Abstract
A nonpathogenic strain of Agrobacterium (=Rhizobium) vitis, ARK-1, limits the development of crown gall of grapevine (Vitis vinifera L.). Co-inoculation of grapevine shoots with ARK-1 and the tumorigenic (Ti) strain VAT03-9 at a 1:1 cell ratio resulted in significantly lower expression of the virulence genes virD2 and virE2 of VAT03-9 1 day after inoculation (dai) compared with expression levels when shoots were inoculated only with VAT03-9. In contrast, nonpathogenic A. vitis strain VAR06-30, which does not limit the development of crown gall of grapevine, co-inoculated with VAT03-9 did not affect expression levels of virD2 and virE2 under the same conditions. ARK-1 began to suppress the VAT03-9 population by seven dai, but no such effect was observed with VAR06-30 during the nine dai study period. Thus, the biological control activity of ARK-1 is likely based on the suppression of essential virulence genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Grapevine (Vitis vinifera L.) crown gall is caused mainly by Agrobacterium vitis (Ti) [=Rhizobium vitis (Ti), A. tumefaciens biovar 3], where “Ti” means “tumor-inducing” or “tumorigenic”. This is one of the most important diseases of grapevine around the world (Burr et al. 1998; Burr and Otten 1999). The virulence (vir) genes and transfer DNA (T-DNA) are located mostly on large tumor-inducing plasmids (pTi) and also chromosomal virulence genes (chv) on chromosomes. Virulent Agrobacterium strains transfer single-strand forms of T-DNA and several Virulence effector proteins through a bacterial type IV secretion system into plant host cells (Gelvin 2012; Nester 2015). The plant molecules acetosyringone and α-hydroxyacetosyringone induce the entire vir regulon in Agrobacterium as well as the formation of T-DNA intermediate molecules (Nester 2015). These plant molecules occur specifically in the exudates from wounded and metabolically-active plant cells and probably allow Agrobacterium to recognize susceptible cells in nature (Nester 2015). T-DNA transfer and processing require products of the vir genes (virA, virB, virC, virD, virE, and virG), which are located outside of the T-DNA coding region (Gelvin 2012; Lacroix and Citovsky 2013; McCullen and Binns 2006; Nester 2015). The expression of virB, virC, virD, and virE is positively regulated at the transcriptional level by plant signal molecules (Gelvin 2012; Lacroix and Citovsky 2013; Nester 2015). Two Agrobacterium proteins, VirD2 and VirE2, directly associate with the T-strand; one VirD2 molecule covalently attaches to the 5′-end of the T-strand, and VirE2, a protein that binds single-stranded DNA, cooperatively coats the rest of the T-strand (Gelvin 2012; Lacroix and Citovsky 2013; Nester 2015).
Several laboratories have attempted to develop some biological measures to control grapevine crown gall. Staphorst et al. (1985) evaluated nonpathogenic A. vitis strain F2/5, which inhibited the growth of most Ti strains of A. vitis in vitro and inhibited crown gall formation in grapevine in greenhouse shoot-wounding experiments. Burr and Reid (1994) reported that F2/5 produces an unidentified agrocin. Wang et al. (2003) reported that the antibacterial compound Ar26 produced by nonpathogenic A. vitis strain E26 inhibited the growth of A. vitis (Ti) on culture plates. Chen et al. (2007) reported that Rahnella aquatilis strain HX2 inhibited the development of crown galls in grapevine.
Previously, we have reported that a nonpathogenic A. vitis strain, VAR03-1, that was isolated from grapevine nursery stock in Japan inhibited tumor formation in grapevine, rose, tomato, sunflower, and apple (Kawaguchi et al. 2008, 2012). Moreover, we identified nonpathogenic A. vitis strain ARK-1, which performed better than VAR03-1 at inhibiting tumor formation in grapevine, as a new antagonistic strain (Kawaguchi 2013; Kawaguchi and Inoue 2012). ARK-1 is endophytic in grapevine, induces no necrosis of the host plant, and controlled grapevine crown gall better than VAR03-1 in field trials (Kawaguchi 2013). Although ARK-1 produced a zone of inhibition against tumorigenic Agrobacterium spp. in in vitro assays, antibiosis was not the main mechanism of biological control, because ARK-1 suppressed tumor formation in grapevine shoots caused by a Ti strain that was insensitive to the antibiosis produced by ARK-1 (Kawaguchi 2014). Although ARK-1 reduced the pathogen population at the wound site through biological control, ARK-1 suppressed the Ti strain population significantly at 2 days after inoculation (dai) but did not suppress the Ti population at 1 and 5 dai (Kawaguchi 2014). It is most likely that transformation by A. vitis would occur prior to 5 days because co-cultivation of Agrobacterium cells and plants has been successfully tried for 2 or 3 days in a general protocol for the Agrobacterium-mediated transformation of plants (Jones et al. 2005). The mechanism by which ARK-1 reduces crown gall might not involve reducing populations of the Ti strains in grapevine plants at an early infection stage, so the main mechanism of ARK-1’s effects remains unknown.
