, 2010, 2012). The prevailing race of P. infestans (1.3.4.7.10.11) in most European populations (Hermansen et al. 2000; Knapova and Gisi 2002; Lehtinen et al. 2008; Hannukkala 2012; Chmielarz et al. 2014; Runno-Paurson et al. 2014) was found only three times in 2011 and once in 2012 in Estonia. In addition, the average number of virulence factors (infected Black’s differentials) per isolate was 6.3, which is lower than those found in other populations from Eastern Europe (Śliwka et al. 2006; Statsyuk et al. 2013; Aav et al. 2015; Runno-Paurson et al. 2015; Michalska et al. 2016) and also from Estonia in previous long-term studies (Runno-Paurson et al. 2012, 2014). It has been shown that P. infestans isolates collected from more resistant varieties have more complex virulence races (Flier et al. 2007; Blandón-Díaz et al. 2012). Although locally bred potato varieties with undetermined R-genes in the genome are quite resistant to late blight in field conditions, these are not grown extensively in Estonia due to limited seed multiplication. However, conventional producers prefer to grow early maturity varieties mostly imported from the Netherlands and Germany, which are quite susceptible to late blight (Runno-Paurson et al. 2013a).
It has earlier been suggested that the difference in the population structure of the late blight pathogen between the Nordic region and most other parts of Europe is caused by climatic conditions (Brurberg et al. 1999). Estonia is located in North-Eastern Europe, where the temperature in the coldest months (December, January, February) is below 0 °C (−4.5 °C to −2.0 °C in average) with absolute minimum below −30 °C (Estonian Weather Service 2016). Before the P. infestans population displacement in the 1980s when sexual reproduction became possible, asexual populations needed a living host for survival to the next growing season (Fry et al. 1993). Because climatic conditions, especially cold winters in this region significantly reduce survival of clones between growing seasons in plant debris, volunteer potatoes and weed hosts (Brurberg et al. 1999; Grönberg et al. 2012), the build-up of blight epidemics was usually delayed until the end of the growing season in Northern Europe (Hannukkala 2012). For last few decades P. infestans populations in Northern Europe commonly reproduce sexually because oospore production benefits the survival and infectiousness of the pathogen (Brurberg et al. 1999; Lehtinen and Hannukkala 2004; Grönberg et al. 2012; Yuen and Andersson 2013). Additionally, highly resilient oospores of P. infestans present in the fields have a relevant role in initiating late blight infections early in the growing season which results in increased use of chemical control (Hannukkala et al. 2007; Widmark et al. 2007; Hannukkala 2012; Runno-Paurson et al. 2013a). Symptoms of soil-borne infection on lower leaves have been noticed in recent years in Estonian potato fields early in the season, especially in fields without or with short rotation of potato crops (personal observation by E. Runno-Paurson). Although oospore presence in the sample fields was not confirmed, mating type ratio, no predominant clonal lineages and high genetic diversity indicated the likely role of sexual reproduction in the P. infestans population in Estonia.
In Estonia, potato is grown under different field management practices. It is still usual to grow potatoes with a low seed quality in small field plots (<1 ha), with limited late blight control and irregular crop rotation (none to three years) (Runno-Paurson et al. 2013b). On the other hand, large conventional producers use certified seed potatoes, apply fungicide as many times as needed and grow potatoes in the same field only every 2–3 years. Study results show that the P. infestans population in Estonia is highly diverse in virulence races and MLGs. Additionally, no genotypes prevailed or persisted through the winter until the next growing season. The complexity of the pathogen population makes late blight management in Estonian potato fields more difficult and, for better results, fungicide treatments should be adjusted according to the population situation. In addition to high population diversity, sexual reproduction also results in oospores which can survive in the soil for several years and with a suitable host in range can induce late blight infection (Drenth et al. 1995). To efficiently control late blight in Estonia, the main aim should be to minimise oospore-derived infections. Therefore, longer crop rotations between growing potatoes in the same field and effective fungicide treatments in conventional fields to prevent infection and stop the late blight from spreading are necessary. Additionally, farmer awareness of P. infestans biology and survival including the infection threat caused by oospores should be raised.
