Abstract
Dual mTORC1/2 inhibitors may be more effective than mTORC1 inhibitor rapamycin. Nevertheless, their metabolic effects on breast cancer cells have not been reported. We compared the anti-proliferative capacity of rapamycin and a novel mTORC1/2 dual inhibitor (AZD8055) in two breast cancer cell lines (MDA-MB-231 and MDA-MB-453) and analyzed their metabolic effects using proton nuclear magnetic resonance (1H NMR) spectroscopy-based metabolomics. We found that AZD8055 more strongly inhibited breast cancer cell proliferation than rapamycin. The half-inhibitory concentration of AZD8055 in breast cancer cells was almost one-tenth that of rapamycin. We identified 22 and 23 metabolites from the 1H NMR spectra of MDA-MB-231 and MDA-MB-453 cells. The patterns of AZD8055- and rapamycin-treated breast cancer cells differed significantly; we then selected the metabolites that contributed to these differences. For inhibiting glycolysis and reducing glucose consumption, AZD8055 was likely to be more potent than rapamycin. For amino acids metabolism, although AZD8055 has a broad effect as rapamycin, their effects in degrees were not exactly the same. AZD8055 and rapamycin displayed cell-specific metabolic effects on breast cancer cells, a finding that deserves further study. These findings help fill the knowledge gap concerning dual mTORC1/2 inhibitors and provide a theoretical basis for their development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer has surpassed lung cancer as the leading cause of global cancer incidence [1]. However, the World Health Organization recommends mammography for women of appropriate age [2], new cases of breast cancer account for 24.5% of all new cancer cases in women [1]. Breast cancer also is the leading cause of cancer death among women worldwide, accounting for 685,000 deaths every year [1]. Given this dire situation, efforts have been made to develop effective treatments. Therapies targeting mTOR [3], EGFR [4], PARP [5], and immune checkpoint inhibitors are prominent among breast cancer treatments and can improve quality of life [6]. Nevertheless, the long-term effect of these treatments on tumor control remains limited, primarily due to the emergence of bypass mechanisms. Therefore, in-depth explorations of the mechanisms of action of these drugs are urgently needed.
mTOR is a highly conserved serine/threonine protein kinase, which belongs to phosphatidylinositol-3-kinase-related kinase protein family. The mTOR signaling pathway participates in cell growth, differentiation, and metabolism and is associated with several cancers' occurrence, progression, and metastasis [7, 8]. There is ample evidence suggesting that mTOR signaling pathway is hyperactive in breast cancer [9]. In addition, mTOR kinase can block autophagy by direct inhibition of the early steps of the process and the control of lysosomal degradative capacity of the cell, which is closely related to the survival and metabolism of tumor cells [10]. That is to say, the upregulated mTOR signal can promote the suppression of autophagy and activation of growth factor receptor signaling, glycolytic metabolism and lipid metabolism to favor tumor growth [11,12,13]. Therefore, restoring mTOR functioning becomes an important factor in breast cancer treatment [14].
mTOR exists its molecular functions via two distinct multiprotein complexes mTORC1 and mTORC2, their cellular functions are well established. The activity of mTORC1 plays key roles in the regulation of cell growth and proliferation, induction of apoptosis and inhibition of autophagy. mTORC2 is involved in the regulation of cell survival, metabolism and cytoskeleton organization [15]. The mTOR inhibitors are classified as first-, second- and third-generation depending on their inhibitory capabilities of mTORC1 and mTORC2. The first-generation inhibitors represented by rapamycin can only inhibit the mTORC1 signaling pathway, it appears some inhibitors have insignificant impact on cancer cell viability because of mTOR escape mechanism [16]. Compared with rapamycin, second- and third-generation inhibitors show much higher efficiency in tumor suppression and are attracting substantial attention. Second-generation mTOR inhibitors represented by AZD8055 refer to small molecule ATP competitive inhibitors, while third-generation inhibitor rapaLink-1 integrate rapamycin and dual mTORC1/2 inhibitor into a new molecule, they all can simultaneously target mTORC1 and mTORC2 [17]. Base on dual inhibition of mTORC1 and mTORC2, AZD8055 has expressed potent inhibitory effects over a wide range of cancers both in vitro and in vivo. Additionally, in combination with other agents, AZD8055 induces synergistic cytotoxic effects against various breast cancer cells [18].