To provide insights into that mechanism, we tested the effect of ARK-1 to suppress the expression of two virulence genes by the Ti strain at the wound site. This report gives that suppression of expression of the virD2 and virE2 virulence genes of the Ti strain by ARK-1 at the grapevine wound site and in vitro; this suppression appears to be responsible for the biological control by this strain. In this paper, the nomenclature for Agrobacterium species proposed by Young et al. (2005) was followed.
Materials and methods
Development of antibiotic-resistant A. vitis strains
Table 1 lists the bacterial strains used in this study. Potato sucrose agar (PSA) medium was used to grow the bacteria in this study (Kawaguchi 2013; Kawaguchi and Inoue 2012). ARK-1sc was a streptomycin (St)- and copper sulfate (CuSO4)–resistant mutant (St-CuSO4 mutant) obtained by growing strain ARK-1 on St-CuSO4-PSA medium, which is PSA amended with 500 μg/ml St (minimum inhibiting concentrations (MIC) =500 μg/ml) and 250 μg/ml CuSO4 (MIC = 250 μg/mL) (Kawaguchi 2013). VAR06-30sc was also a St-CuSO4 mutant obtained by growing on St-CuSO4-PSA medium. VAT03-9n was a nalidixic acid (nal)–resistant mutant (nal-mutant) obtained by growing on nal-PSA medium, which is PSA amended with 50 μg/ml nal (MIC = 50 μg/mL).
Tumor inhibition assay in grapevine seedlings
Grapevine seedlings (V. vinifera cv. ‘Neo Muscat’) were grown from seed. One-year-old grapevine shoots were inoculated using previously established methods (Kawaguchi and Inoue 2012). Cell suspensions of the nonpathogenic strains (ARK-1 and VAR06-30) and of the tumorigenic strain VAT03-9 (Table 1) were prepared from 48-h-old cultures on PSA medium slants, discarded a supernatant of these strains and washed with distilled water, and adjusted to OD600 = 0.1, which corresponds to approximately 108 cells ml−1. A culture filtrate of ARK-1 by filtering with a syringe filter unit (0.2 μm diameter) as “CF of ARK-1” were prepared. Two mixed cell suspensions (ARK-1 plus VAT03-9, VAR06-30 plus VAT03-9), both at a cell ratio of 1:1, and CF of ARK-1 plus VAT03-9 and a VAT03-9 suspension were prepared. A 5-μl drop (containing approximately 5 × 105 cells) of a given cell suspension was dropped onto a needle-prick wound on the stem of a grapevine seedling. Six to seven grapevine seedlings (at one plant per pot) each received 10 inoculations (i.e., a total of 60 to 70 inoculations per treatment). Grapevines inoculated with sterile distilled water were used as the negative control. The seedlings were grown in a greenhouse at 20° to 35 °C for 3 months with natural sunlight (photoperiod; 14 h light: 10 h dark). The experiments were performed three times independently, and one experiment was defined as one replication (i. e., three independent replications). The protection rate provided by co-inoculation was defined as follows:
Protection rate (%) = 100 × [1 – (% of tumor formation for the mixed suspension × 100) ⁄ (% of tumor formation with only VAT03-9)]
Detection of virD2 and virE2 mRNA using the reverse-transcription quantitative polymerase chain reaction
Grapevine seedlings (2 years old, ‘Neo Muscat’) were inoculated with ARK-1, VAR06-30, and VAT03-9 at a cell ratio of 1:1, and CF of ARK-1 plus VAT03-9, as described above. Each grapevine seedling (one plant per pot) was inoculated once with one of the mixed cell suspensions or with only VAT03-9. The seedlings were grown in a greenhouse at 20 to 35 °C with natural sunlight as described previously. Shoot samples that included the one wound site (0.2 g fresh weight per plant, 1 sample per plant) were collected from five plants (i.e., n = 5) at 1, 6, and 12 h after inoculation (hai), and at 1, 2, 3 and 4 dai (i.e., assessed seven times). Table 2 presents the basic information for the reverse-transcription quantitative polymerase chain reaction (RT-qPCR) experiments, which followed the method of Bustin et al. (2009). Table 3 lists the primers used in this analysis. The A and B sets were used to detect the virD2 and virE2 genes of the A. vitis (Ti) strain, respectively (Table 3). The C set was used to amplify the pyrG gene of A. vitis (Ti) strain VAT03-9 for normalization; pyrG is a bacterial housekeeping gene (Table 3). The pyrG gene was used as an internal control after evaluation of the stability