Abstract
Background
TNF-α processing inhibitor-1 (TAPI-1) is a known metalloproteinase inhibitor with potential anti-inflammatory effects. However, its anti-cancer effects on esophageal squamous cell carcinoma (ESCC) have not been uncovered.
Aim
In the present study, the effects of TAPI-1 on ESCC cell viability, migration, invasion, and cisplatin resistance and the underlying molecular mechanisms were investigated in TE-1 and Eca109 cells.
Methods
To this end, TE-1 and Eca109 cells were exposed to TAPI-1 for indicated time intervals. Cell viability was assessed using cell counting kit-8 assay and apoptosis was evaluated using flow cytometry assay. Migration and invasion were assessed using Transwell assays. Gene expressions were analyzed using quantitative reverse transcription polymerase chain reaction. The activation of NF-κB signaling pathway was elucidated via Western blot and chromatin immunoprecipitation assay.
Results
We observed that higher doses (10, 20 μM) of TAPI-1 inhibited ESCC cell viability, while a lower dose (5 μM) of TAPI-1 inhibited ESCC cell migration and invasion and enhanced the chemosensitivity of ESCC cells to cisplatin. Moreover, TAPI-1 suppressed the activation of NF-κB signaling and the target genes expression in the stage of transcription initiation. Furthermore, blocking NF-κB signaling in advance could abolish all the effects of TAPI-1 on ESCC cells.
Conclusion
Overall, these results indicated that TAPI-1 impairs ESCC cell viability, migration, and invasion and facilitates cisplatin-induced apoptosis via suppression of NF-κB signaling pathway. TAPI-1 may serve as a potential adjuvant agent with cisplatin for ESCC therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Esophageal carcinoma is one of the most frequently diagnosed digestive cancers, known as the sixth most common cause of cancer-related death worldwide [1]. Esophageal squamous cell carcinoma (ESCC) is the major histopathological subtype of esophageal carcinoma [2]. Although advances have been made in treatments for ESCC, the poor prognosis remains disappointing due to recurrence, metastasis, and the resistance to chemotherapeutic agents, such as cisplatin [3,4,5]. Therefore, it is necessary to understand the molecular mechanisms of ESCC pathogenesis and develop efficient adjuvant treatment modalities, such as anti-tumor drugs.
TNF-α processing inhibitor-1 (TAPI-1) is a known broad-spectrum metalloproteinase inhibitor, which inhibits the shedding of TNF-α and other cytokine receptors [6, 7]. In recent years, TAPI-1 has been proposed to ameliorate various inflammatory diseases, such as rheumatoid arthritis, kidney injury, and neuroinflammation [8,9,10]. Additionally, several studies showed that TAPI-1 might display anti-tumor efficacy in pancreatic cancer, breast cancer, and hepatoma [11,12,13,14,15]. However, the effect of TAPI-1 on digestive tract tumors remains to be greatly elucidated. Intriguingly, a previous review has concluded that the matrix metalloproteinase (MMP) family, a potential molecular target of TAPI-1, facilitated ESCC progression [16]. Moreover, our previous study revealed that knockdown of TNF-α-converting enzyme (TACE, also named ADAM17), another molecular target of TAPI-1, significantly inhibited ESCC cell viability [17]. Therefore, we hypothesize that TAPI-1 may exhibit anti-tumor effects on ESCC.
In the present study, our data indicated that TAPI-1 could inhibit ESCC cell viability, migration, and invasion and enhanced the chemosensitivity of ESCC cells to cisplatin, while exhibited low toxicity in normal esophageal epithelial cells. Of note, TAPI-1 remarkably inhibited NF-κB signaling pathway in ESCC cells. Moreover, suppression of NF-κB signaling pathway in advance remarkably abolished the anti-tumor effects of TAPI-1, suggesting that TAPI-1 exhibits anti-tumor efficacy in an anti-NF-κB manner in ESCC cells.
Materials and Methods
Cell Lines and Chemicals
Human esophageal epithelial cell line Het-1A and human ESCC cell lines TE-1 and Eca109 were purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). All the cell lines were maintained in RPMI medium (Gibco), supplemented with 10% fetal bovine serum (FBS), 100 units/ml penicillin, and 100-mg/ml streptomycin and incubated at 37 °C in a humidified atmosphere containing 5% CO2.
TAPI-1 (HY-16657) and BAY11-7082 (HY-13453) were purchased from MedChem Express and dissolved in dimethyl sulfoxide (DMSO) for storage at – 80 °C. Cisplatin (PHR1624) was obtained from Sigma-Aldrich and dissolved in phosphate-buffered saline (PBS) for storage at – 20 °C.
Cell Counting Kit-8 (CCK-8) Assay
CCK-8 assay was conducted to assess cell viability. In brief, cells were inoculated in 96-well plates with a density of 1 × 103 cells per well. After chemicals treatment, cells were washed twice in PBS. Cells were then subjected to CCK-8 reagents (Dojindo Laboratories, Kumamoto, Japan) with the recommended protocol. The absorbance was measured at 450 nm using a Synergy H1 Microplate Reader (Bio‐Tek). The average absorbance of six replicates was calculated.