To conclude, our study indicates that in north-eastern Europe, where Estonia is located, the late blight pathogen P. infestans is reproducing sexually, which results in higher population diversity and production of oospores that contribute to better survival through the cold winters. Metalaxyl-sensitive isolates dominate in the pathogen population indicating a sensible and moderate use of metalaxyl-based fungicides. Altogether, the P. infestans population in Estonia is characterised by great virulence race diversity with a high number of virulence pathotypes present. Likewise, the genetic diversity is very high and no clonal lineages are spreading or dominating in the population.
References
Aav, A., Skrabule, I., Bimšteine, G., Kaart, T., Williams, I. H., & Runno-Paurson, E. (2015). The structure of mating type, metalaxyl resistance and virulence of Phytophthora infestans isolates collected from Latvia. Zemdirbyste-Agriculture, 102, 335–342.
Agapow, P.-M., & Burt, A. (2001). Indices of multilocus linkage disequilibrium. Molecular Ecology Notes, 1, 101–102.
Balloux, F., Lehmann, L., & De Meeus, T. (2003). The population genetics of clonal and partially clonal diploids. Genetics, 164, 1635–1644.
Blandón-Díaz, J. U., Widmark, A.-K., Hannukkala, A., Andersson, B., Högberg, N., & Yuen, J. E. (2012). Phenotypic variation within a clonal lineage of Phytophthora infestans infecting both tomato and potato in Nicaragua. Phytopathology, 102, 323–330.
Brurberg, M. B., Hannukkala, A., & Hermansen, A. (1999). Genetic variability of Phytophthora infestans in Norway and Finland as revealed by mating type and fingerprint probe RG57. Mycological Research, 103, 1609–1615.
Brurberg, M. B., Elameen, A., Le, V. H., Naerstad, R., Hermansen, A., Lehtinen, A., et al. (2011). Genetic analysis of Phytophthora infestans populations in the Nordic European countries reveals high genetic variability. Fungal Biology, 115, 335–342.
Bruvo, R., Michiels, N. K., D’Souza, T. G., & Schulenburg, H. (2004). A simple method for the calculation of microsatellite genotype distances irrespective of ploidy level. Molecular Ecology, 13, 2101–2106.
Brylinska, M., Sobkowiak, S., Stefanczyk, E., & Sliwka, J. (2016). Potato cultivation system affects population structure of Phytophthora infestans. Fungal Ecology, 20, 132–143.
Chmielarz, M., Sobkowiak, S., Dębski, K., Cooke, D. E. L., Brurberg, M. B., & Śliwka, J. (2014). Diversity of Phytophthora infestans from Poland. Plant Pathology, 63, 203–211.
Chowdappa, P., Nirmal Kumar, B. J., Madhura, S., Mohan Kumar, S. P., Myers, K. L., Fry, W. E., & Cooke, D. E. L. (2015). Severe outbreaks of late blight on potato and tomato in South India caused by recent changes in the Phytophthora infestans population. Plant Pathology, 64, 191–199.
Cooke, D. E. L., & Lees, A. K. (2004). Markers, old and new, for examining Phytophthora infestans diversity. Plant Pathology, 53, 692–704.
Cooke, D. E. L., Cano, L. M., Raffaele, S., Bain, R. A., Cooke, L. R., Etherington, G. J., Deahl, K. L., Farrer, R. A., Gilroy, E. M., Goss, E. M., Grünwald, N. J., Hein, I., MacLean, D., McNicol, J. W., Randall, E., Oliva, R. F., Pel, M. A., Shaw, D. S., Squires, J. N., Taylor, M. C., Vleeshouwers, V. G. A. A., Birch, P. R. J., Lees, A. K., & Kamoun, S. (2012). Genome analyses of an aggressive and invasive lineage of the Irish potato famine pathogen. PLoS Pathogens, 8, e1002940.