of four candidate reference genes (Table 2 and 3, Supplementary Figure S1; Vandesompele et al. 2002). Relative expression rate of virD2 and virE2 were described as a value based on the degree of expression of each gene at 1 dai as 100 %. These three sets (A to C) were confirmed to amplify genes only from the Ti strain VAT03-9 (Supplementary Figure S2). The amplified PCR products were analyzed using sequencing assay to confirm the specificity of the amplification (data not shown). PCR efficiency was calculated from the standard curves. Amplifications had a PCR efficiency and R 2 value of >88.9 % and 0.980 (virD2), respectively, and of >90.2 % and 0.999 (virE2). Relative quantification of the virD2 and virE2 mRNA concentrations was performed using the ΔΔCt method (Pfaffl et al. 2002) using the pyrG gene’s mRNA values as a reference. Five biological replicates and three technical replicates were used for each assay. The detection assay was independently performed three times.
Detection of virD2 and virE2 mRNA using an in vitro RT-qPCR assay with acetosyringone
Acetosyringone (3′, 5′-dimethoxy-4′-hydroxyacetophenone) (Sigma-Aldrich) was dissolved in 70 % ethanol to produce a 10 mg ml−1 stock solution. After filter sterilization, the stock solution was stored at −20 °C. Four cell suspensions at approximately 108 cells ml−1 (ARK-1 plus VAT03-9, VAR06-30 plus VAT03-9, and CF of ARK-1 plus VAT03-9 at cell ratios of 1:1, and only VAT03-9) were prepared as described above. Mixed cell suspensions (1 ml) or 0.5 ml of only VAT03-9 were added into potato sucrose (PS) medium or PS-AS medium (PS medium with 200 μg/ml acetosyringone) in an Erlenmeyer flask. Each Erlenmeyer flask with 50-mL PS or PS-AS medium (n = 3, flasks per medium) was inoculated with one of the mixed suspensions or with a VAT03-9 cell suspension. Each flask represented one independent technical replicate. The flasks were incubated at 27 °C with shaking (approximately 2.2 × g) for 2 days. Culture solution samples (one 1-ml sample per flask) were collected from each flask at 1 and 2 dai, bacterial RNA was extracted as described above. RT-qPCR for detection of the virD2 and virE2 genes and relative quantification of the virD2 mRNA concentrations were performed as described above. The detection assay was independently performed three times.
Population dynamics during coexistence of a nonpathogenic strain with VAT03-9n in grapevine seedlings
Grapevine seedlings (2 years old, ‘Neo Muscat’) were inoculated with one of three cell suspensions (ARK-1sc plus VAT03-9n and VAR06-30sc plus VAT03-9n at a 1:1 cell ratio, and VAT03-9n alone), as described above. Supplementary Figure S3 illustrates the design of this experiment. Each grapevine seedling (n = 40 for each inoculation mixture, at one plant per pot) was inoculated once with the mixed cell suspension of ARK-1sc or VAR06-30sc plus VAT03-9n. Each grapevine seedling represented one replicate. The seedlings were grown in a greenhouse at 20 to 35 °C with natural sunlight as described previously. To determine the population dynamics of each strain, shoot samples were collected that included the wound site (0.2 g fresh weight per plant, 1 sample per plant) from 5 plants (i.e., n = 5) at 1 hai (0 dai) and at 1, 2, 3, 5, 7, and 9 dai (i.e., assessed at eight times). Each shoot sample was scrubbed by hand, rinsed under tap water for 10 s, and then blotted dry with paper towels. The inoculated stems segments were washed with sterile distilled water, and then crushed in 1 mL of sterile distilled water using an autoclaved mortar and pestle. Ten-fold serial dilutions (100 μl) of the samples were then plated and spread on St-CuSO4-PSA and nal-PSA media, and the plates were incubated at 25 °C for 5 days. Colony growth was then observed on five plates for each dilution, and the numbers of colony-forming units (CFU) of strains ARK-1sc, VAR06-30sc, and VAT03-9n were counted on each medium. The bacterial populations in the wounded sites (CFU g−1 of grapevine shoot) were log10-transformed before statistical analysis. Additional experiments were carried out that each grapevine seedling was inoculated once with the each cell suspension of ARK-1sc, VAR06-30sc or VAT03-9n to confirm the population dynamics of each single strain. This assay was independently performed twice.