RNA Extraction and Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR)
In brief, the total RNA from cultured cells was extracted using TRIzol reagent (Invitrogen). After reverse transcription (TaKaRa, Japan), 2 × QuantiNova SYBR Green PCR Master Mix (QIAGEN) was applied to analyze the relative mRNA levels using a CFX Connect™ Real-Time System (Bio‐Rad). The fold change relative to control was calculated using 2−ΔΔCt method. The mRNA levels were normalized to β-actin. Partial primer sequences were listed as follows: 5′‐CCACGAAACTACCTTCAACTCCATC‐3′ and 5′‐ACTCCTGCTTGCTGATCC- ACAT‐3′ for β-actin, 5′‐TCCCTACAGACAGAGCCACAAG‐3′ and 5′‐CTTCAC- CTCCGTGATTGCCTTC‐3′ for BIM, 5′‐GAGCTGAAGCAGATGCAGGACAA‐3′ and 5′‐TGACGGAGTTGCCACTTGACTTG‐3′ for TRAIL, 5′‐TGTGGATGACTG- AGTACCTGAACC‐3′ and 5′‐CAGAGACAGCCAGGAGAAATCAAAC‐3′ for BCL2, 5′‐AACAGGAACACAGGAGTCATCAGT‐3′ and 5′‐CGGAGGATTATCG- TTGGTGTCAGT‐3′ for CDH1, 5′‐TCCTGCTTATCCTTGTGCTGATGT‐3′ and 5′‐GTCTTCTTCTCCTCCACCTTCTTCA‐3′ for CDH2, 5′‐ACTGTGACAAGG- AATATGTGAGCC‐3′ and 5′‐CAAGGTAATGTGTGGGTCCGAAT‐3′ for SLUG, and 5′‐CCTACACCAAGAACTTCCGTCTGTC‐3′ and 5′‐GTGCCAAGGTCAATGTC- AGGAGAG‐3′ for MMP2. The remaining primer sequences are listed in Table 1.
Flow Cytometry Assay
Apoptosis was evaluated by the Annexin V-propidium iodide (PI) double staining assay. In brief, ESCC cells were digested into single cells and washed twice with ice-cold PBS. After centrifugation, cells were re-suspended and stained with Annexin V and PI using the FITC Annexin V Apoptosis Detection kit I (BD Pharmingen™) according to the manufacturer’s protocols. Apoptosis was analyzed using BD LSRFortessa™.
Migration and Invasion Assays
Cell migration and invasion capacity were evaluated using Transwell inserts (8-µm pores; Corning) in 24‐well plates. In brief, 200 μl of serum‐free RPMI medium with 1 × 105 cells was added to the upper chamber with Matrigel (BD Biosciences) for invasion assay or without Matrigel for migration assay. After chemicals treatment, 500 μl of RPMI medium with 20% FBS was added to the lower chamber, followed by an incubation for 24 h at 37 °C. Nonmigrating cells remaining at the top of the upper chamber were removed with cotton swabs. The cells that migrated to the bottom of the upper chamber were fixed with 4% formaldehyde and stained with crystal violet. Cells were observed under a microscope at ×200 magnification and then lysed with 500 μl of 0.2% Triton X-100 per well. The absorbance was measured at 550 nm using a Synergy H1 Microplate Reader (Bio‐Tek). The average absorbance of three independent experiments was calculated.
Bioinformatics Analysis
The putative transcription factors in the upstream of cisplatin resistance-related genes were speculated via the ChEA3 website (https://maayanlab.cloud/chea3). Ten transcription factors with the highest scores from the Enrichr Queries library are listed in Fig. 4A.
Protein Extraction and Western Blot Analysis
To obtain total protein, cells were lysed in RIPA buffer (Beyotime) containing 0.5-mM PMSF (Sigma) and 1 × phosphatase inhibitor (Roche). Nuclear and Cytoplasmic Protein Extraction Kit (Beyotime) was applied for nuclear protein extraction with the recommended protocol. β-actin and H3 were used as internal controls for the total and nuclear proteins, respectively. The following antibodies were used: anti-β-actin antibody (Proteintech, 66009-1-Ig, 1:5000), anti-H3 antibody (Proteintech, 17168-1-AP, 1:2000), anti-NF-κB p65 antibody (Wanleibio, WL01980, 1:500), anti-P-NF-κB p65 (Wanleibio, WL02169, 1:500), HRP Goat Anti-Mouse IgG (Jackson, 115-035-003, 1:10,000), and HRP Goat Anti-Rabbit IgG (Jackson, 111-035-003, 1:10,000). The gray levels were analyzed using ImageJ software.
Chromatin Immunoprecipitation Assay
Chromatin immunoprecipitation (ChIP) was carried out using Pierce™ Agarose ChIP Kit (Thermo, 26156) according to the manufacturer’s protocol. The ChIP-grade anti-NF-κB p65 antibody was purchased from CST (#6956, 1:50). Immunoprecipitated DNA was analyzed by quantitative polymerase chain reaction (qPCR) with 2 × QuantiNova SYBR Green PCR Master Mix (QIAGEN). The putative NF‐κB‐binding sites in c-Myc, CyclinD1 and MMP2 promoters were speculated via the JASPAR website (https://jaspar.genereg.net) and the primers were generated by TsingKe Biological Technology (Beijing, China). The results from three independent experiments were averaged and plotted as percentage of input. The primer sequences were as followed: 5′‐CTAAGTGATCAGACACCGTCAG‐3′ and 5′‐TGTCCCTGGTCAAGTAGGTTT‐3′ for c-Myc, 5′‐CCAGGCTAGAAGGA- CAAGATGAAGG‐3′ and 5′‐GCGTCGTTGCAAATGCCCAAG‐3′ for CyclinD1, 5′‐GTTAGGCAAGTGACTTCTCAGT‐3′ and 5′‐GTAAGCACCTAAGATTACCT- CTCT‐3′ for MMP2.
Statistical Analysis
Statistical analyses were performed with GraphPad Prism 8.0 software. All quantifications were performed with at least three independent experiments. Comparison of two groups was performed using an unpaired Student’s t test. One-way analysis of variance (ANOVA) was applied when more than two groups were compared. Data were presented as mean ± standard deviation (SD). P < 0.05, P < 0.01, and P < 0.001 represented different degrees of statistical significance.
Results
TAPI-1 Mildly Inhibits ESCC Cell Viability with Low Toxicity
To determine whether exposure to TAPI-1 could influence ESCC cell viability, TE-1 and Eca109 cells were exposed to varying doses of TAPI-1 (0, 1.25, 2.5, 5, 10, and 20 μM) for 24 h, following which, cell viability was determined using CCK-8 assay. Interestingly, following exposure of TAPI-1, TE-1, and Eca109 cell viability was inhibited in a dose-dependent manner. As shown in Fig. 1A, in the presence of 10 and 20 μM of TAPI-1, cell viability decreased significantly in TE-1 and Eca109 cells. Next, we performed the time-course experiments to observe duration of decreased cell viability in TE-1 and Eca109 cells exposed to TAPI-1 (10 μM) for the indicated time points (0, 24, 48, and 72 h). Exposure of ESCC cells to TAPI-1 resulted in remarkably decreased cell viability at 24 and 48 h (Fig. 1B), following which, there was no significant alteration in cell viability at 72 h. Moreover, we carried out the same experiments in human-immortalized esophageal epithelial cell line Het-1A and observed no alteration in cell viability (Fig. 1A and 1B), suggesting that TAPI-1 could inhibit ESCC cell viability without toxicity to normal esophageal epithelial cells.