Metabolic reprogramming is a key function for cancer cell growth [19], which ensure survival and proliferation of cancer cell in a nutrient-scarce situation. Metabolic reprogramming can often mediated by oncogenic signalling. In particular, mTOR signaling is commonly overactivated in tumours and induces cancer cell metabolic alterations [11, 20]. Metabolic alterations affect anabolic pathways for the synthesis of biomass, including aberrant metabolism of amino acids, glucose, nucleotides, fatty acids and lipids. Conversely, metabolic alterations, such as increased glucose or amino acid uptake, may influence mTOR signaling. Thus, an integrated understanding of the crosstalk between mTOR signaling and cancer metabolism can aid in the development of novel therapeutic strategies. In this study, we administered rapamycin and AZD8055 to breast cancer cell lines and measured metabolite changes in cells using proton nuclear magnetic resonance (1H NMR) spectroscopy. Our metabolomic results will provide the public with a deeper understanding of mTOR-mediated metabolic differences in breast cancer cells and lay a solid foundation for future mTORC1/2 dual inhibitor development.
Materials and methods
Cell culture and treatment
Human MDA-MB-231 and MDA-MB-453 cell lines were obtained from the China Infrastructure of Cell Line Resources (Beijing, China). AZD8055 (Selleck, China) and rapamycin (Selleck, China) were dissolved in dimethyl sulfoxide. MDA-MB-231 and MDA-MB-453 cells were grown in high-glucose Dulbecco’s modified Eagle’s medium (Gibco, USA) supplemented with 10% fetal bovine serum (Invitrogen, USA) and 1% penicillin/streptomycin (Invitrogen, USA) at 37 ℃ in a humidified atmosphere containing 5% CO2. After breast cancer cells grew to 50% confluence, the medium was replaced with or without AZD8055 or rapamycin, then incubated at 37 ℃ for 72 h.
CCK-8 assay
Cell proliferation and half-inhibitory concentration (IC50) were measured using CCK-8 reagents (Dojindo, Japan). In brief, the breast cancer cells were seeded at 5000 cells/well in 96-well plates. After the cells were cultured overnight and stabilized, the incubation media in each well were replaced with fresh culture media with or without treatments. The concentrations of AZD8055 were 2, 20, 100, 200, 500, 1000, 2000, and 20,000 nM. Concentrations for rapamycin were 250, 500, 1000, 2000, 3000, 6000, 18,000, and 36,000 nM. After 72 h, 10 μL of CCK-8 reagent was added to each well and incubated for another 4 h at 37 ℃. A microplate reader (Multiskan MK3, Thermo Labsystems, Finland) was used to measure the absorbance at 450 nm. The IC50 was calculated using GraphPad Prism software v7.0 (IBM, USA) from CCK-8 assay data.
Metabolite extraction and preparation of samples
Each group of cells was harvested and extracted using a methanol/chloroform/water extraction method [21]. The pellets obtained from centrifugation were washed three times with Hank’s balanced salt solution (Gibco, USA). Subsequently, the pellets were resuspended in 500 μL of methanol-chloroform (2:1, v/v) mixture before transferring into 1.5 ml Eppendorf tubes. After sonication on ice for 30 min, another 250 μL chloroform-water (1:1, v/v) was added. To obtain the aqueous phase, the mixture was thoroughly homogenized, placed on ice for 15 min, and centrifuged at 10,000 g for 15 min. Finally, the supernatants containing cellular metabolite were lyophilized and stored at –80 ℃ for subsequent experiments.
Acquisition and processing of NMR data
The lyophilized samples were reconstituted in 500 μl of 99.5% D2O (Cambridge Isotope Laboratories, USA) and transferred into 5-mm NMR tubes for NMR. All 1H NMR spectra of cell extracts were measured at 298 K using a Bruker AVANCE III 600 MHz NMR spectrometer (Bruker BioSpin, Germany). Water signal presaturation “ZGPR” (a standard single-pulse sequence) was used to obtain the one-dimensional 1H NMR spectra. The primary acquisition parameters were scans = 512, spectral width = 12,000 Hz, data points = 32 K, relaxation delay = 6 s, and acquisition time = 2.65 s per scan.
All one-dimensional 1H NMR spectra were referenced to the left peak of the lactate methyl peak at 1.33 ppm using Topspin software v2.1 (Bruker BioSpin, Germany) by manual adjustment. The spectral region from 0.5 to 10.0 ppm was intercepted for subsequent processing. The ‘icoshift’ procedure was applied to align NMR spectra in MATLAB software vR2012a (Mathworks, USA) [22]. After removing the region of 4.7–5.2 ppm (water) to eliminate artifacts related to the residual water resonance, the remaining spectral segments were subdivided with a size of 0.01 ppm for multivariate pattern recognition analysis and 0.0015 ppm for quantitative using MATLAB software vR2012a [22]. Then, the subdivided spectral segments were normalized to the total sum of the spectral intensity to minimize differences in the number of cells in each sample. To assign metabolic signals, the Human Metabolome Database (www.hmdb.ca) and Chenomx NMR suite software v7.0 (Chenomx, Canada) were used for identification and validation.