Data analysis
Tukey’s honestly significant difference (HSD) test (after analysis of variance, ANOVA) or the t-test was used to compare the relative gene expression in the various treatments. Ryan’s multiple-comparison test was used to compare the frequency of tumor formation among the treatments. All statistical analyses were performed using version 2.14.0 of the R software (http://www.r-project.org/).
Results
Tumor inhibition assay in grapevine seedlings
To evaluate the inhibition of tumor formation by the nonpathogenic A. vitis strains ARK-1 and VAR06-30, grapevine seedlings were co-inoculated with a mixed cell suspension of one of these strains plus the pathogenic Ti strain VAT03-9. The combination of ARK-1 with VAT03-9 significantly suppressed tumor incidence (P < 0.01) compared with VAT03-9 alone as the control, for a protection rate of 91.5 % (Table 4). In contrast, VAR06-30 and CF of ARK-1 did not significantly inhibit tumor formation compared with the control, with a protection rate of only 12.1 and 7.8 %, respectively (Table 4).
Suppressive effect on expression of the vir genes of the Ti strain by co-inoculation with ARK-1 in grapevine plants
To clarify whether the expression of vir genes of Ti strain VAT03-9 is suppressed by ARK-1 in vivo, the expression of virD2 and virE2 was analyzed in grapevine plants co-inoculated with ARK-1 and VAT03-9. In plants inoculated with only VAT03-9, induction of bacterial virD2 and virE2 expression was detected at 1 dai (Fig. 1). No significant difference in the expression levels of virD2 and virE2 was observed upon co-inoculation with VAR06-30 or CF of ARK-1 and VAT03-9 in comparison with inoculation with only VAT03-9 (Fig. 1). On the other hand, virD2 and virE2 expression was significantly suppressed upon co-inoculation with ARK-1 and VAT03-9 (Fig. 1). These results suggested that treatment with ARK-1, which inhibited tumor formation, could suppress the expression of virD2 and virE2 by the Ti strain in grapevine plants, and that VAR06-30 and CF of ARK-1, which could not inhibit tumor formation, could not suppress expression of these genes.
Relative expression levels of the virD2 and virE2 genes in bacterial cells of the Agrobacterium vitis Ti strain in grapevine plants measured using the reverse-transcription quantitative polymerase chain reaction (RT-qPCR). Values (%) represent the relative expression levels of the vir genes at 1 day after inoculation (dai) compared with the value when only Ti strain VAT03-9 was inoculated (“only VAT03-9”), which had a value of 100 %. Each cell suspension or mixture (ARK-1 plus VAT03-9, VAR06-30 plus VAT03-9, CF of ARK-1 plus VAT03-9, and only VAT03-9) at cell ratios of 1:1 was inoculated onto the stems of grapevine plants after wounding. Stem samples were harvested at 1 dai. Data are means ± standard deviation for samples corresponding to individual plants. Bars labeled with different letters indicate a significant difference from the other bars (P < 0.01, Tukey’s HSD test). CF, culture filtrate
To evaluate the start and end of the suppression of vir gene expression by ARK-1, virD2 expression was analyzed from 1 h after inoculation (hai) to 4 dai in grapevine plants co-inoculated with ARK-1 and VAT03-9. The level of virD2 expression upon inoculation with VAT03-9 was significantly higher at 1 dai than at any other time (Fig. 2). The levels of virD2 expression were significantly lower upon co-inoculation with ARK-1 and VAT03-9 than upon inoculation with only VAT03-9 at 12 hai, 1 dai, and 2 dai (Fig. 2). These results suggest that treatment with ARK-1 could suppress virD2 expression by the Ti strain in grapevine plants during the period from 12 hai to 2 dai.