The effect of TAPI-1 on ESCC cell viability. A CCK-8 analysis of cell viability after exposure of Het-1A, TE-1, and Eca109 cells to DMSO or varying doses of TAPI-1 (1.25, 2.5, 5, 10, and 20 μM). B CCK-8 analysis of cell viability after exposure of Het-1A, TE-1, and Eca109 cells to DMSO or TAPI-1 (10 μM) for 0, 24, 48, and 72 h, respectively. C TE-1 and Eca109 cells were exposed to DMSO or TAPI-1 (10 μM) for 24 h. qRT-PCR quantitative analysis of BIM, TRAIL, and BCL2 expression at mRNA levels. D TE-1 and Eca109 cells were exposed to DMSO or TAPI-1 (10 μM) for 24 h. Apoptosis analysis using flow cytometry and quantitative analysis of apoptotic cells. *P < 0.05, **P < 0.01, and ***P < 0.001, compared with the DMSO group
To further investigate whether exposure to TAPI-1 could influence ESCC cell apoptosis, TE-1 and Eca109 cells were exposed to TAPI-1 (10 μM) or solvent control. Following exposure to TAPI-1 for 24 h, the total RNA was extracted and the relative mRNA levels of apoptosis-related genes were determined using qRT-PCR. As shown in Fig. 1C, we observed no obvious change at the mRNA levels of pro-apoptosis genes BIM and TRAIL, and the anti-apoptosis gene BCL2 after TAPI-1 treatment in Eca109 cells. In TE-1 cells, TRAIL was significantly upregulated in response to TAPI-1 treatment. Moreover, flow cytometry assay showed that neither TE-1 nor Eca109 cell apoptosis was influenced following TAPI-1 treatment for 72 h (P > 0.05) (Fig. 1D). These data suggested that TAPI-1 markedly inhibited ESCC cell viability and showed no effect on cell apoptosis.
TAPI-1 Inhibits ESCC Cell Migration and Invasion
To investigate whether exposure to TAPI-1 could influence ESCC cell migration and invasion, Transwell chambers with or without Matrigel were applied. To rule out the impact of TAPI-1 on ESCC cell numbers, the concentration of 5-μM TAPI-1 chosen to assess the migration and invasion ability of ESCC cells. TE-1 and Eca109 cells were exposed to TAPI-1 (5 μM) or solvent control for 24 h. In Fig. 2A, qRT-PCR showed that the mRNA levels of SLUG and MMP2 expression were remarkably downregulated in response to TAPI-1 treatment in TE-1 and Eca109 cells. Moreover, we observed that CDH1 (E-cadherin) expression was slightly upregulated after TAPI-1 treatment in TE-1 cells. However, no obvious change in CDH2 (N-cadherin) expression was observed in TE-1 or Eca109 cells. Additionally, Transwell assays indicated that TAPI-1 treatment significantly suppressed the invasion and migration of TE-1 and Eca109 cells, respectively (Fig. 2B, C). These data suggested that TAPI-1 could significantly inhibit ESCC cell migration and invasion.
The effect of TAPI-1 on ESCC cell migration and invasion. TE-1 and Eca109 cells were exposed to DMSO or TAPI-1 (5 μM) for 24 h. A qRT-PCR quantitative analysis of CDH1, CDH2, SLUG, and MMP2 expression at mRNA levels. Representative images and absorbance analysis results of Transwell migration assay (B) and invasion assay (C). *P < 0.05, **P < 0.01, and ***P < 0.001, compared with the DMSO group
TAPI-1 Promotes the Sensitivity of ESCC Cells to Cisplatin
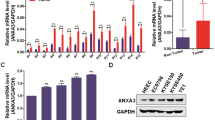
Chemoresistance remains the leading cause of treatment failure in ESCC. Thus, we then evaluated the impact of TAPI-1 on chemoresistance of ESCC cells using the gold-standard chemotherapeutic drug cisplatin. To rule out the impact of TAPI-1 itself on ESCC cell viability, the concentration of 5-μM TAPI-1 chosen to assess the sensitivity of ESCC cells to cisplatin due to no obvious alteration in ESCC cell viability (Fig. 1A). First, to determine the effect of TAPI-1 on cisplatin resistance-related genes, TE-1 and Eca109 cells were exposed to TAPI-1 (5 μM), following which, the total RNA was extract. Numerous studies have documented a variety of cisplatin resistance-related genes over recent years. From 2016 to 2021, 30 members of the genes that have been recently reported to facilitate cisplatin resistance are listed in Tab. 1. As shown in Fig. 3A, the relative mRNA levels of the 30 cisplatin resistance-related genes were determined using qRT-PCR. A Venn diagram was made to compare the differences in gene expressions between TE-1 and Eca109 cells (Fig. 3B). In Fig. 3B, we observed that 29 genes (ABCC10, ACTL6A, AKR1C2, ATP7A, CHD1L, CMTM6, EIF4E, FZD7, GSTP1, HMGN5, HOXB7, ITGA5, ITGB1, LAMP2, MAGED1, MCL1, NFE2L2, NGFR, NSD2, RAC1, RACK1, SFN, SLC2A1, SOD2, SRPX2, UBQLN4, VRK1, YAP1, YBX1) expressions were downregulated in TE-1 cells and 21 genes (ABCC10, AKR1C2, ATP7A, FZD7, GSTP1, ITGA5, ITGB1, LAMP2, MAGED1, MCL1, NFE2L2, RAC1, RACK1, SFN, SLC2A1, SOD2, SRPX2, UBQLN4, YAP1, YBX1, ZFX) expressions were downregulated in Eca109 cells at the mRNA levels in response to TAPI-1 treatment. In summary, 20 genes (ABCC10, AKR1C2, ATP7A, FZD7, GSTP1, ITGA5, ITGB1, LAMP2, MAGED1, MCL1, NFE2L2, RAC1, RACK1, SFN, SLC2A1, SOD2, SRPX2, UBQLN4, YAP1, YBX1) expressions were simultaneously downregulated in TE-1 and Eca109 cells at the mRNA levels after TAPI-1 treatment.