Multivariate pattern recognition analysis
The normalized NMR data were imported into SIMCA-P software v12.0 (Umetrics, Sweden) for multivariate statistical analysis. First, the integral values were mean-centered and Pareto-scaled. Then, partial least squares-discriminant analysis (PLS-DA) was performed to visualize metabolic differences between several groups, while orthogonal projections to latent structures discriminant analysis (OPLS-DA) was performed to maximize the metabolic difference between two groups. Variable importance in the projection (VIP) and S-plots were generated from the corresponding model to select significant metabolites that mainly contributed to the metabolic pattern separations.
Statistical analysis
Data were expressed as mean ± standard deviation. Statistical analyses were performed using SPSS software v16.0 (SPSS, USA). All experimental data were analyzed using the independent sample t-test or one-way analysis of variance. Differences were considered statistically significant at *p < 0.05, **p < 0.01, and ***p < 0.001.
Results
AZD8055 and rapamycin inhibit the growth of breast cancer cells
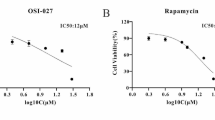
We used MDA-MB-231 and MDA-MB-453 breast cancer cell lines for the CCK-8 assay and found that the required growth inhibition concentrations were lower for AZD8055 than rapamycin. The IC50 values of AZD8055 and rapamycin in MDA-MB-231 cells were 464 nM and 4446 nM, respectively; their values in MDA-MB-453 cells were 107 nM and 1239 nM, respectively (Fig. 1). The IC50 values obtained in this step were used as the treatment concentrations in subsequent metabolic exploration experiments. MDA-MB-453 cells were more sensitive than MDA-MB-231 cells to rapamycin and AZD8055, possibly due to the positive expression of HER2 [23].
Differences in the antiproliferative capacity of AZD8055 and rapamycin in MDA-MB-231 and MDA-MB-453 cell lines. A, B Dose–response curves in MDA-MB-231cell lines after a 72 h exposure to AZD8055 or rapamycin. C, D Dose–response curves in MDA-MB-453 cell lines after a 72 h exposure to AZD8055 or rapamycin. IC50 values were calculated from three independent experiments
NMR-based metabolic profiling of MDA-MB-231 cells and MDA-MB-453 cells
Metabolic abnormalities of tumors are associated with their proliferation capabilities. To explore the metabolic differences between AZD8055 and rapamycin in breast cancer cells, we performed 1H NMR-based metabolomics. Typical 1H NMR spectra of extracts in MDA-MB-231 and MDA-MB-453 cells are shown in Fig. 2A. The spectra of the remaining dosing groups are similar to those in Fig. 2A; therefore, they are not displayed. We identified 22 and 23 metabolites in MDA-MB-231 and MDA-MB-453 cells, respectively. These metabolites were leucine, isoleucine, valine, 3-hydroxyisovalerate, lactate, alanine, acetate, UDP-N-acetylglucosamine (UDP-GlcNAc), glutamate, glutamine, succinate, aspartate, creatine, choline, O-phosphocholine (PCho), glucose, taurine, glycine, myo-inositol, UDP-glucose, inosine, adenosine 5’-monophosphate (AMP), tyrosine, phenylalanine, adenosine 5’-diphosphate (ADP), and nicotinamide adenine dinucleotide (NAD). The chemical shifts of the metabolites are shown in Supplementary Table S1.
NMR-based metabolomic analysis. A Typical 1H NMR spectrum obtained from extracts of MDA-MB-231 and MDA-MB-453 cells. PLS-DA score plots and their corresponding S-plots based on the metabolic profiles of MDA-MB-231 B and MDA-MB-453 cells C. Metabolite assignment: 1, Leucine; 2, Isoleucine; 3, Valine; 4, 3-Hydroxyisovalerate; 5, Lactate; 6, Alanine; 7, Acetate; 8, UDP-GlcNAc; 9, Glutamate; 10, Glutamine; 11, Succinate; 12, Aspartate; 13, Creatine; 14, Choline; 15, PCho; 16, Glucose; 17, Taurine; 18, Glycine; 19, Myo-inositol; 20, UDP-Glucose; 21, Inosine; 22, AMP; 23, Tyrosine; 24, Phenylalanine; 25, ADP; 26, NAD
AZD8055 and rapamycin induce changes in metabolic patterns in breast cancer cells
A PLS-DA model was used to determine the effects of AZD8055 and rapamycin on metabolic patterns in breast cancer cells, and the results are displayed in Figs. 2B and 2C. Results based on PLS-DA demonstrate that the metabolic patterns of the three groups were separated in MDA-MB-231 cells and MDA-MB-453 cells. Then, we obtained their corresponding S-plots from the PLS-DA models to identify the metabolites that contributed to differences in metabolic patterns. In MDA-MB-231 cells, the metabolic difference was ascribed to changes in lactate, leucine, glycine, and PCho (Fig. 2B). Lactate, alanine and PCho played a pivotal role in distinguishing the metabolic patterns among groups in MDA-MB-453 cells (Fig. 2C).