Changes in the relative expression levels of the virD2 gene in grapevine plants measured using the RT-qPCR. Values (%) represent the relative expression levels of the vir genes at 1 day after inoculation (dai) compared with the value when only Ti strain VAT03-9 was inoculated (“only VAT03-9”), which had a value of 100 %. A mixed cell suspension of the nonpathogenic strain ARK-1 and of the Ti strain VAT03-9 at cell ratios of 1:1 or a single cell suspension of VAT03-9 were inoculated onto the stems of wounded grapevine plants. Stem samples were harvested at times ranging from 1 h after inoculation (hai) to 4 dai. Data are means ± standard deviation for samples corresponding to individual plants. Significant differences at a given point in time between the ARK-1 plus Ti mixture and only the Ti strain are indicated by * (t-test, *P < 0.05; **P < 0.01; ns P ≥ 0.05); bars for only the Ti strain that are labeled with different letters differ significantly from other bars for only the Ti strain (Tukey’s HSD test, P < 0.05)
Suppressive effect on vir gene expression by the Ti strain after co-incubation with ARK-1 in culture media with or without acetosyringone
To examine whether expression of the vir genes was suppressed by ARK-1 in culture media, with or without acetosyringone, the kinetics of virD2 and virE2 expression were analyzed. The levels of virD2 and virE2 expression by VAT03-9 incubated in PS-AS medium (which contained acetosyringone) were significantly higher than those in cells incubated in PS medium without acetosyringone (Fig. 3). In PS medium, virD2 and virE2 expression levels in VAT03-9 cells were not significantly affected by co-incubation with ARK-1, VAR06-30 or CF of ARK-1 (Fig. 3). In the PS-AS medium, virD2 and virE2 expression levels in VAT03-9 cells were not significantly affected by co-incubation with VAR06-30 or CF of ARK-1, but were significantly reduced by co-incubation with ARK-1 (Fig. 3). These results suggest that the presence of ARK-1 can suppress virD2 and virE2 expression by the Ti strain in culture medium that contains acetosyringone.
Relative expression levels of the virD2 and virE2 genes in bacterial cells of the tumorigenic (Ti) A. vitis strain on potato sucrose (PS) medium and PS-AS medium, which also includes acetosyringone (AS), measured using RT-qPCR. Values (%) represent the relative expression levels of the vir genes at 1 day after inoculation (dai) compared with the value when only Ti strain VAT03-9 was inoculated (“only VAT03-9”), which had a value of 100 %. Data are means ± standard deviation for samples corresponding to individual incubation flasks. Bars at a given date labeled with different letters differ significantly (Tukey’s HSD test, P < 0.05)
Population dynamics of the nonpathogenic strains and of VAT03-9n after co-inoculation onto grapevine seedlings
Antibiotic-resistant mutants of three strains (ARK-1sc, VAR06-30sc, and VAT03-9n) were used in a survival assay to differentiate the inoculated biological control agents from indigenous agrobacteria (Table 1). The three mutants grew in the St-CuSO4-PSA medium (ARK-1sc and VAR06-30sc) and the nal-PSA medium (VAT03-9n) at rates comparable to the wild-type in PSA medium (data not shown). Populations of ARK-1sc on the plants were significantly higher than those of VAT03-9n at 7 and 9 dai (Fig. 4a). On the other hand, populations of VAR06-30sc and VAT03-9n did not differ significantly up to 9 dai (Fig. 4b), and each single population of ARK-1sc, VAR06-30sc and VAT03-9n did not differ significantly up to 9 dai (Fig. 4c). These results suggest that ARK-1sc could not reduce the pathogen population at the wound site on the grapevine plants during the early period after infection.