The effect of TAPI-1 on the sensitivity of ESCC cells to cisplatin. A TE-1 and Eca109 cells were exposed to DMSO or TAPI-1 (5 μM) for 12 h. qRT-PCR quantitative analysis of ABCC10, ACTL6A, AKR1C2, ATP7A, CHD1L, CMTM6, EIF4E, FZD7, GSTP1, HMGN5, HOXB7, ITGA5, ITGB1, LAMP2, MAGED1, MCL1, NFE2L2, NGFR, NSD2, RAC1, RACK1, SFN, SLC2A1, SOD2, SRPX2, UBQLN4, VRK1, YAP1, YBX1, and ZFX expression at mRNA levels. B Venn diagram analysis of simultaneously downregulated genes in TE-1 and Eca109 cells in response to TAPI-1 treatment. C TE-1 and Eca109 cells were exposed to different concentrations of cisplatin (0, 5, 10, 15, 20, 25 μM) with DMSO or TAPI-1 (5 μM) for 48 h. CCK-8 analysis of cell viability and IC50 values of cisplatin. *P < 0.05, **P < 0.01, and ***P < 0.001, compared with the DMSO group. D TE-1 and Eca109 cells were exposed to cisplatin (10 μM) with DMSO or TAPI-1 (5 μM) for 48 h. Apoptosis analysis using flow cytometry and quantitative analysis of apoptotic cells. *P < 0.05 and **P < 0.01, compared with the DMSO + Cisplatin group
Next, TE-1 and Eca109 cells were exposed to TAPI-1 (5 μM) or solvent control, meanwhile exposed to different concentrations of cisplatin (0, 5, 10, 15, 20, 25 μM) for 48 h. CCK-8 assay revealed that TAPI-1 treatment increased the sensitivity of ESCC cells to cisplatin (Fig. 3C). The IC50 of cisplatin in TE-1 and Eca109 cells with TAPI-1 was significantly lower than that of the control cells (Fig. 3C). To further confirm the effect of TAPI-1 on cisplatin resistance in ESCC cells, TE-1 and Eca109 cells were exposed to TAPI-1 (5 μM) or solvent control, meanwhile exposed to cisplatin (10 μM) for 48 h, following which, flow cytometry assay was carried out. As expected, TAPI-1 (5 μM) significantly aggravated TE-1 and Eca109 cell apoptosis in response to cisplatin (10 μM) to varying degrees, respectively (Fig. 3D). These data indicated that TAPI-1 may be a promising adjuvant agent with cisplatin to therapy ESCC.
TAPI-1 Suppresses NF-κB Signaling Pathway in ESCC Cells
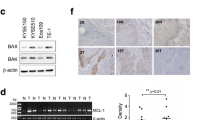
To investigate potential molecular mechanisms behind the anti-tumor efficacy of TAPI-1, we speculated the putative transcription factors in the upstream of cisplatin resistance-related genes using the ChEA3 website. Ten transcription factors with the highest scores from the Enrichr Queries library are listed in Fig. 4A. Thus, we hypothesized that RELA (also named NF-κB p65) signaling may be influenced by TAPI-1. To investigate the role of TAPI-1 in the activation of NF-κB signaling pathway in ESCC cells, TE-1 and Eca109 cells were exposed to TAPI-1 (10 μM) for 12 h, following which, Western blot and ChIP assays were carried out, respectively. As shown in Fig. 4B, TAPI-1 treatment remarkably suppressed the activation of NF-κB signaling, as indicated by the decreased phosphorylation and nuclear translocation of NF-κB p65 in TE-1 and Eca109 cells. Furthermore, ChIP was performed to assess the binding intensity of NF-κB p65 to the promoter regions of target genes. Tumor growth and metastasis-related genes c-Myc, CyclinD1, and MMP2 [17] were previously reported as target genes of NF-κB p65 and were chosen for ChIP-qPCR to evaluate the activation of NF-κB signaling in ESCC cells. ChIP-qPCR revealed that NF-κB p65 pulled fewer promoter segments of c-Myc, CyclinD1, and MMP2 in response to TAPI-1(10 μM) treatment in TE-1 and Eca109 cells, respectively (Fig. 4C).
The effect of TAPI-1 on NF-κB signaling pathway in ESCC cells. A The predicted transcription factors in the upstream of cisplatin resistance-related genes. TE-1 and Eca109 cells were exposed to DMSO or TAPI-1 (10 μM) for 12 h. B Western blot analysis of P-NF-κB p65 and NF-κB p65 protein expression in total lysate and NF-κB p65 protein expression in nuclear lysate. C ChIP-qPCR quantitative analysis of the enrichment of NF-κB p65 at the indicated gene promoter regions. *P < 0.05, **P < 0.01, and ***P < 0.001, compared with the DMSO group
Suppression of NF-κB Signaling Abolishes Anti-tumor Effects of TAPI-1 on ESCC Cells
To further confirm the role of NF-κB signaling pathway in TAPI-1-mediated anti-tumor efficacy in ESCC cells, we blocked NF-κB signaling using BAY11-7082 (10 μM) and observed the decreased phosphorylation of NF-κB p65 in TE-1 and Eca109 cells, respectively (Fig. 5A). CCK-8 assay indicated that the treatment with BAY11-7082 alone or in combination with TAPI-1 could remarkably inhibit ESCC cell viability. However, following the treatment with BAY11-7082, we observed no obvious alteration in ESCC cell viability in the presence or absence of TAPI-1 (Fig. 5B). ESCC cells were pretreated with BAY11-7082 (10 μM) for 6 h, followed by exposure to TAPI-1 (5 μM) for another 24 h. Subsequently, migration and invasion assays were conducted and showed no change of migration and invasion in ESCC cells (Fig. 5C, D). Moreover, CCK-8 assay revealed that a pretreatment with BAY11-7082 (10 μM) could neutralize TAPI-1-enhanced the sensitivity of ESCC cells to different concentrations of cisplatin (0, 5, 10, 15, 20, 25 μM) (Fig. 5E). Taken together, BAY11-7082 abolished anti-tumor effects of TAPI-1 on ESCC cells, which indicated that TAPI-1 exhibited anti-tumor efficacy in ESCC cells in an anti-NF-κB manner.