Cooke, L. R., Schepers, H. T. A. M., Hermansen, A., Bain, R. A., Bradshaw, N. J., Ritchie, F., Shaw, D. S., Evenhuis, A., Kessel, G. J. T., Wander, J. G. N., Andersson, B., Hansen, J. G., Hannukkala, A., Nærstad, R., & Nielsen, B. J. (2011). Epidemiology and integrated control of potato late blight in Europe. Potato Research, 54, 183–222.
Drenth, A., Janssen, E. M., & Govers, F. (1995). Formation and survival of oospores of Phytophthora infestans under natural conditions. Plant Pathology, 44, 86–94.
Estonian Weather Service (2016). Climate normals. http://www.ilmateenistus.ee/kliima/kliimanormid/ohutemperatuur/?lang=en. Accessed 16 July 2016.
EuroBlight (2017). A potato late blight network for Europe. euroblight.net. Accessed 1 March 2017.
Flier, W. G., Kroon, L. P. N. M., Hermansen, A., van Raaij, H. M. G., Speiser, B., Tamm, L., Fuchs, J. G., Lambion, J., Razzaghian, J., Andrivon, D., Wilcockson, S., & Leifert, C. (2007). Genetic structure and pathogenicity of populations of Phytophthora infestans from organic potato crops in France, Norway, Switzerland and the United Kingdom. Plant Pathology, 56, 562–572.
Fry, W. E., Goodwin, S. B., Dyer, A. T., Matuszak, J. M., Drenth, A., Tooley, P. W., et al. (1993). Historical and recent migrations of Phytophthora infestans: Chronology, pathways, and implications. Plant Disease, 77, 653–661.
Fry, W. E., McGrath, M. T., Seaman, A., Zitter, T. A., McLeod, A., Danies, G., Small, I. M., Myers, K., Everts, K., Gevens, A. J., Gugino, B. K., Johnson, S. B., Judelson, H., Ristaino, J., Roberts, P., Secor, G., Seebold Jr., K., Snover-Clift, K., Wyenandt, A., Grünwald, N. J., & Smart, C. D. (2013). The 2009 late blight pandemic in the eastern United States – causes and results. Plant Disease, 97, 296–306.
Gisi, U., & Cohen, Y. (1996). Resistance to phenylamide fungicides: A case study with Phytophthora infestans involving mating type and race structure. Annual Review of Phytopathology, 34, 549–572.
Gisi, U., Walder, F., Resheat-Eini, Z., Edel, D., & Sierotzki, H. (2011). Changes of genotype, sensitivity and aggressiveness in Phytophthora infestans isolates collected in European countries in 1997, 2006 and 2007. Journal of Phytopathology, 159, 223–232.
Goodwin, S. B., Cohen, B. A., & Fry, W. E. (1994). Panglobal distribution of a single clonal lineage of the Irish potato famine fungus. Proceedings of the National Academy of Science of the USA, 91, 11591–11595.
Grönberg, L., Andersson, B., & Yuen, J. (2012). Can weed hosts increase aggressiveness of Phytophthora infestans on potato? Phytopathology, 102, 429–433.
Hannukkala, A. O. (2012). History and consequences of migrations, changes in epidemiology and population structure of potato late blight, Phytophthora infestans, in Finland from 1845 to 2011. Doctoral Dissertation. MTT Science 18. MTT Agrifood Research Finland, Jokioinen, 136 p. Available at http://www.mtt.fi/mtttiede/pdf/mtttiede18.pdf.
Hannukkala, A. O., Kaukoranta, T., Lehtinen, A., & Rahkonen, A. (2007). Late-blight epidemics on potato in Finland, 1933-2002; increased and earlier occurrence of epidemics associated with climate change and lack of rotation. Plant Pathology, 56, 167–176.