Population dynamics of the nonpathogenic strains and VAT03-9n after co-inoculation onto grapevine plants. a Populations of the nonpathogenic strain ARK-1sc and of the tumorigenic (Ti) strain VAT03-9n after co-inoculation on wounded shoots of grapevine seedlings at a 1:1 cell ratio. b Populations of the nonpathogenic strain VAR06-30sc and of the Ti strain VAT03-9n after co-inoculation on wounded shoots of grapevine seedlings at a 1:1 cell ratio. c Populations of the nonpathogenic strain ARK-1sc, of strain VAR06-30sc, and of the tumorigenic (Ti) strain VAT03-9n after each inoculation on wounded shoots of grapevine seedlings. Data are means ± standard deviation for five plants. (a) and (b) Significant differences at a given point in time are indicated by * (t-test, *P < 0.05; **P < 0.01; ns P ≥ 0.05). (c) No significant differences at a given point in time are indicated by ns (Tukey’s HSD test, ns P ≥ 0.05) dai, days after inoculation
Discussion
Two different mechanisms of biological control of plant crown gall disease using antagonistic bacteria have been reported. The first one relates to antibacterial compounds produced by nonpathogenic strains (Chen et al. 2007; Kawaguchi et al. 2008; Kerr 1980; Wang et al. 2003). The second relates to a unique mechanism associated with the quorum-sensing and caseinolytic protease (clp) systems of strain F2/5 (Kaewnum et al. 2013). In the present study, suppression of expression of two vir genes of the Ti strain after treatment with the nonpathogenic A. vitis strain ARK-1 was shown using RT-qPCR both in vitro and in vivo, and these results suggest a different mechanism from those in previous reports. In the co-inoculation assay, ARK-1 suppressed virD2 and virE2 expression by the Ti strain in grapevine plants at 1 dai, even though the population of the ARK-1sc and Ti strains on the plants did not differ significantly at this time. These results suggest that the suppression of virD2 and virE2 expression by the Ti strain was not caused by suppression of growth of the Ti strain by ARK-1. ARK-1 significantly inhibited tumor formation in grapevine plants in this and previous reports (Kawaguchi 2013; Kawaguchi and Inoue 2012), whereas strain VAR06-30, which did not significantly inhibit tumor formation, did not suppress virD2 and virE2 expression or growth of the Ti strain in the plants at 1 dai. These results suggest that the suppression of virD2 and virE2 expression by ARK-1 inhibits tumor formation because the VirD2 and VirE2 proteins are directly associated with transformation of plant cells by Ti strains of Agrobacterium (Gelvin 2012; Lacroix and Citovsky 2013; Nester 2015).
Low levels of virD2 expression by the Ti strain were found at all dates except 1 dai. The peak level of virD2 gene expression by the Ti strain in grapevine plants was observed at 1 dai. On the other hand, the levels of virD2 expression in plants that were co-inoculated with ARK-1 and the Ti strain remained low for 4 dai and were significantly lower than those in plants inoculated with only the Ti strain from 12 hai to 2 dai, indicating that the suppression of virD2 expression by ARK-1 persists for at least 2 dai but becomes unclear at 3 and 4 dai because virD2 expression in plants inoculated with only the Ti strain was also low during this period.
The expression of vir genes by the Agrobacterium Ti strain is strongly activated by the plant molecule acetosyringone, which the bacteria recognize as one of several wound-derived compounds (Nester 2015). In the in vitro study, the levels of virD2 and virE2 expression in Ti cells incubated in PS-AS medium were significantly higher than those in PS medium, which agrees with previous reports (Nester 2015). Moreover, virD2 and virE2 expression levels in the Ti strain co-incubated with ARK-1 were significantly lower than those with only the Ti strain in PS-AS medium, suggesting that similar results were obtained in vitro and in vivo. This evidence supports the hypothesis that ARK-1 suppresses the expression of vir genes in the Ti strain.
Previous research suggested that ARK-1sc suppressed growth of the Ti strain in grapevine plants at 2 dai, but not at 1 and 5 dai (Kawaguchi 2014). In the present study, populations of ARK-1sc on the plants did not differ significantly from those of Ti strain VAT03-9n before 7 dai. Taken together, the present results suggest that ARK-1 cannot strongly suppress growth of the Ti strain in grapevine plants before 7 dai. Kawaguchi (2014) reported that ARK-1 suppressed growth of the Ti strain in grapevines from 9 to 88 dai. In the present study, the ARK-1sc population was significantly higher than that of the Ti strain at 7, 9, and 11 dai. These results suggest that the suppression of growth of the Ti strain by ARK-1 that occurs at 7 or 9 dai may result from suppression of vir gene expression by the Ti strain. This suggests a hypothesis that may explain the relationship between suppression of vir gene expression and growth of the Ti strain: suppression of vir gene expression causes failure of the transformation of the host plant’s cells, thereby preventing the Ti strain from taking up and catabolizing the opines that transformed plant cells should produce before 7 dai, leading to suppression of growth of the Ti strain and a lower population than the ARK-1 population in grapevine plants starting by 7 d. This hypothesis should be tested in future research.