Pretreatment of BAY11-7082 abolishes the effects of TAPI-1 on ESCC cell viability, migration, invasion, and cisplatin resistance. A TE-1 and Eca109 cells were exposed to DMSO or BAY11-7082 (10 μM) for 6 h. Western blot analysis of P-NF-κB p65 and NF-κB p65 protein expression in total lysate. B TE-1 and Eca109 cells were treated with DMSO, BAY11-7082 (10 μM), TAPI-1 (10 μM), or BAY11-7082 (10 μM) in combination with TAPI-1 (10 μM) for 0, 24, 48, and 72 h, respectively. CCK-8 analysis of cell viability. TE-1 and Eca109 cells were pretreated with DMSO or BAY11-7082 (10 μM) for 6 h, followed by an exposure to DMSO or TAPI-1 (5 μM) for another 24 h. Representative images and absorbance analysis results of Transwell migration assay (C) and invasion assay (D). E TE-1 and Eca109 cells were pretreated with DMSO or BAY11-7082 (10 μM) for 6 h, followed by an exposure to cisplatin (10 μM) with DMSO or TAPI-1 (5 μM) for another 48 h. *P < 0.05 and **P < 0.01, compared with the DMSO group
Discussion
As a common malignant tumor of the digestive tract, ESCC makes about 0.15% population afflicted [18]. In spite of the use of advanced surgery combined with radiotherapy and various chemotherapeutic agents, patients with ESCC typically have a poor five-year survival rate [19], largely due to high recurrence rate, high metastatic rate, and chemotherapeutic drug resistance. Despite proved to display anti-tumor efficacy on several tumor deceases, the effect of TAPI-1 on digestive tract tumors remains largely unclear.
In the present study, we demonstrated that TAPI-1 treatment could significantly inhibit ESCC cell viability, migration, invasion, and cisplatin resistance. Furthermore, RELA was determined as an indirect molecular target of TAPI-1 that conferred cisplatin resistance in ESCC. RELA, also named NF-κB p65, known as a critical transcription factor relevant to inflammation and cancer, has been extensively studied in human cancers over recent years. Several studies revealed that NF-κB signaling is constitutively activated in ESCC cells [20,21,22], thus facilitating ESCC tumor growth, metastasis and cisplatin resistance [23]. Silencing NF-κB p65 using small interfering RNA effectively alleviated ESCC, implying that NF-κB may be a potential therapeutic target [21, 22]. Several NF-κB inhibitors have been applied for cancer research, such as BAY11-7082. Unfortunately, BAY11-7082 showed toxicity in human normal esophageal epithelial cells [24]. Nevertheless, no significant changes in the cell viability of Het-1A cells were observed in response to varying doses of TAPI-1 (0, 1.25, 2.5, 5, 10, and 20 μM), implying that TAPI-1 may be a desirable anti-cancer drug with low toxicity. Interestingly, similar to our data, several studies showed that TAPI-1 could attenuate Sjogren’s syndrome, cardiac hypertrophy, and pancreatic cancer via suppression of NF-κB signaling pathway [11, 25, 26].
Despite these data, the molecular mechanism for signal transduction from TACE or MMPs to NF-κB remains enigmatic in ESCC. Previous studies have documented that EGFR and VEGFR2, two typical substrates of TACE, contributed to ESCC progression [27, 28]. Moreover, EGFR and VEGFR2 were proved to activate NF-κB signaling in other disease models [29, 30]. However, the roles of EGFR and VEGFR2 in TACE-activated NF-κB signaling in ESCC remain to be elucidated. Intriguingly, we observed that the molecular targets of TAPI-1, including TACE and MMPs, were regarded as the target genes of NF-κB p65 in ESCC cells [17, 31]. Additionally, our previous data revealed that TACE and NF-κB signaling formed a feedback loop to exacerbate ESCC progression [17]. The interrelationships between MMPs and NF-κB signaling in ESCC await future investigation.
Conclusion
Taken together, in this study, for the first time, we demonstrated that TAPI-1 significantly inhibited ESCC cell viability, migration, and invasion and efficiently enhanced the sensitivity of ESCC cells to cisplatin via suppression of NF-κB signaling pathway (Fig. 6).
Schematic diagram of TAPI-1-mediated anti-tumor efficacy in ESCC cells via suppression of NF-κB signaling pathway. The activation of NF-κB signaling is suppressed in response to TAPI-1 treatment via an unknown pathway, leading to the decrease of ESCC cell viability, migration, and invasion and the enhanced sensitivity of ESCC cells to cisplatin
References
Sung H, Ferlay J, Siegel RL et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA: a cancer journal for clinicians. 2021;71:209–49. https://doi.org/10.3322/caac.21660
Abnet CC, Arnold M, Wei WQ. Epidemiology of Esophageal Squamous Cell Carcinoma. Gastroenterology. 2018;154:360–373. https://doi.org/10.1053/j.gastro.2017.08.023.
Mariette C, Balon JM, Piessen G, Fabre S, Van Seuningen I, Triboulet JP. Pattern of recurrence following complete resection of esophageal carcinoma and factors predictive of recurrent disease. Cancer. 2003;97:1616–1623. https://doi.org/10.1002/cncr.11228.