To understand the metabolic differences between the two groups, we performed pairwise comparisons of the metabolic data. OPLS-DA models and VIPs were generated to maximize the metabolic differences between the groups and identify critical metabolites (Fig. 3). In MDA-MB-231 and MDA-MB-453 cells, we observed a separation of metabolic patterns between each group. This finding suggests the differential effects of AZD8055 and rapamycin on the metabolic patterns of breast cancer cells.
Pairwise comparison of metabolic patterns in MDA-MB-231 and MDA-MB-453 cell models. A-C OPLS-DA score plots and VIP values based on the metabolism data of MDA-MB-231 cell model. D-F OPLS-DA score plots and VIP values based on the metabolism data of MDA-MB-453 cell model. A (R2 = 0.740, Q2 = 0.936), B (R2 = 0.594, Q2 = 0.832), C (R2 = 0.681, Q2 = 0.912), D (R2 = 0.845, Q2 = 0.981), E (R2 = 0.788, Q2 = 0.965), F (R2 = 0.662, Q2 = 0.802)
Metabolites with VIP > 1.0 were considered essential contributors to metabolic separations. In MDA-MB-231 cells, lactate, leucine, taurine, PCho, aspartate glycine, and alanine were selected to compare the control and AZD8055-treated groups (Fig. 3A). Taurine, PCho, glutamate, leucine, glycine, and phenylalanine were selected to compare control and rapamycin-treated groups (Fig. 3B). After focusing on the metabolic differences between AZD8055- and rapamycin-treated MDA-MB-231 cells, we found that lactate, myo-inositol, and aspartate contributed to their metabolic separation (Fig. 3C). However, in MDA-MB-453 cells, the results were somewhat different. The metabolites VIP > 1.0 were lactate, alanine, UDP-glucose, and UDP-GlcNAc when comparing the metabolic patterns of the control group and the AZD8055-treated group (Fig. 3D). Alanine, lactate, PCho, and glycine contributed significantly to the metabolic differences between the control and rapamycin-treated groups. Lactate, UDP-GlcNAc, glycine, alanine, PCho, and UDP-glucose differed significantly between AZD8055- and rapamycin-treated MDA-MB-453 cells (Fig. 3E, F). The metabolites with VIP values > 1.0 are involved in energy metabolism, amino acid metabolism, and lipid metabolism and participate in their respective metabolic pathways.
Effects of AZD8055 and rapamycin on the relative content of metabolites in breast cancer cells
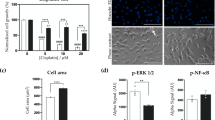
To further understand specific changes in cellular metabolite content, we performed a quantitative analysis of the normalized NMR data. As expected, AZD8055 and rapamycin changed the relative content of metabolites in breast cancer cells; however, the specific changes were not identical (Supplementary Tables S2 and S3). Heat maps were generated from quantitative data for some critical metabolites (Fig. 4). Figure 4A illustrates that AZD8055 and rapamycin reduced branched-chain amino acids (BCAAs) (leucine, isoleucine, and valine) and increased the relative content of glycine, alanine, aspartate, PCho, glutamate, and taurine in MDA-MB-231 cells. It is worth noting that AZD8055 and rapamycin had opposite effects on the content of glucose and glycolysis-related metabolites (lactate and myo-inositol). In MDA-MB-453 cells, changes in metabolite content were equally complex (Fig. 4B). We found that AZD8055-treated MDA-MB-453 cells had the highest glucose levels, while the relative content of most metabolites decreased after AZD8055 treatment.
Discussion
Metabolic reprogramming is ubiquitous in various tumors; the best known are enhanced glycolysis and increased glucose uptake [24, 25]. Gene mutations and overactivation of metabolic pathways are believed to act as drivers [25]. Many investigators have posited that a thorough understanding of the reprogrammed metabolic pathways of cancer cells will help develop novel therapeutic approaches [26, 27]. In this study, we investigated the effects of ADZ8055 and rapamycin on the metabolism of breast cancer cell lines using an NMR-based metabolomic approach. We hope that our findings will lead to a better understanding of dual mTORC1/2 inhibitors and facilitate the development of these inhibitors.