In conclusion, ARK-1 suppressed expression of the virulence genes of the Ti strain at the wound site and suppressed development of the crown gall disease via what appears to be a previously unreported mechanism. Although it remains unclear how ARK-1 suppressed expression of the vir genes by the Ti strain in plants, the CF of ARK-1, which does not inhibit tumor formation, did not suppress virD2 and virE2 expression both in vitro and in vivo, suggesting that ARK-1 might catabolize acetosyringone or produce a very small amount of unknown substances that suppress expression of the vir genes without killing the Ti strain. Moreover, there still remains a possibility of quorum sensing system might have some relation with expression and suppression of vir genes and ARK-1 might affect the quorum sensing system. Additional research will be required to determine whether either of these mechanisms is correct. We are now pursuing these aspects of the bacteria and will report our results in a forthcoming paper.
References
Burr, T. J., & Otten, L. (1999). Crown gall of grape: biology and disease management. Annual Review of Phytopathology, 37, 53–80.
Burr, T. J., & Reid, C. L. (1994). Biological control of grape crown gall with nontumorigenic Agrobacterium vitis F2/5. American Journal of Enology Viticulture, 45, 213–219.
Burr, T. J., Bazzi, C., Süle, S., & Otten, L. (1998). Crown gall of grape: biology of Agrobacterium vitis and the development of disease control strategies. Plant Disease, 82, 1288–1297.
Bustin, S. A., Benes, V., Garson, J. A., Hellemans, J., Huggett, J., Kubista, M., Mueller, R., Nolan, T., & Pfaffl, M. W. (2009). The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clinical Chemistry, 55, 611–622.
Chen, F., Guo, Y. B., Wang, J. H., Li, J. Y., & Wang, H. M. (2007). Biological control of grape crown gall by Rahnella aquatilis HX2. Plant Disease, 91, 957–963.
Gelvin, S. B. (2012). Traversing the cell: Agrobacterium T-DNA’s journey to the host genome. Frontiers Plant Science, 3, 52. doi:10.3389/fpls.2012.00052.
Jones, H. D., Doherty, A., & Wu, H. (2005). Review of methodologies and a protocol for the Agrobacterium-mediated transformation of wheat. Plant Methods, 1, 5–11.
Kaewnum, S., Zheng, D., Reid, C. L., Johnson, K. L., Gee, J. C., & Burr, T. J. (2013). A host-specific biological control of grape crown gall by Agrobacterium vitis strain F2/5; its regulation and population dynamics. Phytopathology, 103, 427–435.
Kawaguchi, A. (2011). Genetic diversity of Rhizobium vitis strains in Japan based on multilocus sequence analysis using the sequences of pyrG, recA and rpoD. Journal of General Plant Pathology, 77, 299–303.
Kawaguchi, A. (2013). Biological control of crown gall on grapevine and root colonization by nonpathogenic Rhizobium vitis strain ARK-1. Microbes and Environments, 28, 306–311.
Kawaguchi, A. (2014). Reduction of pathogen populations at grapevine wound sites is associated with the mechanism of biological control of crown gall by Rhizobium vitis strain ARK-1. Microbes and Environments, 29, 296–302.
Kawaguchi, A., & Inoue, K. (2012). New antagonistic strains of non-pathogenic Agrobacterium vitis to control grapevine crown gall. Journal of Phytopathology, 160, 509–518.
Kawaguchi, A., Inoue, K., & Ichinose, Y. (2008). Biological control of crown gall of grapevine, rose, and tomato by nonpathogenic Agrobacterium vitis strain VAR03-1. Phytopathology, 98, 1218–1225.
Kawaguchi, A., Kondo, K., & Inoue, K. (2012). Biological control of apple crown gall by nonpathogenic Rhizobium vitis strain VAR03-1. Journal of General Plant Pathology, 78, 287–293.