Batra R, Malhotra GK, Singh S, Are C. Managing Squamous Cell Esophageal Cancer. The Surgical clinics of North America. 2019;99:529–541. https://doi.org/10.1016/j.suc.2019.02.006.
Messager M, Warlaumont M, Renaud F et al. Recent improvements in the management of esophageal anastomotic leak after surgery for cancer. European journal of surgical oncology : the journal of the European Society of Surgical Oncology and the British Association of Surgical Oncology. 2017;43:258–269. https://doi.org/10.1016/j.ejso.2016.06.394.
Moss ML, Minond D. Recent Advances in ADAM17 Research: A Promising Target for Cancer and Inflammation. Mediators of inflammation. 2017;2017:9673537. https://doi.org/10.1155/2017/9673537.
Wang Q, Chen X, Feng J et al. Soluble interleukin-6 receptor-mediated innate immune response to DNA and RNA viruses. Journal of virology. 2013;87:11244–11254. https://doi.org/10.1128/jvi.01248-13.
Young J, Yu X, Wolslegel K et al. Lymphotoxin-alphabeta heterotrimers are cleaved by metalloproteinases and contribute to synovitis in rheumatoid arthritis. Cytokine. 2010;51:78–86. https://doi.org/10.1016/j.cyto.2010.03.003.
Bae EH, Kim IJ, Choi HS et al. Tumor necrosis factor α-converting enzyme inhibitor attenuates lipopolysaccharide-induced reactive oxygen species and mitogen-activated protein kinase expression in human renal proximal tubule epithelial cells. The Korean journal of physiology & pharmacology : official journal of the Korean Physiological Society and the Korean Society of Pharmacology. 2018;22:135–143. https://doi.org/10.4196/kjpp.2018.22.2.135.
Martín R, Cordova C, Nieto ML. Secreted phospholipase A2-IIA-induced a phenotype of activated microglia in BV-2 cells requires epidermal growth factor receptor transactivation and proHB-EGF shedding. Journal of neuroinflammation. 2012;9:154. https://doi.org/10.1186/1742-2094-9-154.
Bharadwaj U, Marin-Muller C, Li M, Chen C, Yao Q. Mesothelin overexpression promotes autocrine IL-6/sIL-6R trans-signaling to stimulate pancreatic cancer cell proliferation. Carcinogenesis. 2011;32:1013–1024. https://doi.org/10.1093/carcin/bgr075.
Valdehita A, Bajo AM, Schally AV, Varga JL, Carmena MJ, Prieto JC. Vasoactive intestinal peptide (VIP) induces transactivation of EGFR and HER2 in human breast cancer cells. Molecular and cellular endocrinology. 2009;302:41–48. https://doi.org/10.1016/j.mce.2008.11.024.
Fabre-Lafay S, Garrido-Urbani S, Reymond N, Gonçalves A, Dubreuil P, Lopez M. Nectin-4, a new serological breast cancer marker, is a substrate for tumor necrosis factor-alpha-converting enzyme (TACE)/ADAM-17. The Journal of biological chemistry. 2005;280:19543–19550. https://doi.org/10.1074/jbc.M410943200.
Desbois-Mouthon C, Cacheux W, Blivet-Van Eggelpoël MJ et al. Impact of IGF-1R/EGFR cross-talks on hepatoma cell sensitivity to gefitinib. International journal of cancer. 2006;119:2557–2566. https://doi.org/10.1002/ijc.22221.
Li X, Lv Y, Yuan A, Yi S, Ma Y, Li Z. Gastrin-releasing peptide promotes the growth of HepG2 cells via EGFR-independent ERK1/2 activation. Oncology reports. 2010;24:441–448. https://doi.org/10.3892/or_00000877.
Groblewska M, Siewko M, Mroczko B, Szmitkowski M. The role of matrix metalloproteinases (MMPs) and their inhibitors (TIMPs) in the development of esophageal cancer. Folia histochemica et cytobiologica. 2012;50:12–19. https://doi.org/10.5603/FHC.2012.0002.
Gao L, Liu H, Xu R et al. ADAM17 and NF-κB p65 form a positive feedback loop that facilitates human esophageal squamous cell carcinoma cell viability. International journal of clinical and experimental pathology. 2021;14:845–54 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8339723
Enzinger PC, Mayer RJ. Esophageal cancer. The New England journal of medicine. 2003;349:2241–2252. https://doi.org/10.1056/NEJMra035010.
Kim T, Grobmyer SR, Smith R et al. Esophageal cancer–the five year survivors. Journal of surgical oncology. 2011;103:179–183. https://doi.org/10.1002/jso.21784.
Liu H, Yang J, Yuan Y et al. Regulation of Mcl-1 by constitutive activation of NF-κB contributes to cell viability in human esophageal squamous cell carcinoma cells. BMC cancer. 2014;14:98. https://doi.org/10.1186/1471-2407-14-98.
Tian F, Zang WD, Hou WH, Liu HT, Xue LX. Nuclear factor-kB signaling pathway constitutively activated in esophageal squamous cell carcinoma cell lines and inhibition of growth of cells by small interfering RNA. Acta biochimica et biophysica Sinica. 2006;38:318–326. https://doi.org/10.1111/j.1745-7270.2006.00166.x.
Tian F, Zhang C, Tian W, Jiang Y, Zhang X. Comparison of the effect of p65 siRNA and curcumin in promoting apoptosis in esophageal squamous cell carcinoma cells and in nude mice. Oncology reports. 2012;28:232–240. https://doi.org/10.3892/or.2012.1777.
Li B, Li YY, Tsao SW, Cheung AL. Targeting NF-kappaB signaling pathway suppresses tumor growth, angiogenesis, and metastasis of human esophageal cancer. Molecular cancer therapeutics. 2009;8:2635–2644. https://doi.org/10.1158/1535-7163.MCT-09-0162.
Bus P, Siersema PD, Verbeek RE, van Baal JW. Upregulation of miRNA-143, -145, -192, and -194 in esophageal epithelial cells upon acidic bile salt stimulation. Diseases of the esophagus : official journal of the International Society for Diseases of the Esophagus. 2014;27:591–600. https://doi.org/10.1111/dote.12112.