Several studies reported the in vitro anti-tumor ability of various dual mTORC1/2 inhibitors [28, 29]. Before exploring metabolic changes, we clarified the differences in the anti-proliferative capacity of AZD8055 and rapamycin in breast cancer cell lines. The CCK-8 assay obtained IC50 values for AZD8055 and rapamycin in cell lines. The IC50 values were used as the treatment concentrations to ensure that the metabolic data were derived from cells in the same inhibitory state. AZD8055 exhibited a much stronger anti-proliferative capacity than rapamycin in breast cancer. Numerous studies suggested that HER2 hyperactivity drives tumor growth by upregulation of the mTOR pathway activity and a metabolic shift to glycolysis [30, 31]. These insights also explain our finding that MDA-MB-453 (HER2-positive) cells were more sensitive to these inhibitors than MDA-MB-231 (HER2-negative) cells [32].
NMR-based metabolomics has been applied to identify biomarkers and has provided crucial insights into drug development due to its simple sample preparation and high reproducibility [33, 34]. In the present work, a series of metabolites in two cell lines were identified using this approach, including essential amino acids, semi-essential amino acids, intermediates of glycolysis, the TCA cycle, and cell membrane substrates. Multivariate analysis showed that AZD8055 had a different effect on the metabolic patterns of breast cancer cells than rapamycin. To clarify the metabolic differences between the two groups, OPLS-DA models were generated from metabolic data. R2 (the explained variance of the model) and Q2 (the model's predictive capability) were calculated to evaluate the performance of the OPLS-DA model. There was a clear separation of metabolic patterns across groups. We also identified the critical metabolites that contributed to the differences.
Although quantitative data for important metabolites were presented in heat maps (Fig. 4), we nevertheless believe it is necessary to discuss them in metabolic pathways, as illustrated in Fig. 5. The rapid proliferation of cancer cells requires more glucose for aerobic oxidation and glycolysis to provide energy [24]. The present study demonstrated significantly higher glucose levels in breast cancer cells after treatment with AZD8055 than rapamycin. This finding suggests that AZD8055 is likely to have a stronger ability to inhibit aerobic oxidation and glycolysis. The TCA cycle is involved in the aerobic oxidation of glucose, in which succinate is a vital intermediate [35]. In MDA-MB-231 cells, the succinate concentration in AZD8055-treated cells was significantly lower than in rapamycin-treated cells. This finding suggests that AZD8055 has a stronger ability to inhibit aerobic oxidation than rapamycin in MDA-MB-231 cells.
Metabolic pathway analysis according to the KEGG (www.genome.jp/kegg/) and SMPDB database (www.smpdb.ca/). r.u., relative unit. *p < 0.05, **p < 0.01, ***p < 0.001
Enhanced glycolysis and increased lactate production are common in tumors [36]. Enhanced glycolysis tends to increase the flux of glycolytsis intermediates into biosynthesis pathways that utilize these intermediates as substrates [37]. In our study, AZD8055-treated breast cancer cells had significantly lower lactate content than rapamycin-treated cells. This finding can still prompt us that AZD8055 may have a stronger potential for glycolysis inhibition than rapamycin, after we consider the interfering factors of lactate transport, pyruvate oxidation. Myo-inositol, UDP-glucose, and UDP-GlcNAc are synthesized from glycolytic intermediates through biological pathways [37, 38]. Myo-inositol plays a pivotal function in many metabolic pathways [39], and UDP-GlcNAc promotes the synthesis of hyaluronan, which leads to tumor progression [37, 40]. Changes in their concentrations were observed in our work similar to those in lactate, which strongly supports our conjecture that AZD8055 has a stronger ability to inhibit glycolysis.
Since another aspect of metabolic reprogramming observed in cancer cells is amino acid dysregulation, special attention should be paid to metabolic changes in certain amino acids. Cancer cells typically express higher levels of amino acid transporters on the surface to meet the increased demand for amino acids needed for proliferation [41]. In addition, some functional amino acids (e.g., leucine, arginine and glutamine) can regulate autophagy through the signal transformation of the signal relay station mTORC1 [42]. BCAAs (leucine, isoleucine and valine) are members of the essential amino acid family, which cancer cells must acquire from the circulation or from surrounding tissues [43]. Our study found that AZD8055 and rapamycin have nearly identical roles in regulating BCAAs concentrations in MDA-MB-231 cells, while that was only found in leucine changes in MDA-MB-453 cells. Glutamine is the most consumed amino acid by cancer cells, which can be used to synthesize glutamate and enter the TCA cycle through further transformation [44]. Here, we detected no effect of AZD8055 and rapamycin on glutamine and glutamate concentrations in MDA-MB-453 cells. However, it is worth noting that both AZD8055 and rapamycin increased glutamate content in MDA-MB-231 cells, although AZD8055 was less effective in this regard than rapamycin. Taurine as an antioxidant has strong growth inhibitory effects on various cancers, and its levels are lower in breast cancer patients than in normal people [45]. Based on our data, AZD8055 and rapamycin may equally inhibit the oxidative stress process of MDA-MB-231 cells by increasing the taurine level.