Hermansen, A., Hannukkala, A., Hafskjold Nærstad, R., & Brurberg, M. B. (2000). Variation in populations of Phytophthora infestans in Finland and Norway: Mating type, metalaxyl resistance and virulence phenotype. Plant Pathology, 49, 11–22.
Hu, C.-H., Perez, F. G., Donahoo, R., McLeod, A., Myers, K., Ivors, K., et al. (2012). Recent genotypes of Phytophthora infestans in the eastern United States reveal clonal populations and reappearance of mefenoxam sensitivity. Plant Disease, 96, 1323–1330.
Kamvar, Z. N., Tabima, J. F., & Grünwald, N. J. (2014). poppr: an R package for genetic analysis of populations with mixed (clonal/sexual) reproduction. Peer J, 2, e281.
Knapova, G., & Gisi, U. (2002). Phenotypic and genotypic structure of Phytophthora infestans populations on potato and tomato in France and Switzerland. Plant Pathology, 51, 641–653.
Lees, A. K., Wattier, R., Shaw, D. S., Sullivan, L., Williams, N. A., & Cooke, D. E. L. (2006). Novel microsatellite markers for the analysis of Phytophthora infestans populations. Plant Pathology, 55, 311–319.
Lehtinen, A., & Hannukkala, A. (2004). Oospores of Phytophthora infestans in soil provide an important new source of primary inoculum in Finland. Agricultural and Food Science, 13, 399–410.
Lehtinen, A., Hannukkala, A., Andersson, B., Hermansen, A., Le, V. H., Naerstad, R., et al. (2008). Phenotypic variation in Nordic populations of Phytophtora infestans in 2003. Plant Pathology, 57, 227–234.
Li, Y., van der Lee, T. A. J., Evenhuis, A., van den Bosch, G. B. M., van Bekkum, P. J., Förch, M. G., et al. (2012). Population dynamics of Phytophthora infestans in the Netherlands reveals expansion and spread of dominant clonal lineages and virulence in sexual offspring. G3. Genes Genomes Genetics, 2, 1529–1540.
Li, Y., Cooke, D. E. L., Jacobsen, E., & van der Lee, T. (2013a). Efficient multiplex simple sequence repeat genotyping of the oomycete plant pathogen Phytophthora infestans. Journal of Microbiological Methods, 92, 316–322.
Li, Y., van der Lee, T., Zhu, J. H., Jin, G. H., Lan, C. Z., Zhu, S. X., Zhang, R. F., Liu, B. W., Zhao, Z. J., Kessel, G., Huang, S. W., & Jacobsen, E. (2013b). Population structure of Phytophthora infestans in China – Geographic clusters and presence of the EU genotype Blue_13. Plant Pathology, 62, 932–942.
Malcolmson, J. F., & Black, W. (1966). New R genes in Solanum demissum Lindl. And their complementary races of Phytophthora infestans (Mont.) de Bary. Euphytica, 15, 199–203.
Mazakova, J., Taborsky, V., Zouhar, M., Ryšanek, P., Hausvater, E., & Doležal, P. (2006). Occurrence and distribution of mating types A1 and A2 of Phytophthora infestans (Mont.) de Bary in the Czech Republic. Plant Protection Science, 42, 41–48.
Mayton, H., Smart, C. D., Moravec, B. C., Mizubuti, E. S. G., Muldoon, A. E., & Fry, W. E. (2000). Oospore survival and pathogenicity of single oospore recombinant progeny from 23 a cross involving US-17 and US-8 genotypes of Phytophthora infestans. Plant Disease, 84, 1190–1196.
Michalska, A. M., Sobkowiak, S., Flis, B., & Zimnoch-Guzowska, E. (2016). Virulence and aggressiveness of Phytophthora infestans isolates collected in Poland from potato and tomato plants identified no strong specificity. European Journal of Plant Pathology, 144, 325–336.