Lisi S, Sisto M, Lofrumento DD, D'Amore M. Sjögren's syndrome autoantibodies provoke changes in gene expression profiles of inflammatory cytokines triggering a pathway involving TACE/NF-κB. Laboratory investigation; a journal of technical methods and pathology. 2012;92:615–24. https://doi.org/10.1038/labinvest.2011.190
Miao K, Zhou L, Ba H et al. Transmembrane tumor necrosis factor alpha attenuates pressure-overload cardiac hypertrophy via tumor necrosis factor receptor 2. PLoS biology. 2020;18:e3000967. https://doi.org/10.1371/journal.pbio.3000967.
Shi ZZ, Wang WJ, Chen YX et al. The miR-1224-5p/TNS4/EGFR axis inhibits tumour progression in oesophageal squamous cell carcinoma. Cell death & disease. 2020;11:597. https://doi.org/10.1038/s41419-020-02801-6.
Wu ZY, Chen T, Zhao Q et al. Altered expression of endogenous soluble vascular endothelial growth factor receptor-2 is involved in the progression of esophageal squamous cell carcinoma. The journal of histochemistry and cytochemistry : official journal of the Histochemistry Society. 2013;61:340–347. https://doi.org/10.1369/0022155413480181.
Liu X, Yue C, Shi L et al. MALT1 is a potential therapeutic target in glioblastoma and plays a crucial role in EGFR-induced NF-κB activation. Journal of cellular and molecular medicine. 2020;24:7550–7562. https://doi.org/10.1111/jcmm.15383.
Sisto M, Lisi S, Lofrumento DD, D’Amore M, Frassanito MA, Ribatti D. Sjögren’s syndrome pathological neovascularization is regulated by VEGF-A-stimulated TACE-dependent crosstalk between VEGFR2 and NF-κB. Genes and immunity. 2012;13411–420. https://doi.org/10.1038/gene.2012.9.
Wang X, Li X, Li C, He C, Ren B, Deng Q, Gao W, Wang B. Aurora-A modulates MMP-2 expression via AKT/NF-κB pathway in esophageal squamous cell carcinoma cells. Acta Biochim Biophys Sin (Shanghai). 2016;48:520–527. https://doi.org/10.1093/abbs/gmw030.
Wu K, Yang Y, Zhao J, Zhao S. BAG3-mediated miRNA let-7g and let-7i inhibit proliferation and enhance apoptosis of human esophageal carcinoma cells by targeting the drug transporter ABCC10. Cancer letters. 2016;371:125–133. https://doi.org/10.1016/j.canlet.2015.11.031.
Xiao Y, Lin FT, Lin WC. ACTL6A promotes repair of cisplatin-induced DNA damage, a new mechanism of platinum resistance in cancer. Proceedings of the National Academy of Sciences of the United States of America. 2021;118(3). https://doi.org/10.1073/pnas.2015808118
Zhang ZF, Huang TJ, Zhang XK et al. AKR1C2 acts as a targetable oncogene in esophageal squamous cell carcinoma via activating PI3K/AKT signaling pathway. Journal of cellular and molecular medicine. 2020;24:9999–10012. https://doi.org/10.1111/jcmm.15604.
Li Z, Li S, Wen Y, Chen J, Liu K, Jia J. MiR-495 Inhibits Cisplatin Resistance and Angiogenesis in Esophageal Cancer by Targeting ATP7A. Technology in cancer research & treatment. 2021;20:15330338211039128. https://doi.org/10.1177/15330338211039127.
Li F, Zhang Z, Wang P et al. ALC1 knockdown enhances cisplatin cytotoxicity of esophageal squamous cell carcinoma cells by inhibition of glycolysis through PI3K/Akt pathway. Life sciences. 2019;232:116679. https://doi.org/10.1016/j.lfs.2019.116679.
Mohapatra P, Shriwas O, Mohanty S et al. CMTM6 drives cisplatin resistance by regulating Wnt signaling through the ENO-1/AKT/GSK3β axis. JCI insight. 2021;6(4). https://doi.org/10.1172/jci.insight.143643
Liu T, Li R, Zhao H et al. eIF4E promotes tumorigenesis and modulates chemosensitivity to cisplatin in esophageal squamous cell carcinoma. Oncotarget. 2016;7:66851–64. https://doi.org/10.18632/oncotarget.11694
Liu X, Ma W, Yan Y, Wu S. Silencing HMGN5 suppresses cell growth and promotes chemosensitivity in esophageal squamous cell carcinoma. Journal of biochemical and molecular toxicology. 2017;31(12). https://doi.org/10.1002/jbt.21996
Ogino S, Konishi H, Ichikawa D et al. Glutathione S-transferase Pi 1 is a valuable predictor for cancer drug resistance in esophageal squamous cell carcinoma. Cancer science. 2019;110:795–804. https://doi.org/10.1111/cas.13896.
Liu X, Yan Y, Ma W, Wu S. Knockdown of frizzled-7 inhibits cell growth and metastasis and promotes chemosensitivity of esophageal squamous cell carcinoma cells by inhibiting Wnt signaling. Biochemical and biophysical research communications. 2017;4901112–1118. https://doi.org/10.1016/j.bbrc.2017.06.185.
Zhou T, Fu H, Dong B et al. HOXB7 mediates cisplatin resistance in esophageal squamous cell carcinoma through involvement of DNA damage repair. Thoracic cancer. 2020;11:3071–3085. https://doi.org/10.1111/1759-7714.13142.
Hou S, Jin W, Xiao W et al. Integrin α5 promotes migration and cisplatin resistance in esophageal squamous cell carcinoma cells. American journal of cancer research. 2019;9:2774–88 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6943361
Xu Z, Zou L, Ma G et al. Integrin β1 is a critical effector in promoting metastasis and chemo-resistance of esophageal squamous cell carcinoma. American journal of cancer research. 2017;7:531–42 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5385641
Li L, Wang W, Zhang R et al. High expression of LAMP2 predicts poor prognosis in patients with esophageal squamous cell carcinoma. Cancer biomarkers : section A of Disease markers. 2017;19:305–311. https://doi.org/10.3233/CBM-160469.