Breast cancer cells have varying gene expression levels and thus different metabolic levels [46,47,48]. Unlike MDA-MB-231 cells, MDA-MB-453 cells are HER2-positive [32]; therefore, it makes sense that AZD8055 and rapamycin would have cell-specific metabolic effects on breast cancer cells. For example, the concentration of succinate in AZD8055- and rapamycin-treated MDA-MB-453 cells showed no significant difference, unlike those found in MDA-MB-231 cells. Moreover, the changes of BCAAs, glycine, alanine, PCho, and aspartate in the two cell lines were also inconsistent. This cell-specific metabolic difference requires further study.
In summary, our findings suggest that the ability of the dual mTORC1/2 inhibitor AZD8055 to inhibit the proliferation of breast cancer cells was much stronger than that of rapamycin. NMR-based metabolomic results showed differences in cellular metabolic patterns after AZD8055 and rapamycin treatments. Relative quantitative results and metabolic pathway analysis demonstrated that AZD8055 may inhibit glycolysis and reduce glucose consumption more strongly than rapamycin; both AZD8055 and rapamycin have the ability to broadly affect amino acid metabolism in breast cancer cells, although these effects were not exactly the same. Nevertheless, the cell-specific metabolic effects of AZD8055 and rapamycin on breast cancer cells require further elucidation.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
World Health Organization (WHO) (2014) WHO Position Paper on Mammography Screening. https://paho.org/hq/dmdocuments/2015/WHO-ENG-Mammography-Factsheet.pdf
Lee J, Loh K, Yap Y (2015) PI3K/Akt/mTOR inhibitors in breast cancer. Cancer Biol Med 12:342–354. https://doi.org/10.7497/j.issn.2095-3941.2015.0089
Carey L, Rugo H, Marcom P, Mayer E, Esteva F, Ma C, Liu M, Storniolo A, Rimawi M, Forero-Torres A et al (2012) TBCRC 001: randomized phase II study of cetuximab in combination with carboplatin in stage IV triple-negative breast cancer. J Clin Oncol 30:2615–2623. https://doi.org/10.1200/jco.2010.34.5579
Mateo J, Lord C, Serra V, Tutt A, Balmaña J, Castroviejo-Bermejo M, Cruz C, Oaknin A, Kaye S, de Bono J (2019) A decade of clinical development of PARP inhibitors in perspective. Ann Oncol 30:1437–1447. https://doi.org/10.1093/annonc/mdz192
Hu X, Huang W, Fan M (2017) Emerging therapies for breast cancer. J Hematol Oncol 10:98. https://doi.org/10.1186/s13045-017-0466-3
Tewari D, Patni P, Bishayee A, Sah A, Bishayee A (2022) Natural products targeting the PI3K-Akt-mTOR signaling pathway in cancer: A novel therapeutic strategy. Semin Cancer Biol 80:1–17. https://doi.org/10.1016/j.semcancer.2019.12.008
Wullschleger S, Loewith R, Hall M (2006) TOR signaling in growth and metabolism. Cell 124:471–484. https://doi.org/10.1016/j.cell.2006.01.016
Miller T, Rexer B, Garrett J, Arteaga C (2011) Mutations in the phosphatidylinositol 3-kinase pathway: role in tumor progression and therapeutic implications in breast cancer. Breast Cancer Res 13:224. https://doi.org/10.1186/bcr3039
Deleyto-Seldas N, Efeyan A (2021) The mTOR-Autophagy Axis and the Control of Metabolism. Front Cell Dev Biol 9:655731. https://doi.org/10.3389/fcell.2021.655731
Saxton R, Sabatini D (2017) mTOR Signaling in Growth, Metabolism, and Disease. Cell 168:960–976. https://doi.org/10.1016/j.cell.2017.02.004
Hua H, Kong Q, Zhang H, Wang J, Luo T, Jiang Y (2019) Targeting mTOR for cancer therapy. J Hematol Oncol 12:71. https://doi.org/10.1186/s13045-019-0754-1
Yin Y, Hua H, Li M, Liu S, Kong Q, Shao T, Wang J, Luo Y, Wang Q, Luo T et al (2016) mTORC2 promotes type I insulin-like growth factor receptor and insulin receptor activation through the tyrosine kinase activity of mTOR. Cell Res 26:46–65. https://doi.org/10.1038/cr.2015.133
Zheng Y, Jiang Y (2015) mTOR Inhibitors at a Glance. Mol Cell Pharmacol 7:15–20
Vergès B, Walter T, Cariou B (2014) Endocrine side effects of anti-cancer drugs: effects of anti-cancer targeted therapies on lipid and glucose metabolism. Eur J Endocrinol 170:R43-55. https://doi.org/10.1530/eje-13-0586