Montarry, J., Andrivon, D., Glais, I., Corbiere, R., Mialdea, G., & Delmotte, F. (2010). Microsatellite markers reveal two admixed genetic groups and an ongoing displacement within the French population of the invasive plant pathogen Phytophthora infestans. Molecular Ecology, 19, 1965–1977.
Montes, M. S., Nielsen, B. J., Schmidt, S. G., Bødker, L., Kjøller, R., & Rosendahl, S. (2016). Population genetics of Phytophthora infestans in Denmark reveals dominantly clonal populations and specific alleles linked to metalaxyl-M resistance. Plant Pathology, 65, 744–753.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89, 583–590.
Peakall, R., & Smouse, P. E. (2006). GENALEX 6: Genetic analysis in excel. Population genetic software for teaching and research. Molecular Ecology Notes, 6, 288–295.
Peakall, R., & Smouse, P. E. (2012). GenAlEx 6.5: Genetic analysis in excel. Population genetic software for teaching and research-an update. Bioinformatics, 28, 2537–2539.
Pobedinskaya, M. A., Elansky, S. N., Statsyuk, N. V., & Plyakhnevich, M. P. (2011). Fungicide resistance of Russian Phytophthora infestans strains. PPO – Special Report, 15, 243–248.
Runno-Paurson, E., Fry, W. E., Myers, K. L., Koppel, M., & Mänd, M. (2009). Characterisation of Phytophthora infestans isolates collected from potato in Estonia during 2002-2003. European Journal of Plant Pathology, 124, 565–575.
Runno-Paurson, E., Fry, W. E., Remmel, T., Mänd, M., & Myers, K. L. (2010). Phenotypic and genotypic characterisation of Estonian isolates of Phytophthora infestans in 2004-2007. Journal of Plant Pathology, 92, 375–384.
Runno-Paurson, E., Hannukkala, A., Williams, I., Koppel, M., & Mänd, M. (2012). The structure of mating type, virulence, metalaxyl resistance of Phytophthora infestans in a long-term phenotypic study in distinct location in eastern Estonia. Journal of Plant Diseases and Protection, 119, 45–52.
Runno-Paurson, E., Williams, I. H., Metspalu, L., Kaart, T., & Mänd, M. (2013a). Current potato varieties are too susceptible to late blight to be grown without chemical control under north-east European conditions. Acta Agriculturae Scandinavica, Section B – Soil & Plant Science, 63, 80–88.
Runno-Paurson, E., Hannukkala, A., Williams, I., Koppel, M., & Mänd, M. (2013b). Impact of phytosanitary quality of seed potato and temporal epidemic progress on the phenotypic diversity of Phytophthora infestans populations. American Journal of Potato Research, 90, 245–254.
Runno-Paurson, E., Hannukkala, A., Kotkas, K., Koppel, M., Williams, I. H., & Mänd, M. (2014). Population changes and phenotypic diversity of Phytophthora infestans isolates from Estonia and Finland. Journal of Plant Pathology, 96, 85–95.
Runno-Paurson, E., Ronis, A., Hansen, M., Aav, A., & Williams, I. H. (2015). Lithuanian populations of Phytophthora infestans revealed a high phenotypic diversity. Journal of Plant Diseases and Protection, 122, 57–65.
Runno-Paurson, E., Kiiker, R., Joutsjoki, T., & Hannukkala, A. (2016). High genotypic diversity found among population of Phytophthora infestans collected in Estonia. Fungal Biology, 120, 385–392.
Shattock, R. C., Janssen, B. D., Whitbread, R., & Shaw, D. S. (1977). An interpretation of the frequencies of host specific genotypes of Phytophthora infestans in North Wales. Annals of Applied Biology, 86, 249–260.
Sheldon, A. L. (1969). Equitability indices: Dependence on the species count. Ecology, 50, 466–467.