Yang Q, Pan Q, Li C, Xu Y, Wen C, Sun F. NRAGE is involved in homologous recombination repair to resist the DNA-damaging chemotherapy and composes a ternary complex with RNF8-BARD1 to promote cell survival in squamous esophageal tumorigenesis. Cell death and differentiation. 2016;23:1406–1416. https://doi.org/10.1038/cdd.2016.29.
Yu X, Li W, Xia Z et al. Targeting MCL-1 sensitizes human esophageal squamous cell carcinoma cells to cisplatin-induced apoptosis. BMC cancer. 2017;17:449. https://doi.org/10.1186/s12885-017-3442-y.
Zuo J, Zhao M, Fan Z et al. MicroRNA-153-3p regulates cell proliferation and cisplatin resistance via Nrf-2 in esophageal squamous cell carcinoma. Thoracic cancer. 2020;11:738–747. https://doi.org/10.1111/1759-7714.13326.
Kojima H, Okumura T, Yamaguchi T, Miwa T, Shimada Y, Nagata T. Enhanced cancer stem cell properties of a mitotically quiescent subpopulation of p75NTR-positive cells in esophageal squamous cell carcinoma. International journal of oncology. 2017;51:49–62. https://doi.org/10.3892/ijo.2017.4001.
Xue W, Shen Z, Li L et al. Long non-coding RNAs MACC1-AS1 and FOXD2-AS1 mediate NSD2-induced cisplatin resistance in esophageal squamous cell carcinoma. Molecular therapy Nucleic acids. 2021;23:592–602. https://doi.org/10.1016/j.omtn.2020.12.007.
Zeng RJ, Zheng CW, Gu JE et al. RAC1 inhibition reverses cisplatin resistance in esophageal squamous cell carcinoma and induces downregulation of glycolytic enzymes. Molecular oncology. 2019;13:2010–2030. https://doi.org/10.1002/1878-0261.12548.
Liu B, Wang C, Chen P, Cheng B, Cheng Y. RACKI induces chemotherapy resistance in esophageal carcinoma by upregulating the PI3K/AKT pathway and Bcl-2 expression. OncoTargets and therapy. 2018;11:211–220. https://doi.org/10.2147/OTT.S152818.
Lai KK, Chan KT, Choi MY et al. 14–3-3σ confers cisplatin resistance in esophageal squamous cell carcinoma cells via regulating DNA repair molecules. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine. 2016;37:2127–2136. https://doi.org/10.1007/s13277-015-4018-6.
Sawayama H, Ogata Y, Ishimoto T et al. Glucose transporter 1 regulates the proliferation and cisplatin sensitivity of esophageal cancer. Cancer science. 2019;110:1705–1714. https://doi.org/10.1111/cas.13995.
Zuo J, Zhao M, Liu B et al. TNF-α-mediated upregulation of SOD-2 contributes to cell proliferation and cisplatin resistance in esophageal squamous cell carcinoma. Oncology reports. 2019;42:1497–1506. https://doi.org/10.3892/or.2019.7252.
He F, Wang H, Li Y et al. SRPX2 knockdown inhibits cell proliferation and metastasis and promotes chemosensitivity in esophageal squamous cell carcinoma. Biomedicine & pharmacotherapy. 2019;109:671–678. https://doi.org/10.1016/j.biopha.2018.10.042.
Murakami T, Shoji Y, Nishi T et al. Regulation of MRE11A by UBQLN4 leads to cisplatin resistance in patients with esophageal squamous cell carcinoma. Molecular oncology. 2021;15:1069–1087. https://doi.org/10.1002/1878-0261.12929.
Liu ZC, Cao K, Xiao ZH et al. VRK1 promotes cisplatin resistance by up-regulating c-MYC via c-Jun activation and serves as a therapeutic target in esophageal squamous cell carcinoma. Oncotarget. 2017;8:65642–58. https://doi.org/10.18632/oncotarget.20020
Yan J, Shi L, Lin S, Li Y. MicroRNA-624-mediated ARRDC3/YAP/HIF1α axis enhances esophageal squamous cell carcinoma cell resistance to cisplatin and paclitaxel. Bioengineered. 2021;12:5334–5347. https://doi.org/10.1080/21655979.2021.1938497.
Xu J, Hu Z. Y-box-binding protein 1 promotes tumor progression and inhibits cisplatin chemosensitivity in esophageal squamous cell carcinoma. Biomedicine & pharmacotherapy. 2016;79:17–22. https://doi.org/10.1016/j.biopha.2016.01.037.
Wu J, Zhou Y, Wang T, Jiang C, Gao Y, Wei B. ZFX promotes tumorigenesis and confers chemotherapy resistance in esophageal squamous cell carcinoma. Clinics and research in hepatology and gastroenterology. 2021;45:101586. https://doi.org/10.1016/j.clinre.2020.101586.
Acknowledgments
This study was supported by Postgraduate Research & Practice Innovation Program of Jiangsu Province (SJCX21_1469) and Nantong Science and Technology Project (JC2021001).
Funding
Funding was provided by Graduate Research and Innovation Projects of Jiangsu Province (SJCX21_1469) and Science and Technology Project of Nantong City (JC2021001).
Author information
Authors and Affiliations
Contributions
HL and JQ designed the study and wrote the paper. LG and LL performed the research. JQ analyzed data. DZ revised the paper. All co-authors have reviewed and approved the article before submission.
Corresponding author
Ethics declarations
Conflict of interest
There are no relevant financial or non-financial competing interests to report.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Gao, L., Li, L., Zhang, D. et al. TAPI-1 Exhibits Anti-tumor Efficacy in Human Esophageal Squamous Cell Carcinoma Cells via Suppression of NF-κB Signaling Pathway. Dig Dis Sci 69, 81–94 (2024). https://doi.org/10.1007/s10620-023-08181-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-023-08181-z