Carew J, Kelly K, Nawrocki S (2011) Mechanisms of mTOR inhibitor resistance in cancer therapy. Target Oncol 6:17–27. https://doi.org/10.1007/s11523-011-0167-8
Chen Y, Zhou X (2020) Research progress of mTOR inhibitors. Eur J Med Chem 208:112820. https://doi.org/10.1016/j.ejmech.2020.112820
Li H, Liu L, Chang H, Zou Z, Xing D (2018) Downregulation of MCL-1 and upregulation of PUMA using mTOR inhibitors enhance antitumor efficacy of BH3 mimetics in triple-negative breast cancer. Cell Death Dis 9:137. https://doi.org/10.1038/s41419-017-0169-2
Vander Heiden M, DeBerardinis R (2017) Understanding the Intersections between Metabolism and Cancer Biology. Cell 168:657–669. https://doi.org/10.1016/j.cell.2016.12.039
Zhang Y, Kwok-Shing Ng P, Kucherlapati M, Chen F, Liu Y, Tsang Y, de Velasco G, Jeong K, Akbani R, Hadjipanayis A et al (2017) A Pan-Cancer Proteogenomic Atlas of PI3K/AKT/mTOR Pathway Alterations. Cancer Cell 31:820-832.e823. https://doi.org/10.1016/j.ccell.2017.04.013
Zhang X, Lin Q, Chen J, Wei T, Li C, Zhao L, Gao H, Zheng H (2018) High Glucose-Induced Cardiomyocyte Death May Be Linked to Unbalanced Branched-Chain Amino Acids and Energy Metabolism. Molecules 23. https://doi.org/10.3390/molecules23040807
Savorani F, Tomasi G, Engelsen S (2010) icoshift: A versatile tool for the rapid alignment of 1D NMR spectra. J Magn Reson 202:190–202. https://doi.org/10.1016/j.jmr.2009.11.012
D'Amico P, Reduzzi C, Qiang W, Zhang Y, Gerratana L, Zhang Q, Davis A, Shah A, Manai M, Curigliano G, et al. (2021) Single-Cells Isolation and Molecular Analysis: Focus on HER2-Low CTCs in Metastatic Breast Cancer. Cancers 14. https://doi.org/10.3390/cancers14010079
DeBerardinis R, Chandel N (2016) Fundamentals of cancer metabolism Sci Adv 2:e1600200. https://doi.org/10.1126/sciadv.1600200
Levine A, Puzio-Kuter A (2010) The control of the metabolic switch in cancers by oncogenes and tumor suppressor genes. Science 330:1340–1344. https://doi.org/10.1126/science.1193494
Chakraborty S, Balan M, Sabarwal A, Choueiri T, Pal S (2021) Metabolic reprogramming in renal cancer: Events of a metabolic disease. Biochim Biophys Acta Rev Cancer 1876:188559. https://doi.org/10.1016/j.bbcan.2021.188559
Pavlova N, Zhu J, Thompson C (2022) The hallmarks of cancer metabolism: Still emerging. Cell Metab 34:355–377. https://doi.org/10.1016/j.cmet.2022.01.007
Li S, Sheng J, Liu Z, Fan Y, Zhang C, Lv T, Hu S, Jin J, Yu W, Song Y (2021) Potent antitumour of the mTORC1/2 dual inhibitor AZD2014 in docetaxel-sensitive and docetaxel-resistant castration-resistant prostate cancer cells. J Cell Mol Med 25:2436–2449. https://doi.org/10.1111/jcmm.16155
Zhang Q, Zhang Y, Chen Y, Qian J, Zhang X, Yu K (2019) A Novel mTORC1/2 Inhibitor (MTI-31) Inhibits Tumor Growth, Epithelial-Mesenchymal Transition, Metastases, and Improves Antitumor Immunity in Preclinical Models of Lung Cancer. Clin Cancer Res 25:3630–3642. https://doi.org/10.1158/1078-0432.Ccr-18-2548
Holloway R, Marignani P (2021) Targeting mTOR and Glycolysis in HER2-Positive Breast Cancer. Cancers (Basel) 13. https://doi.org/10.3390/cancers13122922
Paul M, Pan T, Pant D, Shih N, Chen Y, Harvey K, Solomon A, Lieberman D, Morrissette J, Soucier-Ernst D et al (2020) Genomic landscape of metastatic breast cancer identifies preferentially dysregulated pathways and targets. J Clin Invest 130:4252–4265. https://doi.org/10.1172/jci129941
Holliday D, Speirs V (2011) Choosing the right cell line for breast cancer research. Breast Cancer Res 13:215. https://doi.org/10.1186/bcr2889
Luo P, Yin P, Hua R, Tan Y, Li Z, Qiu G, Yin Z, Xie X, Wang X, Chen W et al (2018) A Large-scale, multicenter serum metabolite biomarker identification study for the early detection of hepatocellular carcinoma. Hepatology 67:662–675. https://doi.org/10.1002/hep.29561