Kerr, A. (1980). Biological control of crown gall through production of agrocin 84. Plant Disease, 64, 25–30.
Lacroix, B., & Citovsky, V. (2013). The roles of bacterial and host plant factors in Agrobacterium-mediated genetic transformation. The International Journal of Developmental Biology, 57, 467–481.
McCullen, C. A., & Binns, A. N. (2006). Agrobacterium tumefaciens and plant cell interactions and activities required for interkingdom macromolecular transfer. Annual Review of Cell and Developmental Biology, 22, 101–127.
Nester, E. W. (2015). Agrobacterium: nature’s genetic engineer. Frontiers Plant Science, 5, 730. doi:10.3389/ fpls.2014.00730.
Pfaffl, M. W., Horgan, G. W., & Dempfle, L. (2002). Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Research, 30, e36.
Staphorst, J. L., van Zyl, F. G. H., Strijdom, B. W., & Groenewold, Z. E. (1985). Agrocin-producing pathogenic and nonpathogenic biotype-3 strains of Agrobacterium tumefaciens active against biotype-3 pathogens. Current Microbiology, 12, 45–52.
Vandesompele, J., De Preter, K., Pattyn, F., Poppe, B., Van Roy, N., De Paepe, A., & Speleman, F. (2002). Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biology, 3, research0034.
Wang, H. M., Wang, H. X., Ng, T. B., & Li, J. Y. (2003). Purification and characterization of an antibacterial compound produced by Agrobacterium vitis strain E26 with activity against A. tumefaciens. Plant Pathology, 52, 134–143.
Young, J. M., Kerr, A., & Sawada, H. (2005). Genus II. Agrobacterium. In G. M. Garrity (Ed.), Bergey’s manual of systematic bacteriology (2nd ed., Vol. 2, pp. 340–345). New York: Springer Verlag.
Acknowledgments
This work was supported by a Japan Society for the Promotion of Science KAKENHI Grant (Number 2550038). I thank Associate Prof. Y. Noutoshi (Okayama University Okayama, Japan) for providing useful advice on experimental design.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Stability of expression of four candidate reference genes in grapevine shoots. Stability values (M) for the four reference genes were evaluated with the geNorm algorithm (http://medgen.ugent.be/~jvdesomp/genorm/) in bacterial cells of the tumorigenic Agrobacterium vitis strain inoculated into shoots of the grapevine plant. In the plot, genes are ranked from the least (left) to the most (right) stable. The pyrG gene had the highest expression stability among the reference genes, and was chosen for use in the present study. However, the pairwise variations (V n /V n+1) did not show values (geNorm V ratio < 0.15) for the optimal number of reference genes (data not shown). (JPEG 80 kb)
Fig. S2
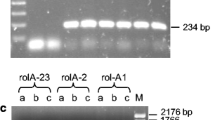
Amplification products of partial sequences of (a) virD2, (b) virE2, and (c) pyrG obtained by RT-qPCR separated by gel electrophoresis. Lanes: M, DNA ladder marker; 1, VAT03-9; 2, ARK-1 plus VAT03-9; 3, VAR06-30 plus VAT03-9; 4, ARK-1; 5, VAR06-30; 6, distilled water. The expected sizes of the RT-PCR products are 61-bp of virD2 (a), 59-bp of virE2 (b), and 69-bp of pyrG (c). (JPEG 170 kb)
Fig. S3
Illustration of the experimental design used to study of the population dynamics during coexistence of the nonpathogenic strains (ARK-1sc or VAR06-30sc) with the tumorigenic strain VAT03-9n in grapevine plants. (a) A 5-μL drop of the mixed cell suspension was dropped onto a needle-prick wound on the grapevine shoot. (b) Shoot samples that included the wound site were collected from 5 plants at each of eight times. (c) After washing and crushing the samples, ten-fold serial dilutions were plated onto St-CuSO4-PSA and nal-PSA media. hai, hours after inoculation; dai, days after inoculation. (JPEG 206 kb)
Rights and permissions
About this article
Cite this article
Kawaguchi, A. Biological control agent Agrobacterium vitis strain ARK-1 suppresses expression of the virD2 and virE2 genes in tumorigenic A. vitis . Eur J Plant Pathol 143, 789–799 (2015). https://doi.org/10.1007/s10658-015-0730-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-015-0730-8