Sjöholm, L., Andersson, B., Högberg, N., Widmark, A.-K., & Yuen, J. (2013). Genotypic diversity and migration patterns of Phytophthora infestans in the Nordic countries. Fungal Biology, 117, 722–730.
Śliwka, J., Sobkowiak, S., Lebecka, R., Avendańo Córcoles, J., & Zimnoch-Guzowska, E. (2006). Mating type, virulence, aggressiveness and metalaxyl resistance of isolates of Phytophthora infestans in Poland. Potato Research, 49, 155–166.
Spielman, L. J., Drenth, A., Davidse, L. C., Sujkowski, L. J., Gu, W., Tooley, P. W., & Fry, W. E. (1991). A second world-wide migration and population displacement of Phtyophthora infestans. Plant Pathology, 40, 422–430.
Statsyuk, N. V., Kozlovskaya, I. N., Koslovsky, B. E., Ulanova, T. I., Morozova, E. V., & Kuznetsova, M. (2013). Changes in phenotypic characteristics of the Moscow Phytophthora infestans population in the period of 2000–2011. Proceedings of the 4th International Symposium “Agrosym 2013” (pp. 607–613). Istočno Sarajevo: Jahorina.
Statsyuk, N. V., Semina, Y. V., Perez, F. G. M., Larsen, M. M., Kuznetsova, M. A., Kozlovskaya, I. N., et al. (2014). Characterization of Russian Phytophthora infestans populations: DNA fingerprinting and SSR analysis. PPO – Special report, 16, 255–266.
Turkensteen, L. J., Flier, W. G., Wanningen, R., & Mulder, A. (2000). Production, survival and infectivity of oospores of Phytophthora infestans. Plant Pathology, 49, 688–696.
USAblight (2017). A national project on tomato & potato late blight in the United States. https://usablight.org/. Accessed 1 March 2017.
Vorobyeva, Y. V., Gridnev, V. V., Bashaeva, E. G., Pospelova, L. A., Kvasnyuk, N. Y., Kuznetsova, L. N., et al. (1991). On the occurrence of the A2 mating type isolates of Phytophthora infestans (Mont.) de Bary in the USSR. Mikologija i fitopatologija, 62–67.
Widmark, A.-K., Andersson, B., Cassel-Lundhagen, A., Sandström, M., & Yuen, J. E. (2007). Phytophthora infestans in a single field in Southwest Sweden early in spring: Symptoms, spatial distribution and genotypic variation. Plant Pathology, 56, 573–579.
Yuen, J. E., & Andersson, B. (2013). What is the evidence for sexual reproduction of Phytophthora infestans in Europe? Plant Pathology, 62, 485–491.
Acknowledgments
Alice Aav, Gerit Dreyersdorff, Kätlin Jõgi, Liis Laane, Helina Nassar, Terje Tähtjärv and Grete Zahkna are thanked for technical support. We are grateful to Asko Hannukkala from the Natural Resources Institute Finland (Luke) for supplying tester isolates for mating type determination. Many thanks too to Dr. Eva Randall at the James Hutton Institute for technical assistance.
Funding
This study was supported by Estonian Foundation grant no 9432, Institutional research funding IUT36–2 of the Estonian Ministry of Education and Research, projects RESIST 3.2.0701.11–0003 and IPMBlight 2.0 8T150054PKTK. The Scottish Government is acknowledged for funding at the James Hutton Institute. The study visit to The James Hutton Institute was supported by the European Social Fund’s Doctoral Studies and Internationalisation Programme DoRa, which is carried out by Foundation Archimedes.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kiiker, R., Hansen, M., Williams, I.H. et al. Outcome of sexual reproduction in the Phytophthora infestans population in Estonian potato fields. Eur J Plant Pathol 152, 395–407 (2018). https://doi.org/10.1007/s10658-018-1483-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-018-1483-y