Yuliana N, Khatib A, Verpoorte R, Choi Y (2011) Comprehensive extraction method integrated with NMR metabolomics: a new bioactivity screening method for plants, adenosine A1 receptor binding compounds in Orthosiphon stamineus Benth. Anal Chem 83:6902–6906. https://doi.org/10.1021/ac201458n
Mills E, O’Neill L (2014) Succinate: a metabolic signal in inflammation. Trends Cell Biol 24:313–320. https://doi.org/10.1016/j.tcb.2013.11.008
Doherty J, Cleveland J (2013) Targeting lactate metabolism for cancer therapeutics. J Clin Invest 123:3685–3692. https://doi.org/10.1172/jci69741
Oikari S, Kettunen T, Tiainen S, Häyrinen J, Masarwah A, Sudah M, Sutela A, Vanninen R, Tammi M, Auvinen P (2018) UDP-sugar accumulation drives hyaluronan synthesis in breast cancer. Matrix Biol 67:63–74. https://doi.org/10.1016/j.matbio.2017.12.015
Koguchi T, Tanikawa C, Mori J, Kojima Y, Matsuda K (2016) Regulation of myo-inositol biosynthesis by p53-ISYNA1 pathway. Int J Oncol 48:2415–2424. https://doi.org/10.3892/ijo.2016.3456
Benvenga S, Antonelli A (2016) Inositol(s) in thyroid function, growth and autoimmunity. Rev Endocr Metab Disord 17:471–484. https://doi.org/10.1007/s11154-016-9370-3
Auvinen P, Tammi R, Kosma V, Sironen R, Soini Y, Mannermaa A, Tumelius R, Uljas E, Tammi M (2013) Increased hyaluronan content and stromal cell CD44 associate with HER2 positivity and poor prognosis in human breast cancer. Int J Cancer 132:531–539. https://doi.org/10.1002/ijc.27707
Saito Y, Soga T (2021) Amino acid transporters as emerging therapeutic targets in cancer. Cancer Sci 112:2958–2965. https://doi.org/10.1111/cas.15006
Liu C, Ji L, Hu J, Zhao Y, Johnston L, Zhang X, Ma X (2021) Functional Amino Acids and Autophagy: Diverse Signal Transduction and Application. Int J Mol Sci 22. https://doi.org/10.3390/ijms222111427
Peng H, Wang Y, Luo W (2020) Multifaceted role of branched-chain amino acid metabolism in cancer. Oncogene 39:6747–6756. https://doi.org/10.1038/s41388-020-01480-z
Geck R, Toker A (2016) Nonessential amino acid metabolism in breast cancer. Adv Biol Regul 62:11–17. https://doi.org/10.1016/j.jbior.2016.01.001
Baliou S, Kyriakopoulos A, Spandidos D, Zoumpourlis V (2020) Role of taurine, its haloamines and its lncRNA TUG1 in both inflammation and cancer progression. On the road to therapeutics? (Review). Int J Oncol 57:631–664. https://doi.org/10.3892/ijo.2020.5100
Yeo S, Guan J (2017) Breast Cancer: Multiple Subtypes within a Tumor? Trends Cancer 3:753–760. https://doi.org/10.1016/j.trecan.2017.09.001
Polyak K (2007) Breast cancer: origins and evolution. J Clin Invest 117:3155–3163. https://doi.org/10.1172/jci33295
Perou C, Sørlie T, Eisen M, van de Rijn M, Jeffrey S, Rees C, Pollack J, Ross D, Johnsen H, Akslen L et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752. https://doi.org/10.1038/35021093
Funding
This research was supported by Medical Health Science and Technology Project of Zhejiang Provincial Health Commission under Grant No. 2019KY448, Project of Wenzhou Science and Technology Bureau under Grant No. Y20210219 and Clinical Scientific Research Project of Zhejiang Medical Association under Grant No. 2017ZYC-A25.
Author information
Authors and Affiliations
Contributions
Xiuhua Zhang and Ziye Zhou design the study and revise the manuscript; Zhitao Ni, Shaolin Xu, Zheng Yu and Huamin Liu perform the experiments; Zhongjiang Ye, Rongqi Li, Chuang Chen and Jianhui Yang analyze the experimental data; Zhitao Ni writes the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants.
Informed consent
For this type of study, informed consent is not required.
Competing interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ni, Z., Xu, S., Yu, Z. et al. Comparison of dual mTORC1/2 inhibitor AZD8055 and mTORC1 inhibitor rapamycin on the metabolism of breast cancer cells using proton nuclear magnetic resonance spectroscopy metabolomics. Invest New Drugs 40, 1206–1215 (2022). https://doi.org/10.1007/s10637-022-01268-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-022-01268-w